Abstract
Interactions between Sphagnum (peat moss) and cyanobacteria play critical roles in terrestrial carbon and nitrogen cycling processes. Knowledge of the metabolites exchanged, the physiological processes involved, and the environmental conditions allowing the formation of symbiosis is important for a better understanding of the mechanisms underlying these interactions. In this study, we used a cross-feeding approach with spatially resolved metabolite profiling and metatranscriptomics to characterize the symbiosis between Sphagnum and Nostoc cyanobacteria. A pH gradient study revealed that the Sphagnum–Nostoc symbiosis was driven by pH, with mutualism occurring only at low pH. Metabolic cross-feeding studies along with spatially resolved matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI-MSI) identified trehalose as the main carbohydrate source released by Sphagnum, which were depleted by Nostoc along with sulfur-containing choline-O-sulfate, taurine and sulfoacetate. In exchange, Nostoc increased exudation of purines and amino acids. Metatranscriptome analysis indicated that Sphagnum host defense was downregulated when in direct contact with the Nostoc symbiont, but not as a result of chemical contact alone. The observations in this study elucidated environmental, metabolic, and physiological underpinnings of the widespread plant–cyanobacterial symbioses with important implications for predicting carbon and nitrogen cycling in peatland ecosystems as well as the basis of general host-microbe interactions.
Similar content being viewed by others
Introduction
Associations between Sphagnum peat mosses and phototrophic microorganisms were first reported more than a century ago, q.v. Limpricht [1], who described the presence of Nostoc within the dead water-filled hyaline cells of plant leaflets [2]. Subsequent work confirmed these associations in sites across the Stordalen mire of Swedish Lapland [3, 4] and the vicinity of Uppsala, Sweden. These sites encompass numerous Sphagnum-containing habitats, including a coniferous forest, a large mire, a nutrient-rich fen, and wetland sites exposed to human activity [5]. These early studies revealed that several algae/cyanobacterial genera are associated with the outside surfaces of Sphagnum as epiphytes, whereas genus Nostoc is most commonly found within plant cells as an endophyte. In the bryophyte hornwort, endophytic cyanobacteria receive sugars from the host plant in exchange for fixed nitrogen (N) in the form of ammonia [6,7,8]. However, the molecules exchanged in the Sphagnum–cyanobacteria symbiosis remain unknown, and it is not clear whether different molecules are exchanged during Sphagnum symbioses with epiphytic and endophytic cyanobacteria.
Cyanobacteria associated with Sphagnum fix N2 at higher rates than those not associated with Sphagnum, and in symbioses, N2 fixation can occur at low pH [2]. It has been hypothesized that Sphagnum hyaline cells have a relatively buffered pH as a consequence of cell wall cation exchange, suggesting that they serve as a “microbiome oasis” under harsh, low-pH pore water conditions [9, 10]. A field study [11] also suggested that pH influences the Sphagnum–cyanobacteria symbiosis. Van den Elzen et al. [11] found that N2 fixation activity was stimulated by the addition of bicarbonate, whereas Sphagnum growth remained unaffected, implying that N2-fixers give N to the host plant in exchange for shelter from low pH.
Sphagnum spp.-dominated peatlands provide a dramatic example of how changes in species interactions can cascade across levels of biological organization. These peatlands occupy just 3% of the Earth’s land surface, yet store ~25–30% of the planet’s soil carbon as dead recalcitrant peat [12]. Together with their associated N2-fixing bacteria, they provide a critical N input to peatland ecosystems [13, 14]. Despite the importance of this unique symbiosis to plant growth, ecosystem productivity, and even ecosystem carbon and nutrient cycling, we lack a basic understanding of the environmental conditions that allow this symbiosis to form, which metabolites are exchanged, and the physiological processes underlying these symbiotic interactions. Endophytic plant—cyanobacteria associations are best characterized and classified as nutritional symbioses where the cyanobacteria provision the plant host with fixed-N products in exchange for host derived reduced C. These mutualistic symbioses are typically facultative, and span a broad plant host phylogeny including hornworts, liverworts, angiosperms and cycads [15, 16]. Epiphytic plant—cyanobacteria symbioses are less studied, yet emerging results from the feathermoss Pleurozium schreberi system is providing an initial characterization. In this association, host nitrogen limitation is similarly necessary for the induction of motile cyanobacteria hormogonia production, taxis and host plant colonization [17]. However, some difference exists. For example, cyanobacteria carbon fixation and photosynthetic gene expression remains high, suggesting that the cyanobacteria may be autotrophic and not receiving reduced C from the plant host [18]. In a follow-up study, Stuart et al. [19] used cyanobacteria targeted mutagenesis coupled with stable isotope probing and high lateral resolution secondary ion mass spectrometry (NanoSIMS) to confirm mutualism as inferred from bidirectional C and N exchange, and also supported a role for organic sulfur. Such studies highlight both similarities and differences among endophytic and epiphytic plant—cyanobacteria associations and brings to question how plants that support both forms of symbiosis, like Sphagnum spp., physiologically operate.
The overarching aim of this study was to provide an initial characterization of the physiological and metabolic reprogramming necessary for the Sphagnum–Nostoc symbiosis and to determine what role, if any, pH plays in the establishment of symbiosis. Specifically, we tested the following hypotheses: (1) that the Sphagnum–Nostoc symbiosis is dependent on the surrounding environmental context, especially regarding pH; and (2) that the dual endophytic and epiphytic nature of the Sphagnum–Nostoc symbiosis results in unique host and bacterium physiology, metabolism, and metabolic exchange relative to other plant—cyanobacteria symbioses. To test these hypotheses, we characterized the Sphagnum–Nostoc symbiosis through metabolic cross-feeding studies, spatially resolved metabolic profiling and metatranscriptomes. The spatially resolved metabolic profiling was optimized to detect disaccharides and sulfur-rich compounds [20, 21], and allowed the placement of symbiotic partners within proximity of each other so metabolites can be sensed and exchanged in an arrangement that complemented the cross-feeding study. Furthermore, Sphagnum has microbially filled hyaline cells resulting in a higher microbe to plant cell ratio compared to other plant-microbe systems, which provides an ideal system for performing metatranscriptomics analysis for both host and Nostoc without the need of biasing RNAseq enrichment protocols.
Materials and methods
Stain selection and culture conditions
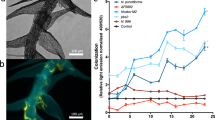
To identify a symbiotically competent Nostoc strain for our symbiosis experiments, preliminary co-culture experiments were performed to test 18 strains from the UTEX Culture Collection of Algae (University of Texas–Austin) for consistent and frequent endophytic colonization. The preliminary evaluation was conducted as described below for growth experiments at pH 5.5. Nostoc spp. strains UTEX 1037, UTEX1632, UTEX 2209, UTEX LB 1833, UTEX 2210, UTEX 1621, UTEX B 1545, UTEX 1038, UTEX B 384, UTEX B EE21, UTEX B EE34, UTEX B EE20, UTEX B EE7, UTEX B 2494, UTEX B EE4, UTEX B EE5, UTEX B 2493, UTEX B 2492 were evaluated for colonization within hyaline cells (i.e., endophytic colonization). Based on this preliminary study, Nostoc muscorum UTEX 1037 was selected as a high-colonizing strain (Fig. S1 image, Supplemental Methods). To further evaluate the suitability of N. muscorum UTEX 1037 as an Sphagnum angustifolium (Russow) C.E.O. cyanobiont, a phylogenetic tree was constructed from 37 NCBI available cyanobacterial genomes including two cyanobacteria known to colonize moss (N. punctiforme and N. KVJ20 [22]) and 3 cyanobacterial genomes isolated from moss [18]. The tree used a concatenated alignment of 31 proteins with dense sampling of the Nostoc punctiforme species group, Anabaena species, and two outgroup taxa Acaryochloris spp. CCMEE 5410 and A. marina MBIC11017 (Fig. S1, Supplemental Methods). The relatively close branching of N. muscorum UTEX 1037 to other moss associated isolates motivated our final selection of this strain for further experimentation.
