Abstract
Moderate soil drying (MSD) is a promising agricultural technique that can reduce water consumption and enhance rhizosheath formation promoting drought resistance in plants. The endophytic fungus Piriformospora indica (P. indica) with high auxin production may be beneficial for rhizosheath formation. However, the integrated role of P. indica with native soil microbiome in rhizosheath formation is unclear. Here, we investigated the roles of P. indica and native bacteria on rice rhizosheath formation under MSD using high-throughput sequencing and rice mutants. Under MSD, rice rhizosheath formation was significantly increased by around 30% with P. indica inoculation. Auxins in rice roots and P. indica were responsible for the rhizosheath formation under MSD. Next, the abundance of the genus Bacillus, known as plant growth-promoting rhizobacteria, was enriched in the rice rhizosheath and root endosphere with P. indica inoculation under MSD. Moreover, the abundance of Bacillus cereus (B. cereus) with high auxin production was further increased by P. indica inoculation. After inoculation with both P. indica and B. cereus, rhizosheath formation in wild-type or auxin efflux carrier OsPIN2 complemented line rice was higher than that of the ospin2 mutant. Together, our results suggest that the interaction of the endophytic fungus P. indica with the native soil bacterium B. cereus favors rice rhizosheath formation by auxins modulation in rice and microbes under MSD. This finding reveals a cooperative contribution of P. indica and native microbiota in rice rhizosheath formation under moderate soil drying, which is important for improving water use in agriculture.
Similar content being viewed by others
Introduction
Rice (Oryza sativa L.) is a major staple food crop for almost half of the global population [1]. Rice cultivation requires large quantities of water input [2, 3]. For instance, to produce 1 kg of rice, 1000–1500 L of irrigation water is needed in the field [4]. Changes in climate have increased the frequency of extreme weather events in the world, such as reduced rainfall [5, 6]. Thus, in order to ensure food security for a rapidly growing human population, it is important to identify beneficial irrigation management techniques to improve crop water-use efficiency [7]. Moderate soil drying (MSD) irrigation is a promising approach for reducing water inputs and maintaining or increasing crop yields [4, 8].
The rhizosheath is commonly used to describe the soil that is around the roots and can be bound by root hairs; in contrast, the rhizosphere is identified as the narrow area of soil influenced by root activity [9,10,11]. The rhizosheath has been observed in many major cereal crops, such as rice, wheat, maize, barley, oat, rye, sorghum and pearl millet [12,13,14]. Compared with the rhizosphere, the rhizosheath is the soil related to root hair cylinder volume and within a 0–0.5 mm radius of rice roots; however, the rhizosphere (a few millimeters wide) is the soil less related to root hair [10, 15]. Many factors can affect rhizosheath formation, including root hair, the soil structure and the mucilage secreted by roots or microbes [10, 16]. For example, barley mutant plants without root hair showed little or no rhizosheath formation [17]. Our previous study also found that rhizosheath can be formed in rice under MSD, but not under continuous flooding [18]. Furthermore, wheat cultivars with larger rhizosheath showed greater drought tolerance than cultivars with smaller rhizosheath [19]. Barley rhizosheath was positively correlated with plant phosphorus accumulation under phosphorus-deficiency stress [17]. For nitrogen, the rhizosheath of 42 plant species had higher nitrogen content than in the bulk soil [20]. In addition, the rhizosheath is also a trait of plant adaptation to soil acidity [21, 22]. Rhizosheath formation is connected with MSD as a drought adaptation in plants [12, 18], and the rhizosheath should be thought of as a mechanism for improving plant tolerance and root protection under drought stress [23,24,25].
Piriformospora indica (P. indica, DSM11827), a cultivable root-colonizing endophytic fungus of the order Sebacinales, can affect plant root growth via auxin production [26,27,28]. P. indica colonizes the roots of a broad host range and promotes their growth [29, 30], enhances nutrient uptake [31,32,33] and improves the host resistance to various biotic and abiotic stresses [34, 35]. For example, P. indica can alleviate the severity of infections by Fusarium oxysporum, Verticillium dahlia and Pepino Mosaic Virus in tomato [36,37,38], and protect tomato against salt stress [39]. In addition, the hormone-mediated signaling pathways of maize can be activated by P. indica under drought stress [40]. Our previous study showed that rhizosheath formation induced by MSD was connected with the root auxin response [12]. Thus, we hypothesized that P. indica may contribute to rice rhizosheath formation under MSD by modulating auxins. Furthermore, recent studies have suggested that fungi can interact with bacteria via the exudation of C-rich compounds (such as sugars, organic acids and amino acids) and the redistribution of soil water under drought conditions [41,42,43,44]. However, it is unclear whether P. indica can interact with native soil bacteria in rhizosheath formation under MSD.
In the present study, we examined rhizosheath formation in different rice varieties and auxin transport mutants inoculated with P. indica under MSD. RNA sequencing (RNA-seq) analysis with or without P. indica inoculation was performed. We further identified the microorganisms associated with P. indica, isolated candidate strains from the Bacillaceae family and finally investigated their role in rhizosheath formation. We propose that the use of P. indica would complement crop rhizosheath formation strategies under soil drying conditions. Our findings provide some evidence that the endophytic fungus interacts with a native bacterium in the soil resulting in the enhancement of rice rhizosheath formation and the improvement of water use under soil drying.
Materials and methods
Plant materials and fungal culture conditions
Three rice (O. sativa) varieties, [‘Nipponbare’ (NIP), ‘Zhonghua11’ (ZH11) and ‘Zhonghan3’ (ZH3)] and OsPIN2 transgenic lines [O. sativa cv Hei-Jing2 (WT), OsPIN2 mutant (ospin2) and three complementation lines with the ospin2 background (C1, C2, and C3)] [45] were surface-sterilized using 1.5% (v/v) NaClO for 20 min, washed with double-distilled water three times, and grown for 3 d in 1/2 MS nutrient medium. Then, the seedlings were transformed to PNM nutrient medium, which contained five fungal discs with P. indica (P. indica; DSM11827; provided by Prof. Kaiwun Yeh from National Taiwan University). One to five rice plants were used per Petri dish and one fungal plague of 5 mm in diameter per rice plant was placed at a distance of 1 cm from the rice root [30]. Control treatments were conducted using media lacking P. indica.
Inoculation and root colonization with P. indica in rice
After 10 d co-cultivation with P. indica, the colonization of P. indica in rice roots was investigated using the marker gene Pitef1 [46] (F, 5’-GATTCCTTCATCGCAGTCA-3’; R, 5’-TTGGTGGTGTCCATCTTGTT-3’) by PCR (Supplementary Fig. S1A). The PCR was performed as follows: 95 °C for 3 min, 30 cycles of 95 °C for 30 s, 55 °C for 30 s, 72 °C for 45 s, 72 °C for 10 min, and ending at 4 °C. The colonization of P. indica in rice roots was also investigated using GFP-labeled P. indica. After 10 d, the roots were observed under a Zeiss LSM710NLO confocal laser scanning microscope at the 488 nm laser line of an argon multiline laser. The genomic DNA was extracted from the samples using the FastDNA Spin Kit for Soil (MP Biomedicals) according to the manufacturer’s instruction. Fifteen rice plants with or without P. indica inoculation were tested for the subsequent experiment.
Water treatments
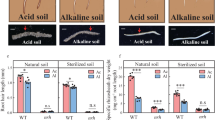
After evaluating the colonization of P. indica using PCR analysis, uniform seedlings with or without P. indica inoculation, were transplanted into pots (12-cm diameter, 14-cm height) containing 1.8 kg dry soil. The soil used in the study was collected from a paddy rice field in the town of Huayang, Jiangxi Province, China (115°09′E, 28°32′N). The air-dried soil with mineral nutrients added was sieved through a 4 mm mesh to remove any coarse material and vegetative matter. Soil chemical factors are shown in Supplementary Table S1. For the water treatments, half of the pots were treated under well-watered (WW) conditions, where a water layer of 3–5 cm above the soil surface was maintained, and the other half were subjected to an MSD regime. For the MSD and severe soil drying (SSD) treatments, rice seedlings were irrigated every 3 d to 80% and 60% (w/w) of field capacity, respectively (Supplementary Fig. S2A–C). The WW, MSD and MSD with auxin efflux inhibitor 1-naphtaylphthalamic acid (NPA) were kept for 2 weeks. The experiments were conducted in the greenhouse under a 14 h light (26 °C)/10 h dark (22 °C), 60% (w/w) relative humidity, and a photosynthetic photon flux density of 300 mmol photons m−2 s−1. Additional alkaline soil from Ronghuashan Town, Liaoning Province, China (122°86′E, 39°93′N) was used to repeat the above experiments to confirm the effect of the fungus and bacterium on rhizosheath formation under MSD.
