Abstract
The access of rhizobia to legume host is a prerequisite for nodulation. Rhizobia are poorly motile in soil, while filamentous fungi are known to grow extensively across soil pores. Since root exudates-driven bacterial chemotaxis cannot explain rhizobial long-distance dispersal, mycelia could constitute ideal dispersal networks to help rhizobial enrichment in the legume rhizosphere from bulk soil. Thus, we hypothesized that mycelia networks act as vectors that enable contact between rhizobia and legume and influence subsequent nodulation. By developing a soil microcosm system, we found that a facultatively biotrophic fungus, Phomopsis liquidambaris, helps rhizobial migration from bulk soil to the peanut (Arachis hypogaea) rhizosphere and, hence, triggers peanut–rhizobium nodulation but not seen in the absence of mycelia. Assays of dispersal modes suggested that cell proliferation and motility mediated rhizobial dispersal along mycelia, and fungal exudates might contribute to this process. Furthermore, transcriptomic analysis indicated that genes associated with the cell division, chemosensory system, flagellum biosynthesis, and motility were regulated by Ph. liquidambaris, thus accounting for the detected rhizobial dispersal along hyphae. Our results indicate that rhizobia use mycelia as dispersal networks that migrate to legume rhizosphere and trigger nodulation. This work highlights the importance of mycelial network-based bacterial dispersal in legume–rhizobium symbiosis.
Similar content being viewed by others
Introduction
The spatial distribution of bacteria significantly influences their ecosystem services, for instance the interaction with host plants. Soil nitrogen-fixing bacteria (rhizobia) can establish beneficial interactions with legume plants and fix atmospheric dinitrogen (N2) by forming a specialized organ, the nodule [1,2,3]. The access of rhizobia to the legume determines nodule initiation [4, 5]. Yet, rhizobia are poorly motile in soil, and nodulating rhizobia, except for seed-borne rhizobia, are derived from soil environment. Especially, root exudates mediate a short-distance recruitment (a few millimeters) of rhizobia, rhizobia, which are separated from legume hosts by a long distance that cannot reach their targets [6]. Moreover, it is impossible for rhizobia to cross the air-filled gaps between soil aggregates via flagellum-mediated active movement without the facilitation of flowing water or other vectors [5, 7]. Thus, it is still unknown how rhizobia migrate to legume rhizosphere from bulk soil.
Beneficial fungi could interact with rhizobia-infected legumes and produce additional effects, such as nutrient acquisition and plant fitness, in a relationship known as tripartite mutualism [8,9,10,11]. Simultaneous colonization of legumes with fungal and rhizobial symbionts is likely common in nature, as fungi and bacteria live together and their synchronous root infection is inevitable [12, 13]. Studies have reported that beneficial fungi enhance legume–rhizobium interactions via nutrient complementarity or plant fitness improvement [9, 14]. Nothing is known about the direct interactions between microbial symbionts before symbiotic establishment or the concomitant effects on subsequent rhizobial associations with legume hosts. Fungal mycelia, representing most of the soil microbial biomass, are demonstrated to grow extensively in soil pores and form a mycelial network, which is known as the “fungal highway” [15]. Such mycelia are also known as an ideal channel for bacterial dispersal in structurally and chemically heterogeneous soil ecosystems. Indeed, it is found that specific bacteria can act as “hitchhikers” traveling along mycelial networks to new niches, where they play important ecological functions, such as bioremediation, foraging, biocontrol, and nutrient cycling [15,16,17,18].
The filamentous ascomycetes Phomopsis spp. are widely distributed in the natural environment, and contribute to ecosystem functions, such as litter decomposition and nutrient cycling [19, 20]. Our previous studies showed that a facultatively biotrophic fungus, Phomopsis liquidambaris, could establish a symbiotic association with peanut (Arachis hypogaea) and enhance nodulation and N2-fixation under controlled and field-realistic conditions [8,9,10,11, 21, 22]. Several mechanisms have been elucidated regarding Ph. liquidambaris-mediated nodulation enhancement, including plant hormone modifications [11], leaf CO2 fixation increasement [9], and soil microflora improvement [22]. Although no direct interactions between two root symbionts and concomitant effects on nodulation are known, Ph. liquidambaris inoculation increased bradyrhizobial diversity in peanut rhizosphere [22], suggesting the possible role of Ph. liquidambaris in rhizobial rhizosphere enrichment. Recent studies demonstrated that mycelia are able to redistribute water and nutrients and release water and carbon (C)-rich compounds to the mycosphere to create a hospitable microhabitat for bacteria [23,24,25]. In addition to its endophytic lifestyle, Ph. liquidambaris is able to degrade soil recalcitrant organic matter and survive in harsh environments [26,27,28,29,30]. Because of the strong saprophytic ability, Ph. liquidambaris could release substantial nutrients to the mycosphere, which may mediate its interaction with rhizobia [31]. Thus, an interaction between Ph. liquidambaris and rhizobia before their symbiosis with legume hosts may exist and participate in Ph. liquidambaris-mediated nodulation enhancement. Here, we hypothesize that mycelial networks can be used by rhizobia as a means to reach the rhizosphere from bulk soil, increase the contact frequency with legumes.
In the present study, using Ph. liquidambaris–peanut–rhizobia tripartite mutualism as a model, we first developed a soil microcosm, where rhizobium was inoculated at some distance away from the plant to study if mycelia of Ph. liquidambaris could transfer rhizobia from bulk soil to the rhizosphere and the concomitant effects on legume–rhizobia interactions. Next, we performed a series of plate experiments to study the dispersal modes of rhizobia along mycelial networks. Last, we performed transcriptome sequencing to reveal the potential genetic mechanisms of rhizobial dispersal on Ph. liquidambaris networks. The obtained results indicate that mycelial networks are important means for rhizobial migration from bulk soil to rhizosphere, and thereby initiate legume–rhizobium interaction.
Materials and methods
Microorganisms and growth media
Experiments were performed with Ph. liquidambaris strain B3 and peanut-nodulating rhizobium, Bradyrhizobium yuanmingense (hereafter Bradyrhizobium) [8, 22]. To mimic soil conditions, soil extraction agar (SEA, 1.5% agar) was used to activate Ph. liquidambaris [32]. A fungal plug was transferred to SEA and placed in an incubator at 28 °C for 7 days. The Bradyrhizobium was activated as in [8].
Soil microcosm design
Peanut (line Gan-Hua) seeds were obtained from the Ecological Experimental Station of Red Soil, China Academy of Science. After surface disinfection, seeds were germinated on 0.5 PDA (1%, w/v) at 28 °C until the radicle reached ~2 cm. The seedlings without microbial contamination were used. The soil was collected from the surface (0–10 cm) of a peanut paddock at Botany Garden of Nanjing Normal University (32° 6.318′ N, 118° 54.88′ E), China, and mixed thoroughly after passing through a 2 mm sieve. The main physicochemical properties and gravimetrical moisture have been described [9]. Each microcosm (12 cm length, 5 cm width, 12 cm height) contained 1.5 kg gramma-sterilized soil, and soil was kept at its original gravimetrical moisture (16%, w/w) with sterile water. Each microcosm contained two compartments, microbial and root, which were separated by a sterile 30 μm mesh [33]. The microcosm experiments contained the following treatments: (1) control setup of the microcosm without microbial inoculation or peanut cultivation; (2) microbial compartment with soil suspension inoculation and root compartment with peanut cultivation; (3) microbial compartment with Ph. liquidambaris and soil suspension inoculation and root compartment with peanut cultivation; (4) microbial compartment with Bradyrhizobial inoculation and root compartment with peanut cultivation; (5) microbial compartment with Ph. liquidambaris and Bradyrhizobial inoculation and root compartment with peanut cultivation; (6) microbial compartment with Bradyrhizobial inoculation and root compartment without peanut cultivation; (7) microbial compartment with Ph. liquidambaris and Bradyrhizobial inoculation and root compartment without peanut cultivation; and (8) microbial compartment without microbial inoculation and root compartment with peanut cultivation and Bradyrhizobial inoculation.
