Abstract
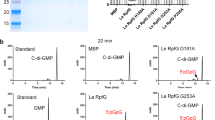
Proteobacteria primarily utilize acyl-homoserine lactones (AHLs) as quorum-sensing signals for intra-/interspecies communication to control pathogen infections. Enzymatic degradation of AHL represents the major quorum-quenching mechanism that has been developed as a promising approach to prevent bacterial infections. Here we identified a novel quorum-quenching mechanism revealed by an effector of the type IVA secretion system (T4ASS) in bacterial interspecies competition. We found that the soil antifungal bacterium Lysobacter enzymogenes OH11 (OH11) could use T4ASS to deliver the effector protein Le1288 into the cytoplasm of another soil microbiome bacterium Pseudomonas fluorescens 2P24 (2P24). Le1288 did not degrade AHL, whereas its delivery to strain 2P24 significantly impaired AHL production through binding to the AHL synthase PcoI. Therefore, we defined Le1288 as LqqE1 (Lysobacter quorum-quenching effector 1). Formation of the LqqE1-PcoI complex enabled LqqE1 to block the ability of PcoI to recognize/bind S-adenosy-L-methionine, a substrate required for AHL synthesis. This LqqE1-triggered interspecies quorum-quenching in bacteria seemed to be of key ecological significance, as it conferred strain OH11 a better competitive advantage in killing strain 2P24 via cell-to-cell contact. This novel quorum-quenching also appeared to be adopted by other T4ASS-production bacteria. Our findings suggest a novel quorum-quenching that occurred naturally in bacterial interspecies interactions within the soil microbiome by effector translocation. Finally, we presented two case studies showing the application potential of LqqE1 to block AHL signaling in the human pathogen Pseudomonas aeruginosa and the plant pathogen Ralstonia solanacearum.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
We are sorry, but there is no personal subscription option available for your country.
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The sequence data from the present study have been submitted to the NCBI GenBank under the following accession numbers: MW052467 (Le0908), MW052468 (Le0989), MW052469 (Le1288), MW052471 (Le1519), MW052472 (Le1841), MW052473 (Le3180), MW052474 (Le3316), MW052488 (Le4230), MW052489 (Le4232), MW052491 (Le4236), MW052493 (Le4253), MW052494 (Le4579), MW052495 (Le4949), OQ192195 (La0125), OQ192196 (Lg4270), OQ179943 (16s rRNA), OQ192197 (GyrB), OQ192199 (RpoB), and OQ192198 (RpoD).
References
Papenfort K, Bassler BL. Quorum sensing signal–response systems in Gram-negative bacteria. Nat Rev Microbiol. 2016;14:576–88.
Deng Y, Wu JE, Tao F, Zhang L. Listening to a new language: DSF-based quorum sensing in Gram-negative bacteria. Chem Rev. 2011;111:160–73.
Galloway WRJD, Hodgkinson JT, Bowden SD, Welch M, Spring DR. Quorum sensing in Gram-negative bacteria: small-molecule modulation of AHL and AI-2 quorum sensing pathways. Chem Rev. 2011;111:28–67.
Whiteley M, Diggle SP, Greenberg EP. Progress in and promise of bacterial quorum sensing research. Nature. 2017;551:313–20.
Miller MB, Bassler BL. Quorum sensing in bacteria. Annu Rev Microbiol. 2001;55:165–99.
González JE, Marketon MM. Quorum sensing in nitrogen-fixing Rhizobia. Microbiol Mol Biol Rev. 2003;67:574–92.
Federle MJ, Bassler BL. Interspecies communication in bacteria. J Clin Investig. 2003;112:1291–9.
Waters CM, Bassler BL. Quorum sensing: cell-to-cell communication in bacteria. Annu Rev Cell Dev Biol. 2005;21:319–46.
Fuqua WC, Winans SC, Greenberg EP. Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators. J Bacteriol. 1994;176:269–75.
Engebrecht J, Nealson K, Silverman M. Bacterial bioluminescence: Isolation and genetic analysis of functions from Vibrio fischeri. Cell. 1983;32:773–81.
Zhang L, Murphy PJ, Kerr A, Tate ME. Agrobacterium conjugation and gene regulation by N-acyl-L-homoserine lactones. Nature. 1993;362:446–8.
de Kievit TR, Iglewski BH. Bacterial quorum sensing in pathogenic relationships. Infect Immun. 2000;68:4839–49.
Parsek MR, Val DL, Hanzelka BL, Cronan JJ, Greenberg EP. Acyl homoserine-lactone quorum-sensing signal generation. Proc Natl Acad Sci USA. 1999;96:4360–5.
