Abstract
Managing the association with microbes is crucial for plants. Evidence is emerging for the plant latent defense response, which is conditionally elicited by certain microbial nonpathogenic factors and thereby guards against potential risks from beneficial or commensal microbes. Latent defense response is an exciting new research area with a number of key issues immediately awaiting exploration. A detailed understanding of latent defense response will underpin the applications of beneficial microbes.
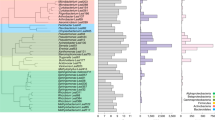
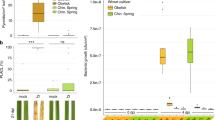
Similar content being viewed by others
Plants naturally live with a wide variety of soil microbes, which can be roughly categorized as pathogenic, commensal, or beneficial based on the microbial impacts on their associated plants, although the impacts caused by the same microbe can be either generalized or plant species-specific while the outcome from a given binary relation is subject to influences by other factors in the complex environment. Unlike commensal microbes that impose neither benefits nor harms to the plants, beneficial microbes promote plant growth or improve plant tolerance to certain stress conditions, whereas pathogens impose threats that in some cases can be deadly to plants [1]. Hence, the detection of and responses to microbial factors are critical for plant survival.
In response to pathogens, the plant immune system is typically activated through microbe-associated molecular pattern (MAMP)-triggered immunity (MTI) and effector-triggered immunity (ETI) [2]. The perception of MAMPs, such as the bacterial flagella epitope flg22, by cell surface-localized pattern recognition receptors (PRRs) initiates MTI without inducing plant cell death. When confronting pathogens that have evolved abilities to subvert MTI through effector proteins released to the apoplast or into the host cells, plants use the evolved nucleotide-binding oligomerization domain-like receptors (NLRs) to detect the effectors directly or indirectly, resulting in the formation of oligomeric sensor complexes that trigger ETI and oftentimes cell death [3, 4]. Plant PRRs also perceives damage-associated molecular patterns (DAMPs), which are plant-derived molecules resulting from cell wall damages and can potentiate MTI [5]. In addition to local defense responses, plants infected by pathogens can produce mobile signals that travel from the infected tissue to distal tissues, resulting in systemic acquired resistance (SAR) that helps protect the plant from future infections [6].
MAMPs are immunogenic signals produced by not only pathogens but also beneficial and commensal microbes [7]. Detection of these extracellular danger signals not only allows for timely defense against pathogens, but also presents obstacles to the establishment of mutualistic association with beneficial microbes. To achieve successful association with plants, microbes have evolved various strategies to avoid or subvert MTI. The evasion of MAMPs from PRR recognition results from alterations in the immunogenic sequences, meanwhile, microbial suppression of MTI can involve extracellular factors that suppress the production of MAMPs, in addition to the intracellular effectors that subvert plant immune signaling [8, 9]. As a result of the suppression of immunity, plant compatibility with pathogens leads to disease, whereas the compatibility with beneficial microbes underlies mutualism. However, in contrast to showing hostility to pathogens, plants show amity with beneficial microbes, because the compatible associations with beneficial microbes are also contributed by plants, as known in rhizobia-legume symbiosis and plant symbiosis with arbuscular mycorrhizal (AM) fungi. Nitrogen-deficient legumes secrete flavonoids that trigger rhizobial synthesis of nodulation factors (Nod factors), which in turn elicit plant signaling that provokes the deformation of root hairs and the formation of nodule primordium [10, 11]; similarly, plants attract AM fungi by root secretion of strigolactones, which trigger extensive hyphal branching and are induced by plant phosphorus deficiency and potentially by nitrogen deficiency in a way that is plant species-specific [12, 13]. The plant’s initiative in establishing mutualism is further supported by plant suppression of MTI, as exemplified by several Medicago truncatula genes that function sequentially in suppressing plant defense against the rhizobial symbionts in nodules [14], as well as by the rice symbiotic receptor OsMYR1 that suppresses OsCERK1-mediated MTI for establishing mutualism with AM fungi [15]. In addition, in the differentiation zone of Arabidopsis thaliana roots, MTI is normally limited due to low expression levels of PRRs but is strongly mounted by local cell damages, thereby providing a way to respond appropriately to pathogens while sparing beneficial and commensal microbes [5].
