Abstract
Chytridiomycosis, caused by the pathogen Batrachochytrium dendrobatidis (Bd), has led to population declines and extinctions of frog species around the world. While it is known that symbiotic skin bacteria can play a protective role against pathogens, it is not known how these defensive bacteria are integrated into the bacterial community on amphibian skin. In this study, we used 16S rRNA gene amplicon sequencing, culturing and Bd inhibition bioassays to characterize the communities of skin bacteria on three Neotropical frog species that persist in a Bd-infected area in Panama and determined the abundance and integration of anti-Bd bacteria into the community. We found that the two treefrog species had a similar bacterial community structure, which differed from the more diverse community found on the terrestrial frog. Co-occurrence networks also revealed differences between frog species such that the treefrogs had a significantly higher number of culturable Bd-inhibitory OTUs with high centrality scores compared with the terrestrial frog. We found that culture-dependent OTUs captured between 21 and 39% of the total relative abundance revealed in culture-independent communities. Our results suggest different ecological strategies occurring within skin antifungal communities on host species that have not succumbed to Bd infections in the wild.
Similar content being viewed by others

Introduction
Fungal diseases of plants and animals are emerging at an increasing rate [1] due to human-caused spread of pathogens to naive hosts and changing environmental conditions that favor lethal infections. Specifically, the chytrid fungal pathogen Batrachochytrium dendrobatidis (Bd), which causes chytridiomycosis, has led to many amphibian declines and extinctions [2, 3]. Chytridiomycosis has mainly affected tropical amphibian populations at high elevations and in cooler seasons [4, 5] since these conditions coincide with the pathogen’s optimal growth range [6]. For instance, in a tropical montane region of Panama, over 53% of the amphibian species declined or went locally extinct once Bd arrived [7, 8]. The effects of these losses extended to the ecosystem level, for example, resulting in increased algal biomass in streams [9]. Alternatively, some amphibian species continue to survive in Bd-infected areas at all elevations without apparent population declines [4, 10,11,12]. The study of amphibian species that are tolerant or resistant to chytridiomycosis can provide insights into how to protect species at risk around the world.
Amphibians have various defensive mechanisms that include adaptive and innate immunity, such as the secretion of antimicrobial peptides (AMPs) [13], and the presence of skin bacteria that produce antifungal metabolites [14]. As one example of the latter, the salamander Plethodon cinereus houses the bacterial species Janthinobacterium lividum, which produces anti-Bd secondary metabolites like violacein and indole-3-carboxaldehyde [15]. These antifungal bacterial species can reach high population densities on the skin and can protect amphibians from chytridiomycosis caused by Bd [14, 16, 17]. Studies show that removal of skin bacteria increased morbidity due to Bd [14, 18], whereas bacterial addition experiments can greatly reduce morbidity and mortality in some amphibian species [16, 17]. In addition, amphibian skin bacterial communities vary depending on host-specific factors, including species [19], habitat [20], and developmental stage [21], as well as environmental factors like seasonality [22] and pathogen presence [23].
To date, the antifungal capacity of skin bacteria in vitro has been described for thousands of isolates obtained from the skin of many different amphibian species [24]. However, little is known about the role these bacteria play within the skin microbial community in vivo. Based on other host-associated microbial systems, the protective role of symbiotic bacteria through the production of secondary metabolites and toxins can be a consequence of bacteria–bacteria interactions such as cooperation and competition within the community [25, 26]. These interactions can be important in structuring ecological communities [27, 28]. In addition, the host can influence the structure of symbiotic communities through several mechanisms including the production of AMPs [26, 29]. In the case of amphibians, innate immunity and skin microbiota can synergize and provide host defense [30].
Our study focused on three frog species that are common members of Neotropical amphibian communities: the treefrogs, Agalychnis callidryas and Dendropsophus ebraccatus, and the terrestrial frog, Craugastor fitzingeri. These three species have survived epizootic events in the highlands [8, 31, 32] and have persisted with Bd in an enzootic infection stage at high and low elevations [12, 31, 33, 34]. These frog species differ in their skin bacterial community structure, but no clear differences have been found between Bd-infected and -uninfected frogs in terms of bacterial community structure and secondary metabolite profiles in Bd-infected regions [34]. Thus, Belden et al. [34] suggested that different bacterial communities might be producing broadly similar sets of metabolites across frog hosts and sites. In addition, the structure of C. fitzingeri skin communities differed between Bd-infected and Bd-naive regions [23]; however, functional redundancy was found with respect to genes involved in bacterial communication and defense functions [35].
In this study, we analyze skin bacterial communities and determine the presence of antifungal bacteria on these three frog species that have co-existed with Bd for several years in a well-studied region in the Panamanian lowlands [12, 32, 34]. We use culturing techniques and Bd inhibition bioassays to identify and describe antifungal bacteria. In addition, we determine their relative abundance in the amphibian skin community by 16S ribosomal RNA (rRNA) gene amplicon (16S amplicon here after) analysis. Previous studies have shown that a surprisingly large fraction of the bacterial species from amphibian skin communities can be cultured [36,37,38], and Bd inhibition bioassays reveal which species secrete products that inhibit Bd growth [39]. In this project, the integration of culturing, Bd inhibition bioassays and 16S amplicon sequencing allow us to describe the antifungal capacity of the community without using purely predictive methods as in previous studies [40,41,42]. In addition, we use co-occurrence network analysis to determine if anti-Bd bacteria are key members of the bacterial community as defined by Operational taxonomic units (OTUs) having high centrality scores in the networks. Hence, these data can help determine if taxa providing a defensive function are well integrated into the skin microbial community of amphibian species that coexist with Bd in the wild.
We hypothesized that if skin bacteria are an important part of these hosts’ defenses, anti-Bd bacteria would have a high relative abundance, as seen in temperate amphibian species [38], and play a role in structuring these communities. This would suggest that the skin microbiota of Bd-tolerant or -resistant frogs could have important defensive functions despite having differences in the antifungal bacterial taxonomic composition and in the overall bacterial community structure. We also hypothesized that since the microhabitats of the treefrogs and the terrestrial species differed, and amphibians are known to house bacteria also found in the environment [23, 43], the taxonomic composition of the treefrogs’ microbiota would be more similar to each other than either was to C. fitzingeri [20, 34]. In addition to testing our hypotheses, in this study we emphasize the need to integrate different approaches such as culturing, bioassays and 16S amplicon sequencing in order to better explore the defensive function skin bacteria play on amphibian hosts.
