Abstract
Microbial communities play important roles in all ecosystems and yet a comprehensive understanding of the ecological processes governing the assembly of these communities is missing. To address the role of biotic interactions between microorganisms in assembly and for functioning of the soil microbiota, we used a top-down manipulation approach based on the removal of various populations in a natural soil microbial community. We hypothesized that removal of certain microbial groups will strongly affect the relative fitness of many others, therefore unraveling the contribution of biotic interactions in shaping the soil microbiome. Here we show that 39% of the dominant bacterial taxa across treatments were subjected to competitive interactions during soil recolonization, highlighting the importance of biotic interactions in the assembly of microbial communities in soil. Moreover, our approach allowed the identification of microbial community assembly rule as exemplified by the competitive exclusion between members of Bacillales and Proteobacteriales. Modified biotic interactions resulted in greater changes in activities related to N- than to C-cycling. Our approach can provide a new and promising avenue to study microbial interactions in complex ecosystems as well as the links between microbial community composition and ecosystem function.
Similar content being viewed by others
Introduction
Microbial communities in nature exist in complex and dynamic consortia of populations that are not only central to all major biogeochemical cycles, but also influence plant, animal, and human welfare [1,2,3]. These communities assemble through neutral processes, as well as through abiotic and biotic filtering [4]. While a large body of research has focused on the importance of abiotic factors [5, 6], there have been relatively less investigations on the importance of biotic factors, and in particular on the interactions between microorganisms, to explain the composition of microbial communities in the environment [7]. Understanding the different processes involved in the assembly of such complex communities is currently receiving attention due to the great potential of translating such knowledge into practical outcomes, e.g., in agroecosystems to increase soil fertility and improve crop production [8, 9].
Various types of positive and negative interactions between microorganisms, ranging from mutualism to competition, have been identified [10,11,12]. For example, in cooperative interactions, microorganisms can divide labor, whereby some individuals specialize to carry out tasks that benefit other individuals [13]. On the other hand, competition can be fierce between microorganisms, with evidence of both indirect exploitative competition, in which an individual consumes the resources required by another member, and direct interference competition, in which an individual inhibits the growth of another through the synthesis of harmful products [12, 14, 15]. Negative interactions between microorganisms also comprise parasitism and predation with diverse predatory viruses, protists, and even bacteria described from a variety of environments [16,17,18,19].
To date, efforts to experimentally identify biotic interactions between microorganisms have typically relied on bottom-up approaches based on synthetic-assemblage experiments conducted in vitro with culturable strains [20,21,22]. These co-culture experiments are based on the assumption that if there is an interaction between two microbial species, the fitness of at least one of them is different when grown together than when grown in the absence of the other species [14]. Such approaches have provided insights into underlying mechanisms by which microorganisms interact, but do not reflect the complexity of natural microbial communities or of their natural habitat. There is therefore little empirical data regarding the extent to which biotic interactions are shaping the composition of complex microbial communities in natural settings.
Here, we take an alternative top-down approach based on microbial community manipulation by targeted removal of various microbial groups in a native soil community to test the role of biotic interactions for microbial community assembly. Specifically, soil microbial communities were first subjected to different biocidal and filtration treatments before being reinoculated in their native, but sterilized soil to allow them to assemble during recolonization. Such removal treatments are intuitively predicted to cause changes in the fitness of the microorganisms interacting with those being depleted by the treatment. We hypothesized that manipulation of the microbial community will lead to changes in the community assembly during the soil colonization process in ways that can unravel the importance of biotic interactions, and their consequences for soil functions.
We show here that 39% of the dominant bacterial taxa across treatments were subjected to competitive interactions during soil recolonization, therefore experimentally showing for the first time the importance of negative interactions between microorganisms for community assembly in a complex environment. This removal approach can also provide a new framework to study microbial interactions in ecosystems as well as links between microbial community composition and ecosystem function based on the analogy to gene-knockout procedures in genomics.
Materials and methods
Soil sampling and experimental design
The soil was collected from the Epoisses site in France (47° 30′ 22.1832′′ N, 4° 10′ 26.4648′′ E) in March 2017. The soil properties were 41.9% clay, 51.9% silt, and 6.2% sand, pH 7.2, and C and N content 15.5 and 1.4 g kg−1 dry soil, respectively. Twenty-five soil subsamples were kept at −20 °C to characterize the initial microbial species pool. The collected soil was sieved through 4 mm before preparing soil suspensions by mixing 100 g equivalent dry mass soil with 150 ml sterile distilled water with a waring blender under sterile conditions. Soil suspensions were diluted ten times, centrifuged at 1000 × g for 2 min and supernatants were then filtrated at 10 µm in order to remove larger microbial communities. Soil suspensions were subjected to ten different treatments aiming at removing various microbial groups: three types of biocidal antibiotics (gentamicin, ramoplanin, and ciprofloxacin), an antimicrobial peptide (RW4), four filtration treatments based on cell size (F ≥ 3 µm, 0.8 ≤ F < 3 µm, 0.4 ≤ F < 0.8 µm, and F < 0.4 µm), a heat shock (0 °C for 5 min/70 °C for 15 min/0 °C for 5 min), and oxidative stress treatments (H2O2 at a final concentration of 50 mM). Each treatment was replicated on 25 soil suspensions and 4.5 mL from each treated soil suspensions was inoculated into 147 mL plasma flasks containing 30 g of the same gamma-sterilized soil (two times 35 kGy; Conservatome, Dagneux, France). All of the 275 soil microcosms were closed with sterile lids and incubated at 20 °C at a soil moisture ranging between 60 and 80% of the soil water-holding capacity for 45 days. Soil microcosms inoculated with non-treated soil suspensions and incubated 45 days (NT control) were used as controls (n = 25; Fig. 1).
a Design of the microcosm experiment to manipulate the soil microbial community by subjecting soil suspensions to a range of removal treatments (three antibiotics, one antimicrobial peptide, four size filtration, one heat shock, and one oxidative stress), targeting various microbial groups (n = 25). Pie charts are symbolizing the composition of microbial communities. Values in parenthesis indicate the number of replicates. b Summary of possible ecological interactions between different species (A, B, and C) in the community (before) and consequences in each case for the other two species (increased or decreased relative fitness represented by the size of the symbols) when species A is depleted by a removal treatment (after). In examples 1–3, depletion of species A caused a decrease in relative fitness of species B, which it is not possible to distinguish from a decrease in relative fitness caused by a direct effect of the removal treatment. Only examples 4–6 causing an increase in the relative fitness are detected in our approach.