Although the plant-microbe mutualism is co-established by both partners, evidence is accumulating that the mutualism may be conditionally switched to incompatibility, suggesting that plants somehow are capable of controlling the compatibility with beneficial microbes. Microbes produce and release a complex array of metabolites that can potentially be perceived by neighboring plants [16]. Unlike MAMPs that are perceived by plants as danger signals [2, 7], many microbial extracellular metabolites are nonpathogenic factors (NPFs) that seemingly do not elicit plant defense responses, and as such, NPFs appear negligible regarding compatibility.
Recent studies showed that plants are capable of controlling the compatibility with beneficial bacteria through the perception of certain NPFs. Bacillus amyloliquefaciens strain GB03 is a plant-beneficial bacterium capable of triggering plant growth-promotion (PGP) through soil inoculation or volatile emissions [17]. The PGP induced by GB03 volatile emissions results from integrated plant responses to the different volatile components. While it remains unclear how the volatile signals initiate plant cellular responses, the known downstream mechanisms include modulation of auxin and abscisic acid signaling [18, 19], enhancement of iron and sulfur assimilation [20, 21], enhancement of photosynthetic apparatus production [19], protection of chlorophylls from stress-induced degradation [22], and the requirement of AGP19 that is important for cell division and expansion [23]. The Arabidopsis plants defected in salicylic acid accumulation (nahG) or signaling (eds1) showed similar fresh weights as the wild type plants [24], indicating that decreasing the basal immunity is not sufficient to cause PGP; in contrast, several NPFs in the GB03 volatile emissions strongly elicited plant immunity under certain conditions and compromised the PGP, showing not only the trade-off between growth and immunity, but also the antagonism against compatibility as evidenced by the immunity-dependent impairment of bacterial colonization [24, 25].
In phosphate (Pi)-sufficient A. thaliana, the bacterial volatile component diacetyl emitted from GB03 suppressed plant immunity in supporting the mutualistic association, whereas in Pi-deficient plants, diacetyl strongly induced plant defense that was mediated by salicylic acid [25]. Phosphorus is an essential macronutrient for both plants and bacteria. Under Pi-deficient conditions, activating immunity could be an optimized strategy for plant Pi acquisition because by doing so, plants can deter bacteria from root colonization and thereby reduce direct competition for Pi; meanwhile, the plants can still take advantage of the bacteria within the rhizosphere if the bacteria are capable of Pi solubilization or producing other plant-beneficial traits. Therefore, the differential responses to diacetyl under different Pi availability suggest a plant’s initiative in controlling compatibility with beneficial microbes through the perception of certain NPFs [25,26,27].
Plant regulation of the compatibility with beneficial microbes through the perception of NPFs was also observed in the Arabidopsis mutant rol1 (regulator of LDR 1), which was isolated from a forward genetic screen for mutants with defective plant growth promotion triggered by B. amyloliquefaciens GB03 [24]. In wild-type plants, GB03 enhanced chloroplastic lipid biosynthesis that consumed oleic acid. In the rol1 mutant, biosynthesis of oleic acid was impaired, resulting in severe disruption in the plant lipidome that was exacerbated by GB03. This exacerbation indicates that GB03 increases the vulnerability of rol1 to potential threats since fatty acids and lipids are important and often essential for various cellular functions [28]. Concomitantly, the rol1 mutant plants responded to GB03 with strong activation of defense, which was mediated through the plant perception of the NPFs 2,3-butanediol, acetoin and 2-methyl-1-propanol in the volatile emissions from GB03.
