Abstract
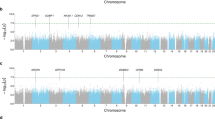
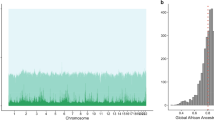
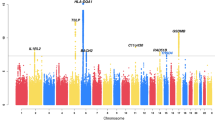
Short-acting β2-adrenergic receptor agonists (SABAs) are the most commonly prescribed asthma medications worldwide. Response to SABAs is measured as bronchodilator drug response (BDR), which varies among racial/ethnic groups in the United States. However, the genetic variation that contributes to BDR is largely undefined in African Americans with asthma. To identify genetic variants that may contribute to differences in BDR in African Americans with asthma, we performed a genome-wide association study (GWAS) of BDR in 949 African-American children with asthma, genotyped with the Axiom World Array 4 (Affymetrix, Santa Clara, CA) followed by imputation using 1000 Genomes phase III genotypes. We used linear regression models adjusting for age, sex, body mass index (BMI) and genetic ancestry to test for an association between BDR and genotype at single-nucleotide polymorphisms (SNPs). To increase power and distinguish between shared vs. population-specific associations with BDR in children with asthma, we performed a meta-analysis across 949 African Americans and 1830 Latinos (total = 2779). Finally, we performed genome-wide admixture mapping to identify regions whereby local African or European ancestry is associated with BDR in African Americans. We identified a population-specific association with an intergenic SNP on chromosome 9q21 that was significantly associated with BDR (rs73650726, p = 7.69 × 10−9). A trans-ethnic meta-analysis across African Americans and Latinos identified three additional SNPs within the intron of PRKG1 that were significantly associated with BDR (rs7903366, rs7070958 and rs7081864, p ≤ 5 × 10−8). Our results failed to replicate in three additional populations of 416 Latinos and 1615 African Americans. Our findings indicate that both population-specific and shared genetic variation contributes to differences in BDR in minority children with asthma, and that the genetic underpinnings of BDR may differ between racial/ethnic groups.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Naqvi M, Thyne S, Choudhry S, Tsai HJ, Navarro D, Castro RA. et al. Ethnic-specific differences in bronchodilator responsiveness among African Americans, Puerto Ricans, and Mexicans with asthma. J Asthma. 2007;44:639–48.
Padhukasahasram B, Yang JJ, Levin AM, Yang M, Burchard EG, Kumar R, et al. Gene-based association identifies SPATA13-AS1 as a pharmacogenomic predictor of inhaled short-acting beta-agonist response in multiple population groups. Pharmacogenomics J 2014;14:365–371.
Palmer LJ, Silverman ES, Weiss ST, Drazen JM. Pharmacogenetics of asthma. Am J Respir Crit Care Med. 2002;165:861–6.
Nelson HS. Beta-adrenergic bronchodilators. N Engl J Med. 1995;333:499–506.
Loza MJ, Penn RB. Regulation of T cells in airway disease by beta-agonist. Front Biosci. 2010;2:969–79.
Salathe M. Effects of beta-agonists on airway epithelial cells. J Allergy Clin Immunol. 2002;110(6 Suppl):S275–81.
Shore SA, Moore PE. Regulation of beta-adrenergic responses in airway smooth muscle. Respir Physiol Neurobiol. 2003;137:179–95.
Jartti T. Asthma, asthma medication and autonomic nervous system dysfunction. Clin Physiol. 2001;21:260–9.
Drake KA, Torgerson DG, Gignoux CR, Galanter JM, Roth LA, Huntsman S, et al. A genome-wide association study of bronchodilator response in Latinos implicates rare variants. J Allergy Clin Immunol. 2014;133:370–8.
Burchard EG, Avila PC, Nazario S, Casal J, Torres A, Rodriguez-Santana JR, et al. Lower bronchodilator responsiveness in Puerto Rican than in Mexican subjects with asthma. Am J Respir Crit Care Med. 2004;169:386–92.
Choudhry S, Ung N, Avila PC, Ziv E, Nazario S, Casal J, et al. Pharmacogenetic differences in response to albuterol between Puerto Ricans and Mexicans with asthma. Am J Respir Crit Care Med. 2005;171:563–70.
Burchard EG, Ziv E, Coyle N, Gomez SL, Tang H, Karter AJ. et al. The importance of race and ethnic background in biomedical research and clinical practice. N Engl J Med. 2003;348:1170–5.
Blake K, Madabushi R, Derendorf H, Lima J. Population pharmacodynamic model of bronchodilator response to inhaled albuterol in children and adults with asthma. Chest. 2008;134:981–9.