The preparation of soil suspension, microbial inoculation, collection of soil samples and determination of 16S rRNA [34], rhizobial symbiotic gene nodC [35], and housekeeping gene recA [36] abundance are described in the Supplementary Appendix. The primers for the target genes are listed in Supplementary Table S1.
Mycelia-based Bradyrhizobial dispersal assays
To monitor the attachment and dispersal of Bradyrhizobium on Ph. liquidambaris networks, a petri plate with a physical barrier was established. A sterile polystyrene ring (Ø 3.5 cm) was placed in the middle of the plate before SEA medium was poured. A fungal plug (Ø 5 mm) was inoculated in the middle of the ring and Bradyrhizobium was inoculated on the inner side of the ring with a bacterial inoculating loop. The plates with only Bradyrhizobial inoculation were also included as controls. The plates were then sealed with parafilm and placed at 28 °C for 7 days. The interactions of Ph. liquidambaris and Bradyrhizobium were observed by scanning electron microscope (JEOL, Japan). The experiments were carried out with three individual replicates, and representative scanning electron micrographs are shown.
Stage-dependent Ph. liquidambaris–Bradyrhizobial interaction assays
To study the interaction between Ph. liquidambaris and Bradyrhizobium at different stages simultaneously, five groups of a fungal plug (Ø 5 mm) and a 5-μl Bradyrhizobial suspension were inoculated on both sides of a square petri dish (13 × 13 cm) with SEA medium in diagonal rows, generating a reverse V-shape of increasingly far inoculation sites [37]. The Bradyrhizobial inoculation sites from near to far away from Ph. liquidambaris plugs were named S1–S5. Plates with only Bradyrhizobial inoculation were also created as controls (S0). The plates were then sealed with parafilm and placed at 28 °C for 7 and 14 days, respectively.
For Bradyrhizobial growth assays, the medium of each site from S0 to S5 was collected at 7 and 14 days after inoculation (dai). The medium was then mixed well with sterile water and spread on the surface of yeast mannitol agar plates (Ø 9 cm) supplemented with 30 mg l−1 natamycin after serial dilutions. Natamycin was used to inhibit the growth of Ph. liquidambaris. The plates were then sealed with parafilm and placed at 28 °C for 7 days. The number of Bradyrhizobial colonies on plates with 5–30 single colonies was counted. The experiment was carried out with three individual replicates.
For Bradyrhizobial motility assays, the medium of different sites was collected at 7 and 14 dai. Images and videos were directly taken with a Zeiss Axio Imager A1 microscope (Zeiss, Jena, Germany) [38]. The experiment was carried out with three individual replicates.
RNA-seq
For RNA-seq, Bradyrhizobia at S0 and S2 were sampled by scraping the medium surface with a sterile razor blade at 14 days after co-inoculation. Harvested cells were immediately frozen with liquid nitrogen and stored at −80 °C. The methods of RNA extraction, sequencing, and annotation are described in the Supplementary Appendix. The experiment was performed with three individual replicates. The RNA-seq reads are deposited in the Sequence Read Archive service of the GenBank database under the accession number SUB6177629.
RNA extraction and qRT-PCR analysis
Bradyrhizobium at each site from S0 to S5 after 14 days cocultivation with Ph. liquidambaris and Bradyrhizobium after fungal exudates (0 and 5 mg ml−1) incubation for 24 h were collected for RNA extraction. The RNA extraction and qRT-PCR procedures are described in the Supplementary Appendix. The Bradyrhizobial 16S rRNA gene [39] was used as reference gene to normalize the expression of selected genes using the 2−ΔΔCt method [40]. The experiment was performed with three individual replicates and three technical replicates, and sterile water was used as a negative control. The selected genes and primers are listed in Supplementary Table S1.
Statistical analyses
All experiments were performed in at least three individual replicates. The data are expressed as mean with standard error (SE). Statistical analyses were performed with SPSS v. 18.0 (SPSS Inc., Chicago, IL, USA) using one-way analysis of variance via Tukey’s honestly significant difference or Student’s t test.
Results
Soil microcosm: Ph. liquidambaris networks facilitate Bradyrhizobial dispersal in soil conditions
To determine whether Ph. liquidambaris can serve as a bridging network for rhizobial migration to peanut rhizosphere, a soil microcosm system was set up (Fig. 1a–c). Soil suspensions with or without Ph. liquidambaris were inoculated into the microbial compartment, which was separated from the peanut plant. In the presence of Ph. liquidambaris, rhizobial marker genes nodC and recA and Ph. liquidambaris-specific ITS were detected in peanut rhizosphere at 15 dai, which were not seen in the absence of Ph. liquidambaris (Fig. 1d; Fig. S1).
a, b A soil microcosm system containing microbial and root compartments was established to determine whether Ph. liquidambaris networks transfer rhizobia to the rhizosphere of peanut. The microbial and root compartments were separated by a sterile 30 μm mesh. Roots were confined to the root soil, but fungi and rhizobia in the microbial soil were able to cross the mesh and enter root soil. The treatments with peanut cultivation contained individual seedling planted in root compartment. Sterile 0.45 μm mesh in soil microcosm was used to observe Ph. liquidambaris–Bradyrhizobium interaction. c Microcosms were placed in transparent boxes with closed lids to minimize water evaporation and avoid microbial contamination. d 16S rRNA, nodC, and recA copies in the rhizosphere soil of peanut. At 15 dai, rhizosphere soil of peanut was collected to determine the copy numbers of 16S rRNA, nodC, and recA when the microbial compartment was inoculated with soil suspension with or without Ph. liquidambaris. Data and error bars are the mean ± SE (n = 4). CFUs colony-forming units; dai days after inoculation; IS inoculation site.
Furthermore, for determining Ph. liquidambaris-mediated rhizobial dispersal rates, a rhizobial species, B. yuanmingense, with or without Ph. liquidambaris was inoculated into the microbial compartment. Bradyrhizobium was quantified in soils at different distances from the inoculation site by colony-forming units (CFUs) and the abundances of nodC and recA, respectively. In the absence of mycelia, no Bradyrhizobium was detected beyond 15 mm away from the inoculation site at 9 dai (Fig. 2a, b; Fig. S2). By contrast, in the presence of mycelia, Bradyrhizobium reached 10–15 and 20–25 mm at 3 and 6 dai, respectively (Fig. 2a). At 9 dai, Bradyrhizobial colonies were detected in rhizosphere soil (Fig. 2a, b; Fig. S2). These results indicate that Ph. liquidambaris helped rhizobial migration to rhizosphere from bulk soil.
a Bradyrhizobial number was obtained by CFU determination from the soil at various collection distances at 3, 6, and 9 dai when the microbial compartment was inoculated with Bradyrhizobium with or without Ph. liquidambaris and the root compartment was planted with or without peanut. b Bradyrhizobial number was obtained by qPCR of nodC from the soil at various collection distances at 3, 6, and 9 dai when the microbial compartment was inoculated with Bradyrhizobium with or without Ph. liquidambaris and the root compartment was planted with or without peanut. Data and error bars are the mean ± SE (n = 4). CFU colony-forming units; dai days after inoculation.
