Abstract
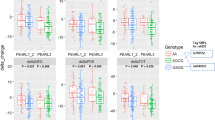
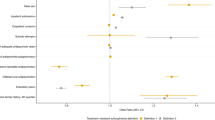
Voltage-gated calcium channels have been implicated in schizophrenia aetiology; however, little is known about their involvement in antipsychotic treatment response. This study investigated variants within the calcium channel subunit genes for association with antipsychotic treatment response in a first episode schizophrenia cohort. Twelve regulatory variants within seven genes were shown to be significantly associated with treatment outcome. Most notably, the CACNA1B rs2229949 CC genotype was associated with improved negative symptomology, where the C allele was predicted to abolish a miRNA-binding site (has-mir-5002-3p), suggesting a possible mechanism of action through which this variant may have an effect. These results implicate the calcium channel subunits in antipsychotic treatment response and suggest that increased activation of these channels may be explored to enhance or predict antipsychotic treatment outcome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–9.
Psychiatric GWAS Consortium Bipolar Disorder Working Group. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat Genet. 2011;43:977–83.
Grebb JA, Shelton RC, Taylor EH, Bigelow LB. A negative, double-blind, placebo-controlled, clinical trial of verapamil in chronic schizophrenia. Biol Psychiatry. 1986;21:691–4.
Price WA. Antipsychotic effects of verapamil in schizophrenia. Hillside J Clin Psychiatry. 1987;9:225–30.
Yamada K, Kanba S, Ashikari I, Ohnishi K, Yagi G, Asai M. Nilvadipine is effective for chronic schizophrenia in a double-blind placebo-controlled study. J Clin Psychopharmacol. 1996;16:437–9.
Suddath RL, Straw GM, Freed WJ, Bigelow LB, Kirch DG, Wyatt RJ. A clinical trial of nifedipine in schizophrenia and tardive dyskinesia. Pharmacol Biochem Behav. 1991;39:743–5.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 4th Edition (DSM-IV-TR). 2000.https://doi.org/10.1176/appi.books.9780890423349
Drogemöller BI, Niehaus DJH, Chiliza B, van der Merwe L, Asmal L, Malhotra AK, et al. Patterns of variation influencing antipsychotic treatment outcomes in South African first-episode schizophrenia patients. Pharmacogenomics. 2014;15:189–99.
Chiliza B, Ojagbemi A, Esan O, Asmal L, Oosthuizen P, Kidd M, et al. Combining depot antipsychotic with an assertive monitoring programme for treating first-episode schizophrenia in a resource-constrained setting. Early Interv Psychiatry. 2016;10:54–62.
Chiliza B, Asmal L, Kilian S, Phahladira L, Emsley R. Rate and predictors of non-response to first-line antipsychotic treatment in first-episode schizophrenia. Hum Psychopharmacol. 2015;30:173–82.
Ovenden ES, Drögemöller BI, van der Merwe L, Chiliza B, Asmal L, Emsley RA, et al. Fine-mapping of antipsychotic response genome-wide association studies reveals novel regulatory mechanisms. Pharmacogenomics. 2017;18:105–20.
Kay SR, Fiszbein A, Opler LA. The positive and negative syndrome scale (PANSS) for schizophrenia. Schizophr Bull. 1987;13:261–76.
Purcell S, Chang C. Plink v1.9. https://www.cog-genomics.org/plink1.9. 2017
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Daya M, van der Merwe L, Galal U, Möller M, Salie M, Chimusa ER, et al. A panel of ancestry informative markers for the complex five-way admixed South African coloured population. PLoS ONE. 2013;8:e82224.
Alexander DH, Novembre J, Lange K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res.2009;19:1655–64.
R Core Team. R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. 2017.
Bate D, Mächler M, Bolker B, Walker S, Fitting linear mixed-effects models using lme4. J Stat Softw. 2015;67:1–48.
Kuznetsova A, Brockhoff P, Bojesen Christensen R. lmerTest: tests in linear mixed effects models. R package version 2.0–33. 2016. https://CRAN.R-project.org/package=lmerTest.
Chelala C, Khan A, Lemoine NR. SNPnexus: a web database for functional annotation of newly discovered and public domain single nucleotide polymorphisms. Bioinformatics. 2009;25:655–61.
Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073–81.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Guo L, Du Y, Chang S, Zhang K, Wang J. rSNPBase: a database for curated regulatory SNPs. Nucleic Acids Res. 2014;42:D1033–1039.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Xia K, Shabalin AA, Huang S, Madar V, Zhou Y-H, Wang W, et al. seeQTL: a searchable database for human eQTLs. Bioinformatics. 2012;28:451–2.
Gamazon ER, Zhang W, Konkashbaev A, Duan S, Kistner EO, Nicolae DL, et al. SCAN: SNP and copy number annotation. Bioinformatics. 2010;26:259–62.
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45:580–5.