Assessment of microbial community composition and diversity
After 45 days of incubation, soil microcosms were used for analyses of total bacterial and fungal diversity and composition by sequencing the 16S rRNA and ITS genes via Illumina Miseq 2 × 250 bp paired-end analysis. First, DNA was extracted from 250 mg from each of the 275 soil microcosms as well as the 25 original soil subsamples using the DNeasy PowerSoil-htp 96-well DNA isolation kit (Qiagen, Hilden, Germany). Amplicons were generated for all 300 DNA extracts in two steps. In the first step, the V3-V4 hypervariable region of the bacterial 16S rRNA gene was amplified by polymerase chain reaction (PCR) using the fusion primers U341F (5’-CCTACGGGRSGCAGCAG-3’) and 805R (5’-GACTACCAGGGTATCTAAT-3’), with overhang adapters (forward: TCGTCGGCAGCGTCAGATGTGTATAAGAGACAG, adapter: GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAG) to allow the subsequent addition of multiplexing index-sequences. Fungal ITS was amplified using the primers ITS3F (5’-GCATCGATGAAGAACGCAGC-3’) and ITS4R (5’-TCCTCSSCTTATTGATATGC-3’). Thermal cycling conditions of the first step PCR were as follows: 98 °C for 3 min followed by 98 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s (25 and 30 cycles for 16S rRNA and ITS genes, respectively) and a final extension for 10 min at 72 °C. Duplicate first step PCR products were pooled and then used as template for the second step PCR. In the second step, PCR amplification added multiplexing index-sequences to the overhang adapters using a unique multiplex primer pair combination for each sample. Thermal cycling conditions were as follows: 8 °C for 3 min followed by 98 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s (8 and 10 cycles for 16S rRNA and ITS genes, respectively) and a final extension for 10 min at 72 °C. Duplicate second step PCR products were pooled then visualized in 2% agarose gel to verify amplification and size of amplicons. The amplicons were cleaned-up and pooled using sequalPrep Normalization plate kit 96-well (Invitrogen, Carlsbad, CA, USA). Sequencing was performed on MiSeq (Illumina, 2 × 250 bp) using the MiSeq reagent kit v2 (500 cycles). Demultiplexing and trimming of Illumina adaptors and barcodes was done with Illumina MiSeq Reporter software (version 2.5.1.3).
Sequencing and bioinformatic analysis
Sequence data from the 300 soil samples were analyzed using an in-house developed Python pipeline (available upon request). Briefly, 16S rDNA and ITS sequences were assembled using PEAR [23] with default settings. Further quality checks were conducted using the QIIME pipeline [24] and short sequences were removed (<400 bp for 16S and <300 bp for ITS). Reference-based and de novo chimera detection, as well as operational taxonomic units (OTUs) clustering were performed using VSEARCH [25] and the adequate reference databases (SILVA representative set of sequences for 16S rRNA and UNITE’s ITS2 reference dynamic dataset for ITS). The identity thresholds were set at 94% for 16S rRNA based on replicate sequencing of a bacterial mock community [26] and 97% for ITS. A total of 4,307,710 bacterial 16S rRNA gene and 15,077,367 fungal ITS region sequences were obtained. Representative sequences for each OTU were aligned using PyNAST [27] and a 16S rDNA phylogenetic tree was constructed using FastTree [28]. Taxonomy was assigned using UCLUST [29] and the SILVA reference database 132 [30]. For ITS, the taxonomy assignment was performed using BLAST [31] and the UNITE reference database (v.7-08/2016 [32]). Raw sequences were deposited at the NCBI under the BioProject PRJNA542862.
Bacterial and fungal α-diversity metrics (i.e., observed species, Simpson’s reciprocal, Shannon, and for bacteria also Faith’s Phylogenetic Diversity PD [33]) and Net Relatedness and Nearest Taxon indices [34] were calculated based on rarefied OTU tables (5000 sequences per sample for 16S rDNA and 8000 sequences per sample for ITS). To assess the contribution of deterministic and stochastic processes to the bacterial community structure, the normalized stochasticity ratio index (NST) was calculated [35]. Weighted UniFrac distance matrix [36] and Bray–Curtis dissimilarity matrix were also computed to detect variations in the structure of microbial communities for 16S rDNA and ITS, respectively.
Low-abundance OTUs were discarded by keeping OTUs with at least 0.5% relative abundance across all samples (353 and 1370 OTUs for 16S rDNA and ITS, respectively). Due to the high proportion of zero counts, fungal OTUs with low prevalence (present in less than 70% of replicates per treatment) were removed (305 OTUs).