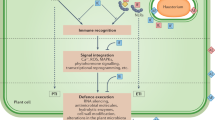
This hidden layer of plant defense highlights the importance of NPFs in mediating plant regulation of mutualism, leading to the concept of the latent defense response (LDR), which is conditionally activated by certain NPFs and antagonizes compatibility [24]. LDR indicates the plant’s initiative in determining the plant-microbe association for optimized benefits, because the conditional activation of LDR avoids unnecessary hostility to beneficial or commensal microbes while enabling plants to deter the same microbes when the microbial association is unfavorable to the plants. The functional importance of LDR is also because it provides a safeguard mechanism for antagonizing mutualism and commensalism, in which MTI is already absent or suppressed in the binary relations. Although the functional importance of LDR is especially highlighted in mutualism and commensalism, it should be noted that LDR would also antagonize the passive compatibility with pathogens that successfully develop diseases. Given that the association between plants and beneficial microbes is as common as plant interactions with pathogens, the plant’s ability of perceiving certain NPFs for controlling mutualism is as important as the ability of perceiving MAMPs for avoiding pathogenicity, i.e., in parallel to MAMP-triggered immunity that promptly defends against pathogens, NPF-triggered LDR protects against potential risks from beneficial or commensal microbes (Fig. 1).
Between plants and the associated beneficial or commensal microbes, the compatibility is co-established by the microbial and plant suppression of MTI. Host-specific recognition of the effectors or defectiveness in the plant MTI suppressors leads to defense against the compatibility. While showing amity to beneficial and commensal microbes, plants deploy LDR as a safeguard mechanism for antagonizing the mutualism and commensalism, in which MTI is absent or suppressed for the compatibility. Activation of LDR is triggered by certain NPFs and occurs only under risky conditions to the plant. The conditional activation of LDR avoids unnecessary hostility to beneficial and commensal microbes while enabling plants to deter the microbial association when it is unfavorable. The new research field of LDR awaits exploration. MAMP microbe-associated molecular pattern, MTI MAMP-triggered immunity, ETI effector-triggered immunity, NPF non-pathogenic factor. The dashed lines pointing to MAMPs and effectors indicate that some compatible microbes may evade immune detection. The dashed lines from NPFs indicate that some NPFs may contribute to compatibility under normal conditions, such as diacetyl that suppresses some immune responses.
The phenomena of plants showing incompatibility with beneficial microbes are accumulating. In particular, legumes and rhizobia form symbiotic relations but incompatibility can occur, often resulting from either failures in MTI suppression or plant ecotype-specific recognition of bacterial effectors that elicit ETI [14]. These types of incompatibility should not be confused with the incompatibility caused by LDR, because the former are caused by danger signals whereas the latter are activated by NPFs only under conditions that are unwelcomed by the plants. Interestingly, mutations in M. truncatula DNF2, a putative phosphatidylinositol phospholipase C-like protein, led to defense reactions and rapid senescence in the nodule when the plants were grown with agar as the medium-solidifying agent; in contrast, the disruption of rhizobial symbiosis in dnf2 mutants was not observed when the agar was replaced by agarose, which is purified agar containing no agaropectins and harboring reduced levels of unidentified impurities [29]. Agar is not known to elicit or prime plant defense, meanwhile, adding flg22 or chitin to the agarose medium did not mimic the effects of the agar [29]. Thus, the conditional activation of defense in dnf2 mutants is reminiscent to LDR, albeit the defense elicitor and the underlying mechanism remain unclear.
