Abstract
Microbial communities in oxygen minimum zones (OMZs) are known to have significant impacts on global biogeochemical cycles, but viral influence on microbial processes in these regions are much less studied. Here we provide baseline ecological patterns using microscopy and viral metagenomics from the Eastern Tropical North Pacific (ETNP) OMZ region that enhance our understanding of viruses in these climate-critical systems. While extracellular viral abundance decreased below the oxycline, viral diversity and lytic infection frequency remained high within the OMZ, demonstrating that viral influences on microbial communities were still substantial without the detectable presence of oxygen. Viral community composition was strongly related to oxygen concentration, with viral populations in low-oxygen portions of the water column being distinct from their surface layer counterparts. However, this divergence was not accompanied by the expected differences in viral-encoded auxiliary metabolic genes (AMGs) relating to nitrogen and sulfur metabolisms that are known to be performed by microbial communities in these low-oxygen and anoxic regions. Instead, several abundant AMGs were identified in the oxycline and OMZ that may modulate host responses to low-oxygen stress. We hypothesize that this is due to selection for viral-encoded genes that influence host survivability rather than modulating host metabolic reactions within the ETNP OMZ. Together, this study shows that viruses are not only diverse throughout the water column in the ETNP, including the OMZ, but their infection of microorganisms has the potential to alter host physiological state within these biogeochemically important regions of the ocean.
Similar content being viewed by others
Introduction
Marine viruses are now recognized to play key roles in marine ecosystems by lysing ~10–40% of bacteria per day, recycling organic matter and nutrients via the ‘viral shunt’, and contributing to microbial niche differentiation through horizontal gene transfer [1,2,3,4]. In addition, some viruses encode “host genes” or auxiliary metabolic genes (AMGs) that metabolically reprogram their hosts during infection, including the well-documented cases of cyanobacterial viruses that express photosynthesis genes during infection and contribute significantly to marine microbial photosynthesis [5,6,7]. The extent of viral-encoded AMGs is just now being revealed, with detected AMGs so far also involved in C, N, P, and S metabolism [4, 8,9,10,11,12].
While viral ecology and the potential influences of viral-encoded AMGs has been examined in global-scale studies (e.g.,[10, 13]), there have been few investigations of viruses in marine oxygen minimum zones [11, 12, 14,15,16]. Marine oxygen minimum zones (OMZs) may be considered as ‘extreme environments’, but they constitute ~7% of oceanic volume and have increased substantially due to global climate change over the past fifty years, especially in the North and Equatorial Pacific where the volume of waters considered functionally anoxic (dissolved O2 below the detection limit [17]) have quadrupled [18, 19]. The expansion of OMZs is predicted to have positive feedbacks on climatologically active trace gases including CH4, N2O and DMS [20, 21] as a result of the chemotrophy performed by the unique microbial assemblages present in these regions [20]. Microbes in these areas deplete bioavailable nitrogen through anaerobic ammonium oxidation (anammox) and denitrification, accounting for 30–50% of oceanic nitrogen removal [21, 22], and also play roles in dissimilatory sulfur oxidation and sulfate reduction [23]. Recent research in OMZs has largely focused on the cycling of these major nutrients, as well as the taxonomically and functionally unique microbes responsible for key metabolic processes [20, 21].
Previous research has shown that oxygen is a driver of viral community structure in the Northeast Subarctic Pacific Ocean OMZ [24] and significantly affects viral infections of the ecologically important SUP05 bacteria in the Saanich Inlet OMZ [8]. Prior work to explicitly examine OMZ viral community structure using non-quantitative methods suggests that viral diversity is low in the Eastern Tropical South Pacific (ETSP) OMZ, but that OMZ viruses contain diverse metabolic genes that can affect biogeochemical cycling [14]. However, more recent work in the ETSP and Cariaco Basin has suggested that viral diversity remains high within the OMZ [15, 16]. Thus, there is critical need for quantitative datasets across diverse OMZs to provide the foundational understanding required to incorporate viruses into OMZ ecosystem models.
Here we investigated viral community structure and potential ecological impacts at two stations in the Eastern Tropical North Pacific (ETNP; Supplementary Fig. S1), which, located south of Baja California, encompasses 41% of global OMZ area and is the largest permanent OMZ [19]. To this end, we combined quantitative microscopic and metagenomic methods to evaluate the influence of environmental parameters, including oxygen concentrations, on (i) viral and bacterial abundances, (ii) lytic viral infection frequency in bacteria, (iii) viral community structure based on morphology and metagenomically-derived viral population abundances, and (iv) the distribution of viral-encoded AMGs. The results from this study indicate that while viral diversity and infection frequency remain high within the ETNP OMZ, the structure of the viral community and the composition of viral-encoded AMGs are substantially altered in the oxycline and the functionally anoxic core of the OMZ.
Results and discussion
Environmental conditions, viral and bacterial abundance, and frequency of infection
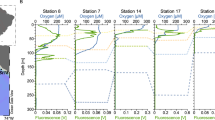
Physiochemical parameters were measured using a Conductivity Temperature Depth profiler equipped with a fluorometer and dissolved oxygen sensor at two stations of the ETNP, one nearshore and one offshore. Examination of environmental conditions revealed similarities and key differences between the nearshore and offshore stations in the ETNP (Supplementary Fig. S1). While both stations exhibited a strong OMZ with oxygen concentrations below detection for ca. 700 m within the water column, the nearshore station revealed a shoaling of the oxycline relative to the offshore station (Fig. 1A, B). Temperature and oxygen profiles at both stations indicated the presence of a diurnal mixed layer modulated by air temperature and wind-driven mixing (Supplementary Fig. S2A, B). We thus use the term “mixed layer” as a depth category indicating the upper, oxygenated portion of the water column, distinct from the oxycline, OMZ, and below the OMZ. Phosphate and nitrate levels increased with depth beginning in the lower portion of the oxycline at both stations (Supplementary Fig. S2C–F) and there were noticeably higher peaks of nitrite and ammonium in the oxycline at the offshore station (Supplementary Fig. S2G–J).
Oxygen concentration and chlorophyll fluorescence (A, B), viral (C, D) and bacterial concentrations (E, F), VMR (G, H), and FIC (I, J) at each station are displayed. Error bars represent standard deviations of the means of triplicate samples. Shaded areas in I–J represent positive and negative 95% confidence intervals for single samples.
Depth profiles of chlorophyll fluorescence also differed markedly between the stations, with a single peak at the offshore station, but two maxima at the nearshore station (Fig. 1A, B). This secondary chlorophyll maximum (SCM) within the OMZ of the nearshore station has been previously documented in both the ETNP [17] and Eastern Tropical South Pacific (ETSP) OMZs [25]. This oxygen-depleted portion of the water column maintains an active photosynthetic community, including Synechococcus and Prochlorococcus [26] that drive a cryptic oxygen cycle in which oxygen is consumed as fast as it is produced [25, 27].
Microbial abundances for each sample were quantified using epifluorescence microscopy. Viral and bacterial concentrations had local maxima in the upper, oxygenated portion of the water column, with positive correlations between them and with oxygen concentration (Fig. 1C–F, Supplementary Fig. S3). While some studies have reported secondary maxima of viral concentrations in marine and freshwater OMZs [14, 28, 29], this was not evident in the ETNP. There were notable differences between stations, including (i) approximately twice the concentration of viruses in the mixed layer at the nearshore station, resulting in a much higher virus-to-microbe ratio (VMR), and (ii) the presence of a secondary maximum in bacterial concentration at the nearshore SCM that was accompanied by only a minor increase in viral concentration (Fig. 1C–H).
In contrast to the VMR, the frequency of infected cells (FIC) as estimated using transmission electron microscopy was similar between stations, reaching local maxima in the mixed layer as well as the OMZ (Fig. 1I, J). VBR was thus not correlated with FIC (Supplementary Fig. S3), reflecting the complex balance between production of viruses and bacteria with removal of viruses through decay [30, 31] and mortality of bacteria resulting from numerous sources [32]. The increased FIC in the OMZ indicated that viruses were still lytically replicating in the OMZ as has been observed in multiple prior studies [28, 29, 33,34,35], resulting in a lack of correlation with oxygen concentration (Supplementary Fig. S3).