Quantification of microbial communities
The abundances of total bacterial and fungal microbial communities as well as that of N-cycle microbial guilds were estimated by real-time quantitative PCR (qPCR) assays. For each treatment, the 25 DNA extracts were used to prepare 5 equimolar DNA mixtures (each corresponding to five different DNA extracts), which were added as templates for the qPCR assays (n = 5). Total bacterial and fungal communities were quantified using 16S rDNA and ITS primers described by Muyzer et al. [37] and White et al. [38], respectively. The nitrification gene amoA and the denitrification genes nirK and nirS were used as molecular markers to quantify the bacterial (AOB) and archaeal (AOA) ammonia-oxidizing and the denitrifying communities, as described previously [39]. qPCR reactions were carried out in a ViiA7 (Life Technologies, Carlsbad, CA, USA) in a 15 µL reaction volume containing 7.5 µL of Takyon Master Mix (Eurogentec, Liège, Belgium), 1 µM of each primer, 250 ng of T4 gene 32 (MP Biomedicals, Santa Ana, CA, USA), and 1 ng of DNA. Two independent runs were performed for each real-time PCR assay. Standard curves were obtained using serial dilutions of linearized plasmids containing appropriated cloned targeted genes from bacterial strains or environmental clones. PCR efficiency for the different assays ranged from 77 to 101%. No-template controls gave null or negligible values. Inhibition in qPCR assay was tested by mixing soil DNA extracts with either control plasmid DNA (pGEM-T Easy Vector, Promega, Madison, WI, USA) or water. No inhibition was detected in any case.
Assessing soil functions related to carbon and nitrogen cycling
Causal effects of microbial community manipulations on soil functioning were assessed by measuring a range of activities related to C and N cycles in replicate soil samples from each treatment (n = 5). The MicroResp method was used to measure microbial respiration rates of different C substrates across different treatments as described by Campbell et al. [40]. Eleven substrates were used: D-(+)-galactose, L-Arginine, Citric acid, L-Alanine, L-Malic acid, L-(+)-Arabinose, N-Acetyl glucosamine, Glucose phosphate, D-(-)-Fructose, D-(+)-Trehalose, and Gallic_acid. Soil nitrogen pools (NO3– and NH4+) were extracted using 50 mL of 1 M KCl that was added to ca. 10 g fresh soil, shaken vigorously (80 rpm for 1 h at room temperature), filtered and kept frozen until quantification according to ISO standard 14256-2. Quantification was performed by colorimetry in a BPC global 240 photometer.
Statistical analyses
Statistical analyses were conducted using R statistical software version 3.4.1 [41]. Differences between treatments in gene copy abundances (16S rRNA, ITS, bacterial and archaeal amoA, nirK, and nirS), ammonium and nitrate concentrations, microbial respiration measurements (n = 5) and the microbial α-diversity indices (n = 25) were tested using ANOVAs followed by Tukey’s honestly significant difference test (p value < 0.05) using the agricolae package [42]. Normality and homogeneity of the residual distribution were verified and log-transformations were performed when necessary.
Evaluating the impact of the removal treatments on beta-diversity
Permutational multivariate analysis of variance (PermANOVA) was carried out on the weighted UniFrac and the Bray–Curtis dissimilarity distance matrices to detect significant differences between treatments (n = 25) in community composition using adonis function implemented in the vegan package [43]. Principal coordinates analysis (PCoA) visualizations were created using the plot3D package using scatter3D function [44].
Identification of OTUs significantly affected by the removal treatments
Differential abundance analysis of microbial community composition was performed by comparing the count matrices between each treatment (n = 25) and the NT control (n = 25) using the default parameters of DESeq2 Bioconductor package (v.1.30.1), which allows for testing for changes in count matrices between conditions based on negative binomial generalized linear models [45, 46]. Significant OTUs result from Benjamini–Hochberg adjusted p values (BH-adjusted p value < 0.00001). Because of the high variability in the distribution of fungal sequences across replicates, only OTUs having a coefficient of variation lower than 200% within a given treatment and within NT control were kept for the DESeq2 analysis.
Bacterial OTUs exhibiting significant changes were used to build pruned trees using the ape package [47] and the trees were visualized using the Interactive Tree of Life [48].
Co-occurrence networks construction
Bacterial networks were constructed based on OTU count data (353 OTUs) using both all treatments plus the NT control for the global network (275 samples). Microbial inter-domain network was constructed using filtered bacterial and fungal OTU tables (353 and 305 OTUs, respectively) from all treatments plus the NT control (275 samples). For the original soil samples (25 samples), raw read count tables of both bacteria and fungi were used after removing OTUs that are not present in all replicates. Networks were inferred using a sparse multivariate Poisson log-normal (PLN) model with a latent Gaussian layer and an observed Poisson layer using the PLNmodels package [49]. The best network was selected using a Stability Approach to Regularization Selection [50]. An inter-domain-specific normalization with the GMPR (geometric mean of pairwise ratios) method was performed to take into account the heterogeneity of sequencing depth [51]. For visualization purpose, only partial correlations with |ρ| > 0.1 were considered. Networks were visualized using the Cytoscape software [52]. Nodes corresponding to the OTUs that showed significant changes in relative abundance based on the DESeq2 analysis were identified using the merge function in R and colored in the network according to log2-fold changes (positive, negative, or both depending on the treatment).
Multivariate integration of microbial activities, gene copy abundances, and community composition
Integration and visualization of bacterial OTUs (353), gene copy abundances (16S rRNA, bacterial, and archaeal amoA, nirK, and nirS), and microbial activity measurements were realized using the mixOmics package [53] using DIABLO (Data Integration Analysis for Biomarker discovery using a Latent component method for Omics studies) in order to identify correlated variables between different data sets (Pearson’s correlation |r| > 0.7) [54].
Results and discussion
Experimental manipulation of microbial community assembly
The experiment takes advantage of the enhanced interactions occurring between microorganisms during their recolonization of sterile soils [55, 56] and we manipulated these interactions using various removal approaches (Fig. 1a). Removal treatments cause depletion of different OTUs, of which some are assumed to have positive or negative interactions with the remaining OTUs. It is therefore expected that the relative fitness of the remaining OTUs that are competing or cooperating with the depleted ones will be modified in the removal treatments during soil recolonization. In this study, it was not possible to distinguish a decrease in relative fitness due to changes in positive interactions from a direct effect of the removal treatment (Fig. 1b). We therefore only focused on negative interactions that caused an increased relative fitness of OTUs that were previously impaired by the depleted ones. Hence, cases of negative interactions for which both competing strains were affected by the removal treatments or causing a decreased relative fitness (example 3, Fig. 1b) could not be determined in our analysis.