Gorina Y. QuickStats: asthma*death rates, by race and age group - United States, 2007–9. In (MMWR) MaMWR (ed). Centers for Disease Control and Prevention. 2012.
Martinez FD, Graves PE, Baldini M, Solomon S, Erickson R. Association between genetic polymorphisms of the beta2-adrenoceptor and response to albuterol in children with and without a history of wheezing. J Clin Invest. 1997;100:3184–8.
Silverman EK, Kwiatkowski DJ, Sylvia JS, Lazarus R, Drazen JM, Lange C, et al. Family-based association analysis of beta2-adrenergic receptor polymorphisms in the childhood asthma management program. J Allergy Clin Immunol. 2003;112:870–6.
Poon AH, Tantisira KG, Litonjua AA, Lazarus R, Xu J, Lasky-Su J, et al. Association of corticotropin-releasing hormone receptor-2 genetic variants with acute bronchodilator response in asthma. Pharm Genom. 2008;18:373–82.
Tantisira KG, Small KM, Litonjua AA, Weiss ST, Liggett SB. Molecular properties and pharmacogenetics of a polymorphism of adenylyl cyclase type 9 in asthma: interaction between beta-agonist and corticosteroid pathways. Hum Mol Genet. 2005;14:1671–7.
Litonjua AA, Lasky-Su J, Schneiter K, Tantisira KG, Lazarus R, Klanderman B, et al. ARG1 is a novel bronchodilator response gene: screening and replication in four asthma cohorts. Am J Respir Crit Care Med. 2008;178:688–94.
Duan QL, Du R, Lasky-Su J, Klanderman BJ, Partch AB, Peters SP, et al. A polymorphism in the thyroid hormone receptor gene is associated with bronchodilator response in asthmatics. Pharm J. 2013;13:130–6.
Reihsaus E, Innis M, MacIntyre N, Liggett SB. Mutations in the gene encoding for the beta 2-adrenergic receptor in normal and asthmatic subjects. Am J Respir Cell Mol Biol. 1993;8:334–9.
Duan QL, Lasky-Su J, Himes BE, Qiu W, Litonjua AA, Damask A, et al. A genome-wide association study of bronchodilator response in asthmatics. Pharm J. 2014;14:41–47.
Israel E, Lasky-Su J, Markezich A, Damask A, Szefler SJ, Schuemann B, et al. Genome-wide association study of short-acting beta2-agonists. A novel genome-wide significant locus on chromosome 2 near ASB3. Am J Respir Crit Care Med. 2015;191:530–7.
Himes BE, Jiang X, Hu R, Wu AC, Lasky-Su JA, Klanderman BJ, et al. Genome-wide association analysis in asthma subjects identifies SPATS2L as a novel bronchodilator response gene. PLoS Genet. 2012;8:e1002824.
Torgerson DG, Ampleford EJ, Chiu GY, Gauderman WJ, Gignoux CR, Graves PE, et al. Meta-analysis of genome-wide association studies of asthma in ethnically diverse North American populations. Nat Genet. 2011;43:887–92.
Galanter JM, Gignoux CR, Torgerson DG, Roth LA, Eng C, Oh SS, et al. Genome-wide association study and admixture mapping identify different asthma-associated loci in Latinos: the Genes-environments & Admixture in Latino Americans study. J Allergy Clin Immunol. 2014;134:295–305.
White MJ, Risse-Adams O, Goddard P, Contreras MG, Adams J, Hu D, et al. Novel genetic risk factors for asthma in African American children: precision medicine and the SAGE II Study. Immunogenetics. 2016;68:391–400.
Winkler CA, Nelson GW, Smith MW. Admixture mapping comes of age. Annu Rev Genom Hum Genet. 2010;11:65–89.
Nishimura KK, Galanter JM, Roth LA, Oh SS, Thakur N, Nguyen EA, et al. Early-life air pollution and asthma risk in minority children. The GALA II and SAGE II studies. Am J Respir Crit Care Med. 2013;188:309–18.
Torgerson DG, Gignoux CR, Galanter JM, Drake KA, Roth LA, Eng C, et al. Case-control admixture mapping in Latino populations enriches for known asthma-associated genes. J Allergy Clin Immunol. 2012;130:76–82 e12.
Gould W, Peterson EL, Karungi G, Zoratti A, Gaggin J, Toma G, et al. Factors predicting inhaled corticosteroid responsiveness in African American patients with asthma. J Allergy Clin Immunol. 2010;126:1131–8.