Measurements of total root length, root hair length and rhizosheath weight
Under MSD, rice roots were carefully collected and shaken after the pots were disassembled. The soil, which tightly adhered to roots upon excavation, was identified as rhizosheath soil. The rhizosheath soil was recovered by washing the root gently in double-distilled water using plastic dishes [12, 14, 18]. The soil and rinsing water were then collected in a tray and dried at 105 °C for 3 d to measure the dry weight of the rhizosheath. An Epson Expression 1640 XL flat-bed scanner (Epson UK, London, UK) and WinRHIZO software (Regent Instruments, Quebec City, QC, Canada) were used to determine the total root length. The specific rhizosheath dry weight was calculated as the total rhizosheath soil dry weight per plant (mg) divided by the total root length (cm) [12, 18, 47]. Root hair length was measured following George et al. [17]. Photos were taken using a SMZ18 stereomicroscope and a DS-U3 camera (Nikon). Ten fully elongated root hairs were determined in each image using Image J software (National Institutes of Health). Four replicates for the rice plants were selected for the experimental measurement, and the experiments were repeated two times.
Assay of auxin concentration
For the measurement of auxin concentrations in the roots and P. indica, IAA was extracted from the P. indica inoculated roots and P. indica. Sample measurement of IAA was conducted by high-performance liquid chromatography (HPLC) according to Hou et al. [48]. Briefly, the fresh weight of the rice roots was determined, and the roots were frozen in liquid nitrogen. Then, the measurement of IAA was carried out using HPLC. The standard IAA sample was bought from Sigma-Aldrich (St Louis, MO, USA). For the measurement of IAA of B. cereus, the supernatant was obtained by passing through filter paper and then used to measure IAA by HPLC according to Yuan et al. [49].
Quantitative real-time PCR (qRT-PCR) and RNA-seq analysis
We harvested the roots to conduct RNA-seq and qRT-PCR analysis under MSD and WW with or without P. indica inoculation. Total RNA was isolated from the rice roots using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Primers for qRT-PCR are shown in Supplementary Table S2. The procedure was conducted according to Xu et al. [50]. For the RNA-seq analysis, RNA was qualified and quantified using a Nano Drop (Thermo Fisher Scientific) and BGISEQ-500 sequencer (BGI, Shenzhen, China). Sequencing libraries were generated according to the method of Zhang et al. [12]. Raw reads were analyzed and cleaned using SOAPnuk (v1.5.2). Gene expression that changed by more than 2-fold with an adjusted P ≤ 0.01 between root samples deemed as significantly different. The treatment groups included the uninoculated seedlings grown under WW (WW), seedlings inoculated with P. indica grown under WW (WW + P. indica), uninoculated seedlings grown under MSD (MSD), and seedlings inoculated with P. indica grown under MSD (MSD + P. indica). The RNA-seq results showed that over 1G of clean bases was obtained from each sample (Supplementary Table S3). In addition, over 97% of the clean reads could be aligned to the reference genome. Pathway enrichment analysis used Kyoto Encyclopedia of Genes and Genomes (KEGG) databases. The sequencing dataset was deposited to the NCBI BioProject database under the accession number PRJNA732534.
Sample preparation and DNA extraction for bacterial community analysis
In order to investigate the bacterial composition and diversity of the rhizosheath and root endosphere under MSD and WW conditions with P. indica inoculation, we collected rhizosheath, root endosphere, and bulk soil samples from the additional three pots in the same rhizosheath formation experiment (Supplementary Figs. S3 and S4A). The different sample types used in this study are as follows: for MSD treatment, a 0.5-mm radius of soil around the root was collected as the rhizosheath soil [18] (Supplementary Fig. S3). Under the WW treatment, we selected the soil around the roots with the same radius (0.5-mm) of rhizosheath under MSD to consider as rhizosheath-like soil under WW owing to that rice is unable to form a rhizosheath under WW conditions. Details of the sample collection methods are described below. Briefly, the root was firstly removed from the pot, following which the soil around the root within a 0.5-mm radius was collected using a Vernier caliper, and other soil around the root was then removed using sterile tweezers. PBS-S buffer (130 mM NaCl, 7 mM Na2HPO4, 3 mM NaH2PO4 (pH 7.0), and 0.02% [v/v] Silwet L-7) was used to wash the root with soil of both MSD and WW treatments in 50-mL tubes [51]. Then, the rhizosheath soil was collected from the washing buffer by centrifuging at 1500 × g for 20 min. Bulk soil was taken from the pot without plant treatments. After collection, soil samples were immediately frozen in liquid nitrogen and stored at −80 °C. The root samples were surface-sterilized using 1.5% (v/v) NaClO for 15 min after they were removed from PBS-S buffer. After sterilizing with NaClO, they were washed with sterilized double-distilled water three times. Thereafter, the final wash water was volume onto Luria-Bertani (LB) medium to verify the efficacy of the sterilization [52]. Root samples were also stored at −80 °C. Total root and soil genomic DNA was extracted from 0.5 g sample using the Mag-Bind Soil DNA Kit (Omega Bio-Tek) following the manufacturer’s instructions. The DNA quality and quantity were determined by gel electrophoresis and NanoDrop ONE spectrophotometry (Thermo Scientific, Waltham, MA, USA). Three biological repeats of each treatment were taken for 16S rRNA gene high-throughput sequencing (Supplementary Table S4).
16S rRNA gene high-throughput sequencing for bacterial community analysis
To determine the changes in the bacterial community composition of the rhizosheath and root endosphere under MSD and WW conditions, we amplified the V5–V7 region of the bacterial 16S rRNA gene using 799F/1193R primers [53]. These primers were selected because they were more suitable for 16S rRNA gene amplification sequencing for plant-associated bacterial communities [54, 55]. The PCR reactions were run in triplicate 50-μL mixtures containing 25 μL Phusion High-Fidelity PCR Master Mix with HF buffer (New England Biolabs), 3 μL of primers (10 μM), 10 μL template DNA, 6 μL double-distilled water, and 3 μL dimethyl sulfoxide. The PCR was performed as follows: 98 °C for 30 s; 25 cycles of 98 °C for 15 s, 58 °C for 15 s, and 72 °C for 15 s; and 72 °C for 60 s. The PCR products were extracted from 2% agarose gels, purified using an AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA), and quantified using Quanti-Fluor-ST (Promega, Madison, WI, USA) according to the manufacturers’ protocols. Then, samples were sequenced on a MiSeq platform (Illumina, San Diego, CA, USA) at Majorbio Bio-Pharm Technology Co. Ltd (Shanghai, China).
The raw 16S rRNA gene sequencing reads were quality-filtered by fastp version 0.20.0 [56]. Paired-end reads were processed in the following steps using USEARCH version 7.0 [57]: joining of paired-end reads, removal of barcodes and primers, filtering of low-quality reads, and finding non-redundant reads. For root endosphere samples, mitochondria- and chloroplast-assigned operational taxonomic units (OTUs) were removed. The OTUs were clustered with a 97% similarity cutoff [58] using UPARSE (version 11.0) [58]. The taxonomy of each 16S rRNA gene sequence was analyzed using the RDP Classifier algorithm [59] against the SILVA (v138) [60] 16S rRNA gene database with a confidence threshold of 70%. The alpha-diversity (observed OTUs, Chao1, Shannon and Good’s coverage) and beta-diversity analyses were performed by Mothur software [61] and the R vegan package (version 4.0.4). The rarefaction curves (based on OTUs) were calculated to access the sampling efficiency of samples. Liner discriminant analysis coupled with an effect size (LEfSe) analysis was conducted to search for significantly different (p < 0.05) taxa among the bulk soil, root endosphere and rhizosheath under MSD and MSD with P. indica inoculation. Ternary plots were conducted using the mean of the log2 transformed relative abundance values per compartment. The function “ternaryplot” was used to plot the point size of each OTU proportional to its weighted sum instead of the total relative abundance [51]. Venn diagrams were created to show the number of common or unique OTUs in multi groups using the R package Venn Diagram setting. All sequencing data were available in the NCBI Sequence Read Archive (SRA) database under accession number PRJNA739870.