Morphological diversity of viral communities
Viral morphology, including morphotype and capsid diameters, was analyzed using quantitative transmission electron microscopy [36]. We found non-tailed viruses to be the most abundant morphotype at both stations and all depths, ranging from 56–93% of each sample (Supplementary Fig. S4), consistent with a global survey in the upper water column [36]. The percent of non-tailed viruses was positively correlated with depth, and negatively with temperature and oxygen (Supplementary Fig. S3). This suggests either (i) an increase with depth of bona fide non-tailed dsDNA viruses such as the Autolykiviridae, which infect among others many widespread Vibrio species [37], (ii) an increased abundance of non-tailed ssDNA viruses [38], or (iii) a loss of tails as an initial step of natural tailed virus decay as has previously been suggested [36].
Viral capsid widths were also similar to those reported for the global upper oceans [36]. While viral capsid width did vary significantly among all samples (global ANOVA p < 0.001), there was not an evident trend in overall capsid widths with depth or between stations (Supplementary Fig. S5). However, capsid width distributions were significantly related to depth, temperature, oxygen, chlorophyll fluorescence, nitrite, and nitrate (p < 0.05, Supplementary Fig. S6A–I). This indicates that viral community structure in the ETNP, based on morphology, is driven directly or indirectly by environmental variables as has previously been shown [13, 36].
Population-level diversity of viral communities
Viral contigs were identified using VirSorter [39] and clustered into 10,601 populations at 95% ANI. Unsurprisingly, less than 1% of observed viral populations could be assigned a taxonomic identification (Supplementary Fig. S7) using RefSeq (version 74), as is common in marine viromes [40]. None of the fifty most abundant viral populations (12.2% of total abundance), which are mostly present in the low-oxygen samples at both stations, could be assigned a specific taxonomic identification. Of the less-abundant populations that could be identified, most were closely related to Synechococcus phages or other cyanophages, which have previously been identified in the mixed layer, oxycline, and even below the oxycline in the ETNP [26, 41]. Diversity (Shannon H’) and evenness (Pielou’s J’) of viral community structure based on relative population abundances in this study were similar to previous results from a global ocean survey [13], with a mean diversity and evenness of 7.12 and 0.899, respectively. Both viral diversity and evenness were consistent among all samples (Fig. 2A, B). This is in contrast to a previous study in the ETSP OMZ, which reported much lower viral alpha diversity and evenness, especially within the OMZ core [14]. However, that study used multiple displacement amplification to amplify nucleic acids for the metagenomes, which was later shown to have substantial biases (reviewed by [42, 43]). A recent study reported that diversity of free-living and particle-associated bacterial communities in the ETNP peaks near the SCM [44]. However, the slight increase in oxycline viral diversity we observed was not significant (Fig. 2A), indicating that extracellular viral community diversity patterns do not mirror those of the bacterial community.
Shannon index H′ (A) and Pielou’s evenness J’ (B). The Euler diagram (C) shows the number of unique and shared viral populations in each group of samples for both stations combined (stress = 0.0011%). Correspondence analyses based on relative abundances of viral populations compare all samples (D) and the subset of samples from low-oxygen environments (E). For panels D and E, the percentage of inertia explained by CA1 and CA2 are reported on the axes.
For both stations combined, viral communities in the surface mixed layer were quite distinct from those in low-oxygen samples, with few shared populations (Fig. 2C, D). Most viral populations were unique to either the mixed layer or the oxycline (n = 5914), with populations in the OMZ core predominantly representing a subset of the oxycline community (Fig. 2C). Hierarchical clustering of the viral populations revealed two clusters comprised of the mixed layer communities and the low-oxygen communities, respectively, and their subtle differences in relative abundance of viral populations was visualized using a heatmap (Supplementary Fig. S8). Mixed layer communities showed clear differences between stations, as well as compared to low-oxygen communities (Fig. 2D), and all communities were significantly related to temperature, salinity, oxygen, and chlorophyll fluorescence (p < 0.05 for all; Supplementary Fig. S9A–I). After removal of the mixed layer communities, the low-oxygen subset of communities clustered by depth category rather than station (Fig. 2E) and were significantly influenced by depth, temperature, salinity, oxygen, chlorophyll fluorescence, nitrite and nitrate concentrations (p < 0.05 for all, Fig. S9J–R). These findings are similar to a previous investigation of viral communities in the subarctic North Pacific OMZ that showed significant differences between photic and aphotic samples, though distance from shore was less influential [24]. In the ETNP, we suspect that coastal upwelling accounted for the differences between mixed layer samples [45], while the strong influences of oxygen and depth overwhelm factors that differ between our stations in low-oxygen samples.
General AMG trends
Viral-encoded AMGs can metabolically reprogram their hosts during infection and significantly impact biogeochemical cycling, even accelerating host niche differentiation [10, 46,47,48]. While most of the genes that could be assigned functions within the ETNP viral metagenomes were related to viral replication, including DNA replication and repair and viral structural proteins, there were 247 genes annotatable as Class I AMGs, and 81 as Class II AMGs (3% of total genes; Supplementary Fig. S10). As with prior studies (reviewed by Roux et al. [10, 41]), we found that AMGs in the ETNP viral community were related to environmental conditions. The majority of AMGs were present at both the offshore and nearshore stations at similar abundances (Fig. 3A). Of the few AMGs that were unique to one station, the most abundant were related to membrane transport and phospholipid metabolism at the offshore station (Fig. 3B). This distribution of AMGs was consistent with the population-level structure of the viral community in which the samples below the mixed layer were highly similar between stations (Fig. 2D, E).
Those found at both stations (A), unique to one station (B), found in both the mixed layer and low-oxygen environments (C), or unique to either the mixed layer or low oxygen environments (D) are displayed. Euler diagram insets in panels A and C represent the number of unique and shared PFAMs by station (within panel A) and depth category (within panel C).
While 52% of the AMGs were present in both mixed layer and low-oxygen samples, the functional categories of these shared AMGs were present at different abundances within depth categories (Fig. 3C). Eleven shared gene categories were more abundant in low-oxygen samples, including amino acid metabolism, carbohydrate metabolism, cell wall and capsule formation, and sporulation genes (Fig. 3C). Of the AMGs unique to the mixed layer, most were related to photosynthesis, respiration, and amino acid metabolism, while those unique to low-oxygen samples were primarily related to amino acid metabolism and membrane transport (Fig. 3D). These larger differences in AMG composition when comparing mixed layer versus low-oxygen samples again support the population-level comparisons of the viral communities (Fig. 2D), in which oxygen concentration is a major driver of viral community composition (Supplementary Fig. S9E).
To further investigate the distribution of AMGs, we examined depth profiles of AMG functional categories at each station (Fig. 4A–G and Supplementary Fig. S11A–L). All AMGs combined exhibited relatively low abundance in the mixed layer at both stations, increased dramatically in the oxycline, and then decreased with depth (Fig. 4A). The number of unique and shared AMGs among depth categories (Fig. 4A, inset) also reflected overall community structure (Fig. 2C), with the majority of AMGs shared across all depth categories, and the most unique AMGs found within the mixed layer and oxycline. This indicates that viruses do not utilize many unique AMGs among the depth categories, but instead that commonly occurring ones are selectively enriched in specific sections of the water column as described below.
Viral influences on photosynthesis, nitrogen, and sulfur metabolism
AMGs related to photosynthesis were unsurprisingly most abundant in the mixed layer (Fig. 4B), where they function to enhance host photosynthetic activity (reviewed by Aldunate et al. [41]). We also observed extremely low abundances of photosynthesis-related AMGs below the OMZ as has been reported in the subarctic North Pacific OMZ, likely due to phage released from sinking organic particles at depth [11]. The most abundant photosynthesis AMGs in the ETNP, psbA/D and psbN, encode for photosynthetic reaction center proteins and have been observed in oceanic datasets previously [11, 14, 49, 50] as they are widely distributed within marine cyanophage isolates. However, while the abundance of these photosynthesis genes was significantly positively correlated with oxygen and chlorophyll concentrations (Supplementary Fig. S12), they were much less abundant at the nearshore station than expected given the high chlorophyll concentration present in the mixed layer and SCM (Fig. 1A, B). Their low abundance at the nearshore station SCM may be due to (i) a high prevalence of uncharacterized viruses that infect the unique cyanobacterial ecotypes in the SCM as found in the ETSP [25, 26], which may not contain these core photosynthesis genes, (ii) a selection for viruses that do not contain these genes, as they may only impart a selective advantage under high light conditions [51, 52], or (iii) a counter selection of these genes because the viruses have short latent periods during which the viral genes could not be sufficiently expressed to boost host photosynthetic potential, as has been suggested with some cyanobacterial isolates previously [50, 52].