Materials and methods
Field collection
Frogs were captured by hand using new nitrile gloves for each individual, and they were placed in individual sterile bags. Once they were all collected separately, frogs were swabbed and then released. Host study species details can be found in Supplementary Materials. Before swabbing, frogs were rinsed with approximately 50 ml of sterile deionized water to remove transient bacteria [44]. To sample the skin bacterial community, each individual was swabbed twice: the first swab was used for 16S amplicon sequencing and the second swab was used for isolating bacteria in culture. Skin swabs were collected according to previously published procedures [34] using sterile rayon swabs (MW113, Medical Wire Equipment & Co. Ltd, Corsham, UK). The swabs used for 16S amplicon sequencing were stored in dry, sterile 1.5 mL microcentrifuge tubes. The swabs for culturing were stored in 1.5 mL microcentrifuge tubes with 1 mL of trypticase soy broth and 40% glycerol (TSB:glycerol). Both types of swabs were placed on ice during field work and then stored at −80 °C in the laboratory until further processing. Bacterial isolation and taxonomic identification methods can be found in Supplementary Materials.
DNA extraction and 16S amplicon sequencing of culture-independent communities
DNA was extracted from all dry swabs using the DNeasy Blood and Tissue kit (Qiagen, Valencia, CA, USA) according to the manufacturer´s instructions, including a pretreatment with lysozyme. All methods were performed as in Rebollar et al. [23]. Briefly, DNA was used as template to amplify the V4 region of the 16S rRNA gene using barcoded primers (F515/R806) and PCR conditions adapted from Caporaso et al. [45]. Amplicons were quantified using the QuantiFluor® dsDNA System (Promega, Madison, WI, USA). Composite samples for sequencing were created by combining equimolar ratios of amplicons from the individual samples, followed by cleaning with the QIAquick PCR clean up kit (Qiagen). Barcoded composite PCR products were sent to the Dana Farber Cancer Institute’s Molecular Biology Core Facilities (Boston, MA, USA) for 250PE MiSeq Illumina sequencing.
16S amplicon data analysis of culture-independent bacterial communities
Raw reads were filtered and processed with the Quantitative Insights Into Microbial Ecology pipeline (QIIME 1.9.0) [46] with assembly and filtering as in Rebollar et al. [23]. Sequences were demultiplexed and clustered into OTUs at a sequence similarity threshold of 97% with the UCLUST method [47]. Sequences were matched against the Greengenes database (May 2013 release) [48], and those that did not match the database were clustered as de novo OTUs. Taxonomy was assigned using the RDP classifier [49]. Representative sequences were aligned to the Greengenes database with PyNAST [50], and a maximum likelihood phylogenetic tree was constructed with FastTree 2 [51]. The OTU table was filtered using a minimum cluster size of 0.001% of the total reads [52], resulting in 34441 to 150379 reads per frog sample. The original OTU table was rarefied according to the sample with the lowest number of reads. The final rarefied OTU table included 2200 OTUs. The raw data (paired end files) were deposited in the NCBI sequence read archive with the BioProject accession number PRJNA521543.
Comparing culture-independent communities with culture-dependent communities
We determined the presence and relative abundance of culture-dependent OTUs in the 16S amplicon community (culture-independent approach) based on a procedure defined by Walke et al. [36]. First, 16S rRNA sequences from the bacterial isolates (N = 359) were clustered at 97% similarity using the UCLUST method [47] in QIIME 1.9.0 [51], obtaining a total of 250 culture-dependent OTUs. The representative sequences of the culture-dependent OTUs were then used as a reference database to cluster the culture-independent 16S amplicon sequences at 97% sequence similarity, using UCLUST [47]. The culture-independent sequences that did not cluster with the 250 culture-dependent OTU reference sequences were clustered as de novo OTUs. All 250 culture-dependent OTUs were identified in the combined OTU table. This table was then filtered using a minimum cluster size of 0.001% of the total reads [52] and samples were rarified according to the sample with the lowest number of reads (35230). After removing OTUs with low abundances [52], this combined analysis resulted in 1891 OTUs that included 78 OTUs present in both culture-dependent and culture-independent sequences, and 1813 OTUs present only in culture-independent sequences. The relative abundance of each culture-dependent OTU was computed as the average across all individuals for each frog species.
Bd inhibition bioassays
A Bd strain from the Global Pandemic Lineage isolated from a Panamanian frog (GPL-JEL 423) was challenged against all 359 bacterial isolates. Bd growth bioassays were designed based on Bell et al. [39] and were performed exactly as in Bletz et al. [53]. Briefly, in 96-well plates, Bd zoospores (2 × 106) were grown in the presence of the cell-free supernatant of each bacterial isolate. Bacteria were previously grown in coculture with Bd zoospores for 3 days. Triplicates of each sample were tested along with the following controls in each assay: (1) 1% tryptone + Bd zoospores (positive control); (2) sterile water + Bd zoospores (nutrient depleted); (3) heat-killed Bd zoospores + 1% tryptone (heat-killed control); and (4) 1% tryptone only (negative control). Assay plates were incubated at 21 °C, and growth was measured as optical density (OD) at 492 nm on a spectrophotometer on days 0, 4, 7, and 10.
To calculate an inhibition score for each isolate, the slope of a regression of OD readings over time was calculated, and triplicate values from each replicate were averaged to generate a mean slope per isolate. The mean slope of each isolate was then divided by the mean slope of the nutrient-depleted control to determine the proportion of growth. The growth proportion was subtracted from 1 to determine the inhibition score so that positive values indicate inhibition and negative values indicate facilitation. Isolates were classified in five categories depending on the degree of Bd growth inhibition or facilitation obtained from the bioassays: (1) strong inhibition when values were equal or greater than 0.75, (2) moderate inhibition when values were <0.75 but ≥0.50, (3) weak inhibition when values were <0.50 but ≥0.25, (4) no inhibition when values were <0.25 but ≥−0.25 and (5) facilitation when values were <−0.25. In the case of the culture-dependent OTUs, the inhibition score associated with each culture-dependent OTU was calculated as the average across the isolates belonging to each OTU (from 1 to 12 isolates per OTU).