Nostoc strains were cultivated in 250-mL Erlenmeyer flasks in 100 mL of BG-110 [23] medium (pH 8.2). The flasks were shaken at 125 rpm at 24 °C with a 16 h/8 h (day/night) cycle at 150 PAR for 21 days. S. angustifolium plants were collected from the SPRUCE study in Marcell Experimental Forest (MN, USA) [24]. S. angustifolium plants were exposed to multiple washes of 0.5–1.0% sodium hypochlorite to generate axenic plant cultures; washes were repeated until there was no visual microbial contamination which was confirmed via genome sequencing (Phytozome v13, S. angustifolium v1.1). Axenic S. angustifolium cultures were maintained on Knop’s [25] medium at pH 5.7 with a 16 h/8 h (day/night) cycle at 150 PAR.
Sphagnum and Nostoc growth experiments
An axenic S. angustifolium individual (upper 2 cm portion) and/or 30 mg of (fresh weight) of N. muscorum UTEX 1037 were used to inoculate 2 ml of BG-110 at pH 3.5, 5.5, or 8.5 in a 2-ml Eppendorf tube. Six replicates were sampled for each treatment after incubation at 24 °C with a 16 h/8 h (day/night) cycle at 150 PAR for 14 days and oven dried for 48 h. Dry weight of S. angustifolium grown individually was compared to the sum of the dry weight N. muscorum UTEX 1037 and S. angustifolium individually grown as well as the dry weight of N. muscorum UTEX 1037 and S. angustifolium grown together using Kruskal-Wallis with multiple testing correction using FDR (False Discovery Rate) cutoff ≤ 0.05 in R v3.6.3.
Cross-feeding experiment
Four replicates of N. muscorum UTEX 1037 (1 g wet weight) and four replicates of axenic S. angustifolium (~1 g wet weight) were each cultured individually in 75 ml of BG-110 at pH 5.5 and shaken at 125 rpm at 24 °C with a 16 h/8 h (day/night) cycle at 150 PAR for 21 days. Cultures were centrifuged at 3360 g for 10 min at 4 °C to pellet organisms. Pellets were equally divided, immediately frozen and stored at −80 °C for exometabolite (n = 4 for each organism) and RNA-seq (n = 4 of each organism) analyses. Supernatants were sterile-filtered through a 0.22-µm filtration device, lyophilized, resuspended in 2 mL 100% methanol, and centrifuged to pellet salts. Supernatants were equally divided for exometabolite analysis or supplementation and dried under vacuum (SpeedVac). Supernatant samples for exometabolite analysis (four replicates of each organism) were stored at −80 °C and the remaining supernatant samples (four replicates of each organism) were pooled, redissolved and supplemented to BG-110 medium. For exometabolite and RNA-seq analysis, four replicates of each organism were cultured for 4 weeks in the supplemented BG-110 media.
Exometabolite analysis from cross-feeding study
Metabolites were extracted from both cell and media samples for LC-MS metabolomics analysis. Pelleted cells in 2 mL Eppendorfs were lyophilized dry (FreeZone 2.5 Plus, Labconco), then bead-beaten with a 3.2 mm stainless steel bead for 5 s (3×) in a bead-beater (Mini-Beadbeater-96, BioSpec Products) to powder. For extraction, 1 mL of 100% MeOH was added to each sample, then each briefly vortexed, sonicated in a water bath for 10 min, then centrifuged for 7 min at 7000 rpm to pellet cellular debris. Supernatant was then transferred to a 2 mL Eppendorf, dried in a SpeedVac (SPD111V, Thermo Scientific), and stored at −80 °C. Extraction controls were prepared similarly but using empty tubes exposed to the same extraction procedures.
In preparation for LC-MS/MS analysis, dried extracts were resuspended by adding 300 µL 100% methanol containing various internal standard compounds (15 µM, mix of 13C-15N labeled amino acids, #767964; 1 µg/mL, 2-Amino-3-bromo-5-methylbenzoic acid, #R435902; 10 µg/mL, d4-lysine, #616192; 5 µg/mL, 13C-15N-phenylalanine, #608017; 2 µg/mL, 9-anthracene carboxylic acid, #A89405; 5 µg/mL, 3,6-dihydroxy-4-methylpyridazine, #668141; 10 µg/mL, d5-benzoic acid, #217158, 10 µg/mL, 4-(3,3-dimethyl-ureido)benzoic acid, #CDS014672—Sigma) vortexed briefly, sonicated in a water bath for 10 min, then centrifuged (5 min at 5000 rpm). 150 µL of resuspended extract was centrifuge-filtered (2.5 min at 2500 rpm) through a 0.22 µm filter (UFC40GV0S, Millipore), then transferred to a glass autosampler vial.
Samples were analyzed via LC-MS on a system consisting of an Agilent 1290 UHPLC coupled to a Thermo QExactive Orbitrap HF (Thermo Scientific, San Jose, CA) mass spectrometer. Normal phase chromatography was performed by injecting 2 µL extract into a zic-HILIC column (Millipore SeQuant ZIC-HILIC, 150 ×2.1 mm, 5 µm, Cat# 50454) warmed to 40 °C. The column was equilibrated with 100% buffer B (95:5 ACN:H2O w/ 5 mM ammonium acetate) for 1.5 min at 0.45 mL/min, diluting buffer B down to 65% with buffer A (H2O w/ 5 mM ammonium acetate) for 13.5 min, down to 0% B over 3 min while increasing flow to 0.6 mL/min, and followed by isocratic elution in 100% buffer A for 5 min. MS and MS/MS data were collected in both positive and negative ion mode using, with full MS spectra acquired ranging from 70 to 1050 m/z at 60,000 resolution, and fragmentation data acquired using an average of stepped collision energies of 10, 20, and 30 eV at 17,500 resolution. Orbitrap instrument parameters included a sheath gas flow rate of 50 (au), auxiliary gas flow rate of 20 (au), sweep gas flow rate of 2 (au), 3 kV spray voltage and 400 °C capillary temperature. Sample injection order was randomized, and an injection blank of methanol only run between each sample.
Metabolites were identified based on exact mass and retention time coupled with comparing MS/MS fragmentation spectra to compound standards. LC-MS data was analyzed using custom Python code [26]. Features (unique m/z coupled with retention time, RT) were assigned a score from 0 to 3, representing the level of confidence in compound identification and then rated according to the Metabolomics Standards Initiative (MSI) confidence levels [27]. A positive level 1 identification was given for compounds detected at m/z < / = 5 ppm or 0.001 Da from theoretical as well as RT < / = 0.5 min compared to a pure standard run using the same LC-MS method. A compound with the highest level of positive identification (score of 3), exceeding level 1, also had matching MS/MS fragmentation spectra in comparison to either an outside database (e.g., METLIN) or internal database generated from standards run and collected on a Q Exactive Orbitrap HF MS. Mismatches in MS/MS invalidated as an identification. Starting total ion count of metabolites in the cross-fed supernatant were compared to the ending supernatant metabolite total ion count using in R v3.6.3. Starting total ion count of metabolites in the cross-fed supernatant were compared to the ending supernatant metabolite total ion count using Kruskal-Wallis with multiple testing correction using FDR cutoff ≤ 0.05 in R v3.6.3.