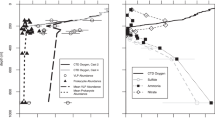
Recent work regarding viruses in the ETSP and Cariaco basin has suggested viral AMGs may influence nitrogen and sulfur cycling within the low-oxygen layer [12, 16]. Although we detected some AMGs related to these processes within the ETNP, they were present at low abundances. Depth profiles of nitrogen metabolism AMGs revealed a peak in the mixed layer and oxycline at the nearshore and offshore stations, respectively (Supplementary Fig. S11H), then decreasing with depth. Sulfur metabolism AMGs were most abundant in the mixed layer at both stations and decreased with depth, though the nearshore station remained slightly higher abundance until below the OMZ (Supplementary Fig. S11K). We detected no genes associated with the processes that dominate nitrogen cycling in the ETNP, including annamox, nitrite oxidation, denitrification, and ammonia oxidation [44, 53,54,55]. We detected draT, which is part of a system that inhibits nitrogenase under high-ammonium conditions or energy depletion [56], though this gene was present at very low levels only in the OMZ core of the offshore station. Additionally, nifU was present in all samples, though as it is often found in organisms not capable of fixing nitrogen [57, 58], we categorized it as an AMG related to iron–sulfur cluster assembly. The most abundant sulfur metabolism gene we detected was tauD, which catalyzes the oxygenolytic release of sulfite from taurine [59] and was present in the mixed layer and oxycline of both stations. SoxY, which encodes part of a thiosulfate oxidizing enzyme complex [60], was also observed in this study at very low abundance in the oxycline and OMZ core of both stations. Sulfur metabolism AMGs previously observed in marine viromes, namely dsrC, soxB, and rdsrA, were not observed here [8, 61,62,63].
Although the ETNP is known to contain a near-complete nitrogen cycle [21, 54, 55, 64] and our chemical profiles indicated the presence of these processes (Supplementary Fig. S2E–J), as well as sulfur oxidation and sulfate reduction [63], we detected very few AMGs related to these processes. This seemingly introduces a paradox in which viral-encoded AMGs do not include genes related to the dominant nitrogen and sulfur cycling processes in the ETNP. However, these results are parsimonious with the hypothesis that viruses will only contain AMGs required to enhance rate-limiting steps in reactions beyond that of which the host requires in order to increase viral replication [10, 47]. Thus, we suggest that AMGs related to nitrogen and sulfur metabolism in the ETNP are so low in abundance because host metabolic processing of these nutrients is sufficient to sustain requirements for viral replication. Our results indicate that viruses influence host physiological state in the ETNP more than manipulate metabolic reactions.
Viral replication strategy and influences on host physiological state
We found that phage integrases increased in relative abundance with depth at the offshore station, while at the nearshore station they peaked in the oxycline and decreased below it (Fig. 4C). This suggests that a higher portion of the extracellular viruses at the offshore station seem to have the ability to integrate into their hosts as lysogens, which has been suggested to increase with depth and environmental stress [29, 65]. The increased potential for lysogeny with elevated stress is consistent with the patterns regarding viral-encoded genes related to host stress described below.
Iron–sulfur cluster synthesis, sporulation, toxin-antitoxin (TA), and antimicrobial resistance (AMR) genes were detected throughout the ETNP. Depth profiles revealed that all three AMG categories were most abundant in the oxycline at both stations and decreased in abundance with depth (Fig. 4D–F), resulting in no correlations with environmental variables (Supplementary Fig. S12). Iron–sulfur clusters are extremely important prosthetic groups required by many microbial enzymes central to metabolism [58, 66,67,68,69], including electron transfer and modulation of gene expression under oxidative and redox stress conditions [67, 70]. The most abundant of these AMGs we detected were involved in cluster assembly or biosynthesis, several of which have been observed in previous viromes [11, 71]. One such assembly gene, sufE, has been shown to enhance the activity of sufS to assemble housekeeping iron–sulfur clusters under oxidative stress [72,73,74,75,76], but cannot function alone [74], so its presence without sufS in the ETNP suggests that host production of sufE is the limiting step in this iron–sulfur cluster formation. The other most common iron–sulfur cluster biosynthesis pathways are the isc [58, 66, 77] and nif [57, 58] operons: of these, we detected nifU as mentioned above. Additionally, we detected several hits to iron–sulfur cluster binding domains commonly found in ferredoxins that mediate electron transfer in various metabolic reactions [78]. The presence of these iron–sulfur cluster genes suggests that viruses may be boosting host response to oxidative and redox stress, especially in the oxycline.
We detected several AMGs related to sporulation, a bacterial survival strategy where the cell produces a hardy endospore form that can persist for extended periods of time [79]. These genes were most abundant in the oxycline and decreased with depth below it, with the offshore remaining more abundant than the nearshore in the OMZ core (Fig. 4E). The most abundant of these, spoVR, along with spoVS, spoIIE, and spoVG, are related to the formation of bacterial spores [80]; of these, spoVS has been seen in soil viromes previously [9]. Phages have been shown to persist in bacterial spores, and some lysogenic phages of Clostridium botulinum and Bacillus species have been shown to encode key virulence genes as well as sporulation genes to modulate host metabolism [81,82,83,84]. Though many spore-producing microbes are obligate or facultative anaerobes, the unique stressors of the unstable oxycline environment may limit these hosts. Thus, ETNP viruses may then help induce sporulation in their hosts to persist similarly to previously characterized phages of Clostridium and Bacillus species [83, 85].
We found that the distribution of AMGs related to TA systems was similar to that of sporulation-related genes, though there was little difference between stations (Fig. 4F). The main driver of this pattern was a gene that encodes zeta toxin, a well-characterized plasmid-borne postsegregational killing system [86, 87]. Some temperate phages that persist as extrachromosomal prophages or integrate into host genomes use genes for plasmid inheritance and persistence, including toxin-antitoxin genes [65], though we did not identify any integrase genes on populations containing TA AMGs. A gene for the SymE toxin, part of a type I TA system that is often found in bacterial chromosomes [88], was present at much lower abundance. In the oxycline, host cells under extreme environmental stress may be more likely to expel plasmids or extrachromosomal prophages because maintenance of them is costly [89,90,91]. We suggest that postsegregational killing systems such as the zeta toxin can deter hosts from expelling viruses while under the unique stressors of the oxycline, allowing the viruses to lyse their hosts.
AMGs related to AMR and virulence peaked in the oxycline at the nearshore station and within the oxycline and OMZ core offshore (Supplementary Fig. S11B, L), resulting in negative and positive correlations with depth and temperature, respectively (Supplementary Fig. S12). Viruses have been shown to help defend their hosts from other microbes in free-living and biofilm settings, as well as influence their bacterial host’s ability to infect their own multicellular hosts and/or establish productive infection [65, 84, 87, 92, 93]. Two of the most abundant AMGs related to AMR and virulence in our dataset are related to the production of alginate, which helps form the capsule of P. aeruginosa that protects the bacterium from antibiotics and is important in biofilm production [94]. Other abundant AMGs include tylF and a tryptophan halogenase, which are involved in the production pathways of a macrolide that functions against Gram positive bacteria, and the broad-spectrum anti-fungal pyrrolnitrin, respectively [95, 96]. The most abundant virulence gene, a glycosyltransferase involved in the biosynthesis of lipopolysaccharide, contains an endotoxin in some Gram-negative bacteria including Pseudomonas [97,98,99]. Boosting the production of antimicrobial compounds and virulence factors in their hosts could allow viruses in the ETNP to keep the hosts alive long enough to facilitate viral replication. Overall, the presence of these genes related to host physiological state is once again parsimonious with the hypothesis that viruses will only contain AMGs required to enhance rate-limiting steps in reactions beyond that of which the host requires in order to increase viral replication [10, 47]. In the ETNP, the survivability of bacterial hosts under stress due to varying nutrient and oxygen conditions may be a more important factor in viral success than speeding up energy or carbon production.