Moore WC, Bleecker ER, Curran-Everett D, Erzurum SC, Ameredes BT, Bacharier L, et al. Characterization of the severe asthma phenotype by the National Heart, Lung, and Blood Institute’s Severe Asthma Research Program. J Allergy Clin Immunol. 2007;119:405–13.
Moore WC, Meyers DA, Wenzel SE, Teague WG, Li H, Li X, et al. Identification of asthma phenotypes using cluster analysis in the Severe Asthma Research Program. Am J Respir Crit Care Med. 2010;181:315–23.
Standardization of Spirometry. 1994 Update. American Thoracic Society. Am J Respir Crit Care Med. 1995;152:1107–36.
Delaneau O, Zagury JF. Haplotype inference. Methods Mol Biol. 2012;888:177–96.
Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009;5:e1000529.
Howie B, Marchini J, Stephens M. Genotype imputation with thousands of genomes. G3. 2011;1:457–70.
Genomes Project C, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Maples BK, Gravel S, Kenny EE, Bustamante CD. RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am J Hum Genet. 2013;93:278–88.
Proceedings of the ATS workshop on refractory asthma: current understanding, recommendations, and unanswered questions. American Thoracic Society. Am J Respir Crit Care Med. 2000;162:2341–51.
Pino-Yanes M, Thakur N, Gignoux CR, Galanter JM, Roth LA, Eng C, et al. Genetic ancestry influences asthma susceptibility and lung function among Latinos. J Allergy Clin Immunol. 2015;135:228–35.
Baran Y, Pasaniuc B, Sankararaman S, Torgerson DG, Gignoux C, Eng C, et al. Fast and accurate inference of local ancestry in Latino populations. Bioinformatics. 2012;28:1359–67.
Kumar R, Nguyen EA, Roth LA, Oh SS, Gignoux CR, Huntsman S, et al. Factors associated with degree of atopy in Latino children in a nationwide pediatric sample: the Genes-environments and Admixture in Latino Asthmatics (GALA II) study. J Allergy Clin Immunol. 2013;132:896–905 e891.
Alexander DH, Novembre J, Lange K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009;19:1655–64.
McGarry ME, Castellanos E, Thakur N, Oh SS, Eng C, Davis A, et al. Obesity and bronchodilator response in black and Hispanic children and adolescents with asthma. Chest. 2015;147:1591–8.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genome-wide association scans. Bioinformatics. 2010;26:2190–1.
PLUMMER MBN, COWLES K, VINES K. CODA: convergence diagnosis and output analysis for MCMC. R News. 2012;6:7–11.
Sobota RS, Shriner D, Kodaman N, Goodloe R, Zheng W, Gao YT, et al. Addressing population-specific multiple testing burdens in genetic association studies. Ann Hum Genet. 2015;79:136–47.
Marcus JH, Novembre J. Visualizing the geography of genetic variants. Bioinformatics 2016;33:594–595.
Consortium GT. The genotype-tissue expression (GTEx) project. Nat Genet. 2013;45:580–5.
Li Z, Xi X, Gu M, Feil R, Ye RD, Eigenthaler M, et al. A stimulatory role for cGMP-dependent protein kinase in platelet activation. Cell. 2003;112:77–86.
Tamura N, Itoh H, Ogawa Y, Nakagawa O, Harada M, Chun TH, et al. cDNA cloning and gene expression of human type I alpha cGMP-dependent protein kinase. Hypertension. 1996;27(3 Pt 2):552–7.
Orstavik S, Natarajan V, Tasken K, Jahnsen T, Sandberg M. Characterization of the human gene encoding the type I alpha and type I beta cGMP-dependent protein kinase (PRKG1). Genomics. 1997;42:311–8.
Burgoyne JR, Madhani M, Cuello F, Charles RL, Brennan JP, Schroder E, et al. Cysteine redox sensor in PKGIa enables oxidant-induced activation. Science. 2007;317:1393–7.
Pfeifer A, Klatt P, Massberg S, Ny L, Sausbier M, Hirneiss C, et al. Defective smooth muscle regulation in cGMP kinase I-deficient mice. EMBO J. 1998;17:3045–51.
Dawes M, Chowienczyk PJ, Ritter JM. Effects of inhibition of the L-arginine/nitric oxide pathway on vasodilation caused by beta-adrenergic agonists in human forearm. Circulation. 1997;95:2293–7.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Lima JJ, Thomason DB, Mohamed MH, Eberle LV, Self TH, Johnson JA. Impact of genetic polymorphisms of the beta2-adrenergic receptor on albuterol bronchodilator pharmacodynamics. Clin Pharmacol Ther. 1999;65:519–25.