Co-occurrence networks
Co-occurrence networks were constructed with the software CoNet v1.1.0 [54], using the combined OTU table obtained from comparing the culture-independent communities with culture-dependent OTUs. Three networks (one network for each frog species) were constructed, so the combined OTU table was split into three separate OTU tables that were used as input for CoNet. Networks were constructed based on pairwise correlations using Spearman correlations tests. Only correlations >0.8 and with P-values < 0.05 were considered as significant (after using the Benjamini–Hochberg test for multiple comparisons). To identify OTUs that likely have an important role in structuring the community, we calculated three centrality indicators that rely on different properties of the networks: degree, closeness centrality and betweenness centrality [55]. The degree of a node in a network is the number of edges (connections) the node has to other nodes. The closeness centrality of a node is the average length of the shortest path between the node and all other nodes in the network; the more central a node, the closer it is to all other nodes. Betweenness centrality is equal to the number of shortest paths from all nodes to all others that pass through that node. Betweenness is a measure of the influence a node has over the spread of information through the network. Central or key OTUs were defined as those that had at least one high centrality value: higher than 20 for degree, 0.25 for closeness centrality, and 0.02 for betweenness centrality. The calculation of centrality indices as well as the visualization and manipulation of networks were completed with Cytoscape v3.3.0 [56].
Results
Skin bacterial diversity using culture-dependent methods
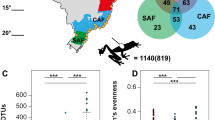
We isolated a total of 359 bacterial isolates from 44 individuals across three frog species (Table 1). C. fitzingeri (terrestrial frog) yielded the largest number of morphologically distinct bacterial isolates, with an average of 11 isolates per individual, whereas A. callidryas and D. ebraccatus (both treefrogs) yielded an average of 7 isolates per individual. Following culture-dependent OTU clustering (Supplementary Methods), C. fitzingeri maintained the highest number of culture-dependent OTUs, followed by D. ebraccatus and A. callidryas (Table 1). Two OTUs were shared across the three species, between 8 and 10 OTUs were shared between pairs of species, and the majority of the OTUs were unique to each species (Fig. 1).
Culture-dependent OTUs belonged to three phyla (Firmicutes, Actinobacteria, and Proteobacteria) and five classes (Bacilli, Actinobacteria, Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria). At the class level, these OTUs were differentially represented across the three frog species: 100% of Betaproteobacteria, 87.5% of Gammaproteobacteria, and 59.6% of Actinobacteria were isolated from C. fitzingeri, whereas 51.5% of Alphaproteobacteria were isolated from A. callidryas and 61.5% of Bacilli were isolated from D. ebraccatus (Fig. 2).
Maximum likelihood phylogenetic tree of the 250 culture-dependent OTUs (clustered at 97% similarity) obtained from C. fitzingeri, A. callidryas, or D. ebraccatus. Colors at the inner part of the phylogeny indicate bacterial classes. Inner circle indicates the average inhibitory score for each OTU and outer circles represent the host species of the OTU (color figure online)
Host species differed in the proportion of anti-Bd bacteria, which belonged to specific bacterial classes
In addition to having the largest number of isolates and culture-dependent OTUs, C. fitzingeri was the species with the highest percentage of culture-dependent Bd-inhibitory OTUs (68.2%) followed by A. callidryas (59.1%) and D. ebraccatus (14.3%) (Table 1). Furthermore, the greatest proportion of strongly inhibitory OTUs, as calculated from the total number of culture-dependent OTUs obtained per species, was also observed in C. fitzingeri (37%), followed by A. callidryas (9.9%) and D. ebraccatus (2.4%) (Fig. 1).
Different bacterial classes contained different proportions of inhibitory OTUs: 100% (6/6) of Betaproteobacteria, 78.12% (25/32) of Gammaproteobacteria, 56.38% (53/94) of Actinobacteria, 42.42% (28/66) of Alphaproteobacteria, and 38.47% (20/52) of Bacilli inhibited Bd growth to some extent. The rest of the OTUs from different bacterial classes either did not affect or facilitated Bd growth (Fig. 2).
Inhibitory scores were significantly different among bacterial classes (analysis of variance (ANOVA) F(4,211) = 10.64, P < 0.001). Tukey post hoc tests showed that Gammaproteobacteria had significantly higher inhibition scores than Alphaproteobacteria, Actinobacteria, and Bacilli (Fig. 3a, Table S1). Also, inhibitory scores of Actinobacteria and Betaproteobacteria were significantly higher than Bacilli (Fig. 3a, Table S1). When we compared inhibition scores of families that had at least three OTUs, we also found significant differences among the 18 bacterial families (ANOVA F(18,197) = 4.08, P < 0.001). Post hoc tests revealed several significant differences between pairs of bacterial families (Table S2). Specifically, the Enterobacteriaceae family had significantly higher inhibition scores in comparison with the Acetobacteraceae and Rhizobiaceae (Alphaproteobacteria), Gordoniae, Micrococcaceae and Mycobacteriaceae (Actinobacteria), and Paenibacillaceae and Bacillaceae (Bacilli) (Fig. 3b, Table S2). In contrast, the Bacillaceae family had significantly lower inhibition scores in comparison with the Microbacteriaceae and Streptomycetaceae (Actinobacteria), Enterobacteriaceae and Moraxellaceae (Gammaproteobacteria), and Comamonadaceae (Betaproteobacteria) (Fig. 3b, Table S2).
Culture-independent methods reveal differences between the skin microbiota of the terrestrial frog and the two treefrog species
The 16S amplicon sequencing yielded a total of 2200 OTUs where 1998, 1855, and 1575 belonged to C. fitzingeri, A. callidryas, and D. ebraccatus, respectively. In contrast to the result obtained with culture-dependent methods, the majority of the culture-independent OTUs were shared among all frog species (62.27% of the total number of OTUs) (Fig. 4a). However, C. fitzingeri had the most unique OTUs, followed by A. callidryas and D. ebraccatus (Fig. 4a).