AMGs related to vitamin metabolism
Vitamin and cofactor metabolism AMGs exhibited relatively low abundance in the mixed layer, increased dramatically in the oxycline, and then decreased with depth at both stations (Fig. 4G), resulting in no correlations with environmental variables (Supplementary Fig. S12). Several of these AMGs, including the most abundant pantoate transferase, are involved in the biosynthesis of pantothenate (vitamin B5), which is required for the synthesis of coenzyme A (CoA, [91, 92]), or are directly involved in CoA synthesis [100, 101]. CoA is an essential coenzyme that plays a key role in fatty acid metabolism and the biosynthesis of peptides [100]; notably, we detected a pantothenate kinase, which is the rate-limiting step in CoA biosynthesis, in all samples except below the OMZ. This suggests that viruses in the ETNP may be boosting pantothenate and CoA levels to bolster host metabolism. We also observed cobT, related to the aerobic production of cobalamin in bacteria [102], mostly in the oxycline of both stations. Recent research suggests the importance of cobalamin-producing archaea, especially Thaumarchaeota [103], in the ETNP as potential drivers of community structure as this highly-sought after coenzyme is produced by few organisms [104, 105], giving them a selective advantage. A study investigating putative archaeal viruses in the ETNP OMZ region did not identify any viruses with cobalamin biosynthesis genes, including cobT, but this may be due to the conservative nature of the analysis [105]. Thus, viruses in the ETNP may be modulating community structure by boosting cobalamin production in their hosts, especially in the oxycline at both stations and in deep offshore areas.
Conclusions
Marine OMZs constitute a relatively large portion of the world’s oceans and harbor unique microbial communities that perform globally-important biochemical processes (reviewed by Paulmier et al. [19,20,21, 27]), yet information regarding the viral communities within these regions remains sparse. Here we demonstrate that there are some similarities, as well as substantial differences, in viral community ecological metrics throughout the water column in the ETNP OMZ region. While viral abundance decreased within the low-oxygen portion of the water column, the frequency of infected cells was approximately equivalent to that found in the upper-ocean mixed layer, indicating that viruses are both present and actively infecting bacteria within the OMZ. We also demonstrate that the diversity of the viral communities was consistently high throughout the water column, but that viral community population structure diverged significantly with oxygen concentration. Given the divergent microbially-driven biogeochemical cycling that occurs as a function of oxygen concentration in marine OMZs, and the importance of AMGs as drivers of viral population structure [4, 8,9,10,11], we expected that viral-encoded AMGs involved in nitrogen and sulfur cycling would drive the differences observed in viral community structure throughout the water column in the ETNP. Instead, we observed that while a high portion of annotated PFAMs were shared among the depth categories, their relative abundances indicate strong selection for different viral-encoded AMGs with varying oxygen concentration that are not related to nitrogen or sulfur cycling (Fig. 5). Our study shows that the AMGs enriched in the upper oxygenated water column are related to increasing host metabolism, including photosynthesis, while viral AMGs within the low-oxygen portion of the water column are primarily related to enhancing the host bacterium’s ability to deal with oxygen-related stress (Fig. 5). We reason that this is due to the requirement of viruses to streamline their genomes and retain only essential genes, as they are limited by physical space in their capsids [106] and the energy and materials required for DNA replication [107]. We speculate that there may only be selective pressure to maintain AMGs within viral populations under certain conditions, such as the presence of nitrogen-related metabolism genes found only within a certain range of nitrogenous nutrient concentrations [10]. Thus, when host metabolism is sufficient to support viral replication, viruses may have no need to encode AMGs related to the predominant biogeochemical reactions being performed by their hosts in that environment, but will instead selectively encode any gene that increases viral replication. In the ETNP OMZ region, our study indicates that viral influences are not predominantly related to directly altering biogeochemical reactions, but instead they encode genes that enhance their bacterial hosts’ ability to sufficiently maintain metabolic functions for the virus to replicate.
These also represent most of the total normalized abundance of identified PFAMs, as represented by the boxes. Of those AMGs that were predominantly found in the mixed layer samples, the most abundant were psbA/D and psbN, which are involved in photosynthesis, and GDCP which is involved in glycine synthesis. The most abundant core low-oxygen AMGs include spoVS and spoVG, involved in sporulation, and hisA/F which is involved in histidine synthesis. This supports the hypothesis that the dominant viral strategy in the ETNP surface mixed layer is to boost host metabolism to facilitate viral replication, whereas those in the oxycline and OMZ are more likely to facilitate host persistence long enough for viral replication to occur.
Methods
Sample collection
Samples were collected from the ETNP OMZ during the OMZ Microbial Biogeochemistry Expedition cruise (R/V New Horizon, 13–28 June 2013) as previously described [105]. Samples were collected from two stations: Station 6 was located on the continental shelf (18°55’12” N and 104°53’24” W), and Station 2 was located ~450 km west of Station 6 (18 °55’12” N and 108°47’60” W). We thus use the terms “nearshore” and “offshore” to describe Stations 6 and 2, respectively. At each station, water samples were collected from eight depths spanning the mixed layer (5 m, 30 m, 60 m at the offshore station; 5 m and 30 m at nearshore station), oxycline (85 m and 130 m at offshore station: 60 m, 85 m, 100 m at nearshore station), OMZ core (300 m and 800 m at both stations), and below the OMZ (1000 m at both stations). Seawater was collected using Niskin bottles on a rosette containing a Conductivity Temperature Depth profiler (Sea-Bird SBE 911plus, Sea-Bird Electronics Inc., Bellevue, WA, USA) equipped with a Seapoint fluorometer (Seapoint Sensors Inc., Exeter, NH, USA) and SBE43- dissolved oxygen sensor (Sea-Bird Electronics Inc., Bellevue, WA, USA). CTD profiles from this study were deposited at the Biological & Chemical Oceanography Data Management Office (BCO-DMO) [108].
Microbial abundances
Triplicate samples (4 ml) for viral and bacterial enumeration were preserved with EM-grade glutaraldehyde (2% final concentration), flash-frozen in liquid nitrogen and stored between –72 °C and −80 °C until analysis. Viral and bacterial concentrations were determined using a previously described method [109] in which thawed samples were filtered onto 0.02-μm- pore-size filters (Anodisc, Whatman, GE Healthcare Life Sciences, Piscataway, NJ, USA), stained with SYBR Gold nucleic acid stain (Invitrogen, Life Technologies, Carlsbad, CA, USA) and enumerated using an epifluorescence microscope (Axio Imager. D2, Zeiss, Jena, Germany). Microbial abundances determined in this study were deposited at the BCO-DMO [110].
Frequency of infected cells
The percentage of cells with active lytic viral infections was determined by transmission electron microscopy to quantify the frequency of visibly infected cells [31] using intact cells [35]. Samples preserved with EM-grade glutaraldehyde (2% final concentration) were flash-frozen and stored at –80 °C until analysis. Samples (12 ml) were centrifuged onto 200-mesh copper grids with carbon-stabilized formvar support (Ted Pella, Redding, CA, USA) made hydrophilic with 20 s of glow discharge with a sputter coater (Hummer 6.2, Anatech, Battle Creek, MI, USA) for 1 h at 55,000 × g using an ultracentrifuge (LM-80, Beckman Coulter, Brea, CA, USA) with a swing-bucket rotor. Grids were then stained with 0.5% uranyl acetate and analyzed as previously described [35] to determine the frequency of visibly infected cells using a transmission electron microscope (FEI Tecnai Spirit). The frequency of infected cells was then calculated from the frequency of visibly infected cells [111]. Frequency of infected cells data generated from this study were deposited at BCO-DMO [112].