Taylor DR, Drazen JM, Herbison GP, Yandava CN, Hancox RJ, Town GI. Asthma exacerbations during long term beta agonist use: influence of beta(2) adrenoceptor polymorphism. Thorax. 2000;55:762–7.
Israel E, Drazen JM, Liggett SB, Boushey HA, Cherniack RM, Chinchilli VM, et al. The effect of polymorphisms of the beta(2)-adrenergic receptor on the response to regular use of albuterol in asthma. Am J Respir Crit Care Med. 2000;162:75–80.
Bustamante CD, Burchard EG, De la Vega FM. Genomics for the world. Nature. 2011;475:163–5.
Popejoy AB, Fullerton SM. Genomics is failing on diversity. Nature. 2016;538:161–4.
Editors PM, Rid A, Johansson MA, Leung G, Valantine H, Burchard EG, et al. Towards equity in health: researchers take stock. PLoS Med. 2016;13:e1002186.
Acknowledgements
This work was supported in part by the Sandler Family Foundation, the American Asthma Foundation, the RWJF Amos Medical Faculty Development Program, National Institutes of Health 1R01HL117004, R01Hl128439, National Institute of Health and Environmental Health Sciences R01 ES015794, R21ES24844, National Institute on Minority Health and Health Disparities 1P60 MD006902, U54MD009523, 1R01MD010443, and the National Institutes of Health National Heart, Lung, and Blood Institute K08 HL118128, U10 HL109164, HL69116, R01 HL69167, HL69170, HL69174 and U10 HL098103. This project has been funded part with federal funds from the National Cancer Institute, National Institutes of Health, under contract HHSN26120080001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products or organizations imply endorsement by the U.S. Government. This research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research. MLS was supported in part by a National Science Foundation Graduate Research Fellowship under grant no. 1144247. MP-Y was funded by the Ramón y Cajal Program (RYC-2015–17205) by the Spanish Ministry of Economy and Competitiveness. MP-Y was also supported by award number AC15/00015 by Instituto de Salud Carlos III thorough AES and EC within AAL framework, and the SysPharmPedia grant awarded from the ERACoSysMed 1st Joint Transnational Call from the European Union under the Horizon 2020; DGT was supported in part by the California Institute for Quantitative Biosciences (QB3). JG was supported in part by NIH Training Grant T32 (5T32GM007546) and career development awards from the NHLBI K23 (5K23HL111636) and NCATS KL2 (5KL2TR000143), as well as the Hewett Fellowship; NT was supported in part by a institutional training grant from the NIGMS (T32GM007546) and career development awards from the NHLBI (5K12HL119997 and K23-HL125551-01A1), Parker B. Francis Fellowship Program, and the American Thoracic Society; RK was supported with a career development award from the NHLBI (5K23HL093023); HJF was supported in part by the GCRC (RR00188); PCA was supported in part by the Ernest S. Bazley Grant. LKW received grant support from the Fund for Henry Ford Hospital, the American Asthma Foundation and the following NIH institutes: NHLBI (R01HL118267, R01HL079055), NIAID (R01AI079139) and NIDDK (R01DK064695). Study accession numbers in dbGaP are phs000921.v2.p1 and phs001180.v1.p1. The authors acknowledge the patients, families, recruiters, health-care providers and community clinics for their participation in SAGE and GALA II. In particular, we thank study coordinator Sandra Salazar and the recruiters who obtained the data: Duanny Alva, MD; Gaby Ayala-Rodriguez; Lisa Caine; Elizabeth Castellanos; Jaime Colon; Denise DeJesus; Blanca Lopez; Brenda Lopez, MD; Louis Martos; Vivian Medina; Juana Olivo; Mario Peralta; Esther Pomares, MD; Jihan Quraishi; Johanna Rodriguez; Shahdad Saeedi; Dean Soto; and Ana Taveras. The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Spear, M.L., Hu, D., Pino-Yanes, M. et al. A genome-wide association and admixture mapping study of bronchodilator drug response in African Americans with asthma. Pharmacogenomics J 19, 249–259 (2019). https://doi.org/10.1038/s41397-018-0042-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-018-0042-4
This article is cited by
-
Epigenomic response to albuterol treatment in asthma-relevant airway epithelial cells
Clinical Epigenetics (2023)
-
Gene expression in African Americans, Puerto Ricans and Mexican Americans reveals ancestry-specific patterns of genetic architecture
Nature Genetics (2023)
-
Tractor uses local ancestry to enable the inclusion of admixed individuals in GWAS and to boost power
Nature Genetics (2021)
-
An epistatic interaction between pre-natal smoke exposure and socioeconomic status has a significant impact on bronchodilator drug response in African American youth with asthma
BioData Mining (2020)