Skin bacterial communities obtained through 16S amplicon sequencing. a Venn diagram of OTUs obtained from three frog species. b Alpha diversity (Shannon index) of skin communities. Letters signify significant differences obtained with post hoc Tukey tests. c Relative abundance of OTUs at the genus level. The legend indicates the six most relatively abundant bacterial taxa. (color figure online)
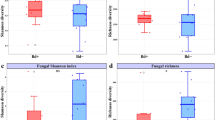
The Shannon index showed significant differences among frog species in OTU alpha diversity (ANOVA F(2, 41) = 7.03, P < 0.005). Tukey post hoc tests indicated that C. fitzingeri individuals had a higher alpha diversity than A. callidryas and D. ebraccatus (C. fitzingeri–A. callidryas P = 0.035, C. fitzingeri–D. ebraccatus P = 0.002, A. callidryas–D. ebraccatus P = 0.516) (Fig. 4b).
Culture-independent OTUs across all frog species belonged to 13 microbial phyla, with Actinobacteria and Proteobacteria being the numerically dominant phyla (41.3 and 57.8%, respectively). These OTUs belonged to 28 classes, with Actinobacteria and Gammaproteobacteria being the numerically dominant classes (41.3 and 50.8%, respectively). Relative abundance of OTUs at the genus level showed that skin bacterial communities of C. fitzingeri were dominated by Pseudomonas (Gammaproteobacteria), whereas A. callidryas and D. ebraccatus communities were dominated by Cellulomonas (Actinobacteria) (Fig. 4c).
Bray–Curtis distances based on OTU relative abundances showed significant differences in bacterial community structure among frog species Permutational multivariate analysis of variance (PERMANOVA): F(2,41) = 4.374, P = 0.001). Specifically, C. fitzingeri were significantly different than A. callidryas (Analysis of similarity (ANOSIM) R(2,30) = 0.212, Padj = 0.003) and D. ebraccatus (ANOSIM R(2,29) = 0.267, Padj = 0.003), whereas the two treefrog species, A. callidryas and D. ebraccatus, did not significantly differ (ANOSIM R(2,29) = 0.0007, Padj = 0.316).
To describe potential relationships occurring among bacteria within the skin communities, we constructed co-occurrence networks of OTUs from each frog species based on significant Spearman correlations. These networks contained 420, 440, and 367 nodes (culture-independent OTUs) on C. fitzingeri, A. callidryas, and D. ebraccatus, respectively. All networks show more positive correlations (co-occurrences) than negative correlations (mutual exclusions) (Table S3, Table S4), both of which are represented as edges within the networks (Fig. 5). Consistent with the most relatively abundant taxa on the skin communities, co-occurrence networks were dominated by Gammaproteobacteria and Actinobacteria classes, followed by Betaproteobacteria and Alphaproteobacteria.
Co-occurrence networks of the skin communities per frog species obtained with 16S amplicon sequencing. Nodes represent OTUs. Edges represent significant correlations. Networks on the left-hand side (a, b, c) show all nodes colored by bacterial class (see lower legend on the left). Networks on the right-hand side (d, e, f) show only the nodes that represent culture-dependent OTUs colored and shaped depending on their inhibitory capacity and their centrality values (see lower legend on the right) (color figure online)
Some network properties differed among frog species: on one hand, the D. ebraccatus network had a higher clustering coefficient, higher average number of neighbors, and higher network density compared with the other two species (Table S3). These three parameters are related to how dense the network is and the tendency of the graph to form clusters. Thus, the D. ebraccatus network has a higher number of connections (correlations/edges) per node and a higher tendency to form clusters. On the other hand, the C. fitzingeri network has a higher network diameter, higher characteristic path length, and higher network heterogeneity compared with the other two species (Table S3). These three parameters are related to the extent or area of the network graph and the heterogeneity of the number of connections associated with each OTU in the network. Thus, C. fitzingeri had a more diffuse network with higher variability in OTU individual connections compared with the other two frog species. In contrast, A. callidryas network had the lowest values for network heterogeneity.
In addition to the species-specific differences in network properties, the correlation structure (edges) in the network between OTUs (nodes) from different bacterial classes also differed mainly between the terrestrial frog (C. fitzingeri) and the treefrogs (A. callidryas and D. ebraccatus). Specifically, negative correlations between Actinobacteria and Betaproteobacteria were 2.5- and 3.2-fold higher in the C. fitzingeri network than in the respective D. ebraccatus and A. callidryas networks. Similarly, negative correlations between Alphaproteobacteria and Betaproteobacteria were two- and threefold higher (Table S5) in the C. fitzingeri network. This increase explains the pattern shown in C. fitzingeri´s network, in which the two main modules (or OTU clusters) are one enriched in Alphaproteobacteria and Actinobacteria and the other one enriched in Betaproteobacteria and Gammaproteobacteria, separated by negative correlations (Fig. 5c). This pattern differs from the treefrog networks where the two main modules of the networks were composed by all bacterial classes (Fig. 5a, b).
Culture-dependent OTUs represent a significant proportion of skin microbial communities
We evaluated the number and relative abundance of culture-dependent OTUs present in the bacterial community obtained with culture-independent methods [36] (Supplementary Methods). Of the 250 culture-dependent OTUs, 76, 63, and 78 OTUs were present in A. callidryas, D. ebraccatus, and C. fitzingeri communities, respectively, representing 4.7, 4.6, and 4.5% of the total number of culture-independent OTUs (Fig. 6a). The majority of the culture-dependent OTUs represented in the culture-independent community were Bd inhibitory: 64.5% in A. callidryas, 71.4% in D. ebraccatus, and 64.1% in C. fitzingeri (Fig. 6a).
Presence and proportion of Bd inhibitory and not inhibitory culture-dependent OTUs in the skin communities obtained with 16S amplicon sequencing. a Culture-dependent OTUs present in the culture-independent communities. b Percentage (relative abundance) of culture-dependent OTUs present in the culture-independent communities
Even though culture-dependent OTUs represented a low percentage of OTUs present in the culture-independent community, their total relative abundance ranged between 21.2 and 39.2% of the total community (Fig. 6b). In the case of the two treefrog species, inhibitory culture-dependent OTUs represented <4% of the relative abundance in the community while in C. fitzingeri, these OTUs represented 10.6% of the total relative abundance. All percentages represent the averages calculated across all individuals for each frog species.