Culturable Bacillus assay, Bacillus isolation and interaction assay
We measured the population densities of culturable Bacillus in the fresh bulk soil, rhizosheath and root endosphere samples, which was conducted using a standard 10-fold dilution plating assay following Wang et al. [62]. For quantification of the Bacillus population, three aliquots of dilution were pretreated at 80 °C for 10 min, and then plated on V8 Bacillus-semiselective medium (326 mL V8 juice, 33 g NaCl, 0.8 g glucose, 45 mg actidione and 22.5 mg polymyxin B per liter; the actidione and polymyxin B were filter-sterilized. In addition, the pH of the medium was adjusted to 5.2 before autoclaving) [63]. After incubation at 30 °C for 2 d, colony counting was performed to calculate the number of colony forming units (CFUs).
Then, the fresh rhizosheath soil or rice roots under MSD with P. indica inoculation were used for the isolation of putative Bacillus. The soil or root suspension was serially diluted and pretreated at 80 °C for 10 min, and plated on V8 Bacillus semiselective medium [55, 64]. After incubation at 30 °C for 2 d, the Bacillus colonies were randomly isolated from the selective plates based on colony morphology. A total of 239 candidate Bacillus strains were isolated with P. indica inoculation under MSD, out of which 129 Bacillus strains were from the rhizosheath and 110 Bacillus strains were from the root endosphere.
The interaction between P. indica and Bacillus was tested as described by Zhang et al. [65]. Candidate Bacillus strains were inoculated in 5 mL of V8 Bacillus-semiselective medium and incubated overnight at 30 °C and 200 rpm shaking. The OD of the bacterial cultures was adjusted to an OD600 of 0.1. Chlamydospores of P. indica were harvested from KM medium using tween water (0.002% Tween 20), scratched with a scalpel, and filtered through Miracloth (Calbiochem, Nottingham, UK) to remove mycelial fragments [66]. Then, the chlamydospore solution was centrifuged in a falcon tube at 3500 rpm for 7 min, and the supernatant was decanted to resuspend the chlamydospore containing pellet with 0.9% NaCl solution, which was sterilized in an autoclave. Next, 2.5 μL P. indica chlamydospore (containing 104 P. indica chlamydospores per μL) was inoculated in a diagonal row on the right side of a Petri dish. After 3 d, 2.5 μL of Bacillus was inoculated on the left side of the Petri dish with a bacterial inoculating loop, creating a V-shape of the inoculation sites with increasing proximity [67]. Plates with P. indica non-inoculation were used as controls. The plates were incubated at 30 °C for 3–5 d. We determined the diameters of candidate Bacillus strains on a line orthogonal to the line dividing the V-shape to select the strain interacting with P. indica. Then, we calculated the relative colony diameter of strains as the ratio of the strain diameter with P. indica inoculation versus the strain diameter under control conditions. Following the assessment of the interaction between P. indica and the entire Bacillus candidates, one strain (Bacillus cereus) isolated from the root endosphere displaying a strong interactive effect was selected for further study. The classification of the interactive isolate was determined by sequencing the 16S rRNA gene using primers 27F and 1492R in Sangon Biotech (Shanghai) Co., Ltd (Sequence accession: 2020023554). The sequencing data have been made available in the NCBI GenBank (MZ433237).
Growth assay of P. indica and B. cereus
Sixty grams of sterilized soil, which was sterilized three times by autoclaving and heat-incubation until completely dehydrated [68], was placed in a 300 mL tissue-culture container and inoculated with 0.4 g of P. indica mycelium [69] and 2 mL of B. cereus suspension at OD600 = 0.5 (~1.5 × 106 CFU ml−1). The B. cereus strain was suspended in 10 mM MgCl2 buffer [70]. The assay only addition of B. cereus without P. indica inoculation was performed as a control. The number of B. cereus cells in the soil was evaluated at 15 d after inoculation. For the quantification of the Bacillus population, three aliquots of dilution were pretreated at 80 °C for 10 min and then plated on the V8 Bacillus-semiselective medium. After incubation at 30 °C for 2 d, colony counting was used to calculate the abundance of B. cereus in the soil.
Rhizosheath formation assay with P. indica and B. cereus inoculation
The experiment was conducted in sterilized or non-sterilized soil under MSD. The sterilized soil was sterilized three times by autoclaving and heat-incubation until completely dehydrated [68]. After the rice plants inoculated or not inoculated with P. indica were transplanted into sterilized or non-sterilized soil as described above, half of the seedlings were inoculated with B. cereus suspension to a concentration of 108 cells g−1 soil, while the uninoculated seedlings were water-treated with sterilized double-distilled water [71]. After 7 d of inoculation, the roots of some seedlings were subjected to MSD treatment for 2 weeks, as described above (Supplementary Fig. S4B). The root length, root hair length, and rhizosheath weight were then measured as described above. Four replicates of the rice plants were selected for each experimental measurement, and experiments were repeated two times.
Statistical analysis
Graphical representations were generated with GraphPad Prism 7 (GraphPad Software, Inc., La Jolla, CA, USA). The means and standard error (SE) of the data were calculated. The overall mean differences between treatments were analyzed using ANOVA (Duncan’s multiple range test) or Student’s t tests, and p values less than 0.05 were considered statistically significant. For principal co-ordinates analysis (PCoA), PERMANOVA (Adonis function, 999 permutations) was used to analyze the bacterial community composition structure on the basis of the weighted-UniFrac distance. The Kruskal–Wallis (KW) sum-rank test was carried out to detect the features displaying significantly different abundances between assigned families in linear discriminant analysis coupled with an effect size (LEfSe) analysis.
Results
Moderate soil drying increased rice rhizosheath formation
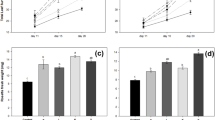
Compared with WW, MSD increased the root dry weight and rhizosheath formation of wild-type NIP rice, while SSD decreased it (Supplementary Fig. S2A–C). Moreover, the exopolysaccharide (EPS) content of rhizosheath soil was significantly increased by 38% under MSD, compared with WW (Supplementary Fig. S2D). The sugars and amino acids of the root exudation were also significantly higher under MSD than that under WW (Supplementary Fig. S2E). Compared with WW, ethylene related genes expressions were also significantly enhanced under MSD (Supplementary Fig. S2F).
The endophytic fungus (P. indica) further increased rice rhizosheath formation under moderate soil drying
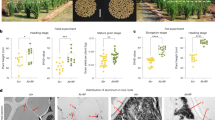
To investigate the effects of P. indica on rice rhizosheath formation, rice plants with and without P. indica inoculation were grown under WW and MSD conditions. We found that Pitef1 was amplified in the PCR-reaction from the root samples inoculated with P. indica (Supplementary Fig. S1A). Furthermore, P. indica-chlamydospores, exhibiting green fluorescence, had formed after colonization (Fig. 1G, H). Then, the total root length and root hair lengths of NIP, ZH11 and ZH3 were increased by 9.6–21% and 51–108% under WW with P. indica inoculation, compared to WW alone (Supplementary Fig. S1B, D). Compared to MSD alone, the total root lengths and root hair length in NIP, ZH11, and ZH3 were 6–32% and 53–92% greater under MSD with P. indica inoculation, respectively (Supplementary Fig. S1C, E). Rice seedlings were able to form rhizosheath under MSD and MSD with P. indica inoculation (MSD + P. indica), but not under WW, or WW with P. indica inoculation (WW + P. indica) (Fig. 1A–D). In addition, the total rhizosheath soil dry weight of NIP, ZH11 and ZH3 increased by ~38–86% under MSD with P. indica inoculation compared to MSD alone (Fig. 1E). A 31–37% increase in the specific rhizosheath soil dry weight was observed in NIP, ZH11, and ZH3 plants grown under MSD with P. indica inoculation compared to MSD alone (Fig. 1F). Moreover, the tiller number, total shoot dry weight, total root dry weight and total root length of rice inoculated with P. indica were significantly higher than those of the non-inoculated rice under field conditions (Supplementary Fig. S5A–D).