Schaefer AL, Val DL, Hanzelka BL, Cronan JE, Greenberg EP. Generation of cell-to-cell signals in quorum sensing: acyl homoserine lactone synthase activity of a purified Vibrio fischeri LuxI protein. Proc Natl Acad Sci USA. 1996;93:9505–9.
Dong S, Frane ND, Christensen QH, Greenberg EP, Nagarajan R, Nair SK. Molecular basis for the substrate specificity of quorum signal synthases. Proc Natl Acad Sci USA. 2017;114:9092–7.
Moré MI, Finger LD, Stryker JL, Fuqua C, Eberhard A, Winans SC. Enzymatic synthesis of a quorum-sensing autoinducer through use of defined substrates. Science. 1996;272:1655–8.
Watson WT, Minogue TD, Val DL, von Bodman SB, Churchill MEA. Structural basis and specificity of acyl-homoserine lactone signal production in bacterial quorum sensing. Mol Cell. 2002;9:685–94.
Raychaudhuri A, Tullock A, Tipton PA. Reactivity and reaction order in acyl homoserine lactone formation by Pseudomonas aeruginosa RhlI. Biochemistry. 2008;47:2893–8.
Pearson JP, Feldman M, Iglewski BH, Prince AA. Pseudomonas aeruginosa cell-to-cell signaling is required for virulence in a model of acute pulmonary infection. Infect Immun. 2000;68:4331–4.
Pirhonen M, Flego D, Heikinheimo R, Palva ET. A small diffusible signal molecule is responsible for the global control of virulence and exoenzyme production in the plant pathogen Erwinia carotovora. The EMBO Journal. 1993;12:2467–76.
Zhang X, Zhang L, Wang L, Zhang H, Xu J, Dong Y. Quenching quorum-sensingdependent bacterial infectionby an N-acyl homoserine lactonase. Nature. 2001;411:813–7.
Sikdar R, Elias M. Quorum quenching enzymes and their effects on virulence, biofilm, and microbiomes: a review of recent advances. Expert Rev Anti Infect Ther. 2020;18:1221–33.
Murugayah SA, Gerth ML. Engineering quorum quenching enzymes: progress and perspectives. Biochem Soc Trans. 2019;47:793–800.
Fetzner S. Quorum quenching enzymes. J Biotechnol. 2015;201:2–14.
Grandclément C, Tannières M, Moréra S, Dessaux Y, Faure D. Quorum quenching: role in nature and applied developments. Fems Microbiol Rev. 2016;40:86–116.
Lade H, Paul D, Kweon JH. Quorum quenching mediated approaches for control of membrane biofouling. Int J Biol Sci. 2014;10:550–65.
Liao J, Shen D, Lin L, Chen H, Jin Y, Chou S, et al. Bacterial quorum sensing quenching activity of Lysobacter leucyl aminopeptidase acts by interacting with autoinducer synthase. Comput Struct Biotechnol J. 2021;19:6179–90.
Liao J, Li Z, Xiong D, Shen D, Wang L, Shao X, et al. A novel and efficient platform for discovering noncanonical quorum-quenching proteins. Microbiol Spectr. 2023;11:e343722.
Dong Y, Xu J, Li X, Zhang L. AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proc Natl Acad Sci USA. 2000;97:3526–31.
Dong Y, Gusti AR, Zhang Q, Xu J, Zhang L. Identification of quorum-quenching N-acyl homoserine lactonases from Bacillus Species. Appl Environ Microbiol. 2002;68:1754–9.
Reina JC, Romero M, Salto R, Cámara M, Llamas I. AhaP, A quorum quenching acylase from Psychrobacter sp. M9-54-1 that attenuates Pseudomonas aeruginosa and Vibrio coralliilyticus virulence. Mar Drugs. 2021;19:16.
Singh BN, Prateeksha, Upreti DK, Singh BR, Defoirdt T, Gupta VK, et al. Bactericidal, quorum quenching and anti-biofilm nanofactories: a new niche for nanotechnologists. Crit Rev Biotechnol. 2017;37:525–40.
Tang K, Su Y, Brackman G, Cui F, Zhang Y, Shi X, et al. MomL, a Novel marine-derived N-acyl homoserine lactonase from Muricauda olearia. Appl Environ Microbiol. 2015;81:774–82.
Lin YH, Xu JL, Hu J, Wang LH, Ong SL, Leadbetter JR, et al. Acyl-homoserine lactone acylase from Ralstonia strain XJ12B represents a novel and potent class of quorum-quenching enzymes. Mol Microbiol. 2003;47:849–60.
Zhang HB, Wang LH, Zhang LH. Genetic control of quorum-sensing signal turnover in Agrobacterium tumefaciens. Proc Natl Acad Sci USA. 2002;99:4638–43.