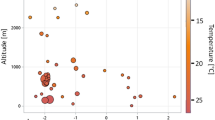
The relative abundance of individual culture-dependent OTUs within the three communities was highly variable, ranging between 1.8 × 10−6 – 3.2 × 10−1 for A. callidryas (Fig. 7a), 2 × 10−6 – 3.5 × 10−1 for D. ebraccatus (Fig. 7b), and 5.6 × 10−6 – 7.9 × 10−2 for C. fitzingeri (Fig. 7c). These results indicate that culturable techniques are capturing OTUs with low, medium, and high abundances within the skin communities.
Relative abundance (proportion) of culture-dependent OTUs within the skin community obtained from 16S amplicon sequencing. Red circles indicate culture-dependent Bd-inhibitory OTUs, green circles indicate culture-dependent non-inhibitory OTUs and stars indicate OTUs that were captured in the co-occurrence networks. a A. callidryas, b D. ebraccatus, and c C. fitzingeri (color figure online)
We evaluated the presence of culture-dependent OTUs in the co-occurrence networks and determined their position in the network according to three centrality indices: degree, betweenness centrality, and closeness centrality [55]. These indices are used to identify nodes (OTUs in this case) that are fundamental for the network structure [55]. A total of 27, 24, and 30 culture-dependent OTUs were present in the networks of A. callidryas, D. ebraccatus and C. fitzingeri, respectively. Most of the culture-dependent OTUs with the highest relative abundances in the community were captured, but all networks also captured some OTUs with low relative abundances (Fig. 7).
We found a significant difference in the proportion of Bd-inhibitory OTUs among the three species networks (χ2 = 6.51, df = 2, P = 0.03). Specifically, C. fitzingeri’s network had a lower proportion of Bd-inhibitory OTUs (53.3%), whereas A. callidryas and D. ebraccatus had a higher proportion (63 and 70.8%, respectively). This difference is more evident when comparing the proportions of the culturable OTUs that had high centrality values (χ2 = 59.97, df = 2, P < 0.001). In this case, the C. fitzingeri network only had one Bd-inhibitory OTU with a high centrality value (25%), and this OTU was not present in A. callidryas and D. ebraccatus networks (Fig. 5f, Table S6), while A. callidryas and D. ebraccatus networks had 11 and 12 inhibitory OTUs with high centrality values (73.3 and 70.3%), respectively (Fig. 5d, e, Table S6).
Discussion
Studies on host–bacterial symbiotic systems have identified bacterial communities with protective functions against pathogens [25, 57, 58]. In the case of amphibians, skin bacteria can be protective against chytridiomycosis caused by Bd [14, 16, 17]. In this study, we used culturing and culture-independent methods to characterize the cutaneous bacterial community of one terrestrial frog species and two treefrog species from a Neotropical lowland site in Panama. Results from both culturing and culture-independent methods showed that the terrestrial frog C. fitzingeri had a higher bacterial diversity and more unique OTUs than did the two treefrog species. Also, C. fitzingeri had a higher number of culture-dependent OTUs that inhibited the growth of the pathogen Bd, than did the treefrog species. In addition, these OTUs represented a higher proportion of bacteria within the culture-independent bacterial community in contrast to the proportions detected in the treefrog species. Co-occurrence network analyses also showed that relationships among culture-independent OTUs differed between C. fitzingeri and the two treefrog species.
Using culturing techniques, we found that Bd-inhibitory culture-dependent OTUs were mainly from the Proteobacteria phylum and in particular from the Betaproteobacteria and Gammaproteobacteria classes. Members of the Gammaproteobacteria had the highest inhibition scores among the culture-dependent OTUs. These results are consistent with previous similar approaches studying the skin bacterial diversity in temperate amphibian species [38]. In contrast, our results differ from a study on Panamanian frogs that found that inhibitory bacteria were found evenly distributed across the phyla Actinobacteria, Bacteroidetes, Firmicutes, and Proteobacteria phyla [59]. However, Becker et al. [59] analyzed 11 frog species with variable susceptibility including highly susceptible species (e.g., species Atelopus limosus and Strabomantis bufoniformis), while we analyzed three Bd non-susceptible (tolerant or resistant) species, which could explain the differences in the phylogenetic profile of inhibitory isolates. Our results suggest that a defensive microbiota can be achieved via a limited number of community structures that are likely influenced by the surrounding environmental pool of bacteria and host-specific factors [60].
Our finding that bacterial communities differed between the terrestrial and the treefrog species suggests that, in addition to host-specific effects, aspects of the microhabitat (perhaps working as a microbial diversity reservoir) could be driving these differences. Previous studies have shown that the skin microbiota differs from the environmental bacterial communities [23, 43, 61]. However, the presence of environmental reservoirs is fundamental to the maintenance of the diversity of skin microbiota in amphibians [40]. In addition, large-scale analyses have shown that host microhabitat (terrestrial, aquatic, or arboreal) can influence amphibian skin microbial diversity [20].
We found that the terrestrial species C. fitzingeri yielded the highest proportion of Bd-inhibitory OTUs in contrast with the treefrogs. However, node centrality scores from the co-occurrence networks suggest that treefrogs have more inhibitory OTUs that could be playing a central role in structuring the community than on the terrestrial frog. Thus, some bacterial species may play key roles in the community and as a result be more connected to other members of the community as reflected by higher centrality values in a co-occurrence network. This situation appears to be the case with the anti-Bd culturable OTUs of the treefrog bacterial communities, while in C. fitzingeri antifungal bacteria appear to not be as well integrated in the community.