MALDI-MSI analysis
Samples were prepared for matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI-MSI) analysis using a modification of a workflow we described previously [21]. Briefly S. angustifolium and N. muscorum UTEX 1037 were positioned 2 cm apart for co-culture or individually in the center (controls) of BG-110 (pH 4.5) 1.5% agar plates and incubated at 24 °C with a 16 h/8 h (day/night) cycle until the interaction zones were analyzed. Agar areas of the N. muscorum UTEX 1037—S. angustifolium interaction and of isolated culture controls were excised from Petri dishes and placed onto double-sided adhesive copper tape adhered to indium tin oxide-coated glass slides (Bruker Daltonics). The copper tape approach enhanced our sensitivity for analysis in negative ionization mode and improved adhesion of agar onto the MALDI target. Notably, the moss protruded from the surface of the agar, so plant tissue was carefully removed prior to dehydration to assist in MS analyses. Samples were dried at 40 °C overnight, after which MALDI matrix was applied using a HTX TM-Sprayer (HTX Technologies). For analysis in positive-ion mode, universal MALDI matrix (1:1 2-5-dihydroxybenzoic acid: α-Cyano-4-hydroxycinnamic acid), 20 mg/mL in 70% MeOH, was applied using the following spraying conditions: 12 passes with a track spacing of 3 mm, flow rate of 0.1 mL/min, spray velocity 1,200 mm/min, spray pressure of 10 psi (N2), and a 40-mm distance from the sprayer nozzle to the sample. For analysis in negative-ion mode, 7 mg/mL of N-(1-naphthyl) ethylenediamine dihydrochloride (NEDC) in 70% MeOH was sprayed with eight passes at 1,200 µL/min, 75 °C, a spray spacing of 3 mm, and a spray velocity of 1200 mm/min. MALDI-MSI was performed on a 15-Tesla Fourier transform ion cyclotron resonance (FTICR)-MS (Bruker Daltonics) equipped with SmartBeam II laser source (355 nm) using 200 shots/pixel with a frequency of 2 kHz and a step size of 200 µm. FTICR-MS was operated to collect m/z 92–700, using a 209-ms transient, which translated to a mass resolution of R ~ 120,000 at 400 m/z. Data were acquired using FlexImaging (v 4.1, Bruker Daltonics), and image processing and visualization were performed using SCiLS (Bruker Daltonics).
RNA sequencing and analysis
RNA was extracted from pelleted N. muscorum UTEX 1037 or S. angustifolium grown in BG-110 and BG-110 supplemented with exometabolites from the corresponding organism (n = 4 for each treatment) by a combined method using CTAB lysis buffer and the Spectrum Total Plant RNA extraction kit (Sigma). Approximately 100 mg of flash-frozen ground tissue was incubated in 850 µl of CTAB buffer (1.0 % β-Me) at 56 °C for 5 min; 600 µl chloroform:isoamylalcohol (24:1) was added, and the samples were centrifuged for 8 min. The supernatant (~730 µl) was transferred to a filter column provided in the Spectrum kit, and the standard Spectrum kit protocol was followed accompanied by DNase treatment. Ribo-zero bacterial (Illumina cat#MRZB12424) and plant (Illumina cat#MRZSR116) were used to enrich the transcripts. The Applied Biosystems SOLiD Total RNA-Seq kit (catalog number 4445374) was used to generate the cDNA template library. Briefly, the mRNA was fragmented by chemical hydrolysis followed by ligation with strand-specific adapters and reverse transcription to generate cDNA. cDNA fragments of 150–250 bp were isolated, and the SOLiD EZ Bead system was used to perform emulsion clonal bead amplification to generate bead templates for SOLiD platform sequencing. Samples were sequenced on the 5500XL SOLiD platform. The 50-base short-read sequences produced by the 5500XL SOLiD sequencer were mapped in color space against the genomes of Sphagnum angustifolium v1.1 (Phytozome v13, phytozome-next.jgi.doe.gov/) and N. muscorum UTEX 1037 draft genome with the SOLiD LifeScope software version 2.5, using default parameters.
Reads from N. muscorum UTEX 1037 were assessed for quality using htseq-qa [28]. Sequence regions showing GC content instability were trimmed using an R script. Reads were assembled using SPAdes version: 3.10.1 [29] using -k 21,33,55,77,99,127. Best assembly quality (as assessed by QUAST [30]) was acquired using 25% of the reads. Gene annotation was performed with Prokka [31], and gene sets for photosynthesis, carbon fixation, and nitrogen fixation were identified using HMMs with HMMER [32]. Plant gene models were assigned to MapMan4 ontology for gene set enrichment. Gene set enrichment analysis (GSEA) of differentially expressed genes in S. angustifolium was performed using MapMan4 gene ontologies on the desktop version of MapMan. MapMan4 gene ontologies were assigned using the Mercator 4 v2.0 web portal [33]. To determine significance, Kruskal–Wallis tests within the MapMan desktop version, with multiple testing correction using FDR cutoff ≤ 0.05, were performed in R v3.6.3. GSEA was performed similarly for N. muscorum UTEX 1037.
Results
Growth study
The Sphagnum–Nostoc symbiosis is dependent on pH
To test the hypothesis that variable pH conditions influence S. angustifolium—N. muscorum UTEX 1037 symbiosis, we grew germ-free S. angustifolium angustifolium and N. muscorum UTEX 1037 at pH 3.5, 5.5, and 8.5. Symbiosis outcome was evaluated by comparing the growth of S. angustifolium in direct contact with N. muscorum UTEX 1037 (designated Nostoc/Sphagnum symbiosis), relative to individual growth in isolation and the sum of individual growth (designated Nostoc + Sphagnum). There was a clear benefit to symbiosis in the low pH (3.5) environment: when the species were grown together under these conditions, the change in weight was greater than the sum of individual growth (Fig. 1; FDR adjusted p = 0.012). However, the growth benefit was not clearly observed at pH of 5.5 or 8.5 (Table S1).
At pH 3.5, growth was highest when both organisms were grown together (Table S1). Growth of Sphagnum grown individually or with Nostoc across a pH gradient were determined by the final dry weight (mg) of organisms grown in BG-110 (n = 6). Samples labeled with “symbiosis” represent the two organisms grown together, and Nostoc + Sphagnum shows the sum of individual growth of each organism alone.
Metabolic cross-feeding study identifies the utilization and release of 225 exometabolites
The growth benefits observed for both N. muscorum UTEX 1037 and S. angustifolium demonstrate a mutualistic symbiosis at low pH and motivated us to characterize the metabolic exchange mediating this interaction. Therefore, we sought to assess the metabolites being released by each partner of the symbiosis when grown in fresh liquid BG-110 medium and the change in abundance of the released exometabolites in spent medium when cross-fed to the other partner (Fig. 2A). Overall, we identified 225 exometabolites from the spent medium of individual partners when cultured in fresh BG-110 medium (Table S2). Of the identified exometabolites, N. muscorum UTEX 1037 53% of exometabolites exuded by S. angustifolium and S. angustifolium depleted 28% of exometabolites exuded by N. muscorum UTEX 1037 (Fig. 2C).
A For the cross-feeding study, each organism was cultured in BG-110 followed by characterization by liquid chromatography-mass spectrometry (LC-MS). The spent medium was then cross-fed in a full factorial design in which the resultant spent medium and cell extracts were profiled using LC-MS. B Experimental design of MALDI-MSI analysis of metabolites produced by each organism alone or when grown together in close proximity. C The number of statistically significant identified cross-fed exometabolites depleted by each organism out of all exometabolites exuded by the symbiosis partner.
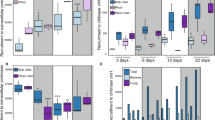
To further characterize metabolite exchange, the exometabolites that significantly (p < 0.05) changed in abundance before and after feeding were categorized into six chemical classes: amino acids, carbohydrates, fatty acids and their conjugates, lipids, nucleotides and nucleosides, and organic acids (Fig. 3). N. muscorum UTEX 1037 depleted large percentages from each chemical class of S. angustifolium exometabolites: 75% of the measured amino acids, 77% of the carbohydrates, 50% of the fatty acids, 68% of the lipids, 35% of the nucleotides/nucleosides, and 62% of the organic acids. Within each class, log-fold change (LFC) analysis revealed that the exometabolites that changed the most after cross-feeding were as follows: the pyrimidine nucleotide sugars uridine 5’-diphosphogalactose (−14.53 LFC, p value = 0.01) and uridine 5′-diphosphoglucose (−14.5 LFC, p value = 0.01); the amino acids arginine (−7.87 LFC, p value = 0.02), acetyl-L-alanine (−5.59 LFC, p value = 0.02), and asparagine (−5.45 LFC, p value = 0.02); the organic acid sulfuric acid ester choline-O-sulfate (−5.62 LFC, p value = 0.02); and the carbohydrate trehalose (−2.7 LFC, p value = 0.02). Thus, these compounds represent S. angustifolium exometabolites depleted by N. muscorum UTEX 1037 (Table S3).