The non-inoculated and P. indica-inoculated rice seedlings (NIP rice) were cultured under well-watered (WW, A, B) and MSD (C, D) conditions. Rice seedlings (NIP rice) were able to form rhizosheath under MSD or MSD with P. indica-inoculation (MSD + P. indica), but not under WW or WW with P. indica-inoculation (WW + P. indica). Bar = 0.5 cm. The total rhizosheath soil dry weight (E) and specific rhizosheath soil dry weight (F) of three rice varieties (NIP, ZH11 and ZH3) under WW, WW + P. indica, MSD and MSD + P. indica. The specific rhizosheath dry weight was calculated as the total rhizosheath soil dry weight per plant (mg) divided by the total root length (cm). N.D. indicated that rhizosheath was not detectable. Data are the means ± SE (n = 8 replicates). Bars with different letters among different treatments are significantly different at p < 0.05 (ANOVA, Duncan’s multiple range test). G, H Fluorescence microscopy of rice seedling (NIP rice) roots showing the presence of intracellular P. indica chlamydospores (spores are in green) in cortical cells using confocal microscopy with a superficial view. The image is an X- and Y-stack reconstruction. Rice roots without GFP-tagged P. indica (P. indica–GFP) inoculation were used as a control. Bar = 150 μm.
Rice root auxin transport was involved in increasing rice rhizosheath formation with P. indica inoculation under moderate soil drying
P. indica can affect plant growth by producing auxin [26]. Here, the IAA concentration (auxin) of the P. indica mycelia was about 270 ng g−1 (Fig. 2A-I). And, the IAA concentration of the rice roots was increased by ~119% with P. indica inoculation compared to the uninoculated plants (Fig. 2A-II). To better understand the possible physiological differences resulting from P. indica inoculation under WW and MSD, we performed transcriptomic comparisons of the seedlings grown under WW and MSD with or without P. indica inoculation. When comparing the MSD samples with the WW samples, there were 1623 significantly up-regulated genes and 1092 significantly down-regulated genes (Supplementary Fig. S6A). There were 3785 differentially expressed genes when comparing MSD and MSD with P. indica (Supplementary Fig. S6A). Moreover, KEGG enrichment analysis showed that the plant hormone signal transduction pathway (including the auxin signaling pathway) was enriched under MSD + P. indica compared with MSD alone (Supplementary Fig. S6B). And, previous study showed that P. indica increased auxin signaling in the root of Chinese cabbage [30]. Thus, auxin related differentially expressed genes were further analyzed (Fig. 2B). The gene expression of auxin efflux carrier component (OsPIN2, Os06g0660200) was enhanced under MSD with P. indica inoculation. Compared to MSD, qRT-PCR revealed that the gene expression of OsPIN2 (an important gene for auxin efflux) in rice root was increased by 2.2-, 2.3- and 3.9-fold with P. indica inoculation, B. cereus inoculation, or a combination of P. indica and B. cereus inoculation under MSD, respectively (Supplementary Fig. S6C).
A IAA concentration of P. indica mycelia (I) or rice roots (II) without P. indica (−PI) and with P. indica (+PI) under axenic conditions. Data are the means ± SE (n = 8 replicates). Significant difference between with (+PI) and without P. indica (−PI) inoculation under WW or MSD were analyzed in figures (D, E) using the Student’s t test (*p < 0.05, **p < 0.01, ***p < 0.001). B Differentially expressed genes related to auxin in rice roots under WW, WW with P. indica inoculation (WW + P. indica), MSD, and MSD with P. indica inoculation (MSD + P. indica). C, D The effect of NPA (an auxin efflux inhibitor) on rice rhizosheath formation with or without P. indica inoculation. The total rhizosheath soil dry weight (C) and specific rhizosheath soil dry weight (D) of NIP rice seedlings with non-inoculation (−PI), NPA (+NPA), P. indica inoculation (+PI), P. indica and NPA treatments (+PI + NPA) under MSD. Data are the means ± SE (n = 8 replicates). Bars with different letters are significantly among different treatments at p < 0.05 (ANOVA, Duncan’s multiple range test). E Auxin efflux carrier OsPIN2 is required for rice rhizosheath formation with or without P. indica inoculation under MSD. The specific rhizosheath soil dry weight of rice seedlings (WT, auxin efflux carrier OsPIN2 mutant ospin2 and its complementation lines) with non-inoculation (−PI) and P. indica inoculation (+PI), under MSD. Data are the means ± SE (n = 8 replicates). Bars with different letters are significantly among different treatments at p < 0.05 (ANOVA, Duncan’s multiple range test).
Under MSD with the addition of auxin efflux inhibitor 1-naphtaylphthalamic acid (NPA), the total rhizosheath soil dry weight, specific rhizosheath soil dry weight and root hair length were significantly lower than those under MSD alone (Fig. 2B, C, Supplementary Fig. S7A). Similarly, the total rhizosheath soil dry weight, specific rhizosheath soil dry weight and root hair length under MSD with inoculation of P. indica and NPA were also decreased compared to the treatment of MSD with inoculation of P. indica (Fig. 2B, C, Supplementary Fig. S7A).
Auxin efflux carrier OsPIN2 was required for rhizosheath formation with P. indica inoculation under moderate soil drying
To further understand whether auxin efflux was involved in rhizosheath formation under MSD with P. indica inoculation, we examined rhizosheath formation with P. indica inoculation under MSD using the auxin efflux carrier OsPIN2 mutant ospin2, complemented OsPIN2 lines and WT rice. Compared to MSD alone, the specific rhizosheath soil dry weight was increased by 21–25% in WT and complemented lines with P. indica inoculation under MSD, while that of ospin2 was only increased by ~9% (Fig. 2E). Furthermore, MSD with P. indica inoculation increased root hair length of 20–33% in the WT and complemented rice lines, compared to MSD alone (Supplementary Fig. S7B). However, no significant difference in root hair length was recorded in ospin2 between MSD and MSD with P. indica inoculation (Supplementary Fig. S7B).
P. indica inoculation increased the abundance of Bacillus in the rhizosheath and root endosphere under moderate soil drying
To investigate whether the P. indica affected rhizosheath or root endosphere microbiota, 16S rRNA gene amplicon sequencing was conducted to analyze the composition of bacterial community in rhizosphere, rhizosheath, root endosphere and bulk soil under MSD and WW (Supplementary Fig. S8D). The gradually flattening rarefaction curves and over 99.1% Good’s coverage showed that the sequencing depth was adequate for capturing the microbial communities (Supplementary Fig. S8E; Supplementary Table S5). Principal coordinate analysis (PCoA) revealed that P. indica inoculation had significant effects on the composition of bacterial communities (based on weighted-UniFrac distance) in the root endosphere under MSD and WW conditions, however, no significant effects on the rhizosheath microbiome (Fig. 3C, D). Venn diagrams showed that P. indica inoculation impacted the bacterial OTUs number in root endosphere under WW and MSD condition (Supplementary Fig. S8A–C). The alpha-diversity (based on Chao and Shannon index) of the endosphere microbiomes significantly increased with P. indica inoculation compared to non-inoculation under MSD conditions, but no significant difference in rhizosheath microbiomes (Supplementary Fig. S8F, G). Moreover, the influence of P. indica on bacterial composition of root endosphere was different between MSD and WW conditions (Fig. 3A, B). For example, the relative abundance of the phylum Firmicutes significantly increased (from 8.3% to 41.6%) in the root endosphere inoculated with P. indica, whereas that of phylum Proteobacteria in the root endosphere (from 90.8% to 51.3%) significantly decreased under MSD (Fig. 3A). These phylum taxonomies changes were mainly due to the increased abundance of the families Bacillaceae, Sporolactobacillaceae and Ruminococcaceae or the decreased abundance of the families Burkholderiaceae, Mitochondria, Enterobacteriaceae and Xanthomonadaceae (Fig. 3B). Under WW, a considerable decrease in Burkholderiaceae (from 64.2% to 19.4%) and a significant increase in Mitochondria (from 13.5% to 58.1%) were observed in the root endosphere with P. indica inoculation compared with the non-inoculation treatment (Fig. 3B). Together, these results showed that P. indica played an important role in the assembly of the bacterial community of the root endosphere under MSD and WW.