What does this mean with respect to the protective role these communities play on their hosts? Additional studies are required to evaluate the degree of protection these microbial communities provide to their host species. However, in a recent experiment where lowland frogs were exposed to Bd at a temperature that favored Bd growth (19 °C), it was found that A. callidryas individuals were infected to a lesser degree (average zoospore genome equivalents (ZGE) = 27, SE = 23) than C. fitzingeri (average ZGE = 102,998, SE = 57,652) individuals after 41 days of being exposed to high Bd doses [62]. In fact, some of the A. callidryas individuals were able to clear the infection by the end of the experiment. In addition to having higher infection intensities, transcriptomic analyses of C. fitzingeri showed that its genes associated with inflammatory response were overexpressed, whereas genes involved in epithelium and skeletal development were downregulated, which did not occur in A. callidryas expression pattern [62]. These data indicate that A. callidryas and C. fitzingeri have different expression patterns in response to Bd infection. Thus, we hypothesize that the differences we found in skin bacterial community structure are associated with differences in expression patterns of genes that code for factors, such as AMPs and cytokines, which favor some bacterial species and community structures over others.
Network analyses allow us to identify patterns in large, complex datasets that are not evident through the use of standard diversity metrics widely used in microbial ecology [63]. Co-occurrence and mutual exclusion patterns can help identify potential biotic interactions (competition or cooperation), but they can also reflect habitat affinities (shared or distinct niches) [64]. In essence, co-occurrence patterns and centrality indices may be helpful to identify bacterial species that are functionally or ecologically important in a community. However, it is important to consider these patterns as hypotheses that can be further tested experimentally. While understanding these limitations, we hypothesize that having anti-Bd bacteria as key members in a community network (i.e., high centrality scores) may lead to resistance to fungal infection and explain, at least in part, why these three species have co-existed with the pathogen and could thus be consider tolerant to Bd infection. This hypothesis could be further tested in vitro with simplified bacterial communities that vary by network centrality values and Bd-inhibitory capacity.
Patterns from the host species in this study suggest two possible mechanisms by which skin bacteria are able to protect their hosts against pathogens: having fewer anti-Bd bacteria that are central to the community network as seen in the treefrog species, or having a greater number of inhibitory bacteria that are less central to the network as seen in the terrestrial frog species. Our results are in agreement with a previous study in which differences in the dominance of skin antifungal bacteria were found among temperate amphibian species [38]. Walke et al. [38] also suggested that different community structures (core vs. peripheral antifungal OTUs) could be maintaining strong disease resistance within these communities. However, it is important to consider that both of these studies were limited to the culturable proportion of anti-Bd bacterial isolates obtained with our specific culturing methods. Thus, our interpretations of the data need to be taken with caution with respect to the underlying mechanisms of host pathogen protection.
While culture-independent characterization of bacterial communities has led to a rapid expansion of our understanding of microbial ecology, culturing continues to play an important role in terms of understanding the biological attributes of bacterial species. Moreover, information from culturing could be more central to our understanding of microbiome ecology if these OTUs represent a high proportion of bacteria from these communities. Previous studies of amphibian skin bacteria found that the relative abundance of culturable OTUs can represent up to 36% of the culture-independent community and most of the dominant OTUs, families, and phyla were represented in culture [36, 38]. In our study, we obtained ~4.6% of OTUs in culture, but these OTUs represented between 21.2 and 39.1% of the total relative abundance of the culture-independent community. We also found that culturing with low nutrient media (under our specific culturing conditions) was successful in capturing OTUs with low, medium, and high relative abundance. In addition, we found that most of the Bd-inhibitory OTUs with the highest relative abundances were captured. Based on our work and the previous studies, it appears that culturing with low nutrient media is capturing many of the numerically dominant and functionally important (i.e., Bd inhibitory) OTUs in the amphibian skin system. It is likely that a greater sampling effort using more than one media would yield an even larger proportion of culturable OTUs.
Overall, in this study we identified antifungal bacteria that likely play an important role within skin microbial communities of three tropical host species. The differences found between the terrestrial frog and the treefrog species suggest that skin communities are structured differently in distinct hosts, and so antifungal bacteria may play different roles within each community. Moreover, our study stresses the importance of integrating several laboratory and molecular techniques (culture-dependent and -independent methods), as well as analytical methods (co-occurrence networks and standard diversity analyses) in order to better describe the dynamics of microbial communities.
References
Fisher MC, Henk DA, Briggs CJ, Brownstein JS, Madoff LC, McCraw SL, et al. Emerging fungal threats to animal, plant and ecosystem health. Nature. 2012;484:186–94.
Berger L, Roberts AA, Voyles J, Longcore JE, Murray KA, Skerratt LF. History and recent progress on chytridiomycosis in amphibians. Fungal Ecol. 2016;19:89–99.
Lips KR. Overview of chytrid emergence and impacts on amphibians. Philos Trans R Soc B Biol Sci. 2016;371:20150465.
Berger L, Speare R, Hines HB, Marantelli G, Hyatt AD, Mcdonald KR, et al. Effect of season and temperature on mortality in amphibians due to chytridiomycosis. Aust Vet J. 2004;82:73–5.
Sapsford SJ, Alford RA, Schwarzkopf L. Elevation, temperature, and aquatic connectivity all influence the infection dynamics of the amphibian chytrid fungus in adult frogs. PLoS ONE. 2013;8:1–12.
Piotrowski JS, Annis SL, Longcore JE. Physiology of Batrachochytrium dendrobatidis, a chytrid pathogen of amphibians. Mycologia. 2004;96:9–15.
Lips KR, Brem F, Brenes R, Reeve JD, Alford RA, Voyles J, et al. Emerging infectious disease and the loss of biodiversity in a Neotropical amphibian community. Proc Natl Acad Sci USA. 2006;103:3165–70.
Crawford AJ, Lips KR, Bermingham E. Epidemic disease decimates amphibian abundance, species diversity, and evolutionary history in the highlands of central Panama. Proc Natl Acad Sci USA. 2010;107:13777–82.
Whiles M, Hall RO, Dodds WK, Verburg P, Huryn AD, Pringle CM, et al. Disease-driven amphibian declines alter ecosystem processes in a tropical stream. Ecosystems. 2013;16:146–57.
Flechas SV, Sarmiento C, Cárdenas ME, Medina EM, Restrepo S, Amézquita A. Surviving chytridiomycosis: differential anti-Batrachochytrium dendrobatidis activity in bacterial isolates from three lowland species of Atelopus. PLoS ONE. 2012;7:e44832.
Kilburn VL, Ibáñez R, Sanjur O, Bermingham E, Suraci JP, Green DM. Ubiquity of the pathogenic chytrid fungus, Batrachochytrium dendrobatidis, in anuran communities in Panamá. Ecohealth. 2010;7:537–48.