Morphological analysis of viral communities
Viral capsid diameters, tail length, and morphotype were determined using the qTEM method described previously [36]. Samples preserved with EM-grade glutaraldehyde (2% final concentration) were flash-frozen and stored at –80 °C until analysis. Viruses were deposited onto TEM grids using an air-driven ultracentrifuge (Airfuge CLS, Beckman Coulter, Brea, CA, USA), followed by positive staining with 2% uranyl acetate (Ted Pella, Redding, CA, USA). TEM grids were dried at room temperature overnight then stored desiccated until analysis using a transmission electron microscope (Philips CM12 FEI, Hilsboro, OR, USA) with 100 kV accelerating voltage. Micrographs of 100 viruses were obtained per sample using a MacroFire Monochrome charge-coupled device camera (Optronics, Goleta, CA, USA) and analyzed using ImageJ software (U.S. National Institutes of Health, Bethesda, MD, USA) [113] to measure the capsid diameter and tail length, and classify their morphotype as previously described [36]. Morphological data generated in this study were deposited at BCO-DMO [114].
Metagenome preparation and analysis
Ten samples were used for metagenomics (30 m, 85 m, 100 m, 300 m, and 1000 m at the nearshore station: 30 m, 60 m, 130 m, 300 m, and 1000 m at the offshore station). For each sample, 20 liters of seawater were filtered through a 0.22 μm-pore-size filter (Steripak; Millipore Sigma, Burlington, MA, USA) and viruses in the filtrate were concentrated using iron chloride flocculation as previously described [115, 116] followed by storage at 4 °C. Viral particles were then resuspended using an ascorbic EDTA buffer and their DNA extracted using a Wizard DNA purification system (Promega, Fitchburg, WI, USA) after treatment with DNase I as previously described [105]. Extracted DNA was then sheared with a Covaris ultra-sonicator (Woburn, MA, USA), gel-purified to select fragments of 160–180 bp in length, and then ligated and amplified using the standard Illumina protocol with PCR amplification of the library. Sequencing was carried out on a HiSEq 2000 system at the DOE Joint Genome Institute (Berkeley, CA, USA), except for the 300 m sample from the offshore station which was sequenced at the University of Arizona Genome Center. Metagenome accession numbers and metadata were deposited at BCO-DMO [117].
Sequencing reads were quality trimmed to remove bases with quality scores lower than two standard deviations from the average score (across sequencing cycles), and bases with a quality score lower than 20. Reads ≥95 bp were assembled using Idba_ud v1.1.2 with default parameters [118]. Assembled contigs were analyzed using the program VirSorter v1.0.2 with the ‘--virome’ option [39], and all contigs ≥10 kb of categories 1 and 2 as well as manually curated contigs and prophage predictions of categories 3–6 were selected for further analysis. Selected viral contigs were clustered with nucmer 3 [119] at ≥95% ANI across ≥80% of their lengths, as in [13], to generate a pool of non-redundant ‘population contigs’. Information about the metagenome assemblies and VirSorter contigs can be found in Table S1.
Taxonomic annotation of the viral populations was determined based on affiliation of >50% of the genes with a reference genome from RefSeq (version 74; using a BLASTp comparison with thresholds of 50 for bitscore and 10−5 for e value). All viral populations generated as described above were used in subsequent analyses regardless of their taxonomic annotation. Functional annotations of all predicted proteins from ETNP viral contigs was based on a comparison to the PFAM domain database v.27 [120] with HmmSearch [121] thresholds of 30 for bit score and 10−3 for e value). PFAMs hits were manually categorized by their general function, including “DNA replication, recombination, repair; nucleotide metabolism”, “lysis”, “metabolism”, “structural”, and “transcription, translation, protein synthesis”. Of these, 270 could not be assigned to a functional category but did match a PFAM domain, whereas 452 did not match any PFAM domain (12.4% and 11.4% bp mapped/kb genome/Mb metagenome, respectively). We included Class I or II AMGs in our analyses based on the established definitions [11], resulting in 329 identified AMGs (16.1% bp mapped/kb genome/Mb metagenome). Of these, 81 could only be identified as peripherally associated with metabolism (8.0% bp mapped/kb genome/Mb metagenome).
Statistical analyses
All statistical analyses were conducted using R version 3.5.1 [122]. Depth profiles, bar plots, stacked bar plots, and the heatmap were created using the ggplot2 package [123]. Shannon index (H’) and Pielou’s evenness (J’) were calculated using ‘diversity’ function in the vegan package [124]. Euler diagrams were visualized using the ‘venneuler’ function in the venneuler package [125].
Correlations were performed using the ‘rcorr’ function of the Hmisc package [126] using Pearson coefficients, and visualized using the ‘corrplot’ function of the corrplot package with an α of p < 0.05 [127]. We used a binomial test to compare the number of significant correlations we identified to the expected false-positive rate as previously described [128]. Briefly, we compared the expected false-positive rate of the number (n) of tests at our significance level α (n x α) to our number of significant correlations (‘successes’ under a binomial distribution). Because our number of determined significant correlations far exceeded the false-positive rate, we considered the correlations to be significant.
Correspondence analysis was performed using the ‘cca’ function in the vegan package [124] to obtain an ordination plot of viral communities based on viral capsid diameters or populations for each sample as in [36]. Average optimal capsid diameter bin size was determined using the method of [129]. Vectors and response surfaces of environmental variables were fitted to the CA ordination plot using the function ‘envfit’ in vegan with 10,000 simulations to estimate p values and the function ‘ordisurf’ in vegan, respectively [124]. Hierarchical clustering was performed using the ‘pvclust’ function of the pvclust package [130] with 100 bootstrap replications to estimate p values.
References
Fuhrman JA. Marine viruses and their biogeochemical and ecological effects. Nature. 1999;399:541–8.
Suttle CA. Viruses in the sea. Nature. 2005;437:356–61.
Breitbart M. Marine viruses: truth or dare. Ann Rev Mar Sci. 2012;4:425–48.
Brum JR, Sullivan MB. Rising to the challenge: accelerated pace of discovery transforms marine virology. Nat Rev Microbiol. 2015;13:147–59.
Breitbart M, Thompson L, Suttle C, Sullivan M. Exploring the vast diversity of marine viruses. Oceanography. 2007;20:135–9.
Puxty RJ, Millard AD, Evans DJ, Scanlan DJ. Shedding new light on viral photosynthesis. Photosynth Res. 2015;126:71–97.
Sharon I, Tzahor S, Williamson S, Shmoish M, Man-Aharonovich D, Rusch DB, et al. Viral photosynthetic reaction center genes and transcripts in the marine environment. ISME J. 2007;1:492–501.
Roux S, Hawley AK, Torres Beltran M, Scofield M, Schwientek P, Stepanauskas R, et al. Ecology and evolution of viruses infecting uncultivated SUP05 bacteria as revealed by single-cell- and meta-genomics. Elife. 2014;3:e03125.
Trubl G, Jang H Bin, Roux S, Emerson JB, Solonenko N, Vik DR, et al. Soil viruses are underexplored players in ecosystem carbon processing. mSystems. 2018;3:e00076–18.
Roux S, Brum JR, Dutilh BE, Sunagawa S, Duhaime MB, Loy A, et al. Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature. 2016;537:689–93.
Hurwitz BL, Brum JR, Sullivan MB. Depth-stratified functional and taxonomic niche specialization in the ‘core’ and ‘flexible’ Pacific Ocean Virome. ISME J. 2015;9:472–84.
Gazitúa MC, Vik DR, Roux S, Gregory AC, Bolduc B, Widner B, et al. Potential virus-mediated nitrogen cycling in oxygen-depleted oceanic waters. ISME J. 2021;15:981–98.
Brum JR, Ignacio-Espinoza JC, Roux S, Doulcier G, Acinas SG, Alberti A, et al. Ocean plankton. Patterns and ecological drivers of ocean viral communities. Science. 2015;348:1261498.
Cassman N, Prieto-Davó A, Walsh K, Silva GGZ, Angly F, Akhter S, et al. Oxygen minimum zones harbour novel viral communities with low diversity. Environ Microbiol. 2012;14:3043–65.
Vik D, Gazitúa MC, Sun CL, Zayed AA, Aldunate M, Mulholland MR, et al. Genome-resolved viral ecology in a marine oxygen minimum zone. Environ Microbiol. 2021;23:2858–74.