A, B The phylum level and family level relative abundance showed >1% relative abundance of all sequences in samples from different root-system compartments (root endosphere and rhizosheath or rhizosphere) of NIP rice with non-inoculation (−PI), P. indica inoculation (+PI) under well-watered (WW) and MSD conditions. C, D Principal-coordinate analysis (PCoA, based on weighted-UniFrac distance) showed the changes in composition of bacterial microbiomes in the bulk soil, rhizosheath (under MSD)/rhizosphere (under WW) and root endosphere of non-inoculated or P. indica inoculated under WW (C) or MSD (D). PERMANOVA (Adonis function, 999 permutations) was used to test differences in the bacterial community composition between treatments in principal co-ordinates analysis (PCoA). Linear discriminant analysis (LDA) for discriminating bacterial taxon (family level) that showed significant differences in abundances among bulk soil, rhizosheath with non-inoculation (−PI), and rhizosheath with P. indica inoculation (+PI) (E; LDA score ≥ 3.2) and among bulk soil, root endosphere with non-inoculation (−PI) and root endosphere with P. indica inoculation (+PI) (F; LDA score ≥ 3.5) under MSD conditions. Significance test p < 0.05. WBS, bulk soil under WW; WRP, rhizosphere soil of NIP rice under WW with non-inoculation; WRP-PI, rhizosphere soil of NIP rice under WW with P. indica inoculation; WRE, root endosphere of NIP rice under WW with non-inoculation; WRE-PI, root endosphere of NIP rice under WW with P. indica inoculation; DBS, bulk soil under MSD; DRH, rhizosheath soil of NIP rice under MSD with non-inoculation; DRH-PI, rhizosheath soil of NIP rice under MSD with P. indica inoculation; DRE, root endosphere of NIP rice under MSD with non-inoculation; DRE-PI, root endosphere of NIP rice under MSD with P. indica inoculation.
P. indica inoculation significantly increased the abundance of the family Bacillaceae, belonging to phylum Firmicutes, in the rhizosheath (from 2.7% to 4.1%) and root endosphere (from 2.9% to 14%) compared to the non-inoculation treatment under MSD (Fig. 3B). We also found that the family Bacillaceae was strongly enriched in the rhizosheath and root endosphere under MSD using the linear discriminant analysis (LDA) effect size tool (LEfSe) analysis (Fig. 3E, F, Supplementary Fig. S8H). Furthermore, the abundance of genus Bacillus (belonging to family Bacillaceae) was 1.5 and 20 times higher in the rhizosheath and root endosphere with P. indica inoculation compared to the non-inoculation treatment under MSD, respectively (Fig. 4A–C). However, there was no significant difference in the relative abundance of Bacillus in the rhizosphere and root endosphere with and without P. indica inoculation under WW (Supplementary Fig. S9A, B). P. indica also significantly increased the abundance of genus Bacillus in root endosphere and rhizosheath using colony counting (Supplementary Fig. S10A). Together, these results showed that P. indica inoculation increased the abundance of Bacillus in the rhizosheath and root endosphere under MSD.
Ternary plot of the bacterial operational taxonomic units (OTUs) in the rhizosheath (A) and root endosphere (B) with non-inoculation (-PI), P. indica inoculation (+PI) and bulk soil under moderate soil drying (MSD). Each circle with color represents different OTUs (relative abundance > 1%), and Bacillus is indicated with red arrows. The size of each circle represents its relative abundance. The position of each circle is determined by the contribution of the OTUs from each of the treatments to the total relative abundance. C Bar plot showing the relative abundance of the reads classified as Bacillus (OTU1256) in the rhizosheath and root endosphere with non-inoculation (−PI), P. indica inoculation (+PI) and bulk soil under MSD. Data are the means ± SE. Bars with different letters are significantly different among different treatments at p < 0.05 (ANOVA, Duncan’s multiple range test). D IAA concentration produced by Bacillus cereus. –BC, medium without Bacillus cereus; +BC, medium with Bacillus cereus. Data are the means ± SE (n = 8 replicates). The asterisk (***) is shown significantly different between without and with Bacillus cereus at p < 0.001 (Student’s t test).
P. indica and B. cereus greatly promoted rice rhizosheath formation under moderate soil drying
To further explore which Bacillus species co-works with P. indica, 239 candidate Bacillus isolates were randomly isolated based on semiselective medium. After that, we found that B. cereus showed the highest relative colony diameter among the candidate isolates in the contact-assay (Supplementary Fig. S10B; Supplementary Table S6). Nextly, P. indica also significantly enhanced the relative diameter of B. cereus in volatile organic compounds interaction assay (Supplementary Fig. S10C). Further, the abundance of B. cereus was significantly increased with P. indica inoculation in sterilized soil, compared with non-inoculation conditions (Supplementary Fig. S10D). Together, the results showed that P. indica enhanced the abundance of B. cereus.
The rice plants with P. indica and B. cereus co-inoculation showed the highest root IAA concentration compared to P. indica inoculation or B. cereus inoculation alone (Supplementary Fig. S11A). To assess the promotional effect of P. indica and B. cereus on rhizosheath formation, WT, ospin2 and the complemented line rice, which were inoculated with P. indica, B. cereus, a combination of P. indica and B. cereus, or the uninoculated control, were treated under MSD using non-sterilized or sterilized soil (Supplementary Fig. S4B). Compared to the non-inoculation treatment, the rhizosheath formation of the WT and complemented line was significantly increased with P. indica inoculation, B. cereus inoculation and co-inoculation of P. indica and B. cereus under MSD using non-sterilized and sterilized soil, while the rhizosheath formation of ospin2 was not significantly different (Fig. 5). In addition, the rhizosheath of the WT and complemented line rice treated with the combination of P. indica and B. cereus was significantly higher than the treatment with P. indica or B. cereus separately. However, no significant difference in the rhizosheath formation was found in the ospin2 mutant with identical treatments (Fig. 5). Furthermore, inoculation of both P. indica and B. cereus also significantly promoted rice rhizosheath formation under MSD using the additional alkaline soil (Ronghuashan Town, Liaoning Province, 122°86′E, 39°93′N) (Supplementary Fig. S11B).
The specific rhizosheath soil dry weight of rice seedlings (WT, auxin efflux carrier OsPIN2 mutant ospin2 and complementation 1) with non-inoculation (Control), P. indica inoculation (+PI), Bacillus cereus inoculation (+BC), or P. indica and Bacillus cereus co-inoculation (+PI+BC) in non-sterilized soil (A) or sterilized soil (B) under MSD. Data are the means ± SE (n = 8 replicates). Bars with different letters are significantly different at p < 0.05 (ANOVA, Duncan’s multiple range test).
Discussion
P. indica can enhance rice rhizosheath formation under moderate soil drying
Rice can form rhizosheath under MSD, but not under WW [18], which is different from dryland-farmed crops. Rhizosheath formation under MSD was significantly higher than that under severe soil drying (SSD), which suggested that MSD was better for rice rhizosheath formation (Supplementary Fig. S2C). Thus, in this study, we investigated the effect of P. indica on rice rhizosheath formation and identified the factors governing rice rhizosheath formation under MSD. Furthermore, the quantity and quality of root exudates can be affected by drought [72]. Carbon in root exudates can provide energy for soil microbiota [14]. In the present study, the sugars and amino acids of the root exudates, which were related to rhizosheath formation [24], were increased under MSD, compared to WW (Supplementary Fig. S2E). Exopolysaccharide (EPS) produced by bacteria significantly enhanced the root-adhering soil per root tissue [73]. Sunflower inoculation of an EPS-producing strain significantly increased root-adhering soil per root mass ratio under drought conditions [74, 75]. Here, the content of rhizosheath EPS under MSD was significantly higher than that of WW (Supplementary Fig. S2D). The results suggest that moderate soil drying can increase rhizosheath formation by regulating root exudation quality and soil EPS extraction.