We then tested the effects of the host plant on mycelia-based Bradyrhizobial dispersal in soil environment. In the absence of peanut, Ph. liquidambaris-mediated Bradyrhizobial dispersal could not be detected beyond 20 mm at 9 dai, which was less than that with peanut cultivation (Fig. 2a, b; Fig. S2), suggesting that host plants acted as attractants that direct mycelia-based Bradyrhizobial dispersal.
Soil microcosm: Ph. liquidambaris-based Bradyrhizobial dispersal triggers nodulation
To directly observe Ph. liquidambaris–Bradyrhizobium interaction in soil environment and on root surface, meshes (0.45 μm) and peanut roots were sampled for scanning electron microscopy at 9 dai. The meshes were used as the soil itself is not suitable for sample preparation procedure of scanning electron microscopy. No fungal network was observed in soil pores without Ph. liquidambaris inoculation (Fig. 3a; Fig. S3a, b). By contrast, Ph. liquidambaris extended vastly in soil pores, generating fungal networks to link soil aggregates after inoculation (Fig. 3b; Fig. S3c). Meanwhile, Ph. liquidambaris hyphae were able to reisolate from soil microcosm with Ph. liquidambaris and bradyrhizobial co-inoculation rather than the microcosms of control and bradyrhizobial inoculation (Fig. S4).
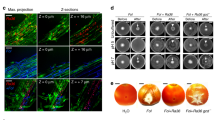
a SEM image showing no fungal network in soil pores of microcosm without Ph. liquidambaris inoculation. b SEM image showing fungal networks formed by Ph. liquidambaris in soil pores of microcosm with Ph. liquidambaris inoculation. c SEM image showing the attachment of Bradyrhizobium on Ph. liquidambaris hyphae. Bradyrhizobium was attached on the hyphal surface and formed fiber-like biofilms in soil. At 9 dai, the meshes (0.45 μm) were sampled for scanning electron microscopy to observe Ph. liquidambaris–Bradyrhizobium interaction in soil environment. d SEM image showing Ph. liquidambaris–Bradyrhizobium interaction on the root surface. e SEM image showing the root invasion of hyphae-attached Bradyrhizobium through crack sites (red arrow). At 9 dai, peanut roots were sampled for scanning electron microscopy to observe Ph. liquidambaris–Bradyrhizobium interaction on root surface. f Nodules on the lateral root of peanut in the presence of Ph. liquidambaris and Bradyrhizobium. No visible nodule was observed on the root in the absence of Ph. liquidambaris. g Toluidine blue-stained transverse section of nodules. h Higher-magnification images from g. The infected cells were well organized in the N2-fixation zone. i, j TEM images showing the symbiosomes and bacteroides. The bacteroides were well organized in symbiosomes. At 45 dai, root nodules were collected and prepared for transmission electron microscopy to observe nodule ultrastructure. The experiments were carried out with three individual replicates, and representative micrographs are shown. SEM scanning electron microscope; TEM transmission electron microscope; dai days after inoculation.
A mass of bradyrhizobium formed dense surface-attached biofilms on the hyphal surface (Fig. 3c). At the root surface, a tripartite interaction, including Bradyrhizobium, Ph. liquidambaris, and plant, was observed (Fig. 3d, e). Dense biofilms were also formed by Bradyrhizobium on hyphae at the root surface (Fig. 3e). Some hyphae began to infect roots via cracks on the root surfaces, which might facilitate the root invasion of attached Bradyrhizobium (Fig. 3e). When Bradyrhizobium was directly inoculated in root compartment, we observed that Bradyrhizobium attached to the cracks on the root surfaces (Fig. S5b). No Ph. liquidambaris hypha or Bradyrhizobium was found on root surface of Bradyrhizobial inoculation in microbial compartment (Fig. S5a).
At 45 dai, we harvested peanut roots and analyzed peanut–Bradyrhizobium nodulation. No visible nodule was formed on the root when Bradyrhizobium was inoculated in the microbial compartment alone. With both inoculation of Ph. liquidambaris and Bradyrhizobium in the microbial compartment, 2 ± 0.5 nodules per plant were formed on lateral roots (Fig. 3f; Supplementary Table S2). When Bradyrhizobium was directly inoculated in peanut rhizosphere or the root surface at the root compartment, 3 ± 0.75 nodules were formed per plant, mainly on the crown root (Fig. S6a; Supplementary Table S2). By histological and transmission electron microscope observations, these nodules showed a normal organization pattern, a central N2-fixation area and vascular bundles surrounding the nodule cortex (Fig. 3g; Fig. S6b). Plant cells that were infected by Bradyrhizobium were assembled tightly in the N2-fixation area (Fig. 3h; Fig. S6c). Moreover, the ultrastructure of symbiosomes and bacteroides was normal, and each symbiosome contained 48.70 ± 12.47 bacteroides (Fig. 3i, j; Supplementary Table S2), indicating that these nodules could fix N2. Similarly, effective nodules with 50.30 ± 10.55 bacteroides per symbiosome were observed when Bradyrhizobium was delivered directly into root compartment (Fig. S6d, e; Supplementary Table S2). These results suggest that Ph. liquidambaris-based Bradyrhizobial dispersal triggered peanut–Bradyrhizobium nodulation.
Plate interaction: Ph. liquidambaris networks help Bradyrhizobium dispersal
To address the direct interactions between Ph. liquidambaris and Bradyrhizobium, three different plate experiments were performed. First, the nested plate system showed that without Ph. liquidambaris inoculation, Bradyrhizobium was not able to cross the ring (Fig. 4a). By contrast, in the presence of Ph. liquidambaris, Bradyrhizobium crossed the ring and reached the outer compartment, i.e., site C (Fig. 4b). Scanning electron microscopy further detail the process: Bradyrhizobium was attached on the hyphal surface at A site, indicating that Bradyrhizobium migrated along established Ph. liquidambaris networks toward fungal plugs (Fig. 4c, d). Then, when the growing hyphae reached the Bradyrhizobial inoculation site, Bradyrhizobium migrated to the surface of hyphae from the growth medium and crossed the ring (Fig. 4e–h). Moreover, we found that Ph. liquidambaris was able to help other three peanut nodule-occupying rhizobia [21], cross the ring (Fig. S7).
a Plate with a sterile polystyrene ring with Bradyrhizobium inoculation alone. b Plate with a sterile polystyrene ring with Ph. liquidambaris and Bradyrhizobium co-inoculation. A–C sites refer to sampling sites for scanning electron microscopy. c, d SEM images showing Bradyrhizobium attached to and dispersed from mycelial networks of Ph. liquidambaris at A site. e, f SEM images showing Bradyrhizobium attached to and dispersal from mycelial networks at B site. g, h SEM images showing Bradyrhizobium attached to and dispersal from mycelial networks at C site. d, f, h Higher-magnification images from c, e, and g. The experiments were carried out with three individual replicates, and representative micrographs are shown. SEM scanning electron microscope.