Mara P, Vik D, Pachiadaki MG, Suter EA, Poulos B, Taylor GT, et al. Viral elements and their potential influence on microbial processes along the permanently stratified Cariaco Basin redoxcline. ISME J. 2020;14:3079–92.
Tiano L, Garcia-Robledo E, Dalsgaard T, Devol AH, Ward BB, Ulloa O, et al. Oxygen distribution and aerobic respiration in the north and south eastern tropical Pacific oxygen minimum zones. Deep Res Part I Oceanogr Res Pap. 2014;94:173–83.
Schmidtko S, Stramma L, Visbeck M. Decline in global oceanic oxygen content during the past five decades. Nature. 2017;542:335–9.
Paulmier A, Ruiz-Pino D. Oxygen minimum zones (OMZs) in the modern ocean. Prog Oceanogr. 2009;80:113–28.
Wright JJ, Konwar KM, Hallam SJ. Microbial ecology of expanding oxygen minimum zones. Nat Rev Microbiol. 2012;10:381–94.
Bertagnolli AD, Stewart FJ. Microbial niches in marine oxygen minimum zones. Nat Rev Microbiol. 2018;1:723–9.
Codispoti LA, Friedrich GE, Packard TT, Glover HE, Kelly PJ, Spinrad RW, et al. High nitrite levels off northern Peru: a signal of instability in the marine denitrification rate. Science. 1986;233:1200 LP–1202.
Canfield DE, Stewart FJ, Thamdrup B, De Brabandere L, Dalsgaard T, Delong EF, et al. A cryptic sulfur cycle in oxygen-minimum-zone waters off the Chilean coast. Science. 2010;330:1375–8.
Hurwitz BL, Westveld AH, Brum JR, Sullivan MB. Modeling ecological drivers in marine viral communities using comparative metagenomics and network analyses. Proc Natl Acad Sci USA. 2014;111:10714 LP–10719.
Garcia-Robledo E, Padilla CC, Aldunate M, Stewart FJ, Ulloa O, Paulmier A, et al. Cryptic oxygen cycling in anoxic marine zones. Proc Natl Acad Sci USA. 2017;114:8319–24.
Lavin P, González B, Santibáñez JF, Scanlan DJ, Ulloa O. Novel lineages of Prochlorococcus thrive within the oxygen minimum zone of the eastern tropical South Pacific. Environ Microbiol Rep. 2010;2:728–38.
Ulloa O, Canfield DE, DeLong EF, Letelier RM, Stewart FJ. Microbial oceanography of anoxic oxygen minimum zones. Proc Natl Acad Sci USA. 2012;109:15996–6003.
Bettarel Y, Sime-Ngando T, Amblard C, Dolan J. Viral activity in two contrasting lake ecosystems. Appl Environ Microbiol. 2004;70:2941–51.
Weinbauer MG, Brettar I, Höfle MG. Lysogeny and virus-induced mortality of bacterioplankton in surface, deep, and anoxic marine waters. Limnol Oceanogr. 2003;48:1457–65.
Heldal M, Bratbak G. Production and decay of viruses in aquatic environments. Mar Ecol Prog Ser. 1991;72:205–12.
Proctor LM, Fuhrman JA. Viral mortality of marine bacteria and cyanobacteria. Nature. 1990;343:60–62.
Brum JR, Morris J, Décima M, Stukel M. Mortality in the oceans: causes and consequences. In Eco-DAS IX Symposium Proceedings. Association for the Sciences of Limnology and Oceanography; 2014.
Colombet J, Sime-Ngando T. Seasonal depth-related gradients in virioplankton: lytic activity and comparison with protistan grazing potential in Lake Pavin (France). Micro Ecol. 2012;64:67–78.
Colombet J, Sime-Ngando T, Cauchie HM, Fonty G, Hoffmann L, Demeure G. Depth-related gradients of viral activity in Lake Pavin. Appl Environ Microbiol. 2006;72:4440–5.
Brum J, Steward G, Jiang S, Jellison R. Spatial and temporal variability of prokaryotes, viruses, and viral infections of prokaryotes in an alkaline, hypersaline lake. Aquat Micro Ecol. 2005;41:247–60.
Brum JR, Schenck RO, Sullivan MB. Global morphological analysis of marine viruses shows minimal regional variation and dominance of non-tailed viruses. ISME J. 2013;7:1738–51.
Kauffman KM, Hussain FA, Yang J, Arevalo P, Brown JM, Chang WK, et al. A major lineage of non-tailed dsDNA viruses as unrecognized killers of marine bacteria. Nature. 2018;554:118–22.
Székely AJ, Breitbart M. Single-stranded DNA phages: from early molecular biology tools to recent revolutions in environmental microbiology. FEMS Microbiol Lett. 2016;363:27.
Roux S, Enault F, Hurwitz BL, Sullivan MB. VirSorter: Mining viral signal from microbial genomic data. PeerJ. 2015;2015:e985.
Hurwitz BL, Sullivan MB. The Pacific Ocean Virome (POV): a marine viral metagenomic dataset and associated protein clusters for quantitative viral ecology. PLoS One. 2013;8:e57355.
Aldunate M, Henríquez-Castillo C, Ji Q, Lueders-Dumont J, Mulholland MR, Ward BB, et al. Nitrogen assimilation in picocyanobacteria inhabiting the oxygen-deficient waters of the eastern tropical North and South Pacific. Limnol Oceanogr. 2020;65:437–53.
Solonenko SA, Ignacio-Espinoza JC, Alberti A, Cruaud C, Hallam S, Konstantinidis K, et al. Sequencing platform and library preparation choices impact viral metagenomes. BMC Genom. 2013;14:320.
Duhaime MB, Deng L, Poulos BT, Sullivan MB. Towards quantitative metagenomics of wild viruses and other ultra-low concentration DNA samples: a rigorous assessment and optimization of the linker amplification method. Environ Microbiol. 2012;14:2526–37.
Ganesh S, Bristow LA, Larsen M, Sarode N, Thamdrup B, Stewart FJ. Size-fraction partitioning of community gene transcription and nitrogen metabolism in a marine oxygen minimum zone. ISME J. 2015;9:2682–96.
Allen LZ, Allen EE, Badger JH, McCrow JP, Paulsen IT, Elbourne LD, et al. Influence of nutrients and currents on the genomic composition of microbes across an upwelling mosaic. ISME J. 2012;6:1403–14.
Hurwitz BL, U’Ren JM. Viral metabolic reprogramming in marine ecosystems. Curr Opin Microbiol. 2016;31:161–8.
Breitbart M, Bonnain C, Malki K, Sawaya NA. Phage puppet masters of the marine microbial realm. Nat Microbiol. 2018;3:754–66.
Ignacio-Espinoza JC, Sullivan MB. Phylogenomics of T4 cyanophages: lateral gene transfer in the ‘core’ and origins of host genes. Environ Microbiol. 2012;14:2113–26.
Crummett LT, Puxty RJ, Weihe C, Marston MF, Martiny JBH. The genomic content and context of auxiliary metabolic genes in marine cyanomyoviruses. Virology. 2016;499:219–29.
Sullivan MB, Lindell D, Lee JA, Thompson LR, Bielawski JP, Chisholm SW. Prevalence and evolution of core photosystem II genes in marine cyanobacterial viruses and their hosts. PLoS Biol. 2006;4:e234.
Bragg JG, Chisholm SW. Modeling the fitness consequences of a cyanophage-encoded photosynthesis gene. PLoS One. 2008;3:e3550.
Puxty RJ, Evans DJ, Millard AD, Scanlan DJ. Energy limitation of cyanophage development: implications for marine carbon cycling. ISME J. 2018;12:1273–86.
White AE, Foster RA, Benitez-Nelson CR, Masqué P, Verdeny E, Popp BN, et al. Nitrogen fixation in the Gulf of California and the Eastern Tropical North Pacific. Prog Oceanogr. 2013;109:1–17.
Jayakumar A, Chang BX, Widner B, Bernhardt P, Mulholland MR, Ward BB. Biological nitrogen fixation in the oxygen-minimum region of the eastern tropical North Pacific ocean. ISME J. 2017;11:2356–67.