A previous study suggests that the hyphae of arbuscular mycorrhizal fungi show a minor involvement in rhizosheath formation [76]. P. indica, which can produce auxin and be involved in root development [26, 30], significantly increased rice rhizosheath formation in two different soil types (acid soil or alkaline soil) under MSD (Fig. 1F, Supplementary Fig. S11B). The probable reason for this finding is that P. indica can increase the rice root hair growth to search for water under MSD via auxins modulation, which also provides an important physical framework for rice rhizosheath formation [10, 17, 77].
Auxins are important for the P. indica-promoted rhizosheath formation in rice under moderate soil drying
P. indica-induced stimulation of root architecture changes is involved with auxin metabolism or signaling in roots [26, 30]. Rice root IAA concentration was enhanced after P. indica-inoculation (Fig. 2A, Supplementary Fig. S11A), which is consistent with results in Chinese cabbage [29] and barley [78]. Further, the rice roots co-inoculated with P. indica and B. cereus showed the highest IAA concentration (Supplementary Fig. S11A), which suggests that auxins can be produced by the rice plant, fungi and bacteria after inoculation. Auxin plays an important role in root growth for searching of water under MSD [12, 79]. The relative expression of the auxin efflux carrier gene OsPIN2 in the rice roots was significantly increased with P. indica inoculation under MSD, compared to MSD alone (Fig. 2B, Supplementary Fig. S6C), suggesting that auxins are important for rhizosheath formation. Moreover, ethylene related genes were also significantly increased under MSD (Supplementary Fig. S2F), which is consistent with our previous study [18]. In this study, we focused on auxins and rhizosheath formation with P. indica inoculation under MSD. Plants tend to rearrange their root architecture in search of water by mediating root auxin under drought conditions [80]. Compared to WW, the relative expression of some auxin related genes under MSD was also significantly increased (Fig. 2B).
When auxin transport was inhibited by NPA (an auxin efflux inhibitor), the total rhizosheath soil dry weight and specific rhizosheath soil dry weight was significantly decreased (Fig. 2C, D). The probable reason for this decrease was the low root hair length under NPA treatment (Supplementary Fig. S7B), which is similar to the findings in our previous study [12]. Numerous studies have shown that PIN2 is mainly asymmetrically localized at the upper side in the epidermis of the root meristem and elongation zones [81, 82], which in turn results in the suppression of root hair growth. When rice plants (WT and complementation rice) were inoculated with P. indica, rhizosheath formation significantly increased, but not in ospin2 because auxins are not sensed (Fig. 5). The results suggested that auxins are important for P. indica-promoted rhizosheath formation in rice under MSD.
The fungus P. indica can co-work with the native soil bacterium Bacillus cereus for rice rhizosheath formation under moderate soil drying
Bacteria-fungi interactions are common in soil environments [83, 84], and their interactions are significant drivers of important ecosystem functions and services [84]. These interactions allow the respective partners to utilize sparse carbonaceous nutrients, which is beneficial for either or both of the partners [85,86,87]. Moreover, bacterial communities have a positive effect on plant rhizosheath formation [11, 13]. Here, the relative abundance of Bacillus was significantly increased in the rhizosheath and root endosphere with P. indica inoculation under MSD, compared with MSD alone (Fig. 4A–C). This is probably because fungal exudates are a source of nutrients for bacteria in the mycosphere under drought stress [42, 65, 88]. B. cereus grew toward P. indica and grew more in the presence of P. indica under sterilized soil and Petri plate conditions (Supplementary Fig. S10C, D), which suggests that P. indica may interact with B. cereus. Furthermore, rhizosheath formation in WT and the OsPIN2 complemented rice line inoculated with P. indica under non-sterilized soil conditions was significantly higher than under sterilized soil conditions (Fig. 5B). The WT and OsPIN2 complemented rice line with a combine of P. indica and B. cereus inoculation showed the greatest rhizosheath formation under MSD (Fig. 5). The results suggest that the fungus P. indica can co-work with the native soil bacterium Bacillus cereus for rice rhizosheath formation under MSD.
In conclusion, we demonstrate that the cooperation of P. indica (fungus) and B. cereus (bacterium) can contribute to rhizosheath formation in rice plants under MSD through the involvement of the auxin efflux carrier OsPIN2 (Supplementary Fig. S12). These results improve our understanding of how plant roots integrate microbial action into drought responses, which will help to identify novel research avenues for improving plant adaptation to changing climate.
References
Fageria N. Yield physiology of rice. J Plant Nutr. 2007;30:843–79.
Wang Z, Zhang W, Beebout S, Zhang H, Liu L, Yang J, et al. Grain yield, water and nitrogen use efficiencies of rice as influenced by irrigation regimes and their interaction with nitrogen rates. Field Crops Res. 2016;193:54–69.
Zhang H, Xue Y, Wang Z, Yang J, Zhang J. An alternate wetting and moderate soil drying regime improves root and shoot growth in rice. Crop Sci. 2009;49:2246–60.
Bouman B, Tuong T. Field water management to save water and increase its productivity in irrigated lowland rice. Agr Water Manag. 2001;49:11–30.
Harrison M, Tardieu F, Dong Z, Messina C, Hammer G. Characterizing drought stress and trait influence on maize yield under current and future conditions. Glob Change Biol. 2014;20:867–78.
Lesk C, Rowhani P, Ramankutty N. Influence of extreme weather disasters on global crop production. Nature. 2016;529:84–87.
Thorup-Kristensen K, Kirkegaard J. Root system-based limits to agricultural productivity and efficiency: the farming systems context. Ann Bot. 2016;118:573–92.
Yao F, Huang J, Cui K, Nie L, Xiang J, Liu X, et al. Agronomic performance of high-yielding rice variety grown under alternate wetting and drying irrigation. Field Crops Res. 2012;126:16–22.
Danin A. Plant adaptations to environmental stresses in desert dunes. In: Danin A (ed). Plants of desert dunes. (Springer, Berlin, 1996), pp 133–152.
Pang J, Ryan M, Siddique K, Simpson R. Unwrapping the rhizosheath. Plant Soil. 2017;418:129–39.
Marasco R, Mosqueira M, Fusi M, Ramond J, Merlino G, Booth J, et al. Rhizosheath microbial community assembly of sympatric desert speargrasses is independent of the plant host. Microbiome. 2018;6:215.
Zhang Y, Du H, Gui Y, Xu F, Liu J, Zhang J, et al. Moderate water stress induces rice rhizosheath formation associated with ABA and auxin responses. J Exp Bot. 2020;71:2740–51.
Duell R, Peacock G. Rhizosheaths on mesophytic grasses. Crop Sci. 1985;25:880–3.
Ndour P, Gueye M, Barakat M, Ortet P, Bertrand-Huleux M, Pablo A, et al. Pearl millet genetic traits shape rhizobacterial diversity and modulate rhizosphere aggregation. Front Plant Sci. 2017;8:1288.
Philippot L, Raaijmakers J, Lemanceau P, van der Putten W. Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol. 2013;11:789–99.
Ndour P, Heulin T, Achouak W, Laplaze L, Cournac L. The rhizosheath: from desert plants adaptation to crop breeding. Plant Soil. 2020;456:1–13.
George T, Brown L, Ramsay L, White P, Newton A, Bengough A, et al. Understanding the genetic control and physiological traits associated with rhizosheath production by barley (Hordeum vulgare). N Phytol. 2014;203:195–205.
Zhang Y, Du H, Xu F, Ding Y, Gui Y, Zhang J, et al. Root-bacterial associations boost rhizosheath formation in moderately dry soil through ethylene responses. Plant Physiol. 2020;183:780–92.
Basirat M, Mousavi S, Abbaszadeh S, Ebrahimi M, Zarebanadkouki M. The rhizosheath: a potential root trait helping plants to tolerate drought stress. Plant Soil. 2019;445:565–75.