The dispersal of Bradyrhizobium on mycelial networks detected in above-mentioned experiment could be due to the passive dispersal when Bradyrhizobium is pushed horizontally across surface by the growing hyphae as filamentous fungi extend by apical growth [38, 41]. Thus, by developing synthetic glass fiber networks simulating mycelial networks, we further determined whether Bradyrhizobium used active motility or passive dispersal along mycelia [38]. At 7 and 14 dai, Bradyrhizobium migrated out from the inoculation spot and followed the topology of the glass fiber networks (Fig. S8a, b). Although we could not rule out the possibility that capillary forces contribute to this process [42], these results suggested the involvement of bacterial active motility during Bradyrhizobial dispersal along mycelial networks. Third, existing fungal networks of Ph. liquidambaris was created to determine whether Bradyrhizobium could disperse on established fungal networks. At 7 dai, Bradyrhizobium rapidly dispersed across the existing fungal networks (Fig. S8c), with a 5.71-fold increase in dispersal distance, which further confirmed the involvement of bacterial active motility during Bradyrhizobial dispersal along mycelial networks.
Stage-dependent Ph. liquidambaris–Bradyrhizobial interaction assays
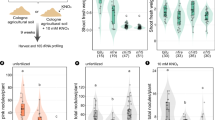
Ph. liquidambaris and Bradyrhizobium were inoculated on a square petri dish in a diagonal row, which enabled the simultaneous sampling of fungi and bacteria at different interactive stages (Fig. 5a). We first quantified the Bradyrhizobial CFUs at 7 and 14 dai. These two time points were chosen because Ph. liquidambaris and Bradyrhizobium began to interact at S2 and S3 at 7 and 14 dai, respectively (Fig. 5a; Fig. S9). Ph. liquidambaris networks increased Bradyrhizobial growth at S1 and S2 at 7 dai, and this positive effect at S1 was decreased at 14 dai, while the cell number of Bradyrhizobium at S3 was increased when Ph. liquidambaris reached the Bradyrhizobial colony at 14 dai. By contrast, no significant difference in Bradyrhizobial growth was found at S4 or S5 in relative to S0 (Fig. 5b).
a Ph. liquidambaris and Bradyrhizobium were inoculated on a square plate in a diagonal row. At 14 dai, Ph. liquidambaris began to interact with Bradyrhizobium at S3. At S4, Bradyrhizobium was attracted by Ph. liquidambaris before physical contact. The sketch refers to sampling sites of square plate above for microscopic observation. The big circle represents fungal plug and the small circle represents Bradyrhizobial colony. a–f in the sketch represent the sampling sites. a, b in the sketch represent the sampling sites at left and right of Bradyrhizobial colony. b Effects of Ph. liquidambaris on Bradyrhizobial growth at S0–S5 at 7 and 14 dai. Bradyrhizobial number was determined by CFU. Data and error bars are the mean ± SE (n = 3) and different letters indicate significant differences among different sites at the same dai (one-way analysis of variance with Tukey’s test, P < 0.05). c Different stages of Ph. liquidambaris–Bradyrhizobial interaction at 14 dai showing Bradyrhizobial dispersal along mycelial networks. Samples were collected from a and directly observed with a Zeiss Axio Imager A1 microscope. Bars, 50 μm. d Average number of Bradyrhizobium on hyphae of different sampling site of c. e Average ratio of motile and stationary Bradyrhizobium on hyphae of different sampling site of c. The number of total, motile, and stationary Bradyrhizobium on 20 μm hyphae was counted from images and time-lapse videos, respectively. The experiments were performed with three individual replicates and each replicate contained 15 images or time-lapse videos over 60 s. Data and error bars are the mean ± SE (n = 3) and different letters indicate significant differences among different sites (one-way analysis of variance with Tukey’s test, P < 0.05). dai days after inoculation.
We then captured the interactions of Ph. liquidambaris and Bradyrhizobium at macroscopic and microscopic scales at 14 dai. At this time, the Bradyrhizobium at the colony edge of S4 was stimulated toward the fungal plug (Fig. 5a), suggesting that Bradyrhizobium was attracted by Ph. liquidambaris before physical contact. To directly observe the Bradyrhizobial dispersal along mycelial networks, we used time-lapsed microscopy of Bradyrhizobium and Ph. liquidambaris at S1–S3. The interaction between Ph. liquidambaris and Bradyrhizobium at S3 was expected to represent the initial stage of interaction establishment, as fungal colony had just come into physical contact with the Bradyrhizobium [43]. At S3, Bradyrhizobium at the edge of the colony began to swim along hyphae toward Ph. liquidambaris (Fig. 5c; Movie S1). At S1 and S2, Ph. liquidambaris networks had already established interactions with Bradyrhizobium and facilitated their long-distance dispersal (Fig. 5a, c). When analyzing bacterial motility along the direction of mycelial growth, we found that pioneer Bradyrhizobium on the growing hyphae was still motile (Movies S2 and S3), but most Bradyrhizobium on the established mycelial networks had converted to stationary state (Movie S4). The Bradyrhizobial dispersal mode was supported by the quantitative data of motile and stationary Bradyrhizobial number on Ph. liquidambaris surface at S1 and S2. Along the direction of mycelial growth, the Bradyrhizobial number and ratio of stationary Bradyrhizobium decreased gradually (Fig. 5d, e), while the ratio of motile Bradyrhizobium was gradually increased (Fig. 5e).
Bradyrhizobium was attracted by Ph. liquidambaris at the “no-physical-contact” stage and Bradyrhizobial number was increased at the early interactive stage, indicating that hyphal exudates might be involved in this process. We then collected hyphal exudates of Ph. liquidambaris and tested their effects on Bradyrhizobial growth, biofilm formation, and chemotactic behaviors. The addition of hyphal exudates at 5 and 10 mg ml−1 showed higher Bradyrhizobial optical densities (ODs) than those in the control (Fig. 6a). Meanwhile, Bradyrhizobial biofilm formation and chemotaxis were significantly enhanced in the presence of hyphal exudates at 1, 5, and 10 mg ml−1 (Fig. 6b, c).
a Ph. liquidambaris exudates increased Bradyrhizobial growth. The OD590 values were recorded every 4 h for 78 h. b Ph. liquidambaris exudates increased Bradyrhizobial biofilm formation. c Chemotactic response of Bradyrhizobium toward Ph. liquidambaris exudates. The chemotactic response of Bradyrhizobium toward Ph. liquidambaris exudates was evaluated by capillary assay. Data and error bars are the mean ± SE (n = 3) and asterisks indicate significant differences between control and fungal exudates treatments (Student’s t test, *P < 0.05, **P < 0.01, ***P < 0.001). CFU colony-forming units.
In brief, these results indicated that Bradyrhizobial motility, growth, and proliferation were involved in their dispersal along mycelial networks while fungal exudates might contribute to this process.