Christensen P, Cook FD. Lysobacter, a new genus of nonfruiting, gliding bacteria with a high base ratio. Int J Syst Bacteriol. 1978;28:367–93.
Puopolo G, Tomada S, Pertot I. The impact of the omics era on the knowledge and use of Lysobacter species to control phytopathogenic micro-organisms. J Appl Microbiol. 2018;124:15–27.
Lou L, Qian G, Xie Y, Hang J, Chen H, Zaleta-Rivera K, et al. Biosynthesis of HSAF, a tetramic acid-containing macrolactam from Lysobacter enzymogenes. J Am Chem Soc. 2011;133:643–5.
Yu F, Zaleta-Rivera K, Zhu X, Huffman J, Millet JC, Harris SD, et al. Structure and biosynthesis of heat-stable antifungal factor (HSAF), a broad-spectrum antimycotic with a novel mode of action. Antimicrob Agents Chemother. 2006;51:64–72.
Shen X, Wang B, Yang N, Zhang L, Shen D, Wu H, et al. Lysobacter enzymogenes antagonizes soilborne bacteria using the typeIV secretion system. Environ Microbiol. 2021;23:4673–88.
Souza DP, Oka GU, Alvarez-Martinez CE, Bisson-Filho AW, Dunger G, Hobeika L, et al. Bacterial killing via a type IV secretion system. Nat Commun. 2015;6:6453.
Bayer-Santos E, Cenens W, Matsuyama BY, Oka GU, Di Sessa G, Mininel IDV, et al. The opportunistic pathogen Stenotrophomonas maltophilia utilizes a type IV secretion system for interbacterial killing. Plos Pathog. 2019;15:e1007651.
Alegria MC, Souza DP, Andrade MO, Docena C, Khater L, Ramos CHI, et al. Identification of new protein-protein interactions involving the products of the chromosome-and plasmid-encoded Type IV secretion loci of the phytopathogen Xanthomonas axonopodis pv. citri. J Bacteriol. 2005;187:2315–25.
Russell AB, Peterson SB, Mougous JD. Type VI secretion system effectors: poisons with a purpose. Nat Rev Microbiol. 2014;12:137–48.
Dong TG, Ho BT, Yoder-Himes DR, Mekalanos JJ. Identification of T6SS-dependent effector and immunity proteins by Tn-seq in Vibrio cholerae. Proc Natl Acad Sci USA. 2013;110:2623–8.
Gao G, Yin D, Chen S, Xia F, Yang J, Li Q, et al. Effect of biocontrol agent Pseudomonas fluorescens 2P24 on soil fungal community in cucumber rhizosphere using T-RFLP and DGGE. Plos One. 2012;7:e31806.
Qian G, Wang Y, Qian D, Fan J, Hu B, Liu F. Selection of available suicide vectors for gene mutagenesis using chiA (a chitinase encoding gene) as a new reporter and primary functional analysis of chiA in Lysobacter enzymogenes strain OH11. World J Microbiol Biotechnol. 2012;28:549–57.
Zhu J, Chai Y, Zhong Z, Li S, Winans SC. Agrobacterium bioassay strain for ultrasensitive detection of N-acyl homoserine lactone-type quorum-sensing molecules: detection of autoinducers in Mesorhizobium huakuii. Appl Environ Microbiol. 2003;69:6949–53.
McClean KH, Winson MK, Fish L, Taylor A, Chhabra SR, Camara M, et al. Quorum sensing and Chromobacterium violaceum: exploitation of violacein production and inhibition for the detection of N-acyl homoserine lactones. Microbiology. 1997;143:3703–11.
Yan J, Li P, Wang X, Zhu M, Shi H, Yu G, et al. RasI/R Quorum sensing system controls the virulence of Ralstonia solanacearum Strain EP1. Appl Environ Microbiol. 2022;88:e0032522.
Liao L, Schaefer AL, Coutinho BG, Brown PJB, Greenberg EP. An aryl-homoserine lactone quorum-sensing signal produced by a dimorphic prosthecate bacterium. Proc Natl Acad Sci USA. 2018;115:7587–92.
Wei H, Zhang L. Quorum-sensing system influences root colonization and biological control ability in Pseudomonas fluorescens 2P24. Antonie Van Leeuwenhoek. 2006;89:267–80.
Muteeb G, Sen R. Random mutagenesis using a mutator strain. In: Braman J, editor. In vitro mutagenesis protocols. 3rd ed., vol. 634. Totowa, NJ: Humana Press; 2010. p. 411–9.
Coudert E, Gehant S, de Castro E, Pozzato M, Baratin D, Neto T, et al. Annotation of biologically relevant ligands in UniProtKB using ChEBI. Bioinformatics. 2022;39:btac793.