Rows show metabolites depleted by each partner, columns indicate the class of the metabolite, and colors indicate the source of the metabolite (green = Sphagnum, blue=Nostoc). Boxes span from the 25th to the 75th percentile. The horizontal line indicates the median, the error bars are the range of data or 1.5 interquartile range, and the points represent individual metabolites; n = 4.
When fed N. muscorum UTEX 1037 exometabolites, S. angustifolium depleted 30% of the identified amino acids, 54% of the carbohydrates, and 76% of the nucleosides. Among the depleted metabolites within each class, LFC analysis revealed that the most depleted exometabolites were as follows: the nucleosides cytidine (−17.7 LFC, p value = 0.01), xanthosine (−5.67 LFC, p value = 0.02), adenine (−3.98 LFC, p value = 0.02), 2’-deoxyguanosine (−2.43 LFC, p value = 0.02), and uridine (−2.66 LCF, p value = 0.02); the amino acids amino acid derivatives aspartate (−4.49 LFC, p value = 0.02), acetyl-methionine (−2.77 LFC, p value = 0.02), arginine (−2.6 LFC, p value = 0.02), glutamic acid (−1.8 LFC, p value = 0.02); and the carbohydrates and carbohydrate derivatives phosphoglyceric acid (−3.12 LFC, p value = 0.02) and gluconate (−2.6 LFC, p value = 0.02) (Table S4).
Spatial metabolite profiling corroborates the exchange of trehalose and xanthosine in the Sphagnum–Nostoc symbiosis
We further investigated the results of the cross-feeding study using ultrahigh mass resolution and mass accuracy MALDI-MSI. This approach allowed us to place the symbiotic partners within proximity of each other, allowing metabolites to be sensed and exchanged in an arrangement that complemented the cross-feeding study. We hypothesized that metabolites critical for symbiosis would increase in abundance when the symbiosis partner was present.
To test this hypothesis, we first targeted candidate metabolites identified from the cross-feeding study and investigated their spatially resolved abundance profiles across the interaction zone. Using this approach, we found that N. muscorum UTEX 1037 and S. angustifolium produced small amounts of xanthosine when grown individually; however, when the species were grown in proximity to each other, N. muscorum UTEX 1037 increased xanthosine production, whereas S. angustifolium did not (Fig. 4D). Furthermore, a cross-feeding experiment showed that S. angustifolium depleted N. muscorum UTEX 1037 provided xanthosine (from ave. ion count 1.4 × 105, sd 0.8 × 105 to ave. ion count 0.02 × 105, sd 0.05 × 105), whereas N. muscorum UTEX 1037 did not deplete the S. angustifolium -provided xanthosine (from ave. ion count 1.0 × 107, sd 0.37 × 107 to ave. ion count 1.0 × 107, sd 0.4 × 107) (Fig. 4A). Similar results were observed for adenine (Fig. 4E): cross-feeding revealed that S. angustifolium depleted adenine released from N. muscorum UTEX 1037 from ave. ion count 14.9 × 107, sd 8.9 × 107 to ave. ion count 0.9 × 107, sd 0.3 × 107), and N. muscorum UTEX 1037 released elevated levels of adenine when supplemented with S. angustifolium exudates from ave. ion count 1.0 × 107, sd 0.38 × 107 to ave. ion count 2.4 × 107, sd 1.0 × 107) (Fig. 4C). Trehalose distribution was investigated similarly. Cross-feeding analysis revealed a clear trend in production of putative trehalose by S. angustifolium, with N. muscorum UTEX 1037 consuming up to 85% of the exometabolite (Fig. 5A). The abundance of putative trehalose being produced by S. angustifolium was similar between individual S. angustifolium cultures in fresh BG-110 medium (ave. ion count 2.8 × 106, sd .94 × 106) or after being fed N. muscorum UTEX 1037 exometabolites (ave. ion count 16.0 × 106, sd 8.3 × 106). The MALDI-MSI data support these findings: ion abundance was similar in isolated S. angustifolium and S. angustifolium grown in proximity of N. muscorum UTEX 1037. However, we observed a notable increase in the abundance of the disaccharide pool in N. muscorum UTEX 1037 when co-cultured with S. angustifolium. (Fig. 5B).
Boxplots of exometabolite cross-feeding of Sphagnum angustifolium (Sphagnum) and Nostoc muscorum UTEX 1037 (Nostoc) spent media demonstrate an exchange of xanthosine (A), choline O-sulfate (B), and adenine (C) between Nostoc and Sphagnum. Boxes span from the 25th to the 75th percentile n = 4. The horizontal line indicates the median, the lines denote the max and min values, outliers are displayed as dots. Exchange of choline O-sulfate by Sphagnum for xanthosine provided by Nostoc (D) and an increase in production of adenine by Nostoc in the presence of Sphagnum (E) were also confirmed by MALDI-MSI.
Boxplots of exometabolite cross-feeding of Sphagnum angustifolium (Sphagnum) and Nostoc muscorum UTEX 1037 (Nostoc) spent media demonstrate significant exudation of trehalose by moss fed cyanobacteria exometabolites and depletion of trehalose by Nostoc when fed Sphagnum exometabolites (A). Boxes span from the 25th to the 75th percentile n = 4. The horizontal line indicates the median, the lines denote the max and min values, outliers are displayed as dots. Negligible exudation of trehalose was detected by MALDI-MSI in moss alone, but increased exudation was detected when Nostoc (N) and Sphagnum (S) were grown together (B).
Spatial metabolite profiling corroborates choline-O-sulfate exchange and identifies additional sulfur-rich taurine and sulfoacetate as exometabolites contributing to the symbiosis
Choline-O-sulfate was also identified in our cross-feeding study and visualized by MALDI-MSI. Ion images indicated that S. angustifolium produces choline-O-sulfate regardless of the presence of the N. muscorum UTEX 1037 partner (Fig. 4D) and that the metabolite was not detected in N. muscorum UTEX 1037. Cross-feeding results revealed that on average, 98% of choline-O-sulfate provided by S. angustifolium was depleted by N. muscorum UTEX 1037 (Fig. 4B). In addition, MALDI-MSI analysis showed that S. angustifolium produced sulfoacetate, which was excreted at high levels in the agar medium only when N. muscorum UTEX 1037 and S. angustifolium were grown together. Furthermore, MALDI-MSI revealed evidence that taurine was metabolized to sulfoacetaldehyde via 2-oxyglutarate and glutamate, and that the sulfoacetaldehyde was further converted to acetylphosphate, likely feeding into pyruvate metabolism (Fig. S2) within N. muscorum UTEX 1037. The induction of those metabolites, with the exception of acetylphosphate, was only observed when N. muscorum UTEX 1037 was cultured in proximity to S. angustifolium.
Transcriptional responses of Nostoc and Sphagnum provide insight into symbiotic interactions
Metatranscriptomes were collected at the conclusion of our cross-feeding experiment. Samples were collected from S. angustifolium and Nostoc grown separately, S. angustifolium and Nostoc cross-fed with exometabolites from the other partner, and S. angustifolium and Nostoc grown together in direct contact (Fig. 2A). Given a priori knowledge of other plant–cyanobacterium symbiosis, we hypothesized that the S. angustifolium—N. muscorum UTEX 1037 symbiosis would result in the downregulation of N. muscorum UTEX 1037 photosynthesis, as carbohydrates are supplied by the plant host in exchange for N-rich metabolites from the N. muscorum UTEX 1037 partner via enhanced N2 fixation [34].
The results from the N. muscorum UTEX 1037 RNA-seq analysis targeting the N2 fixation pathway did not support this hypothesis (Fig. 6). Instead, genes involved in N2 fixation exhibited a decreasing trend, with the exception of glutamine synthetase (glnA; Fig. 6). In accordance with expectations, expression of N. muscorum UTEX 1037 genes involved in photosynthesis generally decreased for both photosystem I and II when the cyanobacteria were supplemented with S. angustifolium exometabolites, although expression of photosystem I genes tended in increase when N. muscorum UTEX 1037 was grown directly with S. angustifolium.