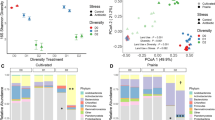
A total of 5551 bacterial and 6949 fungal OTUs were found across the different treatments and the original soil samples. To capture the effects of our removal approach on bacterial and fungal communities, differentially abundant OTUs were tested after soil recolonization by pairwise comparisons between the removal treatments and the control without removal treatment (NT control) using DESeq2 [45]. The removal approach effectively led to the depletion of bacterial groups (i.e., with significant decrease in relative abundances in a removal treatment compared to the NT control, BH-adjusted p value < 0.00001) that were different between treatments (Fig. 2). For example, we found that members of the Actinomycetes and Bacteroides were affected by the ramoplanin, which is an actinomycete-derived antibiotic. By contrast, ciprofloxacin mostly affected members of the Proteobacteria. Proteobacteria and Bacteroides were also the groups that exhibited the sharpest decline after the oxidative stress and the heat shock treatments. Overall, the depleted OTUs represented between 0.02 and 25% of the total bacterial community in the NT control (and <0.0003–2.3% in the original soil samples in which the five most abundant OTUs represented <5% of the community), which indicates that the selected treatments successfully affected both dominant and rare taxa (Supplementary Fig. 1).
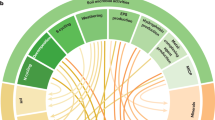
Log2-fold change, as calculated by DESeq2 analysis, of significantly increasing and decreasing bacterial OTUs in the removal treatments when compared to the NT control are represented by the blue-to-red color gradient. The white color indicates OTUs that are not affected by the treatments. The affiliation of OTUs at the phylum or class levels is indicated by different colors on the internal ring. Bootstrap values >80 are indicated by black circles.
Phylogenetic diversity of OTUs that declined significantly in any of the removal treatments compared to the NT control had a higher degree of relatedness than the total bacterial community (Tukey’s test, p value < 0.05; Supplementary Fig. 2). This shows that, as expected, bacterial taxa were mainly non-randomly depleted. UniFrac analyses demonstrated that our manipulation experiment resulted in differences in bacterial community structure between treatments with various degrees of dissimilarity to the NT control (PermANOVA, p value < 0.01; Fig. 3a). Major changes were observed in the heat shock and oxidative stress treatments and to a lesser extent in the ciprofloxacin treatments, while the bacterial community structure remained more similar to the control in the ramoplanin and two filtration treatments (Fig. 3 and Supplementary Figs. 3 and 4; Tukey’s test, p value < 0.05). The strong clustering by treatment suggests a limited stochasticity during bacterial community assembly despite a few treatments exhibiting more random effects than others (Fig. 3a). This was supported by a NST below 8% for all treatments and NRI/NTI indices higher than 2, which indicates that the coexisting OTUs within treatments were phylogenetically more closely related than expected by chance (Supplementary Figs. 5 and 6). Since the manipulated communities were inoculated in identical soil microcosms, differences in bacterial community assembly were mainly governed by the deterministic effects of the removal treatments in combination with altered species interactions rather than abiotic filtering (i.e., soil properties) or stochastic processes. By contrast, a high variability in the distribution of fungal OTUs was observed between replicates for the manipulated communities, but not in the original soil samples (Supplementary Fig. 7). This was likely due to the breakdown of hyphae during the experimental procedure causing the stochastic distribution of fungal OTUs. We found weak to non-significant effects of the removal treatments on fungal communities with only zero to two fungal OTUs depleted in all treatments but the heat shock treatment, in which nine OTUs mostly belonging to the Hypocreales significantly decreased. Therefore, even if fungi are not excluded, we consider that our experimental approach mainly allows detection of interactions involving bacteria.
a Principal coordinates analysis (PCoA) of the weighted UniFrac distance matrix of 16S rRNA gene amplicons showing shifts in the bacterial community structures between the original soil, NT control, and removal treatments. The different treatments are represented by different colors as specified in the legend. b Bacterial community composition between the original soil samples, NT control, and the different treatments. Relative abundances are shown at the phylum and class levels and expressed as a percentage of the total number of OTUs.
Changes in bacterial community composition were concomitant with an increase in bacterial abundance, as determined by qPCR, by at least 1–2 orders of magnitude during soil recolonization. Thus, using the 16S rRNA gene copy number in the original soil to calculate a maximal bacterial density in the inoculum before treatment, we estimated that the inoculation level was less than 107 16S rRNA gene copies g−1 dry soil, while we detected at least 5 × 108 16S rRNA gene copies g−1 dry soil after 45 days (Supplementary Fig. 8). Ecological theory suggests that expansion competition, in which strains race to utilize resources and occupy uninhabited space, is the dominant process determining the outcome of colonization of the sterile soil microcosms by the inoculated communities [57]. Accordingly, the three over-dominant OTUs that were the best at recolonizing the sterile soil in absence of biocidal treatment (NT control) belonged to Bacteroidetes and γ-Proteobacteria, which have previously been reported as copiotrophs (i.e., fast growing in nutrient rich environments) [58,59,60]. These fast-growing OTUs represented 53% of the total bacterial community in the NT control, but only 2% in the original soil samples despite being among the 25 most dominant OTUs (out of 4057). By contrast, members of the Acidobacteria were common in the original soil, but incapable of thriving in the sterilized soil. Given that soil sterilization by gamma irradiation causes release of organic carbon compounds [61], the disappearance of Acidobacteria supports their proposed oligotrophic lifestyle [58, 59].