Fuchsman CA, Devol AH, Saunders JK, McKay C, Rocap G. Niche partitioning of the N cycling microbial community of an offshore oxygen deficient zone. Front Microbiol. 2017;8:2384.
Zhang Y, Pohlmann EL, Halbleib CM, Ludden PW, Roberts GP. Effect of P(II) and its homolog GlnK on reversible ADP-ribosylation of dinitrogenase reductase by heterologous expression of the Rhodospirillum rubrum dinitrogenase reductase ADP-ribosyl transferase-dinitrogenase reductase-activating glycohydrolase regula. J Bacteriol. 2001;183:1610–20.
Tong W-H. Distinct iron-sulfur cluster assembly complexes exist in the cytosol and mitochondria of human cells. EMBO J. 2000;19:5692–5700.
Py B, Barras F. Building Feg-S proteins: bacterial strategies. Nat Rev Microbiol. 2010;8:436–46.
Eichhorn E, van der Ploeg JR, Kertesz MA, Leisinger T. Characterization of alpha-ketoglutarate-dependent taurine dioxygenase from Escherichia coli. J Biol Chem. 1997;272:23031–6.
Friedrich CG, Bardischewsky F, Rother D, Quentmeier A, Fischer J. Prokaryotic sulfur oxidation. Curr Opin Microbiol. 2005;8:253–9.
Anantharaman K, Duhaime MB, Breier JA, Wendt KA, Toner BM, Dick GJ. Sulfur oxidation genes in diverse deep-sea viruses. Science. 2014;344:757 LP–760.
Callbeck CM, Lavik G, Ferdelman TG, Fuchs B, Gruber-Vodicka HR, Hach PF, et al. Oxygen minimum zone cryptic sulfur cycling sustained by offshore transport of key sulfur oxidizing bacteria. Nat Commun. 2018;9:1729.
Carolan MT, Smith JM, Beman JM. Transcriptomic evidence for microbial sulfur cycling in the eastern tropical North Pacific oxygen minimum zone. Front Microbiol. 2015;6:334.
Ganesh S, Bertagnolli AD, Bristow LA, Padilla CC, Blackwood N, Aldunate M, et al. Single cell genomic and transcriptomic evidence for the use of alternative nitrogen substrates by anammox bacteria. ISME J. 2018;1:2706–22.
Howard-Varona C, Hargreaves KR, Abedon ST, Sullivan MB. Lysogeny in nature: mechanisms, impact and ecology of temperate phages. ISME J. 2017;11:1511–20.
Lill R, Dutkiewicz R, Elsässer HP, Hausmann A, Netz DJA, Pierik AJ, et al. Mechanisms of iron-sulfur protein maturation in mitochondria, cytosol and nucleus of eukaryotes. Biochim Biophys Acta Mol Cell Res. 2006;1763:652–67.
Fontecave M. Iron-sulfur clusters: ever-expanding roles. Nat Chem Biol. 2006;2:171–4.
Roche B, Aussel L, Ezraty B, Mandin P, Py B, Barras F. Iron/sulfur proteins biogenesis in prokaryotes: formation, regulation and diversity. Biochim Biophys Acta Bioenerg. 2013;1827:455–69.
Xu XM, Møller SG. Iron-sulfur clusters: Biogenesis, molecular mechanisms, and their functional significance. Antioxid Redox Signal. 2011;15:271–307.
Miller HK, Auerbuch V. Bacterial iron-sulfur cluster sensors in mammalian pathogens. Metallomics. 2015;7:943–56.
Sharon I, Battchikova N, Aro E-M, Giglione C, Meinnel T, Glaser F, et al. Comparative metagenomics of microbial traits within oceanic viral communities. ISME J. 2011;5:1178–90.
Loiseau L, Ollagnier-de-Choudens S, Nachin L, Fontecave M, Barras F. Biogenesis of Fe-S cluster by the bacterial suf system. SufS and SufE form a new type of cysteine desulfurase. J Biol Chem. 2003;278:38352–9.
Outten FW, Wood MJ, Muñoz FM, Storz G. The SufE protein and the SufBCD complex enhance SufS cysteine desulfurase activity as part of a sulfur transfer pathway for Fe-S cluster assembly in Escherichia coli. J Biol Chem. 2003;278:45713–9.
Ayala-Castro C, Saini A, Outten FW. Fe-S cluster assembly pathways in bacteria. Microbiol Mol Biol Rev. 2008;72:110–25.
Shepard EM, Boyd ES, Broderick JB, Peters JW. Biosynthesis of complex iron-sulfur enzymes. Curr Opin Chem Biol. 2011;15:319–27.
Lill R. Function and biogenesis of iron–sulphur proteins. Nature. 2009;460:831–8.
Seidler A, Jaschkowitz K, Wollenberg M. Incorporation of iron-sulphur clusters in membrane-bound proteins. Biochem Soc Trans. 2001;29:418–21.
Buchanan BB, Schürmann P, Wolosiuk RA, Jacquot J-P. The ferredoxin/thioredoxin system: from discovery to molecular structures and beyond. Photosynth Res. 2002;73:215–22.
Dubnau D, Losick R. Bistability in bacteria. Mol Microbiol. 2006;61:564–72.
Resnekov O, Driks A, Losick R. Identification and characterization of sporulation gene spoVS from Bacillus subtilis. J Bacteriol. 1995;177:5628–35.
Sonenshein AL. Bacteriophages: how bacterial spores capture and protect phage DNA. Curr Biol. 2006;16:R14–R16.
Sullivan MB, Coleman ML, Weigele P, Rohwer F, Chisholm SW. Three Prochlorococcus cyanophage genomes: signature features and ecological interpretations. PLoS Biol. 2005;3:0790–806.
Fortier L-C, Sekulovic O. Importance of prophages to evolution and virulence of bacterial pathogens. Virulence. 2013;4:354–65.
Mobberley J, Nathan Authement R, Segall AM, Edwards RA, Slepecky RA, Paul JH. Lysogeny and sporulation in Bacillus isolates from the Gulf of Mexico. Appl Environ Microbiol. 2010;76:829–42.
Brüssow H, Canchaya C, Hardt W-D. Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol Mol Biol Rev. 2004;68:560–602.
Meinhart A, Alonso JC, Strater N, Saenger W. Crystal structure of the plasmid maintenance system /: functional mechanism of toxin and inactivation by 2 2 complex formation. Proc Natl Acad Sci. 2003;100:1661–6.
Schuster CF, Bertram R. Toxin-antitoxin systems are ubiquitous and versatile modulators of prokaryotic cell fate. FEMS Microbiol Lett. 2013;340:73–85.
Kawano M. Divergently overlapping cis -encoded antisense RNA regulating toxin-antitoxin systems from E. coli. RNA Biol. 2012;9:1520–7.
Smith MA, Bidochka MJ. Bacterial fitness and plasmid-loss: the importance of culture conditions and plasmid size. Can J Microbiol. 1998;44:351–5.
Summers DK. The kinetics of plasmid loss. Trends Biotechnol. 1991;9: 273–8.
Persad AK, Williams ML, LeJeune JT. Rapid loss of a green fluorescent plasmid in Escherichia coli O157:H7. AIMS Microbiol. 2017;3:872–84.
Modi SR, Lee HH, Spina CS, Collins JJ. Antibiotic treatment expands the resistance reservoir and ecological network of the phage metagenome. Nature. 2013;499:219–22.
Hargreaves KR, Kropinski AM, Clokie MR. Bacteriophage behavioral ecology. Bacteriophage. 2014;4:e29866.
Naught LE, Gilbert S, Imhoff R, Snook C, Beamer L, Tipton P. Allosterism and cooperativity in Pseudomonas aeruginosa GDP-mannose dehydrogenase. Biochemistry. 2002;41:9637–45.
Dong C, Flecks S, Unversucht S, Haupt C, van Pee K-H, Naismith JH. Tryptophan 7-halogenase (PrnA) structure suggests a mechanism for regioselective chlorination. Science. 2005;309:2216–9.
Fouces R, Mellado E, Diez B, Barredo JL. The tylosin biosynthetic cluster from Streptomyces fradiae: genetic organization of the left region. Microbiology. 1999;145:855–68.