Likewise, S. angustifolium RNA-seq analysis with MapMan4 ontology enrichment did not support the hypothesis that photosynthesis genes would be increased during symbiosis. However, enrichment analysis identified downregulation of genes involved in S. angustifolium host defense when the host was in direct contact with the N. muscorum UTEX 1037 (Fig. 7). For example, genes associated with cysteine rich peptides (CRP) (−1.48 LFC, p = 0.006), glutathione S-transferase activities (GST) (−1.4 LFC, p = 0.00006), and jasmonic acid synthesis (JAS) (−1.7 LFC, p = 0.03) were expressed at lower levels in S. angustifolium grown in direct contact with N. muscorum UTEX 1037 (Table S5) than in S. angustifolium fed N. muscorum UTEX 1037 exometabolites (Table S6). Furthermore, the plant defense–related gene phenylalanine ammonia lyase (PAL) was induced (1.49 LFC, p = 0.00004) when S. angustifolium was fed N. muscorum UTEX 1037 exometabolites, but the expression of the gene did not change when the organisms were in in direct contact (Fig. 7).
Discussion
The overarching aim of this study was to provide an initial characterization of the physiological and metabolic reprogramming necessary for the Sphagnum–Nostoc symbiosis while testing the following hypotheses: 1) the Sphagnum–Nostoc symbiosis is dependent on the surrounding environmental context, particularly in regard to pH, and, 2) the dual endophytic and epiphytic nature of the Sphagnum–Nostoc symbiosis [22] results in unique host and bacterial physiology, metabolism, and metabolic exchange relative to other plant—cyanobacteria symbioses. The first hypothesis was based on field observations showing that raising the pH by addition of bicarbonate benefited the N2-fixing partners of the symbiosis over the Sphagnum host [11], leading the authors of that study to suggest that the cyanobacteria trade growth for protection under specific environmental conditions. The second hypothesis stems from the unique Sphagnum-Nostoc association that can take place epiphytically on leaflet and stem surfaces similar to the feathermoss-cyanobacteria symbiosis [18] or endophytically within Sphagnum host hyaline cells analogous to the enclosed cavities found in some liverwort and hornworts or the intracellular Gunnera system (reviewed in [16, 35]). Below, we address each hypothesis within the context of the current results and the broader plant–cyanobacteria literature.
The Sphagnum–Nostoc symbiosis is dependent on pH
In support of our first hypothesis, we found that pH was a critical factor in determining both the formation of the symbiosis and its outcome. We observed mutualism, in which both the S. angustifolium host and N. muscorum UTEX 1037 symbiont increased their growth, only at a low pH (3.5). As pH increased to 5.5 and to 8.5, N. muscorum UTEX 1037 growth continued to increase growth, whereas S. angustifolium growth declined steeply at high pH. This finding is supported by studies that have investigated S. angustifolium and cyanobacteria individually. For example, previous reports on Sphagnum growth to variable pH levels and other environmental factors [36] revealed that growth increased from a low pH of 3.5 to a peak around 5.5, followed by a sharp decline at pH 7.5. Indeed, high pH is associated with Sphagnum die-off [11, 37]; this is in direct contrast to our observation of cyanobacteria, for which the same high pH conditions were associated with peak growth. This trend is consistent with field observations showing that a pH of 8.1 was associated with the highest percentage of heterocystous cyanobacteria [38]. Taken together, these findings support the notion proposed by van den Elzen et al. [11] that the N2-fixing diazotrophic community and Sphagnum host have different niche preferences, and that at low pH and sub-optimal acidic conditions the diazotrophs trade growth in exchange for protection from predation [39].
The Sphagnum—Nostoc metabolic exchange includes amino acids, purine metabolism, and trehalose
Plant–cyanobacteria interactions are commonly characterized as nutrient exchange symbioses in which carbohydrates are produced by the plant in exchange for N-rich compounds produced by the cyanobacterial symbiont (e.g., [1, 2]). This general statement encompasses significant variation, and uncertainty remains regarding the identities of the molecules being exchanged and how this compares to other plant–cyanobacteria symbioses ranging from ephemeral epiphytic interactions (e.g., Pleurozium spp. [18]) to endophytic interactions such as those of Gunnera. Therefore, we sought to identify the metabolites being released by each partner of the symbiosis when grown in fresh liquid BG-110 medium, as well as the change in abundance of those exometabolites, using both a static cross-feeding approach and a MALDI-MSI approach in which symbiotic partners were able to interact (Fig. 2A).
Nostoc in symbiosis tends to decrease conversion of fixed N into amino acids through down-regulation of glnA (glutamine synthetase, GS) and gltB (glutamate synthase, GOGAT). This allows the release of fixed N as ammonium, 40–95% of which is then taken up by the plant host (reviewed in [40]). Cycad associations are an exception to this, as amino acids seem to be the main N currency [41], and the lack of change in glnA and gltB transcript levels in the epiphytic feathermoss–Nostoc association led Warshan et al. [18] to hypothesize that amino acids are also the main N currency for that system. In the current study, N. muscorum UTEX 1037 gltB expression was reduced only when the cyanobacteria were cultured in direct contact with S. angustifolium, but not when they were fed S. angustifolium exometabolites, suggesting that N. muscorum UTEX 1037 derived ammonium may act as a currency that can be exchanged only during direct symbiosis (Fig. 8). However, we must be circumspect regarding this possibility: the ammonium ion was too small for detection in the design of the current study, and our cross-feeding analyses clearly identified the transfer of the amino acids arginine, alanine, and asparagine from N. muscorum UTEX 1037 to the moss host, suggesting that the metabolic currency comprises multiple forms of N. Future this symbiosis supports both endophytic and epiphytic associations, thus studies should test the hypothesis that direct endophytic colonization favors inorganic ammonium as the main N currency, whereas epiphytic associations favor organic forms such as amino acids.
Blue boxes indicate the metabolite was confirmed with MALDI-MSI, red boxes indicate the metabolite was identified in the cross-feeding experiment, and red/blue boxes represent metabolites confirmed with both MALDI-MSI and cross-feeding. Black boxes represent generalized metabolic pathways that were not evidenced by MALDI-MSI or from the cross-feeding study but have some support via metatranscriptomics (Tables S5, S6). For simplicity, metabolic processes specific to Nostoc heterocyst (i.e., N2 fixation) and vegetative cells are not separated.
Cross-feeding and MALDI-MSI analysis also suggest multiple roles for purine metabolism within the S. angustifolium—N. muscorum UTEX 1037 symbiosis (Fig. 8). First, N. muscorum UTEX 1037 significantly increased release of adenine when cross-fed S. angustifolium exometabolites, whereas S. angustifolium depleted adenine when fed N. muscorum UTEX 1037 exometabolites. Purines, and in particular adenine, play key roles in the interactions between several nodule-forming rhizobacteria and their hosts. For example, the Sinorhizobium fredii purine metabolism mutant purL- or strains with reduced purL expression levels fail to produce viable nodules on soybean (Glycine max) [42, 43]. Nodulation is also inhibited on the Aeschynomene host in Bradyrhizobium purL mutants and mutants in other purine synthesis genes [44]. Although the mechanism is not entirely understood, it is interesting to note that modifications of purine biosynthesis also inhibit symbiosis with non-plant hosts, including the Burkholderia–stinkbug [45] and Photorhabdus–nematode [46] associations, suggesting a role for purines that extend well beyond moss–Nostoc interactions.
We also observed a role for purine metabolism within the S. angustifolium - N. muscorum UTEX 1037 symbiosis involving xanthosine (Fig. 8). Similar to adenine, the abundance of exuded xanthosine increased when N. muscorum UTEX 1037 was cross-fed S. angustifolium exometabolites, whereas S. angustifolium depleted xanthosine when fed N. muscorum UTEX 1037 exometabolites. Xanthosine is a catabolite of purine nucleotides and a key constituent of RNA. It also plays a well-known role in soybean–rhizobium symbiosis in which xanthosine forms the purine base xanthine, leading to the formation of uric acid via xanthine dehydrogenase. Uricase then converts uric acid to allantoin (reviewed in [47]), which along with allantoic acid is the major transport form of N through plant host xylem [48]. By contrast, in non–N2-fixing soybean plants, amino acids constitute the major form of N [49]. Furthermore, recent studies extended the role of allantoin to abiotic stress tolerance through the involvement of abscisic acid and jasmonic acid [50]. The role of purine metabolism in the S. angustifolium – N. muscorum UTEX 1037 symbiosis, and more specifically allantoin as a form of N currency and its involvement in the stress response, deserves further investigation.