Othman A, Amer W, Fayez M, Hegazi N. Rhizosheath of sinai desert plants is a potential repository for associative diazotrophs. Microbiol Res. 2004;159:285–93.
Haling R, Richardson A, Culvenor R, Lambers H, Simpson R. Root morphology, root-hair development and rhizosheath formation on perennial grass seedlings is influenced by soil acidity. Plant Soil. 2010;335:457–68.
Delhaize E, James R, Ryan P. Aluminium tolerance of root hairs underlies genotypic differences in rhizosheath size of wheat (Triticum aestivum) grown on acid soil. N Phytol.2012;195:609–19.
Liu T, Ye N, Song T, Cao Y, Gao B, Zhang D, et al. Rhizosheath formation and involvement in foxtail millet (Setaria italica) root growth under drought stress. J Integr Plant Biol. 2019;61:449–62.
Liu T, Chen M, Zhang Y, Zhu F, Liu Y, Tian Y, et al. Comparative metabolite profiling of two switchgrass ecotypes reveals differences in drought stress responses and rhizosheath weight. Planta. 2019;250:1355–69.
Brown L, George T, Neugebauer K, White P. The rhizosheath–a potential trait for future agricultural sustainability occurs in orders throughout the angiosperms. Plant Soil. 2017;418:115–28.
Sirrenberg A, Göbel C, Grond S, Czempinski N, Ratzinger A, Karlovsky P, et al. Piriformospora indica affects plant growth by auxin production. Physiol Plant. 2007;131:581–9.
Weiβ M, Waller F, Zuccaro A, Selosse M. Sebacinales-one thousand and one interactions with land plants. N Phytol. 2016;211:20–40.
Vadassery J, Ranf S, Drzewiecki C, Mithoer A, Mazars C, Scheel D, et al. A cell wall extract from the endophytic fungus Piriformospora indica promotes growth of Arabidopsis seedlings and induces intracellular calcium elevation in roots. Plant J. 2009;59:193–206.
Lee Y, Johnson J, Chien C, Sun C, Cai D, Lou B, et al. Growth promotion of Chinese cabbage and Arabidopsis by Piriformospora indica is not stimulated by mycelium-synthesized auxin. Mol Plant Microbe Interact. 2011;24:421–31.
Dong S, Tian Z, Chen P, Senthil Kumar R, Shen C, Cai D, et al. The maturation zone is an important target of Piriformospora indica in Chinese cabbage roots. J Exp Bot. 2013;64:4529–40.
Rani M, Raj S, Dayaman V, Kumar M, Dua M, Johri A. Functional characterization of a hexose transporter from root endophyte Piriformospora indica. Front Microbiol. 2016;7:1083.
Prasad D, Verma N, Bakshi M, Narayan O, Singh A, Dua M, et al. Functional characterization of a magnesium transporter of root endophytic fungus Piriformospora indica. Front Microbiol. 2018;9:3231.
Narayan O, Verma N, Jogawat A, Dua M, Johri A. Sulfur transfer from the endophytic fungus Serendipita indica improves maize growth and requires the sulfate transporter SiSulT. Plant Cell. 2021;33:1268–85.
Baltruschat H, Fodor J, Harrach B, Niemcayk E, Barna B, Gullner G, et al. Salt tolerance of barley induced by the root endophyte Piriformospora indica is associated with a strong increase in antioxidants. N Phytol. 2008;180:501–10.
Jogawat A, Saha S, Bakshi M, Dayaman V, Kumar M, Dua M, et al. Piriformospora indica rescues growth diminution of rice seedlings during high salt stress. Plant Signal Behav. 2013;8:e26891.
Fakhro A, Andrade-Linares D, von Bargen S, Bandte M, Buttner C, Grosch R. Impact of Piriformospora indica on tomato growth and on interaction with fungal and viral pathogens. Mycorrhiza. 2010;20:191–200.
Sarma M, Kumar V, Saharan K, Srivastava R, Sharma A, Prakash A, et al. Application of inorganic carrier-based formulations of fluorescent pseudomonads and Piriformospora indica on tomato plants and evaluation of their efficacy. J Appl Microbiol. 2011;111:456–66.
Sun C, Shao Y, Vahabi K, Lu J, Bhattacharya S, Dong S, et al. The beneficial fungus Piriformospora indica protects Arabidopsis from Verticillium dahliae infection by downregulation plant defense responses. BMC Plant Biol. 2014;14:268.
Abdelaziz M, Abdelsattar M, Abdeldaym E, Atia M, Mahmoud A, Saad M, et al. Piformospora indica alters Na+/K+ homeostasis, antioxidant enzymes and LeNHX1 expression of greenhouse tomato grown under salt stress. Sci Hortic. 2019;256:108532.
Zhang W, Wang J, Xu L, Wang A, Huang L, Du H, et al. Drought stress responses in maize are diminished by Piriformospora indica. Plant Signal Behav. 2017;13:e1414121.
Pion M, Spangenberg J, Simon A, Bindschedler S, Flury C, Chatelain A, et al. Bacterial farming by the fungus Morchella crassipes. Proc R Soc B. 2013;280:20132242.
Guhr A, Borken W, Spohn M, Matzner E. Redistribution of soil water by a saprotrophic fungus enhances carbon mineralization. Proc Natl Acad Sci USA. 2015;112:14647–51.
Warmink J, Nazir R, van Elsas J. Universal and species-specific bacterial ‘fungiphiles’ in the mycospheres of different basidiomycetous fungi. Environ Microbiol. 2009;11:300–12.
Nazir R, Warmink J, Boersma H, van Elsas J. Mechanisms that promote bacterial fitness in fungal-affected soil microhabitats. FEMS Microbiol Ecol. 2010;71:169–85.
Wang L, Guo M, Li Y, Ruan W, Mo X, Wu Z, et al. LARGE ROOT ANGLE1, encoding OsPIN2, is involved in root system architecture in rice. J Exp Bot. 2018;69:385–97.
Bütehorn B, Rhody D, Franken P. Isolation and characterization of Pitef1 encoding the translation elongation factor EF-1α of the root endophyte Piriformospora indica. Plant Biol. 2008;2:687–92.
Haling R, Brown L, Bengough A, Young I, Hallett P, White P, et al. Root hairs improve root penetration, root-soil contact, and phosphorus acquisition in soils of different strength. J Exp Bot. 2013;64:3711–21.
Hou M, Luo F, Wu D, Zhang X, Lou M, Shen D, et al. OsPIN9, an auxin efflux carrier, is required for the regulation of rice tiller bud outgrowth by ammonium. N Phytol 2021;229:935–49.
Yuan J, Ruan Y, Wang B, Zhang J, Waseem R, Huang Q, et al. Plant growth-promoting rhizobacteria strain Bacillus amyloliquefaciens NJN-6-enriched bio-organic fertilizer suppressed Fusarium wilt and promoted the growth of banana plants. J Agr Food Chem. 2013;61:3774–80.
Xu F, Wang K, Yuan W, Xu W, Liu S, Kronzucker H, et al. Overexpression of aquaporin OsPIP1;2 in rice improves yield by enhancing mesophyll CO2 conductance and phloem sucrose transport. J Exp Bot. 2019;70:671–81.
Bulgarelli D, Rott M, Schlaeppi K, Ver Loren van Themaat E, Ahmadinejad N, Assenza F, et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature. 2012;488:91–95.
Marasco R, Rolli E, Ettoumi B, Vigani G, Mapelli F, Borin S, et al. A drought resistance-promoting microbiome is selected by root system under desert farming. PLoS One. 2012;7:e48479.
Bodenhausen N, Horton M, Bergelson J. Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. PLoS One. 2013;8:e56329.
Schlaeppi K, Dombrowski N, Oter R, Themaat E, Schulze-Lefert P. Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc Natl Acad Sci USA. 2014;111:585–92.
Han Q, Ma Q, Chen Y, Tian B, Xu L, Bai Y, et al. Variation in rhizosphere microbial communities and its association with the symbiotic efficiency of rhizobia in soybean. ISME J. 2020;14:1915–28.
Chen S, Zhou Y, Chen Y, Gu J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–90.