Rebollar EA, Hughey MC, Harris RN, Domangue RJ, Medina D, Ibáñez R, et al. The lethal fungus Batrachochytrium dendrobatidis Is present in lowland tropical forests of far eastern Panamá. PLoS ONE. 2014;9:e95484.
Rollins-Smith LA, Conlon JM. Antimicrobial peptide defenses against chytridiomycosis, an emerging infectious disease of amphibian populations. Dev Comp Immunol. 2005;29:589–98.
Becker MH, Brucker RM, Schwantes CR, Harris RN, Minbiole KPC. The bacterially produced metabolite violacein is associated with survival of amphibians infected with a lethal fungus. Appl Environ Microbiol. 2009;75:6635–8.
Brucker RM, Harris RN, Schwantes CR, Gallaher TN, Flaherty DC, Lam Ba, et al. Amphibian chemical defense: antifungal metabolites of the microsymbiont Janthinobacterium lividum on the salamander Plethodon cinereus. J Chem Ecol. 2008;34:1422–9.
Harris RN, Brucker RM, Walke JB, Becker MH, Schwantes CR, Flaherty DC, et al. Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus. ISME J. 2009;3:818–24.
Kueneman JG, Woodhams DC, Harris R, Archer HM, Knight R, Mckenzie VJ. Probiotic treatment restores protection against lethal fungal infection lost during amphibian captivity. Proc R Soc B Biol Sci. 2016;283:20161553.
Walke JB, Becker MH, Loftus SC, House LL, Teotonio TL, Minbiole KPC, et al. Community structure and function of amphibian skin microbes: an experiment with bullfrogs exposed to a chytrid fungus. PLoS ONE. 2015;10:e0139848.
McKenzie VJ, Bowers RM, Fierer N, Knight R, Lauber CL. Co-habiting amphibian species harbor unique skin bacterial communities in wild populations. ISME J. 2012;6:588–96.
Bletz MC, Archer H, Harris RN, McKenzie VJ, Rabemananjara FCE, Rakotoarison A, et al. Host ecology rather than host phylogeny drives amphibian skin microbial community structure in the biodiversity hotspot of Madagascar. Front Microbiol. 2017;8:1530.
Prest TL, Kimball AK, Kueneman JG, Mckenzie VJ. Host-associated bacterial community succession during amphibian development. Mol Ecol. 2018;27:1–15.
Longo AV, Zamudio KR. Environmental fluctuations and host skin bacteria shift survival advantage between frogs and their fungal pathogen. ISME J. 2017;11:349–61.
Rebollar EA, Hughey MC, Medina D, Harris RN, Ibáñez R, Belden LK. Skin bacterial diversity of Panamanian frogs is associated with host susceptibility and presence of Batrachochytrium dendrobatidis. ISME J. 2016;10:1–14.
Woodhams DC, Alford RA, Antwis RE, Archer H, Becker MH, Belden LK, et al. Antifungal isolates database of amphibian skin-associated bacteria and function against emerging fungal pathogens. Ecology. 2015;96:595.
Fraune S, Anton-Erxleben F, Augustin R, Franzenburg S, Knop M, Schröder K, et al. Bacteria–bacteria interactions within the microbiota of the ancestral metazoan Hydra contribute to fungal resistance. ISME J. 2014;9:1–14.
Foster KR, Schluter J, Coyte KZ, Rakoff-Nahoum S. The evolution of the host microbiome as an ecosystem on a leash. Nat Publ Gr. 2017;548:43–51.
Stubbendieck RM, Vargas-Bautista C, Straight PD. Bacterial communities: interactions to scale. Front Microbiol. 2016;7:1234.
Estrela S, Brown SP. Community interactions and spatial structure shape selection on antibiotic resistant lineages. PLoS Comput Biol. 2018;14:1–21.
Franzenburg S, Walter J, Künzel S, Wang J, Baines JF, Bosch TCG, et al. Distinct antimicrobial peptide expression determines host species-specific bacterial associations. Proc Natl Acad Sci USA. 2013;110:E3730–8.
Myers JM, Ramsey JP, Blackman AL, Nichols AE, Minbiole KPC, Harris RN. Synergistic inhibition of the lethal fungal pathogen Batrachochytrium dendrobatidis: the combined effect of symbiotic bacterial metabolites and antimicrobial peptides of the frog Rana muscosa. J Chem Ecol. 2012;38:958–65.
Brem FMR, Lips KR. Batrachochytrium dendrobatidis infection patterns among Panamanian amphibian species, habitats and elevations during epizootic and enzootic stages. Dis Aquat Organ. 2008;81:189–202.
Woodhams DC, Kilburn VL, Reinert LK, Voyles J, Medina D, Ibáñez R, et al. Chytridiomycosis and amphibian population declines continue to spread eastward in Panama. Ecohealth. 2008;5:268–74.
Perez R, Richards-Zawacki CL, Krohn AlexanderR, Robak M, Griffith EdgardoJ, Ross Heidi, et al. Field surveys in Western Panama indicate populations of Atelopus varius frogs are persisting in regions where Batrachochytrium dendrobatidis is now enzootic. Amphib Reptile Conserv. 2014;8:30–5.
Belden LK, Hughey MC, Rebollar EA, Umile TP, Loftus SC, Burzynski EA, et al. Panamanian frog species host unique skin bacterial communities. Front Microbiol. 2015;6:1171.
Rebollar EA, Gutiérrez-Preciado A, Noecker C, Eng A, Hughey MC, Medina D, et al. The skin microbiome of the Neotropical frog Craugastor fitzingeri: inferring potential bacterial-host-pathogen interactions from metagenomic data. Front Microbiol. 2018;9:466.
Walke JB, Becker MH, Hughey MC, Swartwout MC, Jensen RV, Belden LK. Most of the dominant members of amphibian skin bacterial communities can be readily cultured. Appl Environ Microbiol Aem. 2015;81:6589–600.
Medina D, Walke JB, Gajewski Z, Becker MH, Swartwout MC, Belden LK. Culture media and individual hosts affect the recovery of culturable bacterial diversity from amphibian skin. Front Microbiol. 2017;8:1574.