Ecological importance of biotic interactions for microbial community assembly
The successful removal treatments should result in an increased relative abundance of the microorganisms that were subjected to negative interactions with the depleted ones. Due to the compositional nature of the community data, the depletion of certain taxa is also expected to result in an increase in the relative abundance of the other OTUs; with the proportional increase being the same for all the remaining OTUs and corresponding to the fraction of OTUs depleted. However, we observed fold increase in relative abundance that spanned over orders of magnitude for the same removal treatment, which indicates that the relative fitness of some OTUs was stimulated by the depletion of others. Thus, to identify the OTUs showing a significant increase in relative fitness after recolonization compared to the control NT (i.e., higher increase than what would be expected based on just the fraction of OTUs depleted following the removal treatment), a differential abundance analysis of microbial community composition was performed with DESeq2 (negative binomial generalized linear model, BH-adjusted p value < 0.00001) (Fig. 1b). Although differential abundance analysis can result in the identification of false positives, DESeq2 has been shown to be conservative and to control well the false positive rate [45]. Furthermore, having 25 replicates per treatment as well as including OTUs with at least 0.5% relative abundance across all samples and removing fungal OTUs with low prevalence, and thereby reducing zero counts, also helped minimizing the number of false positives. Only four fungal OTUs, all belonging to the genus Trichoderma, showed a significantly increased relative fitness across the heat shock and oxidative stress treatments. In contrast to fungi, we found a greater number of negative interactions during bacterial community assembly. Thus, out of the 353 most abundant bacterial OTUs across all treatments (i.e., relative abundance of at least 0.5% in any sample), the relative fitness of 139 OTUs was significantly stimulated in at least one of the removal treatments with 3- to more than 5000-fold changes compared to the NT control.
Because larger microorganisms were eliminated by filtering all soil suspensions at 10 μm before treatment and no known bacterivore bacterium was identified among the depleted OTUs, it is likely that exploitative or interference competition was responsible for the inferred negative interactions even though predation and parasitism cannot be ruled out [62]. Some of these bacterial OTUs were abundant in one or more removal treatments, whereas they were barely detected in the NT control (Fig. 2). For example, a quarter of the most dominant OTUs in the oxidative stress treatment belonged to the Bacillales, which accounted for 21% of the bacterial community, whereas Bacillales were rare in both the NT control and the original soil samples (0.57% and 0.14%, respectively). This was the case even when considering that the total bacterial abundance was two to six times lower in the oxidative stress treatment than in the original soil samples or in the NT control (Supplementary Fig. 8). This increase in the proportion of Bacillales in the community, usually a low-abundance taxa in soils [63], supports the idea that microbial rank abundance curves may be highly dynamic and that even rare subordinate species can become dominant when conditions turn out to be more favorable [64, 65]. Given that environmental factors and resources were the same for all microcosms before inoculation, our findings indicate that competitive exclusion rather than abiotic constraints prevents the rise of rare populations during soil recolonization. One interesting question is why high fitness inequalities have not driven the weak competitors to extinction in the original soil samples, a contradiction known as the biodiversity paradox. In the case of the Bacillales, it is likely their ability to enter a dormant state as spores and therefore to persist in a non-interactive state that explains their coexistence over time via the soil spore bank as previously suggested using theoretical models [64]. Overall, our findings in complex soil systems show that 39% of the dominant bacterial taxa across treatments were subjected to competitive interactions during soil recolonization. Although our approach may have inflated biotic interactions, for example by depriving bacteria from their natural shelters, the results extend previous evidence from in vitro and theoretical studies suggesting that competition is a common type of biotic interaction between bacteria [22, 62, 66]. Despite being beyond the scope of this work, efforts to characterize the mechanistic forces behind the observed interactions will allow deciphering whether exploitative or interference competition was prevalent in the studied soil microbiota.
Ecological network inference recapitulates biotic interactions
We reconstructed different networks, to infer significant associations between microorganisms, using a recently developed sparse multivariate PLN model [49]. First, the inferred global network across domains from all manipulation treatments and the NT control shows no association between bacteria and fungi, while significant edges between both domains were observed in the original soil samples (Supplementary Fig. 9a, b, respectively). This supports the results of the DESeq2 analysis, suggesting that bacteria rather than fungi were engaged in interactions unraveled by our removal approach. Next, we inferred a bacterial network including all treatments. To identify bacterial OTUs whose depletion actually leads to a significant increased relative fitness of the remaining ones, the nodes were colored according to the results of the DESeq2 analysis (Fig. 4). Out of the 180 OTUs present in this network, 90 were among those that significantly increased in at least one of the removal treatments compared to the NT control. Moreover, 50 out of 55 negative edges in the network are connecting depleted OTUs to OTUs with increased relative fitness. Thus, the interactions unraveled by our microbial community manipulation approach closely matched the inferred nodes and edges, which provides a cross-validation of our network analysis. Using a removal approach allowed going beyond simply testing significant associations in natural communities, as we could determine which OTUs were outcompeted. Thus, we found conserved negative interactions indicating that Bacillales were capable to grow only when α− and/or γ- Proteobacteria were depleted by the removal treatments (e.g., heat shock and oxidative stress; Figs. 2 and 4). Similar antagonistic patterns have been observed in previous studies showing segregated spatial co-occurrences or checkerboard patterns between Firmicutes and γ- Proteobacteria [55, 67]. Interestingly, in the global bacterial network, several γ-Proteobacteria (Xanthomonas sp.) have negative edges with another γ-Proteobacteria (Burkholderia sp.) and two Bacillus hubs, which were themselves connected by negative edges. Since the fitness of these Bacillus and Burkholderia strains was significantly increased by a removal treatment, this suggests that these strains were thriving, but in competition when their competitors were depleted. Altogether these findings can be interpreted as evidence for predictable rule of bacterial community assembly. Such understanding of what govern community assembly would be instrumental for steering the soil microbiome toward a community that enhance ecosystem services [9, 20].