Transcriptomic analysis of Bradyrhizobium to Ph. liquidambaris
To reveal the underlying mechanisms driving Bradyrhizobial dispersal on Ph. liquidambaris networks, Bradyrhizobium at S0 (B) and S2 (E + B) was sampled for RNA-seq at 14 dai. Bradyrhizobium at S2 was chosen, as bacteria at this stage showed high reproduction and motility in the presence of mycelial networks (Fig. 5b, c). Transcriptome analysis showed that Bradyrhizobium underwent a global gene expression shift in the presence of mycelia, with 831 differentially expressed genes (expression change of greater than twofold, P < 0.05), including 368 genes with increased expression levels and 463 genes with decreased expression levels (Supplementary Table S3). Gene Ontology (GO) enrichment analysis of differentially expressed genes was first performed. Most of differentially expressed genes with predicted function annotations were assigned to metabolism, environmental information processing, and cellular processes (Fig. 7a). Then, we performed KEGG pathway enrichment analysis for the differentially expressed genes. KEGG pathway analysis showed that amino acid and derivatives metabolism, carbohydrate metabolism, energy metabolism membrane transport, signal transduction, and cell motility were enriched (Fig. 7b). Within the significantly upregulated genes, 55 genes were associated with metabolism, including amino acids and derivatives (24), carbohydrates (23), cofactors and vitamins (8), nucleotides (6), and lipids (5). In total, 29 genes with significantly decreased expression were associated with metabolism, mainly that of carbohydrates (12), amino acids and derivatives (9), cofactors and vitamins (5), and lipids (3) (Fig. 7b; Supplementary Table S4). Consistently, several genes related to transport of nutrients in Bradyrhizobium were regulated, for instance glucose/mannose transport system substrate-binding protein, multiple sugar transport system ATP-binding protein, branched-chain amino acid transport system ATP-binding system, and phospholipid/cholesterol/gamma-HCH transport system permease protein (Supplementary Table S3).
a Number of differentially expressed genes of GO categories related to biological process, cellular component, and molecular function in Bradyrhizobium inoculation alone (B) vs Bradyrhizobium and Ph. liquidambaris co-inoculation (E + B). b KEGG analysis of differentially expressed genes in B vs E + B. c Expression of Bradyrhizobial differentially expressed genes associated with bacterial division at S0-S5 at 14 dai by RNA-seq and qRT-PCR. The responses of these differentially expressed genes to fungal exudates were also included. d Expression of Bradyrhizobial differentially expressed genes associated with bacterial chemotaxis, flagellar biosynthesis, and motility at S0–S5 at 14 dai by RNA-seq and qRT-PCR. The responses of these differentially expressed genes to fungal exudates were also included. e Bacterial swimming chemotaxis pathway, which includes sensor, transduction, and actuator modules. dai days after inoculation; FE fungal exudates.
Giving that the Bradyrhizobium at S2 had a high cell number and motility compared with those at S0, we expected the genes associated with cell division, the chemosensory system, flagellar biosynthesis and assembly, and motility would be differently regulated by Ph. liquidambaris networks. An increase in nucleotide metabolism and protein synthesis of Bradyrhizobium in cocultivations was determined in the presence of Ph. liquidambaris (Fig. 7c; Supplementary Table S5). The expression of genes associated with purine and pyrimidine metabolism, bacterial peptidoglycan biosynthesis and large-subunit ribosomal protein components was increased in cocultivations (Fig. 7c; Supplementary Table S5). Among the differentially expressed genes involved in bacterial chemotaxis and motility, methyl-accepting chemotaxis proteins (MCPs) (GA0061099_1001674, GA0061099_1003438, GA0061099_1002562), and chemotaxis signaling proteins CheA (GA_0061099_10071, GA0061099_10021), CheR (GA0061099_1006673), CheY (GA0061099_1006671), and FliC (GA0061099_1001941) were significantly upregulated, while genes encoding flagellar M-ring protein FliF (GA0061099_1001939), flagellar P-ring protein precursor Flgl (GA0061099_1001946), flagellar biosynthetic protein FlhB (GA0061099_1001952), and ribose transport system substrate-binding protein rbsB (GA0661099_1010150) were downregulated in the presence of Ph. liquidambaris (Fig. 7d, e; Supplementary Table S5). In addition, genes associated with biofilm formation (GA0061099_1001800, GA0061099_1007256, GA0061099_1002199) were increased in cocultivations (Supplementary Table S3).
To understand the dynamic transcriptional regulation of Bradyrhizobial motility and proliferation at different stages of interactions, we employed qRT-PCR to analyze the relative expression of genes associated with motility and proliferation at S1–S5 with respect to S0. The transcriptional responses underlying Bradyrhizobial motility and proliferation to fungal exudates were also included. The expression of selected genes of Bradyrhizobium at S2 using qRT-PCR showed similar trends to RNA-seq (Fig. S10; Supplementary Table S5). Most of cell division-related genes are induced at S1–S4 (Fig. 7c). The Bradyrhizobial chemosensory system had already been primed in cocultivations before physical contact, as some of the tested mcp and che genes, especially mcp (GA0061099_1003438) and cheR (GA0061099_1006673), were upregulated at S4 (Fig. 7d). The tested flagellar biosynthesis- and motility-related genes showed a stage-dependent expression pattern. The flagellar biosynthesis gene fliF, flgl and motility-related gene rbsB were upregulated at S3–S4, but decreased at S1–S2. The fliC was induced at S1–S4 (Fig. 7d). Most of the tested genes were significantly induced when Bradyrhizobium was incubated with Ph. liquidambaris exudates (5 mg ml−1) in respect to 0 mg ml−1 exudates (Fig. 7c, d), suggesting that fungal exudates might play an important role in the Ph. liquidambaris-mediated Bradyrhizobial gene expression shift.
The Bradyrhizobial detoxification process was not enhanced when the bacteria were grown with Ph. liquidambaris. Genes encoding glutathione synthase (GA0061099_1002255), glutathione S-transferase (GA0061099_1003480), most of multidrug efflux pumps and the NodT family of efflux pumps were decreased in cocultivations (Supplementary Table S3).
Discussion
Mycelia are effective dispersal networks that contribute to the maintenance of bacterial ecological functions, such as bioremediation and predation [15,16,17]. However, little is known about the role of mycelia-based bacterial dispersal in legume–rhizobium symbiosis. Previous studies showed that Ph. liquidambaris enhanced nodulation of peanut and increased population of Bradyrhizobium sp. in rhizosphere under continuous cropping system that is rhizobium-deficient [9, 22], suggesting the possible role of Ph. liquidambaris in rhizobial rhizosphere enrichment.
Here, we developed a microcosm system to test Bradyrhizobial migration in soil in the presence or absence of Ph. liquidambaris, and revealed that Ph. liquidambaris helped Bradyrhizobial migration to peanut rhizosphere and, hence, triggered nodulation. In addition, Ph. liquidambaris might facilitate the root invasion of hyphae-attached Bradyrhizobium through crack sites. Mycelia-based Bradyrhizobial crack infection has potential to increase subsequent nodulation as crack entry is essential for peanut–Bradyrhizobium nodulation [44, 45]. To gain insight on Ph. liquidambaris–Bradyrhizobium interaction, we performed Bradyrhizobial dispersal modes assays and transcriptomic analysis. The results indicated that bacterial proliferation and motility mediated Bradyrhizobial dispersal along Ph. liquidambaris, while fungal exudates might contribute to this process.
Dispersal mode analyses indicated that Bradyrhizobial cells on the growing and established hyphal networks shared different functions in new niche colonization. The ratio of motile and static Bradyrhizobium showed opposite trends along the direction of mycelial growth: the ratio of motile Bradyrhizobium was decreased and the ratio of static Bradyrhizobium was gradually increased. This suggested that Bradyrhizobium changed behavior along Ph. liquidambaris networks. The pioneer Bradyrhizobium on the growing hyphae was motile, which explored new niche with the growing hyphae. While a large number of Bradyrhizobium on the established hyphae was static and detached from hyphal surface for niche occupation. Thus, Bradyrhizobial cell proliferation and motility mediated its dispersal along mycelia.