Walke JB, Becker MH, Hughey MC, Swartwout MC, Jensen RV, Belden LK. Dominance-function relationships in the amphibian skin microbiome. Environ Microbiol. 2017;19:3387–97.
Bell SC, Alford RA, Garland S, Padilla G, Thomas AD. Screening bacterial metabolites for inhibitory effects against Batrachochytrium dendrobatidis using a spectrophotometric assay. Dis Aquat Organ. 2013;103:77–85.
Loudon AH, Woodhams DC, Parfrey LW, Archer H, Knight R, McKenzie V, et al. Microbial community dynamics and effect of environmental microbial reservoirs on red-backed salamanders (Plethodon cinereus). ISME J. 2014;8:830–40.
Kueneman JG, Woodhams DC, Van Treuren W, Archer HM, Knight R, McKenzie VJ. Inhibitory bacteria reduce fungi on early life stages of endangered Colorado boreal toads (Anaxyrus boreas). ISME J. 2015;10:1–11.
Bletz MC, Perl RGB, Bobowski BT, Japke LM, Tebbe CC, Dohrmann AB, et al. Amphibian skin microbiota exhibits temporal variation in community structure but stability of predicted Bd-inhibitory function. ISME J. 2017;11:1521–34.
Walke JB, Becker MH, Loftus SC, House LL, Cormier G, Jensen RV, et al. Amphibian skin may select for rare environmental microbes. ISME J. 2014;8:2207–17.
Lauer A, Simon M, Banning JL, Andre E, Duncan K, Harris RN. Common cutaneous bacteria from the eastern red-backed salamander can inhibit pathogenic fungi. Copeia 2007;3:630–40.
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA. 2011;108(Suppl):4516–22.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012;6:610–8.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–7.
Caporaso JG, Bittinger K, Bushman FD, Desantis TZ, Andersen GL, Knight R. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics. 2010;26:266–7.
Price MN, Dehal PS, Arkin AP. FastTree 2 - approximately maximum-likelihood trees for large alignments. PLoS ONE 2010. https://doi.org/10.1371/journal.pone.0009490.
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon I, Knight R. et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. 2013;10:57–9.
Bletz MC, Myers J, Woodhams DC, Rabemananjara FCE, Rakotonirina A, Weldon C, et al. Estimating herd immunity to amphibian chytridiomycosis in Madagascar based on the defensive function of amphibian skin bacteria. Front Microbiol. 2017;8:1751.
Faust K, Raes J. Microbial interactions: from networks to models. Nat Rev Microbiol. 2012;10:538–50.
Koschützki D, Schreiber F. Centrality analysis methods for biological networks and their application to gene regulatory networks. Gene Regul Syst Bio. 2008;2008:193–201.
Cline MS, Smoot M, Cerami E, Kuchinsky A, Landys N, Workman C, et al. Integration of biological networks and gene expression data using Cytoscape. Nat Protoc. 2007;2:2366–82.
Berg G. Plant-microbe interactions promoting plant growth and health: perspectives for controlled use of microorganisms in agriculture. Appl Microbiol Biotechnol. 2009;84:11–8.
Hoyt JR, Cheng TL, Langwig KE, Hee MM, Frick WF, Kilpatrick AM. Bacteria isolated from bats inhibit the growth of Pseudogymnoascus destructans, the causative agent of white-nose syndrome. PLoS ONE. 2015;10:e0121329.
Becker MH, Walke JB, Murrill L, Woodhams DC, Reinert LK, Rollins-Smith LA, et al. Phylogenetic distribution of symbiotic bacteria from Panamanian amphibians that inhibit growth of the lethal fungal pathogen Batrachochytrium dendrobatidis. Mol Ecol. 2015;24:1628–41.
Walke JB, Belden LK. Harnessing the microbiome to prevent fungal infections: lessons from amphibians. PLoS Pathog. 2016;12:e1005796.
Fitzpatrick BM, Allison AL. Similarity and differentiation between bacteria associated with skin of salamanders (Plethodon jordani) and free-living assemblages. FEMS Microbiol Ecol. 2014;88:482–94.
Ellison AR, Tunstall T, DiRenzo GV, Hughey MC, Rebollar EA, Belden LK, et al. More than skin deep: functional genomic basis for resistance to amphibian chytridiomycosis. Genome Biol Evol. 2014;7:286–98.
Proulx SR, Promislow DEL, Phillips PC. Network thinking in ecology and evolution. Trends Ecol Evol. 2005;20:345–53.
Barberán A, Bates ST, Casamayor EO, Fierer N. Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 2012;6:343–51.
Acknowledgements
We thank Molly C. Bletz for her assistance in the laboratory. Scientific collection permits were provided by the Panamanian authorities (Autoridad Nacional del Ambiente): permits SE/A-47-12, SEX/A-65-12, SEX/A-77-12 and SEX/A-89-12. Animal care protocols were approved by the Smithsonian Tropical Research Institute’s Animal Care Committee: protocol 2011-1110-2014 and by Virginia Tech’s Animal Care Committee: protocol 11-105-BIOL. This project was supported by the National Science Foundation Grants: DEB-1136602 (to RNH) and DEB-1136640 (to LKB).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Rebollar, E.A., Bridges, T., Hughey, M.C. et al. Integrating the role of antifungal bacteria into skin symbiotic communities of three Neotropical frog species. ISME J 13, 1763–1775 (2019). https://doi.org/10.1038/s41396-019-0388-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0388-x
This article is cited by
-
Bacteria from the skin of amphibians promote growth of Arabidopsis thaliana and Solanum lycopersicum by modifying hormone-related transcriptome response
Plant Molecular Biology (2024)
-
Associations of Batrachochytrium dendrobatidis with skin bacteria and fungi on Asian amphibian hosts
ISME Communications (2023)
-
Linking microbiome and stress hormone responses in wild tropical treefrogs across continuous and fragmented forests
Communications Biology (2023)
-
Signatures of functional bacteriome structure in a tropical direct-developing amphibian species
Animal Microbiome (2022)
-
Variation of soil nitrate and bacterial diversity along soil profiles in manure disposal maize field and adjacent woodland
Journal of Soils and Sediments (2020)