Bhattacharya A, Ziebarth JD, Cui Y. PolymiRTS Database 3.0: linking polymorphisms in microRNAs and their target sites with human diseases and biological pathways. Nucleic Acids Res. 2014;42:D86–D91.
Ziebarth JD, Bhattacharya A, Chen A, Cui Y. PolymiRTS Database 2.0: linking polymorphisms in microRNA target sites with human diseases and complex traits. Nucleic Acids Res. 2012;40:D216–D221.
Agarwal V, Bell GW, Nam J-W, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs. eLife. 4, e05005 https://doi.org/10.7554/eLife.05005.
Cahill ME, Xie Z, Day M, Photowala H, Barbolina MV, Miller CA, et al. Kalirin regulates cortical spine morphogenesis and disease-related behavioral phenotypes. Proc Natl Acad Sci USA. 2009;106:13058–63.
Sommer JE, Budreck EC. Kalirin-7: linking spine plasticity and behavior. J Neurosci J Soc Neurosci. 2009;29:5367–9.
Xie Z, Srivastava DP, Photowala H, Kai L, Cahill ME, Woolfrey KM, et al. Kalirin-7 controls activity-dependent structural and functional plasticity of dendritic spines. Neuron. 2007;56:640–56.
Wishart DS, Knox C, Guo AC, Shrivastava S, Hassanali M, Stothard P, et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 2006;34:D668–672.
Uzbay T, Goktalay G, Kayir H, Eker SS, Sarandol A, Oral S, et al. Increased plasma agmatine levels in patients with schizophrenia. J Psychiatr Res. 2013;47:1054–60.
Gambardella A, Labate A, Colosimo E, Ambrosio R, Quattrone A. Monotherapy for partial epilepsy: focus on levetiracetam. Neuropsychiatr Dis Treat. 2008;4:33–38.
Bhat S, Dao DT, Terrillion CE, Arad M, Smith RJ, Soldatov NM, et al. CACNA1C (Cav1.2) in the pathophysiology of psychiatric disease. Prog Neurobiol. 2012;99:1–14.
Gershon ES, Grennan K, Busnello J, Badner JA, Ovsiew F, Memon S, et al. A rare mutation of CACNA1C in a patient with bipolar disorder, and decreased gene expression associated with a bipolar-associated common SNP of CACNA1C in brain. Mol Psychiatry. 2014;19:890–4.
Wolf C, Mohr H, Schneider-Axmann T, Reif A, Wobrock T, Scherk H, et al. CACNA1C genotype explains interindividual differences in amygdala volume among patients with schizophrenia. Eur Arch Psychiatry Clin Neurosci. 2014;264:93–102.
Heyes S, Pratt WS, Rees E, Dahimene S, Ferron L, Owen MJ, et al. Genetic disruption of voltage-gated calcium channels in psychiatric and neurological disorders. Prog Neurobiol. 2015;134:36–54.
Green EK, Grozeva D, Jones I, Jones L, Kirov G, Caesar S, et al. The bipolar disorder risk allele at CACNA1C also confers risk of recurrent major depression and of schizophrenia. Mol Psychiatry. 2010;15:1016–22.
Majumdar D, Nebhan CA, Hu L, Anderson B, Webb DJ. An APPL1/Akt signaling complex regulates dendritic spine and synapse formation in hippocampal neurons. Mol Cell Neurosci. 2011;46:633–44.
Wang Y, Wang J, Wang S, Liu S-S, Cao J, Li X, et al. Adaptor protein APPL1 couples synaptic NMDA receptor with neuronal prosurvival phosphatidylinositol 3-kinase/Akt pathway. J Neurosci J Soc Neurosci. 2012;32:11919–29.
Keefe RSE, Harvey PD. Cognitive impairment in schizophrenia. In: Geyer MA, Gross G, editors. Novel antischizophrenia treatments. Berlin Heidelberg: Springer; 2012. p. 11–37.
Leucht S, Cipriani A, Spineli L, Mavridis D, Orey D, Richter F, et al. Comparative efficacy and tolerability of 15 antipsychotic drugs in schizophrenia: a multiple-treatments meta-analysis. Lancet. 2013;382:951–62.
Acknowledgements
The work reported here was supported by the following grants provided by the South African National Research Foundation (NRF), i.e., KSO'C was funded by the Scarce Skills Post-Doctoral Fellowship (grant no. 96833) and LW was funded by the Competitive Program for Rated Researchers (grant no. 93498) and the Bioinformatics and Functional Genomics Program (grant no. 93681). The opinions expressed and conclusions arrived at are those of the authors and are not necessarily attributed to these funding sources.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
O’Connell, K.S., McGregor, N.W., Malhotra, A. et al. Variation within voltage-gated calcium channel genes and antipsychotic treatment response in a South African first episode schizophrenia cohort. Pharmacogenomics J 19, 109–114 (2019). https://doi.org/10.1038/s41397-018-0033-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-018-0033-5