Similar to the roles of root exudates in plant–microbe interactions, hyphae release a substantial amount of C-rich compounds, including sugars, organic acids, amino acids, and polyols to the mycosphere, where they mediate fungal–bacterial interactions [25, 31, 33, 46]. For example, arbuscular mycorrhizal fungus exudates stimulate the growth and phytate mineralization of phosphate-solubilizing bacteria [33, 46]. During Bradyrhizobial dispersal along mycelia, fungal exudates might contribute to this process with dual roles. Before physical contact, Bradyrhizobium was attracted by Ph. liquidambaris, suggesting that fungal exudates functioned as chemo-attractants providing information about a nearby nutrient-rich niches. At early interactive stage, Bradyrhizobial number was increased, indicating that Ph. liquidambaris provided nutrients for Bradyrhizobial reproduction. This conclusion was further supported by analyzing the responses of Bradyrhizobial growth and chemotactic behavior to Ph. liquidambaris exudates. As we did not quantify the Ph. liquidambaris exudates, we could not unambiguously identify the primary chemo-attractants and nutrients from Ph. liquidambaris that Bradyrhizobium utilized in the present study. Further works with a combination of transcriptomics and exometabolomics at different interactive stages would help to reveal the chemical exchange between interactive partners [47].
The transcriptomic analysis underpinned the results from soil microcosm experiment and dispersal mode assays. Bradyrhizobial genes associated with cell division were induced by Ph. liquidambaris, explaining the increase in cell number of Bradyrhizobium during its interaction with Ph. liquidambaris. Bradyrhizobial genes associated with carbohydrate, amino acid, and derivatives metabolism were induced by Ph. liquidambaris, suggesting that mycelia provide nutrients to support Bradyrhizobial proliferation, which is also reported in other fungal–bacterial interactions [46, 48]. In addition, most genes with functions assigned to production and conversion of energy were increased by mycelia, confirming that Bradyrhizobium utilized nutrients derived from fungi to raise their bioenergetic status [32, 49]. Biofilms are important for the fitness of bacteria [50] and have been reported to enable bacterial biomass accumulation during microbial interspecies interaction [51]. Correspondingly, genes associated with bacterial biofilm formation were induced when Bradyrhizobium was cocultivated with Ph. liquidambaris and biofilms were formed on the surface of mycelia by Bradyrhizobium and Ph. liquidambaris exudates stimulated Bradyrhizobial biofilm formation, suggesting that biofilm might contribute to Bradyrhizobial cell increase.
Bradyrhizobial genes associated with chemosensory system, flagellar biosynthesis, and motility were induced at the early interactive stage, highlighting the role of chemotaxis and flagellum-mediated motility in Bradyrhizobial dispersal along Ph. liquidambaris hyphae [32]. This combined with Bradyrhizobial dispersal along glass fiber networks and established mycelial networks suggested that Bradyrhizobium swim and/or swarm along mycelial networks. The transmembrane chemoreceptors MCPs were induced before physical contact, suggesting that Bradyrhizobium had already sensed Ph. liquidambaris. This notion was further confirmed by the activation of chemotactic signal transduction system and flagellar biosynthesis process, especially methyltransferase CheR, histidine kinase CheA, and FliC. CheR catalyzes the conversion of specific glutamyl residues to glutamyl methyl esters on the cytosolic side of MAPs and induces the autophosphorylation of CheA, which increases bacterial tumbling [51,52,53].
Unlike one study of [48], which reported that the detoxification process of Burkholderia was activated when cocultivated with A. alternate or F. solani, our results showed that genes associated with antioxidant system and drug efflux pumps of Bradyrhizobium were not enhanced in the presence of Ph. liquidambaris. One possible explanation for the differences is that Ph. liquidambaris did not produce any antimicrobial compounds during its interactions with Bradyrhizobium. In addition, we cannot rule out the involvement of other detoxification pathways. Compared with other soil bacteria, the rhizobial genome, especially that of Bradyrhizobium, is large and diverse, contributing to their lifestyle switches and environmental adaptations [54, 55]. Thus, the genomic traits of rhizobia probably confer their ability to degrade and utilize fungal-derived secondary metabolites, which are detrimental to other soil bacteria. In addition, detoxification is an energy-consuming process and might be shut off, which saves the resources for Bradyrhizobial growth and motility. Together, these results suggest that Bradyrhizobial chemosensory system, flagellum-mediated motility, and cell proliferation participated in their dispersal and growth on Ph. liquidambaris networks, which might be mediated by hyphal exudates.
The dispersal of rhizobia in soil environment is poor [56], and rhizobia are deficient and not uniform in degraded soil, for instance, acidic soil, saline soil, and arid soil [57]. Thus, the nodulation of legumes under these soil environment is normally deficient [57, 58]. To increase the symbiotic potential of legumes in rhizobially deficient environment, rhizobial inoculants have been introduced to legumes via on-seed (seed-applied) or in-furrow (soil-applied) methods [4, 59, 60]. On-seed inoculation can deliver rhizobia to the rhizosphere of the emerging root, but in-furrow inoculation has more advantages, including separating the rhizobia from seed-applied bactericides and insecticides, avoiding the damage to seeds, and increasing the rhizobial number in the soil of legumes, which germinate epigeally, such as peanut, soybean, and subterranean clover [59, 60]. However, in-furrow inoculation increases the average distance between rhizobia and legume hosts. Our result that mycelia transfer rhizobia to the rhizosphere and, hence, trigger peanut–rhizobium symbiosis provide evidence that this drawback could be effectively solved by in-furrow co-inoculation of rhizobia and beneficial fungi. Moreover, filamentous fungi are better adapted to harsh soil environment than bacteria [23, 25, 61]. Thus, co-application of rhizobia and beneficial fungi could improve the nodulation competitiveness of inoculated rhizobia over native soil rhizobia, as fungal–bacterial interactions increase bacterial adaptive plasticity and survival in the soil environment by providing microhabitats and nutrients [18, 24].
Conclusions
Our study indicates that mycelial networks transfer rhizobia to the rhizosphere from bulk soil and, hence, initiate legume–rhizobium interaction. Mycelia-based enrichment of rhizobia in the rhizosphere is probably common in the soil environment, as a relatively high proportion of saprotrophic fungi also have facultative biotrophic capacity and mycosphere is a hospitable microhabitat for rhizobia. A recent study of 201 species of wood-decay basidiomycetes and a microcosm system showed that 16.9% (34) of tested fungi could enter into facultative biotrophic relationships with plant roots [62, 63]. In addition to nutrients provision, the mycosphere is a weakly acidic environment, which favors the colonization of rhizobia, especially the Bradyrhizobium genus [64, 65]. Our study reveals long-distance rhizobial enrichment to the rhizosphere, which cannot be mediated by root exudates. Moreover, our study provides a theoretical basis for the practical use of beneficial fungi and rhizobia in combinations to improve the symbiotic efficiency of leguminous crops in degraded lands.
References
Zipfel C, Oldroyd GED. Plant signalling in symbiosis and immunity. Nature. 2017;543:328–36.
Mus F, Crook MB, Garcia K, Costas AG, Geddes BA, Kouri ED, et al. Symbiotic nitrogen fixation and the challenges to its extension to nonlegumes. Appl Environ Microbiol. 2016;82:3698–710.
Burchill W, James EK, Li D, Lanigan GJ, Williams M, Iannetta PPM, et al. Comparisons of biological nitrogen fixation in association with white clover (Trifolium repens L.) under four fertilizer nitrogen inputs as measured using two 15N techniques. Plant Soil. 2014;385:287–302.