The nodes represent individual OTUs with the corresponding phylum. The nodes are also colored according to the DESeq2 results: green nodes represent OTUs with decreasing fitness, blue nodes represent OTUs with increasing fitness, gray nodes represent OTUs with both increasing and decreasing fitness (depending on the treatment), and orange nodes represent OTUs that were neither increasing nor decreasing. Edge thickness is proportional to partial correlations between nodes and represents associative (black, ρ > 0.1) or exclusionary relationships (red, ρ < −0.1).
Shifts in soil functions associated with microbial community manipulation
Soil functions related to the C and N cycles were investigated using microbial respiration rates of different C substrates (MicroResp), inorganic nitrogen pools as well as the abundance of ammonia-oxidizer and denitrifier genes as proxies for these specific processes. We found significant differences in the emergent functions of the manipulated soil microbial communities (Tukey’s test, p value < 0.05; Supplementary Fig. 10). Notably, microbial communities that were subjected to ciprofloxacin exhibited a significantly higher ability to respire glucosamine and fructose than all other removal treatments and controls (Tukey’s test, p value < 0.05; Supplementary Fig. 11), whereas respiration rates of many C substrates were lower in microbial communities that assembled after heat shock and oxidative stress treatments. Among the soil functions assessed, the greatest changes were observed for N cycling. The soil nitrate content declined in all treatments except one (0.8 µm filtration) when compared to the original soil samples (Tukey’s test, p value < 0.05; Supplementary Fig. 10b), which is in accordance with a previous manipulation experiment [55].
To further explore the relationship between the composition of the manipulated microbial community membership and soil functions, we used a multivariate dimension reduction discriminant analysis method that builds on Projection to Latent Structure models [54] (Fig. 5). The red and blue modules are showing OTUs (mostly Firmicutes and Actinobacteria and Actinobacteria, Chloroflexi and α-Proteobacteria, respectively) that were highly correlated with each other and with the total bacterial abundance but without any correlation to a soil function. However, several correlations that emerged in the other modules agreed with empirical knowledge. For example, Actinobacteria, which have a well-described role in carbon cycling [68], are representing three out of the five bacterial OTUs that were grouped in the module containing all the C-substrate respiration rates but gallic acid. In line with previous studies describing soil nitrification—the oxidation of ammonia to nitrate—as usually limited by its first step performed by ammonia oxidizers [69], the nitrate pool was correlated with the proportion of ammonia-oxidizing bacteria in the total bacterial community (AOB/16S rRNA). The proportion of AOB was included in a fully connected module comprising the proportion of nirK- and nirS-denitrifiers and eight OTUs, one of them belonging to the Nitrospiracea, a well-known group of nitrite oxidizers [70]. Interestingly, the nitrate pool was also correlated with another module of seven OTUs, one of them belonging to the Chloroflexi, which have been identified as a novel group of nitrite oxidizing bacteria [71] (Fig. 5). Since the nitrate pool and the abundance of AOB were the highest both in the original soil samples and in the 0.8 µm filtration treatment, while 16S rRNA gene sequences from known bacterial nitrifiers (Nitrospira and Nitrosomonas) in the 0.8 µm filtration treatment was about eight times lower than in original soil samples, one can speculate that one or several of these OTUs in this module could be also be capable of ammonia oxidation.
a Visualization of associated bacterial OTUs (blue nodes), N- and C-cycling activities (green nodes), and abundances of microbial groups (red nodes). The taxonomic identities of the OTUs are indicated at the phylum or class level. The N- and C-cycling activities are based on inorganic N pools and respiration rates of different substrates, respectively. Abundances of microbial groups are corresponding to the total bacterial community abundance (16S), the proportion of ammonia-oxidizing bacteria, and denitrifiers in the total bacterial community (AOB/16S, nirK/16S, and nirS/16S, respectively). Edges indicate positive (blue) or negative correlations (red) as defined by Pearson’s correlation r > 0.7 or r < −0.7, respectively. Different modules are represented by different colors and numbered (I–V). Correlation matrices of variables across data types in b modules I and II (green and yellow) and c within module III (purple).
Conclusions
By exploring biotic interactions within a community of naturally co-occurring soil microorganisms during the recolonization of their original habitat, we demonstrated that 39% of the dominant bacteria across treatments were subjected to negative interactions during community assembly. The approach used here allowed us to tie correlation patterns inferred by network analysis to ecological interactions revealed by experimental manipulation of the microbial community. We found evidence for competitive interactions between members of the low-abundance Bacillales and the dominant Proteobacteriales and suggest that competition-driven niche segregation rather than habitat features prevents the rise of rare populations in soil. Differences in the emergent functions of the manipulated communities were detected, with more pronounced shifts in functions related to N- rather than C-cycling. Thus, microbial community manipulation by removal, in addition to being informative about biotic interactions during assembly, may represent an alternative avenue to better understand the links between microbial community composition and ecosystem functioning based on the analogy to gene-knockout procedures in genomics [72]. Overall, our results suggest that some simple rules of bacterial community assembly can be identified, which has potential for predicting and steering the soil microbiota to promote or suppress certain functions in managed ecosystems. However, whether these empirically observed interactions can be generalized to other environments remains to be elucidated.
References
Falkowski PG, Fenchel T, Delong EF. The microbial engines that drive Earth’s biogeochemical cycles. Science. 2008;320:1034–9.
Le Chatelier E, Nielsen T, Qin JJ, Prifti E, Hildebrand F, Falony G, et al. Richness of human gut microbiome correlates with metabolic markers. Nature. 2013;500:541–6.
Philippot L, Raaijmakers JM, Lemanceau P, van der Putten WH. Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol. 2013;11:789–99.