Heacock-Kang Y, Zarzycki-Siek J, Sun Z, Poonsuk K, Bluhm AP, Cabanas D, et al. Novel dual regulators of Pseudomonas aeruginosa essential for productive biofilms and virulence. Mol Microbiol. 2018;109:401–14.
Kurtov D, Kinghorn JR, Unkles SE. The Aspergillus nidulans panB gene encodes ketopantoate hydroxymethyltransferase, required for biosynthesis of pantothenate and Coenzyme A. Mol Gen Genet. 1999;262:115–20.
Huisjes R, Card DJ. Methods for assessment of pantothenic acid (Vitamin B5). In: Harrington D, editor. Laboratory assessment of vitamin status. London, UK; San Diego, CA, USA; Cambridge, MA, USA; Oxford, UK : Elsevier Inc.; 2019. p. 265–299. https://doi.org/10.1038/s41396-021-01143-1.
Leonardi R, Jackowski S. Biosynthesis of pantothenic acid and coenzyme A. EcoSal Plus. 2007;2:2.
Begley TP, Kinsland C, Strauss E. The biosynthesis of coenzyme a in bacteria. Vitam Horm. 2001;61:157–71.
Cameron B, Guilhot C, Blanche F, Cauchois L, Rouyez MC, Rigault S, et al. Genetic and sequence analyses of a Pseudomonas denitrificans DNA fragment containing two cob genes. J Bacteriol. 1991;173:6058–65.
Doxey AC, Kurtz DA, Lynch MD, Sauder LA, Neufeld JD. Aquatic metagenomes implicate Thaumarchaeota in global cobalamin production. ISME J. 2015;9:461–71.
Heal KR, Qin W, Amin SA, Devol AH, Moffett JW, Armbrust EV, et al. Accumulation of NO2-cobalamin in nutrient-stressed ammonia-oxidizing archaea and in the oxygen deficient zone of the eastern tropical North Pacific. Environ Microbiol Rep. 2018;10:453–7.
Vik DR, Roux S, Brum JR, Bolduc B, Emerson JB, Padilla CC, et al. Putative archaeal viruses from the mesopelagic ocean. PeerJ. 2017;5:e3428.
Streisinger G, Emrich J, Stahl MM. Chromosome structure in phage T4, III. Terminal redundancy and length determination. Proc Natl Acad Sci USA. 1967;57:292–5.
Mahmoudabadi G, Milo R, Phillips R. Energetic cost of building a virus. Proc Natl Acad Sci USA. 2017;114:E4324–E4333.
Brum J. 5m intervals of CTD profiles from R/V New Horizon cruise NH1315 in the Eastern Tropical North Pacific (ETNP) during June 2013. Biological and Chemical Oceanography Data Management Office (BCO-DMO). (Version 1) Version Date 2020-08-31 (2020). https://doi.org/10.26008/1912/bco-dmo.822818.1.
Noble RT, Fuhrman JA. Use of SYBR Green I for rapid epifluorescence counts of marine viruses and bacteria. Aquat Micro Ecol. 1998;14:113–8.
Brum J. Estimated abundances of viruses and bacteria determined in samples collected in the Eastern Tropical North Pacific (ETNP) on R/V New Horizon cruise NH1315 during June 2013. Biological and Chemical Oceanography Data Management Office (BCO-DMO). (Version 1) Version Date 2020-09-02 (2020). https://doi.org/10.26008/1912/bco-dmo.823094.1.
Binder B. Reconsidering the relationship between vitally induced bacterial mortality and frequency of infected cells. Aquat Micro Ecol. 1999;18:207–15.
Brum J. Estimated frequency of lytic viral infection from samples collected in the Eastern Tropical North Pacific oxygen minimum zone region (ETNP OMZ) on R/V New Horizon cruise NH1315 during June 2013. Biological and Chemical Oceanography Data Management Office (BCO-DMO). (Version 1) Version Date 2020-09-01 (2020). https://doi.org/10.26008/1912/bco-dmo.822914.1.
Abramoff MD, Magalhaes PJ, Ram SJ. Image processing with ImageJ. Biophotonics Int 2004;11:36–42.
Brum J. Morphotypes, capsid widths, and tail lengths of viruses from samples collected in the Eastern Tropical North Pacific oxygen minimum zone region (ETNP OMZ) on R/V New Horizon cruise NH1315 during June 2013. Biological and Chemical Oceanography Data Management Office (BCO-DMO). (Version 1) Version Date 2020-09-02 (2020). https://doi.org/10.26008/1912/bco-dmo.823131.1.
John SG, Mendez CB, Deng L, Poulos B, Kauffman AKM, Kern S, et al. A simple and efficient method for concentration of ocean viruses by chemical flocculation. Environ Microbiol Rep. 2011;3:195–202.
Duhaime MB, Sullivan MB. Ocean viruses: Rigorously evaluating the metagenomic sample-to-sequence pipeline. Virology. 2012;434:181–6.
Brum J. Accession numbers of viral metagenomes from samples collected in the Eastern Tropical North Pacific oxygen minimum zone region (ETNP OMZ) on R/V New Horizon cruise NH1315 during June 2013. Biological and Chemical Oceanography Data Management Office (BCO-DMO). (Version 1) Version Date 2020-09-04 (2020) https://doi.org/10.26008/1912/bco-dmo.823295.1.
Peng Y, Leung HCM, Yiu SM, Chin FYL. IDBA-UD: A de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics. 2012;28:1420–8.
Delcher AL, Salzberg SL, Phillippy AM. Using MUMmer to identify similar regions in large sequence sets. Curr Protoc Bioinforma. 2003; Chapter 10: Unit 10.3.
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, et al. Pfam: the protein families database. Nucleic Acids Res. 2014;42:D222–D230.
Eddy SR. Accelerated profile HMM searches. PLoS Comput Biol. 2011;7:1002195.
Team RCR. A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2018.
Wickham H. ggplot2: elegant graphics for data analysis. Springer-Verlag, New York: 2016.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB et al. vegan: Community Ecology Package. R package version 2.5-2. 2013 http://RForge.R-project.org/projects/vegan/.
Wilkinson L. venneuler: Venn and Euler diagrams. R package version 1.1-0. 2011 https://CRAN.Rproject.org/package=venneuler.
Harrell FE, With contributions from Charles Dupont and many others. Hmisc: Harrell Miscellaneous. R package version 4.3-0. 2019 https://CRAN.R-project.org/package=Hmisc.
Wei T, Simko V. R package “corrplot”: Visualization of a Correlation Matrix (Version 0.84). 2017. Available from https://github.com/taiyun/corrplot.
Emerson JB, Roux S, Brum JR, Bolduc B, Woodcroft BJ, Jang H Bin, et al. Host-linked soil viral ecology along a permafrost thaw gradient. Nat Microbiol. 2018;3:870–80.
Sturges HA. The choice of a class interval. J Am Stat Assoc. 1926;21:65–66.
Suzuki R, Shimodaira H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics. 2006;22:1540–2.
Acknowledgements
We thank the captain and crew of the R/V New Horizon and participants on the OMZ Microbial Biogeochemistry Expedition cruise. This work was funded by awards from the National Science Foundation (OCE #1658040) to JRB; the National Science Foundation (OCE #1536989) and the Gordon and Betty Moore Foundation (GBMF #3790) to MBS; National Science Foundation (1151698, 1558916, 1564559), Sloan Foundation grant (RC944) and Simons Foundation award (346253) to FJS; as well as the National Science Foundation Graduate Research Fellowship Program (1746902) to SKJ. The work conducted by the U.S. Department of Energy Joint Genome Institute (SR) is supported by the Office of Science of the U.S. Department of Energy under contract no. DE-AC02-05CH11231. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation.
Author information
Authors and Affiliations
Contributions
JRB, MBS, and FJS performed the project planning; JRB, SMS, and SKJ performed the experimental work; JRB, SR, SMS, and SKJ analyzed the data; JRB, SKJ, SR, SMS, MBS, and FJS wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jurgensen, S.K., Roux, S., Schwenck, S.M. et al. Viral community analysis in a marine oxygen minimum zone indicates increased potential for viral manipulation of microbial physiological state. ISME J 16, 972–982 (2022). https://doi.org/10.1038/s41396-021-01143-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01143-1