LDR is characterized by the conditional activation, in which nutrient availability can be a key determinant. In A. thaliana, the transcriptional regulator PHR1 not only positively regulates phosphate starvation responses (PSR), but also negatively regulates plant defense through promoter occupancy at its target genes [30]. Under Pi deficient conditions, the recruitment of PHR1 to PSR gene loci likely would decrease the promoter occupancy by PHR1 at the defense-related genes, since PHR1 gene expression is only weakly responsive to Pi starvation [31]. In such a scenario, Pi deficiency would create a permissive environment for transcriptional activities at the PHR1-suppressed defense-related genes, thereby allowing for diacetyl-mediated elicitation of LDR. Pi deficiency also impairs the symbiosis between legumes and the nitrogen (N)-fixing rhizobia. As shown in common bean (Phaseolus vulgaris), the reduction of nodule numbers under Pi deficiency was mediated through PvPHR1-dependent activation of the autoregulation of nodulation (AON) pathway, which controls nodule numbers for optimal balance between the costs for and the benefits from the mutualism with rhizobia [32, 33]. As shown in soybean (Glycine max), the nodule autoregulation receptor kinase (GmNARK) potentially links AON with defense [34]. It is unclear whether PHR1 in legume species play dual and opposite roles in regulating PSR and defense as in A. thaliana.
In some cases, the conditional activation of plant defense can also result from nutrient repleteness. For instance, the compatible association between A. thaliana with Colletotrichum tofieldiae, an endophytic fungus that can transfer Pi to the plant, occured only under Pi deficient conditions; whereas under Pi sufficient conditions, plants detered fungal colonization through the accumulation of antifungal metabolites [35]. Similarly, high Pi availability suppressed plant symbiosis with AM fungi, which facilitate Pi acquisition in approximately 80% of land plants; the suppression, as shown in rice, was due to the repression of OsPHR2 that governed the symbiosis gene regulatory network [36, 37]. It is interesting but unclear whether defense mechanisms contribute to the suppression of plant symbiosis with AM fungi under high Pi availability. Similar to Pi suppression of plant-fungi symbiosis, the symbiosis between legumes and rhizobia can be inhibited by N fertilizers especially nitrate [32, 38]. The negative regulatory pathway induced by nitrate converges with the AON pathway and affects each step of nodulation [32]. In a split-root study of M. truncatula symbiosis with Sinorhizobium medicae, mature symbiotic nodules responded to whole-plant N-satiety signaling with nodule senescence and defense activation [39]. Nitrate stimulated M. truncatula nodule production of nitric oxide (NO) that provoked early nodule senescence [40]. NO is an important mediator of plant defense, thus it may play a role in connecting immunity and senescence in the nodule, where the relationship between the two processes remained obscure. During symbiotic interactions, rhizobial Nod factor repressed and induced the production of reactive oxygen species at early and later stages, respectively, as exemplified in M. truncatula roots treated with Sinorhizobium meliloti Nod factor [41, 42]. Both induction and repression of defense-related gene expression were observed in Nod factor-treated M. truncatula root epidermis [43]. The ability of inducing host defense responses makes Nod factors somewhat reminiscent to NPFs that are capable of inducing LDR, since Nod factors induce nodule formation and are therefore commonly not considered as danger signals. It is intriguing whether nitrate-replete legume roots mount persistent defense in response to Nod factors. The nutrient repleteness-induced defense showed target specificity, in that the targeted microbes carried functions tightly related to the nutrients and were no longer needed for the plants, thereby pointing to the involvement of the yet-to-be identified microbial factors in the conditional activation of defense.
LDR is an exciting new research area with a number of key issues immediately awaiting exploration (Fig. 1). For each LDR event, the sensing of NPF signals and the signal transition that switches the compatibility to defense are the two core processes to be disclosed. In addition, it is an interesting question for each LDR event whether the function of the LDR is accurate or indiscriminate in repelling the plant-associated microbes that impose the risk to different degrees. Mutualistic relations between plants and microbes are commonly investigated as pairwise interactions, however, plants in natural environments are inevitably interacting with microbial communities consisting of different species in the same or different niches. Once LDR is activated by a particular microbial partner, how will the plant reshape the microbiome and rebalance the complex interactive network of all the plant-microbe interactions for optimal benefits? Particularly, it is intriguing whether LDR is less prone to be elicited in species-specific mutualism than generalist mutualism, since the former is an obligate relation whereas the latter may be reconstituted by other microbial partners.