Nemergut DR, Schmidt SK, Fukami T, O’Neill SP, Bilinski TM, Stanish LF, et al. Patterns and processes of microbial community assembly. Mol Biol Rev. 2013;77:342–56.
Jones RT, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N. A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J. 2009;3:442–53.
Rasche F, Knapp D, Kaiser C, Koranda M, Kitzler B, Zechmeister-Boltenstern S, et al. Seasonality and resource availability control bacterial and archaeal communities in soils of a temperate beech forest. ISME J. 2011;5:389–402.
Goberna M, Garcia C, Verdu M. A role for biotic filtering in driving phylogenetic clustering in soil bacterial communities. Glob Ecol Biogeogr. 2014;23:1346–55.
Zhou JZ, Ning DL. Stochastic community assembly: does it matter in microbial ecology? Mol Biol Rev. 2017;81:e00002–17.
Fierer N. Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol. 2017;15:579–90.
Faust K, Raes J. Microbial interactions: from networks to models. Nat Rev Microbiol. 2012;10:538–50.
Griffin AS, West SA, Buckling A. Cooperation and competition in pathogenic bacteria. Nature. 2004;430:1024–7.
Hibbing ME, Fuqua C, Parsek MR, Peterson SB. Bacterial competition: surviving and thriving in the microbial jungle. Nat Rev Microbiol. 2010;8:15–25.
West SA, Cooper GA. Division of labour in microorganisms: an evolutionary perspective. Nat Rev Microbiol. 2016;14:716–23.
Foster KR, Bell T. Competition, not cooperation, dominates interactions among culturable microbial species. Curr Biol. 2012;22:1845–50.
Garcia-Bayona L, Comstock LE. Bacterial antagonism in host-associated microbial communities. Science. 2018;361:eaat2456.
Braga LPP, Spor A, Kot W, Breuil MC, Hansen LH, Setubal JC, et al. Impact of phages on soil bacterial communities and nitrogen availability under different assembly scenarios. Microbiome. 2020;8:52.
Saleem M, Fetzer I, Harms H, Chatzinotas A. Diversity of protists and bacteria determines predation performance and stability. ISME J. 2013;7:1912–21.
Nair RR, Vasse M, Wielgoss S, Sun L, Yu YTN, Velicer GJ. Bacterial predator-prey coevolution accelerates genome evolution and selects on virulence-associated prey defences. Nat Commun. 2019;10:4301.
Perez J, Moraleda-Munoz A, Marcos-Torres FJ, Munoz-Dorado J. Bacterial predation: 75 years and counting! Environ Microbiol. 2016;18:766–79.
Friedman J, Higgins LM, Gore J. Community structure follows simple assembly rules in microbial microcosms. Nat Ecol Evol. 2017;1:109.
Goldford JE, Lu NX, Bajic D, Estrela S, Tikhonov M, Sanchez-Gorostiaga A, et al. Emergent simplicity in microbial community assembly. Science. 2018;361:469–74.
Russel J, Roder HL, Madsen JS, Burmolle M, Sorensen SJ. Antagonism correlates with metabolic similarity in diverse bacteria. Proc Natl Acad Sci USA. 2017;114:10684–8.
Zhang JJ, Kobert K, Flouri T, Stamatakis A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics. 2014;30:614–20.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Rognes T, Flouri T, Nichols B, Quince C, Mahe F. VSEARCH: a versatile open source tool for metagenomics. PeerJ. 2016;4:e2584.
Engelhardt IC, Welty A, Blazewicz SJ, Bru D, Rouard N, Breuil MC, et al. Depth matters: effects of precipitation regime on soil microbial activity upon rewetting of a plant-soil system. ISME J. 2018;12:1061–71.
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics. 2010;26:266–7.
Price MN, Dehal PS, Arkin AP. FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS One. 2010;5:e9490.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–6.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Abarenkov K, Nilsson RH, Larsson KH, Alexander IJ, Eberhardt U, Erland S, et al. The UNITE database for molecular identification of fungi—recent updates and future perspectives. N Phytol. 2010;186:281–5.
Faith DP. Conservation evaluation and phylogenetic diversity. Biol Conserv. 1992;61:1–10.
Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD, et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics. 2010;26:1463–4.
Ning DL, Deng Y, Tiedje JM, Zhou JZ. A general framework for quantitatively assessing ecological stochasticity. Proc Natl Acad Sci USA. 2019;116:16892–8.
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R. UniFrac: an effective distance metric for microbial community comparison. ISME J. 2011;5:169–72.
Muyzer G, Dewaal EC, Uitterlinden AG. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol. 1993;59:695–700.
White TJ, Bruns TD, Lee SB, Taylor JWI. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR-protocols and applications: a laboratory manual. New York, NY: Academic Press; 1990. p. 315–22.
Bru D, Ramette A, Saby NPA, Dequiedt S, Ranjard L, Jolivet C, et al. Determinants of the distribution of nitrogen-cycling microbial communities at the landscape scale. ISME J. 2011;5:532–42.
Campbell CD, Chapman SJ, Cameron CM, Davidson MS, Potts JM. A rapid microtiter plate method to measure carbon dioxide evolved from carbon substrate amendments so as to determine the physiological profiles of soil microbial communities by using whole soil. Appl Environ Microbiol. 2003;69:3593–9.
R Development Core Team. R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2018.
de Mendiburu F. Agricolae: statistical procedures for agricultural research. R Package Version. 2017;1:2–8.
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, et al. vegan: community ecology package. 2018.
Soetaert K. plot3D: plotting multi-dimensional data. R package version 1.0. 2013.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Huber W, Carey VJ, Gentleman R, Anders S, Carlson M, Carvalho BS, et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat Methods. 2015;12:115–21.
Paradis E, Claude J, Strimmer K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics. 2004;20:289–90.