Edgar R. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
Edgar R. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods. 2013;10:996–8.
Wang Q, Garrity G, Tiedje J, Cole J. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb. 2007;73:5261–7.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–6.
Schloss P, Westcott S, Ryabin T, Hall J, Hartmann M, Hollister E, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microb. 2009;75:7537–41.
Wang B, Yuan J, Zhang J, Shen Z, Zhang M, Li R, et al. Effects of novel bioorganic fertilizer produced by Bacillus amyloliquefaciens W19 on antagonism of Fusarium wilt of banana. Biol Fertil Soils. 2013;49:435–46.
Turner J, Backman P. Factors relating to peanut yield increases after seed treatment with Bacillus subtilis. Plant Dis. 1991;75:347–53.
Wei Z, Gu Y, Friman V, Kowalchuk G, Xu Y, Shen Q, et al. Initial soil microbiome composition and functioning predetermine future plant health. Sci Adv. 2019;5:eaaw0759.
Zhang W, Li X, Sun K, Tang M, Xu F, Zhang M, et al. Mycelial network-mediated rhizobial dispersal enhances legume nodulation. ISME J. 2020;14:1015–29.
Mela F, Fritsche K, de Boer W, van Veen J, de Graaff L, van den Berg M, et al. Dual transcriptional profiling of a bacterial/fungal confrontation: Collimonas fungivorans versus Aspergillus niger. ISME J. 2011;5:1494–504.
Berendsen R, Vismans G, Yu K, Song Y, de Jonge R, Burgman W, et al. Disease-induced assemblage of a plant-beneficial bacterial consortium. ISME J. 2018;12:1496–507.
Zhang J, Liu Y, Zhang N, Hu B, Jin T, Xu H, et al. NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat Biotechnol. 2019;37:676–84.
Waller F, Achatz B, Baltruschat H, Fodor J, Becker K, Fischer M, et al. The endophytic fungus Piriformospora indica reprograms barley to salt-stress tolerance, disease resistance, and higher yield. Proc Natl Acad Sci USA. 2005;102:13386–91.
Chen T, Nomura K, Wang X, Sohrabi R, Xu J, Yao L, et al. A plant genetic network for preventing dysbiosis in the phyllosphere. Nature. 2020;580:653–7.
Rolli E, Marasco R, Vigani G, Ettoumi B, Mapelli F, Deangelis M, et al. Improved plant resistance to drought is promoted by the root-associated microbiome as a water stress-dependent trait. Environ Microbiol. 2015;17:316–31.
Preece C, Peñuelas J. Rhizodeposition under drought and consequences for soil communities and ecosystem resilience. Plant Soil. 2016;409:1–17.
Bezzate S, Aymerich S, Chambert R, Czarnes S, Berge O, Heulin T. Disruption of the Paenibacillus polymyxa levansucrase gene impairs its ability to aggregate soil in the wheat rhizosphere. Environ Microbiol. 2000;2:333–42.
Alami Y, Achouak W, Marol C, Heulin T. Rhizosphere soil aggregation and plant growth promotion of sunflowers by an exopolysaccharide-producing Rhizobium sp. strain isolated from sunflower roots. Appl Environ Microbiol. 2000;66:3393–8.
Berge O, Lodhi A, Brandelet G, Santaella C, Roncato M, Christen R, et al. Rhizobium alamii sp. nov., an exopolysaccharide-producing species isolated from legume and non-legume rhizospheres. Int J Syst Evol Microbiol. 2009;59:367–72.
Moreno-Espíndola I, Rivera-Becerril F, de Jesús F-GM, De León-González F. Role of root-hairs and hyphae in adhesion of sand particles. Soil Biol Biochem. 2007;39:2520–6.
Watt M, Mccully M, Canny M. Formation and stabilization of rhizosheaths of Zea mays L. (effect of soil water content). Plant Physiol. 1994;106:179–86.
Schafer P, Pfiffi S, Voll L, Zajic D, Chandler P, Waller F, et al. Manipulation of plant innate immunity and gibberellin as factor of compatibility in the mutualistic association of barley roots with Piriformospora indica. Plant J. 2009;59:461–74.
Xu W, Jia L, Shi W, Liang J, Zhou F, Li Q, et al. Abscisic acid accumulation modulates auxin transport in the root tip to enhance proton secretion for maintaining root growth under moderate water stress. N Phytol. 2013;197:139–50.
Uga Y, Sugimoto K, Ogawa S, Rane J, Ishitani M, Hara N, et al. Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat Genet. 2013;45:1097–102.
Luschnig C, Gaxiola R, Grisafi P, Fink G. EIR1, a root-specific protein involved in auxin transport, is required for gravitropism in Arabidopsis thaliana. Genes Dev. 1998;12:2175–87.
Müller A, Guan C, Gälweiler L, Tänzler P, Huijser P, Marchant A, et al. AtPIN2 defines a locus of Arabidopsis for root gravitropism control. EMBO J. 1998;17:6903–11.
de Boer W, Folman R, Summerbell R, Boddy L. Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol Rev. 2005;29:795–811.
Hogan D, Kolter R. Pseudomonas-Candida interactions: an ecological role for virulence factors. Science. 2002;296:2229–32.
Ravnskov S, Nybroe O, Jakobsen I. Influence of an arbuscular mycorrhizal fungus on Pseudomonas fluorescens DF57 in rhizosphere and hyphosphere soil. N Phytol. 1999;142:113–22.
Torsvik V, Øvreas L, Thingstad T. Prokaryotic diversity-magnitude, dynamics, and controlling factors. Science. 2002;296:1064–6.
Wamberg C, Christensen S, Jakobsen I, Müller A, Sørensen S. The mycorrhizal fungus (Glomus intraradices) affects microbial activity in the rhizosphere of pea plants (Pisum sativum). Soil Biol Biochem. 2003;35:1349–57.
van Hees P, Rosling A, Essen S, Godbold D, Jones D, Finlay R. Oxalate and ferricrocin exudation by the extrametrical mycelium of an ectomycorrhizal fungus in symbiosis with Pinus sylvestris. N Phytol. 2006;169:367–78.
Acknowledgements
We thank Prof. Chuanzao Mao (Zhejiang University, Hangzhou, China) for donating the ospin2, Complementation lines and cv. Hei-Jing2 rice seeds, Prof. Faxing Chen (Fujian Agriculture and Forestry University, Fuzhou, China) for P. indica-GFP strain and Dr. Chengyuan Tao, Dr. Guan Pang (Nanjing Agricultural University, Nanjing, China) for sharing microbial technology. We are grateful for the financial support from the National Key Research and Development Program of China (2017YFE0118100), National Natural Science Foundation of China (31761130073 and 31872169) and Postdoctoral Science Foundation of China (2020M671920).
Author information
Authors and Affiliations
Contributions
WX and FX planned and designed the research. FX, YZ, JL, LS, XZ, JY, KW, XW, and YD conducted most of the experiments. FX, HL, YZ, MY and CL analyzed the data. FX, HL, CR, JZ, KY and WX wrote the article. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Xu, F., Liao, H., Zhang, Y. et al. Coordination of root auxin with the fungus Piriformospora indica and bacterium Bacillus cereus enhances rice rhizosheath formation under soil drying. ISME J 16, 801–811 (2022). https://doi.org/10.1038/s41396-021-01133-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01133-3
This article is cited by
-
Nutrient and mycoremediation of a global menace ‘arsenic’: exploring the prospects of phosphorus and Serendipita indica-based mitigation strategies in rice and other crops
Plant Cell Reports (2024)
-
Auxin-producing bacteria promote barley rhizosheath formation
Nature Communications (2023)
-
Hyphosphere microorganisms facilitate hyphal spreading and root colonization of plant symbiotic fungus in ammonium-enriched soil
The ISME Journal (2023)
-
Halophyte functional groups influence seasonal variations in rhizosphere microbial necromass and enzyme activities in an inland saline ecosystem
Biology and Fertility of Soils (2023)
-
Bacillus sp. LC390B from the Maize Rhizosphere Improves Plant Biomass, Root Elongation, and Branching and Requires the Phytochromes PHYA and PHYB for Phytostimulation
Journal of Plant Growth Regulation (2023)