In general during plant–cyanobacteria symbiosis, cyanobacterial photosynthesis is downregulated, and the reduced C-fixing capacity is compensated by carbohydrates supplied by the plant. The carbohydrates exchanged in plant–cyanobacteria symbioses is usually sucrose [8], but bryophytes tend to use trehalose as a major form of carbohydrate. Indeed, our earlier work [20] in conjunction with the findings reported here clearly identify a role for S. angustifolium produced trehalose as the main C currency during symbiosis (Fig. 8). Consistent with this, MALDI-MSI revealed that trehalose was the only disaccharide released from S. angustifolium. Trehalose is a non-reducing carbohydrate, it is relatively chemically inert and very stable at low pH; moreover, it has long been recognized as an important signaling molecule in vascular plant symbiosis. Numerous studies have documented its role as a cellular osmoprotectant [51,52,53,54] leading to stress tolerance [55, 56] and even enhanced N2 fixation [57]. In our experiments, the abundance of S. angustifolium-released trehalose increased in the presence of N. muscorum UTEX 1037. The exudation of trehalose may simply be the consequence of a metabolic byproduct, as trehalose is a major carbohydrate form in mosses. However, the stability of trehalose within low-pH environments, along with its contribution to stress tolerance at harsh high-latitude ecosystems, may have driven co-evolutionary events that made trehalose the major form of carbohydrate in the metabolic currency.
A possible role for sulfur metabolism in the Sphagnum–Nostoc symbiosis
Sulfur was recently suggested to play a role in the cyanobacteria–feather moss symbiosis [18]. To explore this notion in the S. angustifolium—N. muscorum UTEX 1037 system, we mined MALDI-MSI spectra for S-rich compounds. Choline-O-sulfate (Fig. 4D) and taurine (Fig. S2) were produced in large abundance by the moss host and depleted by N. muscorum UTEX 1037 (Fig. 8), e.g., 98% of choline-O-sulfate provided by S. angustifolium was depleted by N. muscorum UTEX 1037. Although choline-O-sulfate itself can serve as a compatible solute for osmotic stress regulation in bacteria and plants [58,59,60], some bacteria are able to metabolize this compound to yield sulfate for use as a sulfur source and choline that can further be transformed to glycine betaine, which can in turn be further metabolized to ammonia and pyruvate. Hence, it is not surprising that choline-like moieties are present in lipid head groups of bacteria occupying Sphagnum-dominated northern peatlands, suggesting that these compounds contribute to membrane stability under acidic conditions [17, 19, 61]. Similarly, taurine is considered a major C source for marine prokaryotes [62] and along with other organosulfur molecules is considered to be a key exchange molecule in a cosmopolitan marine diatom–bacteria symbiosis [63]. Such findings led Warshan et al. [18] to suggest that cyanobacteria in epiphytic associations do not obligately rely on the moss host for fixed C, but rather receive C as a reward from aliphatic sulfonate compounds such as taurine, as revealed in this current study. Cyanobacteria require considerable amounts of S to support the Fe–S clusters necessary for mature nitrogenase synthesis [16, 18]. Therefore, the provision of choline-O-sulfate and taurine by the moss host may benefit N. muscorum UTEX 1037 in multiple ways, including access to limiting nutrients for N2 fixation and amino acid synthesis, osmotic protection from low-pH bog conditions, and even by providing a C source.
Host defense response to cyanobacteria colonization
Our metatranscriptome analysis revealed that S. angustifolium host defense was downregulated when the plant was in direct contact with the N. muscorum UTEX 1037 symbiont, but not as a result of chemical contact alone (Fig. 7). This conclusion was supported by reductions in expression of genes associated with cysteine rich peptides, glutathione S-transferases, and jasmonic acid biosynthesis. Furthermore, phenylalanine ammonia lyase (PAL) was induced when S. angustifolium was fed N. muscorum UTEX 1037 exometabolites, but not when S. angustifolium was in direct contact with N. muscorum UTEX 1037. Multiple studies have shown that PAL expression responds to environmental stressors including pathogen infection, wounding, nutrient depletion and extreme temperature change [64, 65]. In Arabidopsis, the quadruple mutant pal1/pal2/pal3/pal4 is extremely susceptible to pathogenic bacteria, lower levels of salicylic acid, and reduced levels of lignin relative to wild-type plants [66]. Given the role of PAL in plant defense, it is not surprising that this enzyme would also play a role in symbiotic outcomes, as beneficial microbes must bypass host defense systems. In Lotus japonicus, for example, PAL expression is significantly induced 2 days after inoculation with Mesorhizobium loti inoculation, but dramatically suppressed after prolonged inoculation [67]. A comprehensive study by Chen et al. [68], demonstrated that PAL influences multiple processes in the Lotus japonicus–Rhizobium symbiosis, including lignin modification and salicylic acid signaling and biosynthesis. A note of caution must be placed on the current results as altered N status, and other abiotic factors could influence gene expression. Nonetheless, N. muscorum UTEX 1037 bypasses the S. angustifolium host defense system as evidenced by colonization, and how this happens remains to be determined.
Conclusion
These observations expand our knowledge of the environmental, metabolic, and physiological underpinnings of the S. angustifolium—N. muscorum UTEX 1037 mutualism. Our findings show that mutualism is dependent on environmental context, and in particular is driven by acidic pH. Moreover, we showed that trehalose is the main disaccharide provided by the S. angustifolium host in exchange for N-rich nucleosides and amino acids, and their derivatives, including cytidine, xanthosine, adenine, 2’-deoxyguanosine, uridine aspartate, acetyl-methionine, arginine, glutamic acid. Although this suggests that the N provided by N. muscorum UTEX 1037 is organic rather than ammonium, as is often the case in plant—cyanobacteria associations, we cannot draw a strong conclusion on this issue because neither the cross-feeding or MALDI-MSI approaches we used in this study were capable of identifying ammonium. Notably in this regard, however, our metatranscriptome analysis revealed downregulation of N. muscorum UTEX 1037 GS-GOGAT, which is associated with the release of fixed N as ammonium. This effect was observed only when the organisms were in direct contact and not cross-fed. Thus, the environmental context of the symbiosis (i.e., pH) may not only drive the type of symbiosis, from commensal epiphytic symbiosis to mutualistic endophytic symbiosis, but may also influence the form of N currency that is exchanged. In addition to identifying C- and N-rich exometabolites, we also clearly demonstrated exchange of exometabolites for host-provided S-rich molecules such as taurine, choline-O-sulfate, and sulfoacetate. How sulfur metabolism is involved in the C and N nutrient exchange remains to be fully elucidated. It is reasonable to hypothesize that cyanobacteria need S to support the synthesis of Fe-S clusters for nitrogenase, and also benefit from the protective effects of choline-O-sulfate in harsh, low-pH peat bogs. Our conceptual model (Fig. 8) details these newly described metabolic interactions along with key hypothesized interactions the testing of which will improve our understanding of plant–cyanobacterial interactions. We note that Sphagnum hyaline cells can be occupied by a number of interacting micro-organisms [10, 69, 70] and that the current conceptual model will need to include additional members to ultimately enhance predictions of C and N cycling in peatland ecosystems.
References
Limpricht KG. Die laubmoose. In: Rabenhorst L (ed). Kryptogamen-Flora von Deutschland, Oesterreich und der Schweiz, Zweite Auflage. 1890. Kummer, Leipzig.
Basilier K. Fixation and uptake of nitrogen in Sphagnum blue-green algal associations. Oikos. 1980;34:239.
Granhall U, Selander H. Nitrogen fixation in a subarctic mire. Oikos. 1973;24:8.