Letunic I, Bork P. Interactive Tree of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res. 2011;39:W475–8.
Chiquet J, Mariadassou M, S. R. Variational inference for sparse network reconstruction from count data. ICML. 2018;97:1162–71.
Liu H, Roeder K, Wasserman L. Stability Approach to Regularization Selection (StARS) for high dimensional graphical models. Adv Neural Inf Process Syst. 2010;31:1432–40.
Chen L, Reeve J, Zhang LJ, Huang SB, Wang XF, Chen J. GMPR: a robust normalization method for zero-inflated count data with application to microbiome sequencing data. PeerJ. 2018;6:e4600.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Rohart F, Gautier B, Singh A, Le Cao KA. mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol. 2017;13:e1005752.
Singh A, Gautier B, Shannon CP, Rohart F, Vacher M, Tebutt SJ, et al. DIABLO: from multi-omics assays to biomarker discovery, an integrative approach. Bioinformatics. 2019;35:3055–62.
Calderon K, Spor A, Breuil MC, Bru D, Bizouard F, Violle C, et al. Effectiveness of ecological rescue for altered soil microbial communities and functions. ISME J. 2017;11:272–83.
Hol WHG, de Boer W, de Hollander M, Kuramae EE, Meisner A, van der Putten WH. Context dependency and saturating effects of loss of rare soil microbes on plant productivity. Front Plant Sci. 2015;6:485.
Weber MF, Poxleitner G, Hebisch E, Frey E, Opitz M. Chemical warfare and survival strategies in bacterial range expansions. J Royal Soc Interface. 2014;11:20140172.
Fierer N, Bradford MA, Jackson RB. Toward an ecological classification of soil bacteria. Ecology. 2007;88:1354–64.
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R. Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J. 2012;6:1007–17.
Kurm V, van der Putten WH, de Boer W, Naus-Wiezer S, Hol WHG. Low abundant soil bacteria can be metabolically versatile and fast growing. Ecology. 2017;98:555–64.
Berns AE, Philipp H, Narres HD, Burauel P, Vereecken H, Tappe W. Effect of gamma-sterilization and autoclaving on soil organic matter structure as studied by solid state NMR, UV and fluorescence spectroscopy. Eur J Soil Sci. 2008;59:540–50.
Ghoul M, Mitri S. The ecology and evolution of microbial competition. Trends Microbiol. 2016;24:833–45.
Delgado-Baquerizo M, Oliverio AM, Brewer TE, Benavent-Gonzalez A, Eldridge DJ, Bardgett RD, et al. A global atlas of the dominant bacteria found in soil. Science. 2018;359:320–5.
Jones SE, Lennon JT. Dormancy contributes to the maintenance of microbial diversity. Proc Natl Acad Sci USa. 2010;107:5881–6.
Kurm V, Geisen S, Hol WHG. A low proportion of rare bacterial taxa responds to abiotic changes compared with dominant taxa. Environ Microbiol. 2019;21:750–8.
Garbeva P, Hordijk C, Gerards S, de Boer W. Volatile-mediated interactions between phylogenetically different soil bacteria. Front Microbiol. 2014;5:289.
Karimi B, Terrat S, Dequiedt S, Saby NPA, Horriguel W, Lelievre M, et al. Biogeography of soil bacteria and archaea across France. Sci Adv. 2018;4:eaat1808.
Lewin GR, Carlos C, Chevrette MG, Horn HA, McDonald BR, Stankey RJ, et al. Evolution and ecology of actinobacteria and their bioenergy applications. Annu Rev Microbiol. 2016;70:235–54.
Prosser JI, Nicol GW. Archaeal and bacterial ammonia-oxidisers in soil: the quest for niche specialisation and differentiation. Trends Microbiol. 2012;20:523–31.
Daims H, Lebedeva EV, Pjevac P, Han P, Herbold C, Albertsen M, et al. Complete nitrification by Nitrospira bacteria. Nature. 2015;528:504–9.
Sorokin DY, Luecker S, Vejmelkova D, Kostrikina NA, Kleerebezem R, Rijpstra WIC, et al. Nitrification expanded: discovery, physiology and genomics of a nitrite-oxidizing bacterium from the phylum Chloroflexi. ISME J. 2012;6:2245–56.
Bell T. Next-generation experiments linking community structure and ecosystem functioning. Environ Microbiol Rep. 2019;11:20–2.
Acknowledgements
This research was funded through the 2016 ERA-NET BiodivERsA COFUND call for research proposals, with the national funders Agence Nationale de la Recherche (ANR, France; grant ANR-16-EBI3-0004-01) and the Swedish Research Council (FORMAS, Sweden). The authors would like to thank Noah Fierer for helpful discussion and valuable comments on the manuscript.
Author information
Authors and Affiliations
Contributions
LP, SR, SH, and AS designed the experiments. SR, M-CB, DB, and MT carried out the experiments. JA, SO, AM, AS, and SR analyzed the data. SR and LP drafted the manuscript and all the authors contributed to revisions and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Romdhane, S., Spor, A., Aubert, J. et al. Unraveling negative biotic interactions determining soil microbial community assembly and functioning. ISME J 16, 296–306 (2022). https://doi.org/10.1038/s41396-021-01076-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01076-9
This article is cited by
-
Manipulating the physical distance between cells during soil colonization reveals the importance of biotic interactions in microbial community assembly
Environmental Microbiome (2024)
-
Lose-lose consequences of bacterial community-driven invasions in soil
Microbiome (2024)
-
The interplay between microbial communities and soil properties
Nature Reviews Microbiology (2024)
-
Advances in Microbe-Driven Ecological Restoration on Bauxite Residue Disposal Areas
Reviews of Environmental Contamination and Toxicology (2024)
-
Modeling Microbial Community Networks: Methods and Tools for Studying Microbial Interactions
Microbial Ecology (2024)