Some pathogens may also produce NPFs capable of activating LDR, which can then reinforce plant disease resistance in addition to the contribution by the forefront MAMP-triggered immunity. Because NPFs do not behave like MAMPs in directly activating plant immunity, it is unlikely that NPFs are perceived through the same mechanisms as MAMPs. However, the plant immunity network functions as a concerted system such that different segments are inherently connected and coordinated, as demonstrated by the mutual potentiation between the MTI and ETI pathways [44, 45]. It is an important question as how LDR is connected with and coordinate with the other segments of the immunity network. A clear answer to this question would be built on knowledge about how plants perceive NPF signals, in addition to genetic dissections of the defense outputs from the plant-microbe interactions. Similarly, how different LDR pathways may interact with each other is also to be explored. For instance, the diacetyl-triggered LDR is mediated through a mechanism different from the LDR in rol1, because the latter is not induced by diacetyl [24, 25]. Moreover, the activation of LDR by its eliciting NPFs may be interfered indirectly by the plant developmental regulation or environmental responses, as well as by other factors from the NPF-producing microbes or the other microbes present in the same plant-associated community.
The diversity of LDR mechanisms and the conservativeness of LDR phenomena are two other key issues awaiting to be addressed. On one hand, there are numerous species and combinations of beneficial microbes and their host organisms in nature, so species-based diversity in LDR mechanisms can be expected. For plants, Pi deficiency and oleic acid deficiency are not the only two threats that can be affected by microbes; thus, the diversity of LDR mechanisms likely would also come from different high-risk conditions that allow for LDR activation. In addition, while beneficial microbes may help plants combat environmental stressors, the compatibility may be abandoned by plants when it becomes unnecessary. On the other hand, environmental stressors, such as Pi deficiency, are common threats to different plant species and thereby may commonly influence plant relations with commensal or beneficial microbes. This commonality may confer conservativeness of LDR mechanisms across different plant species. The conservativeness of certain LDR mechanisms is likely also contributed by the fact that the same NPF can be produced by different microbes. A detailed understanding of these and other questions in the field of LDR will underpin the applications of beneficial microbes.
References
Delaux P, Schornack S. Plant evolution driven by interactions with symbiotic and pathogenic microbes. Science. 2021;371:eaba66.
Jones JD, Dangl JL. The plant immune system. Nature. 2006;444:323–9.
Thomma BPHJ, Nürnberger T, Joosten MHAJ. Of PAMPs and effectors: the blurred PTI-ETI dichotomy. Plant Cell. 2011;23:4–15.
Bi G, Su M, Li N, Liang Y, Dang S, Xu J, et al. The ZAR1 resistosome is a calcium-permeable channel triggering plant immune signaling. Cell. 2021;184:3528–41.
Zhou F, Emonet A, Tendon VD, Marhavy P, Wu D, Lahaye T, et al. Co-incidence of damage and microbial patterns controls localized immune responses in roots. Cell. 2020;180:440–53.
Klessig DF, Choi HW, Dempsey DA. Systemic acquired resistance and salicylic acid: past, present, and future. Mol Plant Microbe Interact. 2018;31:871–88.
Ausubel FM. Are innate immune signaling pathways in plants and animals conserved? Nat Immunol. 2005;6:973–9.
Zipfel C, Oldroyd GE. Plant signaling in symbiosis and immunity. Nature. 2017;543:328–36.
Wang Y, Pruitt RN, Nürnberger T, Wang Y. Evasion of plant immunity by microbial pathogens. Nat Rev Microbiol. 2022;20:449–64.
Relić B, Perret X, Estrada-García MT, Kopcinska J, Golinowski W, Krishnan HB, et al. Nod factors of rhizobium are a key to the legume door. Mol Microbiol. 1994;13:171–9.