Bateman A, Martin M, Orchard S, Magrane M, Agivetova R, Ahmad S, et al. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 2021;49:D480–D489.
Callaway E. ‘It will change everything’: DeepMind’s AI makes gigantic leap in solving protein structures. Nature. 2020;588:203–4.
Xu G, Han S, Huo C, Chin K, Chou S, Gomelsky M, et al. Signaling specificity in the c-di-GMP-dependent network regulating antibiotic synthesis in Lysobacter. Nucleic Acids Res. 2018;46:9276–88.
Yang M, Ren S, Shen D, Yang N, Wang B, Han S, et al. An intrinsic mechanism for coordinated production of the contact-dependent and contact-independent weapon systems in a soil bacterium. Plos Pathog. 2020;16:e1008967.
Luo Z, Isberg RR. Multiple substrates of the Legionella pneumophila Dot/Icm system identified by interbacterial protein transfer. Proc Natl Acad Sci USA. 2004;101:841–6.
Lee DJ, Jo AR, Jang MC, Nam J, Choi HJ, Choi G, et al. Analysis of two quorum sensing-deficient isolates of Pseudomonas aeruginosa. Microb Pathog. 2018;119:162–9.
Koch AK, Käppeli O, Fiechter A, Reiser J. Hydrocarbon assimilation and biosurfactant production in Pseudomonas aeruginosa mutants. J Bacteriol. 1991;173:4212–9.
Redzej A, Ukleja M, Connery S, Trokter M, Felisberto Rodrigues C, Cryar A, et al. Structure of a VirD4 coupling protein bound to a VirB typeIV secretion machinery. EMBO J. 2017;36:3080–95.
Hersch SJ, Lam L, Dong TG. Engineered Type Six secretion systems deliver active exogenous effectors and Cre recombinase. mBio. 2021;12:e0111521.
Lee J, Zhang L. The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell. 2015;6:26–41.
Brint JM, Ohman DE. Synthesis of multiple exoproducts in Pseudomonas aeruginosa is under the control of RhlR-RhlI, another set of regulators in strain PAO1 with homology to the autoinducer-responsive LuxR-LuxI family. J Bacteriol. 1995;177:7155–63.
Aoun N, Tauleigne L, Lonjon F, Deslandes L, Vailleau F, Roux F, et al. Quantitative disease resistance under elevated temperature: genetic basis of new resistance mechanisms to Ralstonia solanacearum. Front Plant Sci. 2017;8:1387.
Gould TA, Schweizer HP, Churchill MEA. Structure of the Pseudomonas aeruginosa acyl-homoserinelactone synthase LasI. Mol Microbiol. 2004;53:1135–46.
Grohmann E, Christie PJ, Waksman G, Backert S. Type IV secretion in Gram-negative and Gram-positive bacteria. Mol Microbiol. 2018;107:455–71.
Voth DE, Broederdorf LJ, Graham JG. Bacterial Type IV secretion systems: versatile virulence machines. Future Microbiol. 2012;7:241–57.
Hersch SJ, Manera K, Dong TG. Defending against the type six secretion system: beyond immunity genes. Cell Rep. 2020;33:108259.
Han Y, Wang Y, Tombosa S, Wright S, Huffman J, Yuen G, et al. Identification of a small molecule signaling factor that regulates the biosynthesis of the antifungal polycyclic tetramate macrolactam HSAF in Lysobacter enzymogenes. Appl Microbiol Biotechnol. 2015;99:801–11.
Acknowledgements
This study was funded by the Natural Key Research and Development Program (2022YFD1400200 to GQ), followed by the National Natural Science Foundation of China (U22A20486, 32072470, and 31872016 to GQ), Science and technology project of Shanxi Branch of China National Tobacco Corporation (KJ-2022-04), the Fundamental Research Funds for the Central Universities (KJJQ202001, KYT201805, and KYTZ201403 to GQ), and the Jiangsu University advantage discipline construction project (80900246 to XS).
Author information
Authors and Affiliations
Contributions
GQ and JL conceived the project and designed experiments. JL carried out experiments. JL, ZL, DX, DS, LW, L Liao, and XS analyzed data and prepared figures and tables. JL, GQ, and XS wrote the manuscript. GQ, XS, L Lin, PL, LZ, and HW revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liao, J., Li, Z., Xiong, D. et al. Quorum quenching by a type IVA secretion system effector. ISME J 17, 1564–1577 (2023). https://doi.org/10.1038/s41396-023-01457-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01457-2
This article is cited by
-
Bacterial communication mediated by T4SS effectors
Phytopathology Research (2023)