Basilier K, Granhall U, Stenström T-A. Nitrogen fixation in wet minerotrophic moss communities of a subarctic mire. Oikos. 1978;31:236.
Basilier K. Moss-associated nitrogen fixation in some mire and coniferous forest environments around Uppsala, Sweden. Lindbergia. 1979;5:84–88.
Meeks JC. Physiological adaptations in nitrogen-fixing Nostoc–plant symbiotic associations. In: Pawlowski K (ed). Prokaryotic symbionts in plants. 2007. Springer, Berlin, Heidelberg, pp 181–205.
Adams DG. Cyanobacteria in symbiosis with hornworts and liverworts. Cyanobacteria in symbiosis. 2002. Springer, Dordrecht, pp 117-35.
Meeks JC, Elhai J. Regulation of cellular differentiation in filamentous cyanobacteria in free-living and plant-associated symbiotic growth states. Microbiol Mol Biol Rev. 2002;66:94–121.
Kostka JE, Weston DJ, Glass JB, Lilleskov EA, Shaw AJ, Turetsky MR. The Sphagnum microbiome: new insights from an ancient plant lineage. N Phytol. 2016;211:57–64.
Granhall U, Hofsten AV. Nitrogenase activity in relation to intracellular organisms in Sphagnum mosses. Physiol Plant. 1976;36:88–94.
van den Elzen E, Kox MAR, Harpenslager SF, Hensgens G, Fritz C, Jetten MSM, et al. Symbiosis revisited: phosphorus and acid buffering stimulate N2 fixation but not Sphagnum growth. Biogeosciences. 2017;14:1111–22.
Yu Z, Loisel J, Brosseau DP, Beilman DW, Hunt SJ. Global peatland dynamics since the last glacial maximum. Geophys Res Lett. 2010;37:1–5.
Lindo Z, Nilsson MC, Gundale MJ. Bryophyte-cyanobacteria associations as regulators of the northern latitude carbon balance in response to global change. Glob Chang Biol. 2013;19:2022–35.
Carrell AA, Kolton M, Glass JB, Pelletier DA, Warren MJ, Kostka JE, et al. Experimental warming alters the community composition, diversity, and N2 fixation activity of peat moss (Sphagnum fallax) microbiomes. Glob Chang Biol. 2019;25:2993–3004.
Rai AN, Söderbäck E, Bergman B. Tansley Review No. 116. N Phytol. 2000;147:449–81.
Adams DG, Duggan PS. Cyanobacteria-bryophyte symbioses. J Exp Bot. 2008;59:1047–58.
Bay G, Nahar N, Oubre M, Whitehouse MJ, Wardle DA, Zackrisson O, et al. Boreal feather mosses secrete chemical signals to gain nitrogen. N Phytol. 2013;200:54–60.
Warshan D, Espinoza JL, Stuart RK, Richter RA, Kim S-Y, Shapiro N, et al. Feathermoss and epiphytic Nostoc cooperate differently: expanding the spectrum of plant–cyanobacteria symbiosis. ISME J. 2017;12:1–13.
Stuart RK, Pederson ERA, Weyman PD, Weber PK, Rassmussen U, Dupont CL. Bidirectional C and N transfer and a potential role for sulfur in an epiphytic diazotrophic mutualism. ISME J. 2020;14:3068–78.
Veličković D, Chu RK, Carrell AA, Thomas M, Paša-Tolić L, Weston DJ, et al. Multimodal MSI in conjunction with broad coverage spatially resolved MS2 increases confidence in both molecular identification and localization. Anal Chem. 2018;90:702–7.
Nagy G, Veličković D, Chu RK, Carrell AA, Weston DJ, Ibrahim YM, et al. Towards resolving the spatial metabolome with unambiguous molecular annotations in complex biological systems by coupling mass spectrometry imaging with structures for lossless ion manipulations. Chem Commun. 2019;55:306–9.
Warshan D, Liaimer A, Pederson E, Kim S-Y, Shapiro N, Woyke T, et al. Genomic changes associated with the evolutionary transitions of Nostoc to a plant symbiont. Mol Biol Evol. 2018;35:1160–75.
Rippka R, Deruelles J, Waterbury JB, Herdman M, Stanier RY. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol. 1979;111:1–61.
Hanson PJ, Riggs JS, Robert Nettles W, Phillips JR, Krassovski MB, Hook LA, et al. Attaining whole-ecosystem warming using air and deep-soil heating methods with an elevated CO2 atmosphere. Biogeosciences. 2017;14:861–83.
Frank W, Decker EL, Reski R. Molecular tools to study Physcomitrella patens. Plant Biol. 2005;7:220–7.
Yao Y, Sun T, Wang T, Ruebel O, Northen T, Bowen BP. Analysis of metabolomics datasets with high-performance computing and metabolite atlases. Metabolites. 2015;5:431–2.
Sumner LW, Amberg A, Barrett D, Beale MH, Beger R, Daykin CA, et al. Proposed minimum reporting standards for chemical analysis. Metabolomics. 2007;3:211–21.
Anders S, Pyl PT, Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19:455–77.
Gurevich A, Saveliev V, Vyahhi N, Tesler G. QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013;29:1072–5.
Seemann T. Genome analysis Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30:2068–9.
Finn RD, Clements J, Eddy SR. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 2011;39:W29–W37.
Schwacke R, Ponce-Soto GY, Krause K, Bolger AM, Arsova B, Hallab A, et al. MapMan4: a refined protein classification and annotation framework applicable to multi-omics data analysis. Mol Plant. 2019;12:879–92.
Black K, Osborne B. An assessment of photosynthetic downregulation in cyanobacteria from the Gunnera-Nostoc symbiosis. N Phytol. 2004;162:125–32.
Santi C, Bogusz D, Franche C. Biological nitrogen fixation in non-legume plants. Ann Bot. 2013;111:743–67.
Clymo RS. The growth of Sphagnum: some effects of environment. Br Ecol Soc. 1973;61:849–69.
Lamers LPM, Farhoush C, Van Groenendael JM, Roelofs JGM. Calcareous groundwater raises bogs; the concept of ombrotrophy revisited. J Ecol. 1999;87:639–48.
Nayak S, Prasanna R. Soil pH and its role in cyanobacterial abundance and diversity in rice field soils. Appl Ecol Environ Res. 2007;5:103–13.
Jassey VEJ, Meyer C, Dupuy C, Bernard N, Mitchell EAD, Toussaint ML, et al. To what extent do food preferences explain the trophic position of heterotrophic and mixotrophic microbial consumers in a Sphagnum peatland? Micro Ecol. 2013;66:571–80.
Meeks JC. Symbiosis between nitrogen- fixing cyanobacteria and plants. Symbiosis. 1998;48:266–76.
Pate S, Lindblad P, Atkins A. Planta in coralloid roots of cycad-Nostoc symbioses. Planta. 1988;176:461–71.
Xie B, Chen DS, Zhou K, Xie YQ, Li YG, Hu GY, et al. Symbiotic abilities of Sinorhizobium fredii with modified expression of purL. Appl Microbiol Biotechnol. 2006;71:505–14.
Xie B, Chen D, Cheng G, Ying Z, Xie F, Li Y, et al. Effects of the purl gene expression level on the competitive nodulation ability of Sinorhizobium fredii. Curr Microbiol. 2009;59:193–8.
Giraud E, Moulin L, Vallenet D, Barbe V, Cytryn E, Avarre JC, et al. Legumes symbioses: absence of Nod genes in photosynthetic bradyrhizobia. Science. 2007;316:1307–12.
Kim JK, Jang HA, Won YJ, Kikuchi Y, Heum Han S, Kim CH, et al. Purine biosynthesis-deficient Burkholderia mutants are incapable of symbiotic accommodation in the stinkbug. ISME J. 2014;8:552–63.
An R, Grewal PS. Molecular mechanisms of persistence of mutualistic bacteria Photorhabdus in the entomopathogenic nematode host. PLoS ONE. 2010;5:e13154.
Zrenner R, Stitt M, Sonnewald U, Boldt R. Pyrimidine and purine biosynthesis and degradation in plants. Annu Rev Plant Biol. 2006;57:805–36.