Coronado C, Zuanazzi JAS, Sallaud C, Quirion JC, Esnault R, Husson HP, et al. Alfalfa root flavonoid production is nitrogen regulated. Plant Physiol. 1995;108:533–42.
Akiyama K, Matsuzaki K, Hayashi H. Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature. 2005;435:824–7.
Yoneyama K, Xie X, Kim HI, Kisugi T, Nomura T, Sekimoto H, et al. How do nitrogen and phosphorus deficiencies affect strigolactone production and exudation? Planta. 2012;235:1197–207.
Benezech C, Doudement M, Gourion B. Legumes tolerance to rhizobia is not always observed and not always deserved. Cell Microbiol. 2020;22:e13124.
Zhang C, He J, Dai H, Wang G, Zhang X, Wang C, et al. Discriminating symbiosis and immunity signals by receptor competition in rice. Proc Natl Acad Sci USA. 2021;118:e2023738118.
Gámez-Arcas S, Baroja-Fernández E, García-Gómez P, José Muñoz F, Almagro G, Bahaji A, et al. Action mechanisms of small microbial volatile compounds in plants. J Exp Bot. 2022;73:498–510.
Paré PW, Zhang H, Aziz M, Xie X, Kim M, Shen X, et al. Beneficial rhizobacteria induce plant growth: mapping signaling networks in Arabidopsis. In: Witzany G, editor. Biocommunication in soil microorganisms, soil biology, 23. Springer-Verlag Berlin Heidelberg; 2011.
Zhang H, Kim M, Krishnamachari V, Payton P, Sun Y, Grimson M, et al. Rhizobacterial volatile emissions regulate auxin homeostasis and cell expansion in Arabidopsis. Planta. 2007;226:839–51.
Zhang H, Xie X, Kim M, Kornyeyev DA, Holaday S, Paré PW. Soil bacteria augment Arabidopsis photosynthesis by decreasing glucose sensing and abscisic acid levels in planta. Plant J. 2008;56:264–73.
Zhang H, Sun Y, Xie X, Kim M, Dowd SE, Paré PW. A soil bacteria regulates plant acquisition of iron via deficiency-inducible mechanisms. Plant J. 2009;58:568–77.
Aziz M, Nadipalli RK, Xie X, Sun Y, Surowiec K, Zhang J, et al. Augmenting sulfur metabolism and herbivore defense in Arabidopsis by bacterial volatile signaling. Front Plant Sci. 2016;7:458.
Singh SK, Sun Y, Yang Y, Zuo Z, Wu X, Shao C, et al. Bacterial diacetyl suppresses abiotic stress-induced senescence in Arabidopsis. J Integr Plant Biol. 2022;64:1135–9.
Lv S, Yang Y, Yu G, Peng L, Zheng S, Singh SK, et al. Dysfunction of histone demethylase IBM1 in Arabidopsis causes autoimmunity and reshapes the root microbiome. ISME J. 2022;16:2513–24.
Yang Y, Chen S, Wu X, Peng L, Vílchez JI, Kaushal R, et al. Plant latent defense response to microbial non-pathogenic factors antagonizes compatibility. Natl Sci Rev 2022;9:nwac109.
Morcillo RJL, Singh SK, He D, An G, Vílchez JI, Tang K, et al. Rhizobacterium-derived diacetyl modulates plant immunity in a phosphate-dependent manner. EMBO J. 2020;39:e102602.
Zuccaro A. Plant phosphate status drives host microbial preferences: a trade-off between fungi and bacteria. EMBO J. 2020;39:e104144.
Singh SK, Wu X, Shao C, Zhang H. Microbial enhancement of plant nutrient acquisition. Stress Biol. 2022;2:3.
Li-Beisson Y, Shorrosh B, Beisson F, Andersson MX, Arondel V, Bates PD, et al. Acyl-lipid metabolism. Arabidopsis Book. 2013;11:e0161.