López-García SL, Perticari A, Piccinetti C, Ventimiglia L, Arias N, De Battista JJ, et al. In-furrow inoculation and selection for higher motility enhances the efficacy of Bradyrhizobium japonicum nodulation. Agron J. 2009;101:357–63.
Vicario JC, Dardanelli MS, Giordano W. Swimming and swarming motility properties of peanut-nodulating rhizobia. FEMS Microbiol Lett. 2015;362:1–6.
Bais HP, Park SW, Weir TL, Callaway RM, Vivanco JM. How plants communicate using the underground information superhighway. Trends Plant Sci. 2004;9:26–32.
Horiuchi J, Prithiviraj B, Bais HP, Kimball BA, Vivanco JM. Soil nematodes mediate positive interactions between legume plants and rhizobium bacteria. Planta. 2005;222:848–57.
Zhang W, Wang HW, Wang XX, Xie XX, Siddikee MA, Xu RS, et al. Enhanced nodulation of peanut when co-inoculated with fungal endophyte Phomopsis liquidambari and bradyrhizobium. Plant Physiol Biochem. 2016;98:1–11.
Zhang W, Wang XX, Yang Z, Ashaduzzaman SM, Kong MJ, Lu LY, et al. Physiological mechanisms behind endophytic fungus Phomopsis liquidambari-mediated symbiosis enhancement of peanut in a monocropping system. Plant Soil. 2017;416:325–42.
Xie XG, Fu WQ, Zhang FM, Shi XM, Zeng YT, Li H, et al. The endophytic fungus Phomopsis liquidambari increases nodulation and N2 fixation in Arachis hypogaea by enhancing hydrogen peroxide and nitric oxide signalling. Microb Ecol. 2017;74:427–40.
Zhang W, Sun K, Shi RH, Yuan J, Wang XJ, Dai CC. Auxin signalling of Arachis hypogaea activated by colonization of mutualistic fungus Phomopsis liquidambari enhances nodulation and N2-fixation. Plant Cell Environ. 2018;41:2093–108.
van Overbeek LS, Saikkonen. Impact of bacterial-fungal interactions on the colonization of the endosphere. Trends Plant Sci. 2016;21:230–42.
Deveau A, Bonito G, Uehling J, Paoletti M, Becker M, Bindschedler S, et al. Bacterial-fungal interactions: ecology, mechanisms and challenges. FEMS Microbiol Rev. 2018;42:335–52.
Ossler JN, Zielinski CA, Heath KD. Tripartite mutualism: facilitation or trade-offs between rhizobial and mycorrhizal symbionts of legume hosts. Am J Bot. 2015;102:1332–41.
Kohlmeier S, Smits THM, Ford RM, Keel C, Harms H, Wick LY. Taking the fungal highway: mobilization of pollutant-degrading bacteria fungi. Environ Sci Technol. 2005;39:4640–6.
Worrich A, König S, Miltner A, Banitz T, Centler F, Frank K, et al. Mycelium-like networks increase bacterial dispersal, growth, and biodegradation in a model ecosystem at various water potentials. Appl Environ Microbiol. 2016;82:2902–8.
Otto S, Bruni EP, Harms H, Wick LY. Catch me if you can: dispersal and foraging of Bdellovibrio bacteriovorus 109J along mycelia. ISME J. 2017;11:386–93.
Hassani MA, Durán P, Hacquard S. Microbial interactions within the plant holobiont. Microbiome. 2018;6:58.
Porras-Alfaro A, Bayman P. Hidden fungi, emergent properties: endophytes and microbiomes. Annu Rev Phytopathol. 2011;49:291–315.
Zhou J, Li X, Huang PW, Dai CC. Endophytism or saprophytism: decoding the lifestyle transition of the generalist fungus Phomopsis liquidambari. Microbiol Res. 2018;206:99–112.
Wu JR, Xu FJ, Cao W, Zhang W, Guan YX, Dai CC. Fungal endophyte Phomopsis liquidambaris B3 enriches the diversity of nodular culturable endophytic bacteria associated with continuous cropping of peanut. Arch Agronom Soil Sci 2019;65:240–52.
Xie XG, Zhang FM, Yang T, Chen Y, Li XG, Dai CC. Endophytic fungus drives nodulation and N2 fixation attributable to specific root exudates. mBi.o 2019;10:e00728–19.
Guhr A, Borken W, Spohn M, Matzner E. Redistribution of soil water by a saprotrophic fungus enhances carbon mineralization. Proc Natl Acad Sci USA. 2015;112:14647–51.
Nazir R, Warmink JA, Boersma H, Van Elsas JD. Mechanisms that promote bacterial fitness in fungal-affected soil microhabitats. FEMS Microbiol Ecol. 2009;71:169–85.
Worrich A, Stryhanyuk H, Musat N, König S, Banitz T, Centler F, et al. Mycelium-mediated transfer of water and nutrients stimulates bacterial activity in dry and oligotrophic environments. Nat Commun. 2017;8:15472.
Chen Y, Peng Y, Dai CC, Ju Q. Biodegradation of 4-hydroxybenzoic acid by Phomopsis liquidambari. Appl Soil Ecol. 2011;51:102–10.
Chen Y, Wang HW, Li L, Dai CC. The potential application of the endophyte Phomopsis liquidambaris to the ecological remediation of long-term cropping soil. Appl Soil Ecol. 2013;67:20–26.
Xie XG, Dai CC. Degradation of a model pollutant ferulic acid by the endophytic fungus Phomopsis liquidambari. Bioresour Technol. 2015;179:35–42.
Wang HW, Zhang W, Su CL, Zhu H, Dai CC. Biodegradation of the phytoestrogen luteolin by the endophytic fungus Phomopsis liquidambari. Biodegradation. 2015;26:197–210.
Sun K, Cao W, Hu LY, Fu WQ, Gong JH, Kang N, et al. Symbiotic fungal endophyte Phomopsis liquidambari-rice system promotes nitrogen transformation by influencing below-ground straw decomposition in paddy soil. J Appl Microbiol. 2019;126:191–203.
de Boer W, Folman LB, Summerbell RC, Boddy L. Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol Rev. 2005;29:795–811.
Haq IU, Dini-Andreote F, van Elsas JD. Transcriptional responses of the bacterium Burkholderia terrae BS001 to the fungal host Lyophyllum sp. strain Karsten under soil-mimicking conditions. Microb Ecol. 2017;73:236–52.
Zhang L, Xu M, Liu Y, Zhang F, Hodge A, Feng G. Carbon and phosphorus exchange may enable cooperation between an arbuscular mycorrhizal fungus and a phosphate-solubilizing bacterium. N. Phytol. 2016;210:1022–32.
Jiang Y, Liu M, Zhang J, Chen Y, Chen L, Li H, et al. Nematode grazing promotes bacterial community dynamics in soil at the aggregate level. ISME J. 2017;11:2705–17.
Sarita S, Sharma PK, Priefer UB, Prell J. Direct amplification of rhizobial nodC sequences from soil total DNA and comparison to nodC diversity of root nodule isolates. FEMS Microbiol Ecol. 2005;54:1–11.
Stępkowski T, Moulin L, Krzyżańska A, Mclnnes A, Law IJ, Howieson J. European origin of Bradyrhizobium populations infecting lupins and serradella in soils of western Australia and South Africa. Appl Environ Microbiol. 2005;71:7041–52.