Atkins CA, Smith PMC. Translocation in legumes: assimilates, nutrients, and signaling molecules. Plant Physiol. 2007;144:550–61.
Ueda S, Ikeda M, Yamakawa T. Provision of carbon skeletons for amide synthesis in non-nodulated soybean and pea roots in response to the source of nitrogen supply. Soil Sci Plant Nutr. 2008;54:732–7.
Kaur H, Chowrasia S, Gaur VS, Mondal TK. Allantoin: emerging role in plant abiotic stress tolerance. Plant Mol Biol Rep. 2021;39:648–61.
Paul MJ, Primavesi LF, Jhurreea D, Zhang Y. Trehalose metabolism and signaling. Annu Rev Plant Biol. 2008;59:417–41.
Iturriaga G, Suárez R, Nova-Franco B. Trehalose metabolism: from osmoprotection to signaling. Int J Mol Sci. 2009;10:3793–810.
John R, Raja V, Ahmad M, Jan N, Majeed U, Ahmad S, et al. Trehalose: metabolism and role in stress signaling in plants. Stress signaling in plants: genomics and proteomics perspective, Volume 2. 2016. Springer International Publishing, pp 261–75.
Sharma K, Palatinszky M, Nikolov G, Berry D, Shank EA. Transparent soil microcosms for live-cell imaging and non-destructive stable isotope probing of soil microorganisms. Elife. 2020;9:1–28.
Streeter JG. Effect of trehalose on survival of Bradyrhizobium japonicum during desiccation. J Appl Microbiol. 2003;95:484–91.
Sugawara M, Cytryn EJ, Sadowsky MJ. Functional role of Bradyrhizobium japonicum trehalose biosynthesis and metabolism genes during physiological stress and nodulation. Appl Environ Microbiol. 2010;76:1071–81.
Suárez R, Wong A, Ramírez M, Barraza A, Orozco MDC, Cevallos MA, et al. Improvement of drought tolerance and grain yield in common bean by overexpressing trehalose-6-phosphate synthase in rhizobia. Mol Plant-Microbe Interact. 2008;21:958–66.
Mackay MA, Norton RS, Borowitzka LJ. Organic osmoregulatory solutes in cyanobacteria. J Gen Microbiol. 1984;130:2177–91.
Reed RH, Richardson DL, Warr SRC, Stewart WDP. Carbohydrate accumulation and osmotic stress in cyanobacteria. J Gen Microbiol. 1984;130:1–4.
Csonka LN. Physiological and genetic responses of bacteria to osmotic stress. Microbiol Rev. 1989,53:121–47.
Moore EK, Hopmans EC, Rijpstra WIC, Villanueva L, Dedysh SN, Kulichevskaya IS, et al. Novel mono-, di-, and trimethylornithine membrane lipids in northern wetland planctomycetes. Appl Environ Microbiol. 2013;79:6874–84.
Clifford EL, Varela MM, De Corte D, Bode A, Ortiz V, Herndl GJ, et al. Taurine is a major carbon and energy source for marine prokaryotes in the North Atlantic ocean off the Iberian Peninsula. Micro Ecol. 2019;78:299–312.
Amin SA, Hmelo LR, van Tol HM, Durham BP, Carlson LT, Heal KR, et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature. 2015;522:98–101.
Dixon RA, Paiva NL. Stress-induced phenylpropanoid metabolism. Plant Cell. 1995,7:1085–97.
Payyavula RS, Navarre DA, Kuhl JC, Pantoja A, Pillai SS. Differential effects of environment on potato phenylpropanoid and carotenoid expression. BMC Plant Biol. 2012;12:39.
Huang J, Gu M, Lai Z, Fan B, Shi K, Zhou YH, et al. Functional analysis of the Arabidopsis PAL gene family in plant growth, development, and response to environmental stress. Plant Physiol. 2010;153:1526–38.
Kouchi H. Large-scale analysis of gene expression profiles during early stages of root nodule formation in a model legume, Lotus japonicus. DNA Res. 2004;11:263–74.
Chen Y, Li F, Tian L, Huang M, Deng R, Li X, et al. The phenylalanine ammonia lyase gene LjPAL1 is involved in plant defense responses to pathogens and plays diverse roles in Lotus japonicus -rhizobium symbioses. Mol Plant-Microbe Interact. 2017;30:739–53.
Bragina A, Berg C, Cardinale M, Shcherbakov A, Chebotar V, Berg G. Sphagnum mosses harbour highly specific bacterial diversity during their whole lifecycle. ISME J. 2012;6:802–13.
Bragina A, Berg C, Müller H, Moser D, Berg G. Insights into functional bacterial diversity and its effects on Alpine bog ecosystem functioning. Sci Rep. 2013;3:1955.
Acknowledgements
We thank the editor and anonymous reviewers for detailed and thoughtful comments that improved the manuscript. Collection of starting Sphagnum angustifolium was made possible through the SPRUCE project, which is supported by Office of Science; Biological and Environmental Research (BER); US Department of Energy (DOE), Grant/Award Number: DE-AC05–00OR22725. Experimentation, sample collection, and analyses were supported by the DOE BER Early Career Research Program. Oak Ridge National Laboratory is managed by UT-Battelle, LLC, for the US DOE under contract no DE-AC05–00OR22725. AJS was supported by NSF DEB-1737899, 1928514. A portion of this research was performed under the Facilities Integrating Collaborations for User Science (FICUS) program and used resources at the DOE Joint Genome Institute and the Environmental Molecular Sciences Laboratory (grid.436923.9), which are DOE Office of Science User Facilities. Both facilities are sponsored by the Biological and Environmental Research program and operated under Contract Nos. DE-AC02-05CH11231 (JGI) and DE-AC05-76RL01830 (EMSL). Figs. 2 and 8 were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiment: AAC, DAP, DJW; Performed growth and crossfeeding experiments: AAC; Performed exometabolite crossfeeding profiling: BPB, KBL, TRN; Performed spatial metabolic profiling and analysis: DV, RKC, CRA; Performed metatranscriptomics sequencing and analysis: HDM, GO, LMM, SSJ; Developed plant genomic resources for metatranscriptomic analysis: JG, AJS, JS; Performed statistical analyses and data synthesis: AAC, DV, TJL, DLC,TRN, CRA, DAP, DJW; All authors prepared, edited and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no conflict of interests. This manuscript has been authored by UT-Battelle, LLC under Contract No. DE-AC05-00OR22725 with the U.S. Department of Energy. The United States Government retains and the publisher, by accepting the article for publication, acknowledges that the United States Government retains a non-exclusive, paid-up, irrevocable, world-wide license to publish or reproduce the published form of this manuscript, or allow others to do so, for United States Government purposes. The Department of Energy will provide public access to these results of federally sponsored research in accordance with the DOE Public Access Plan (http://energy.gov/downloads/doe-public-access-plan).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Carrell, A.A., Veličković, D., Lawrence, T.J. et al. Novel metabolic interactions and environmental conditions mediate the boreal peatmoss-cyanobacteria mutualism. ISME J 16, 1074–1085 (2022). https://doi.org/10.1038/s41396-021-01136-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01136-0
This article is cited by
-
Unraveling microbial processes involved in carbon and nitrogen cycling and greenhouse gas emissions in rewetted peatlands by molecular biology
Biogeochemistry (2024)
-
An overview of the roles of critical insiders and outsiders for reciprocal plant–microbe interaction: Heterotrimeric G-proteins, small RNAs, pollinators, microalgae
Symbiosis (2024)
-
The Feather Moss Hylocomium splendens Affects the Transcriptional Profile of a Symbiotic Cyanobacterium in Relation to Acquisition and Turnover of Key Nutrients
Microbial Ecology (2024)
-
Structure and Functions of Endophytic Bacterial Communities Associated with Sphagnum Mosses and Their Drivers in Two Different Nutrient Types of Peatlands
Microbial Ecology (2024)
-
Chemical Stimulation of Heterocyte Differentiation by the Feather Moss Hylocomium splendens: a Potential New Step in Plant-Cyanobacteria Symbioses
Microbial Ecology (2023)