Berrabah F, Bourcy M, Cayrel A, Eschstruth A, Mondy S, Ratet P, et al. Growth conditions determine the DNF2 requirement for symbiosis. PLoS ONE. 2014;9:e91866.
Castrillo G, Teixeira PJPL, Paredes SH, Law TF, Lorenzo L, Feltcher ME, et al. Root microbiota drive direct integration of phosphate stress and immunity. Nature. 2017;543:513–8.
Rubio V, Linhares F, Solano R, Martín AC, Iglesias J, Leyva A, et al. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 2001;15:2122–33.
Nishida H, Suzaki T. Two negative regulatory systems of root nodule symbiosis: how are symbiotic benefits and costs balanced? Plant Cell Physiol. 2018;59:1733–8.
Isidra-Arellano MC, Pozas-Rodríguez EA, Reyero-Saavedra MDR, Arroyo-Canales J, Ferrer-Orgaz S, Sánchez-Correa MDS, et al. Inhibition of legume nodulation by Pi deficiency is dependent on the autoregulation of nodulation (AON) pathway. Plant J. 2020;103:1125–39.
Kinkema M, Gresshoff PM. Investigation of downstream signals of the soybean autoregulation of nodulation receptor kinase GmNARK. Mol Plant Microbe Interact. 2008;21:1337–48.
Hiruma K, Gerlach N, Sacristán S, Nakano RT, Hacquard S, Kracher B, et al. Root endophyte Colletotrichum tofieldiae confers plant fitness benefits that are phosphate status dependent. Cell. 2016;165:464–74.
Das D, Paries M, Hobecker K, Gigl M, Dawid C, Lam H, et al. PHOSPHATE STARVATION RESPONSE transcription factors enable arbuscular mycorrhizal symbiosis. Nat Commun. 2022;13:477.
Shi J, Zhao B, Zheng S, Zhang X, Wang X, Dong W, et al. A phosphate starvation response-centered network regulates mycorrhizal symbiosis. Cell. 2021;184:5527–40. e18
Moreau C, Gautrat P, Frugier F. Nitrate-induced CLE35 signaling peptides inhibit nodulation through the SUNN receptor and miR2111 repression. Plant Physiol. 2021;185:1216–28.
Lambert I, Pervent M, Queré AL, Clément G, Tauzin M, Severac D, et al. Responses of mature symbiotic nodules to the whole-plant systemic nitrogen signaling. J Exp Bot. 2020;71:5039–52.
Cam Y, Pierre O, Boncompagni E, Hérouart D, Meilhoc E, Bruand C. Nitric oxide (NO): a key player in the senescence of Medicago truncatula root nodules. N Phytol. 2012;196:548–60.
Ramu SK, Peng H, Cook DR. Nod factor induction of reactive oxygen species production is correlated with expression of the early nodulin gene rip1 in Medicago truncatula. Mol Plant Microbe Interact. 2002;15:522–8.
Shaw SL, Long SR. Nod factor inhibition of reactive oxygen efflux in a host legume. Plant Physiol. 2003;132:2196–204.
Jardinaud MF, Boivin S, Rodde N, Catrice O, Kisiala A, Lepage A, et al. A laser dissection-RNAseq analysis highlights the activation of cytokinin pathways by Nod factors in the Medicago truncatula root epidermis. Plant Physiol. 2016;171:2256–76.
Ngou BPM, Ahn H, Ding P, Jones JDG. Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature. 2021;592:110–5.
Yuan M, Jiang Z, Bi G, Nomura K, Liu M, Wang Y, et al. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature. 2021;592:105–9.
Acknowledgements
The author apologizes to those colleagues whose work is not cited owing to space constraints. The work of HZ laboratory is supported by the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
All contributions were from HZ.
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, H. Plant latent defense response against compatibility. ISME J 17, 787–791 (2023). https://doi.org/10.1038/s41396-023-01399-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-023-01399-9