Berendsen RL, Vismans G, Yu K, Song Y, Jonge R, Burgman WP, et al. Disease-induced assemblage of a plant-beneficial bacterial consortium. ISME J. 2018;12:1496–507.
Zhang Y, Kastman EK, Guasto JS, Wolfe BE. Fungal networks shape dynamics of bacterial dispersal and community assembly in cheese rind microbiomes. Nat Commun. 2018;9:336.
Liu Y, Jiang X, Guan D, Zhou W, Ma M, Zhao B, et al. Transcriptional analysis of genes involved in competitive nodulation in Bradyrhizobium diazoefficiens at the presence of soybean root exudates. Sci Rep. 2017;7:10946.
Schmittgen TD, Livak KJ. Analysing real-time PCR data by the comparative C (T) method. Nat Protoc. 2008;3:1101–8.
Riquelme M. Tip growth in filamentous fungi: a road trip to the apex. Annu Rev Microbiol. 2013;67:587–607.
Jost D, Winter J, Gallert C. Distribution of aerobic motile and non-motile bacteria within the capillary fringe of silica sand. Water Res. 2010;44:1279–87.
Mondo SJ, Lastovetsky OA, Gaspar ML, Schwardt NH, Barber CC, Riley R, et al. Bacterial endosymbionts influence host sexuality and reveal reproductive genes of early divergent fungi. Nat Commun. 2017;8:1843.
Boogerd FC, van Rossum D. Nodulation of groundnut by Bradyrhizobium: a simple infection process by crack entry. FEMS Microbiol Rev. 1997;21:5–27.
Ibáñez F, Wall L, Fabra A. Starting points in plant-bacteria nitrogen-fixing symbioses: intercellular invasion of the roots. J Exp Bot. 2017;68:1905–18.
Zhang L, Feng G, Declerck S. Signal beyond nutrient, fructose, exuded by an arbuscular mycorrhizal fungus triggers phytate mineralization by a phosphate solubilizing bacterium. ISME J. 2018;12:2339–51.
Zhalnina K, Louie KB, Hao Z, Mansoori N, da Rocha UN, Shi S, et al. Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat Microbiol. 2018;3:470.
Stopnisek N, Zühlke D, Carlier A, Barberán A, Fierer N, Becher D, et al. Molecular mechanisms underlying the close association between soil Burkholderia and fungi. ISME J. 2016;10:253–64.
Deveau A, Barret M, Diedhiou AG, Leveau J, de Boer W, Martin F, et al. Pairwise transcriptomic analysis of the interactions between the ectomycorrhizal fungus laccaria bicolor S238N and three beneficial, neutral and antagonistic soil bacteria. Micro Ecol. 2015;69:146–59.
Flemming HC, Wingender J, Szewzyk U, Steinberg P, Rice SA, Kjelleberg S. Biofilms: an emergent form of bacterial life. Nat Rev Microbiol. 2016;14:563–75.
Hansen SK, Rainey PB, Haagensen JA, Molin S. Evolution of species interactions in a biofilm community. Nature. 2007;445:533–6.
Sampedro I, Parales RE, Krell T, Hill JE. Pseudomonas chemotaxis. FEMS Microbiol Rev. 2014;39:17–46.
Springer WR, Koshland DE. Identification of a protein methyltransferase as the cheR gene product in the bacterial sensing system. Proc Natl Acad Sci USA. 1997;74:533–7.
Tian CF, Zhou YJ, Zhang YM, Li QQ, Zhang YZ, Li DF, et al. Comparative genomics of rhizobia nodulating soybean suggests extensive recruitment of lineage-specific genes in adaptations. Proc Natl Acad Sci USA. 2012;109:8629–34.
Poole P, Ramachandran V, Terpolilli J. Rhizobia: from saprophytes to endosymbionts. Nat Rev Microbiol. 2018;16:291.
Lowther WL, Patrick HN. Spread of Rhizobium and Bradyrhizobium in soil. Soil Biol Biochem. 1993;25:607–12.
de la Peña TC, Pueyo JJ. Legume in the reclamation of marginal soils, from cultivar and inoculant selection to transgenic approaches. Agron Sustain Dev. 2012;32:65–91.
Bordeleau LM, Prévost D. Nodulation and nitrogen fixation in extreme environments. Plant Soil. 1994;161:115–25.
Denton MD, Phillips LA, Peoples MB, Pearce DJ, Swan AD, Mele PM, et al. Legume inoculant application methods: effects on nodulation patterns, nitrogen fixation, crop growth and yield in narrow-leaf lupin and faba bean. Plant Soil. 2017;419:25–39.
Deaker R, Roughley RJ, Kennedy IR. Legume seed inoculation technology-a review. Soil Biol Biochem. 2004;36:1275–88.
Schimel J, Balser TC, Wallenstein M. Microbial stress-response physiology and its implications for ecosystem function. Ecology. 2007;88:1386–94.
Smith GR, Finlay RD, Stenlid J, Vasaitis R, Menkis A. Growing evidence for facultative biotrophy in saprotrophic fungi: data from microcosm tests with 201 species of wood-decay basidiomycetes. N. Phytol. 2017;215:747–55.
Baldrian P, Kohout P. Interactions of saprotrophic fungi with tree roots: can we observe the emergence of novel ectomycorrhizal fungi? N. Phytol. 2017;215:747–55.
Man CX, Wang H, Chen WF, Sui XH, Wang ET, Chen WX. Diverse rhizobia associated with soybean grown in the subtropical and tropical regions of China. Plant Soil. 2008;310:77–87.
Zhang YM, Li Y Jr, Chen WF, Wang ET, Tian CF, Li QQ, et al. Biodiversity and biogeography of rhizobia associated with soybean plants grown in the North China Plain. Appl Environ Microbiol. 2011;77:6331–42.
Acknowledgements
We thank Yin-Ping Zhang and Chao-Chao Hu of Analytical & Testing Center, Nanjing Normal University, for their help with the microscopic imaging. This work was financially supported by the National Natural Science Foundation of China (NSFC no. 31870478; 31370507); the Priority Academic Program Development of Jiangsu Higher Education Institutions; and Doctor Breeding Project of Nanjing Normal University (1812000006317).
Author information
Authors and Affiliations
Contributions
WZ, XGL, and CCD designed the experiments, analyzed the data, and wrote and revised the manuscript. WZ, KS, MJT, FJX, and MZ performed experiments. All of authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, W., Li, XG., Sun, K. et al. Mycelial network-mediated rhizobial dispersal enhances legume nodulation. ISME J 14, 1015–1029 (2020). https://doi.org/10.1038/s41396-020-0587-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-0587-5
This article is cited by
-
Intercropping-induced leaf metabolic changes increase plant resistance to herbivory
Plant and Soil (2024)
-
Microbiota-mediated nitrogen fixation and microhabitat homeostasis in aerial root-mucilage
Microbiome (2023)
-
Hyphosphere microorganisms facilitate hyphal spreading and root colonization of plant symbiotic fungus in ammonium-enriched soil
The ISME Journal (2023)
-
Mechanisms of Epichloë bromicola to Promote Plant Growth and Its Potential Application for Coix lacryma-jobi L. Cultivation
Current Microbiology (2023)
-
Coordination of root auxin with the fungus Piriformospora indica and bacterium Bacillus cereus enhances rice rhizosheath formation under soil drying
The ISME Journal (2022)