Abstract
The microorganisms in the gastrointestinal (GI) tract can influence the metabolism, immunity, and behavior of animal hosts. Increasing evidence suggests that communication between the host and the microbiome also occurs in the opposite direction, with hormones and other host-secreted compounds being sensed by microorganisms. Here, we addressed one key aspect of the host–microbe communication by studying chemotaxis of a model commensal bacterium, Escherichia coli, to several compounds present abundantly in the GI tract, namely catecholamines, thyroid hormones, and polyamines. Our results show that E. coli reacts to five out of ten analyzed chemicals, sensing melatonin, and spermidine as chemorepellents and showing mixed responses to dopamine, norepinephrine and 3,4-dihydroxymandelic acid. The strongest repellent response was observed for the polyamine spermidine, and we demonstrate that this response involves the low-abundance chemoreceptor Trg and the periplasmic binding protein PotD of the spermidine uptake system. The chemotactic effects of the tested compounds apparently correlate with their influence on growth and their stability in the GI tract, pointing to the specificity of the observed behavior. We hypothesize that the repellent responses observed at high concentrations of chemoeffective compounds might enable bacteria to avoid harmful levels of hormones and polyamines in the gut and, more generally, antimicrobial activities of the mucous layer.
Similar content being viewed by others
Introduction
Humans and other animals share a mutualistic relationship with numerous resident microorganisms, collectively known as the microbiome. Over the past two decades, the role of the host–microbiome interactions in a number of physiological processes became increasingly clear [1,2,3], and it is likely that the gut microorganisms evolved specific mechanisms to detect multiple compounds that are released into the lumen of the gastrointestinal (GI) tract by the endocrine and immune systems of the host [4,5,6,7].
Homeostasis of the mucous layer of intestinal epithelial cells (IECs) in the mammalian GI tract is regulated by a variety of signals including cellular polyamines and hormonal signals [8,9,10]. Due to the abundance of catecholamines in the GI tract, most gut-brain axis studies have focused on the interactions between gut bacteria and the most abundant catecholamines – epinephrine, norepinephrine (NE; also known as noradrenaline, NA) and dopamine [11,12,13,14]. Recently, more attention has also been drawn to the thyroid hormones, mainly serotonin and melatonin (5-methoxy-N-acetyltryptamine), that are synthesized from tryptophan in the IECs and involved in the regulation of the GI tract function and of circadian cycles [15,16,17,18,19]. Moreover, polyamines such as putrescine and spermidine may likewise have a role in microbial endocrinology [10]. Polyamines are introduced with the diet, but also produced by microorganisms and host cells and regulate many distinct cellular functions in eukaryotes and prokaryotes, which include proliferation and differentiation of intestinal cells [20, 21]. The concentrations of polyamines in the GI lumen can approach millimolar levels, and they are known to affect the microbiome composition [10, 22]. Enteric bacteria, such as Escherichia coli, possess specific systems for polyamine uptake [23, 24].
Chemotaxis is one way in which bacteria could react to the compounds present in the gut. It allows bacteria to navigate in environmental gradients of various chemicals [25,26,27] in order to locate conditions that are beneficial for growth [28]. Chemotaxis to self-secreted signals can also mediate collective behaviors, such as the autoinducer-2 dependent autoaggregation and biofilm formation by E. coli [29, 30]. The chemotactic signal transduction pathway of E. coli includes two high-abundance (or major) receptors, Tsr and Tar, as well as three low-abundance (or minor) receptors, Trg, Tap, and Aer. These receptors form mixed complexes in the membrane together with the histidine kinase CheA, where the autophosphorylation activity of CheA is inhibited by the increased exposure to attractants. Low-kinase activity leads to reduced phosphorylation of the motor regulator CheY, which promotes counter-clockwise (CCW) flagellar rotation and thus smooth swimming up the attractant gradient. In contrast, the exposure to repellents activates CheA and elevates CheY phosphorylation, inducing the clockwise (CW) rotation and swimming reorientation. Dephosphorylation of CheY is catalyzed by the phosphatase CheZ, which is essential to quickly readjust bacterial behavior. Additionally, the chemotaxis pathway includes an adaptation system that gradually offsets the initial stimulation by attractants or repellents through changes in receptor methylation.
Several previous studies have suggested that the animal pathogens Helicobacter pylori, Campylobacter jejuni, and Salmonella enterica exhibit chemotaxis toward several compounds derived from the human gastric epithelium [31,32,33,34,35] and that such chemotaxis plays an important role in bacterial invasion, as well as survival in the intestine [36,37,38]. Moreover, both commensal E. coli K-12 and enterohemorrhagic E. coli (EHEC) can sense NE as a chemoattractant [39,40,41]. This response requires conversion of NE to 3,4-dihydroxymandelic acid (DHMA) by the monoamine oxidase TynA and the aromatic aldehyde dehydrogenase FeaB of E. coli, with DHMA then serving as the chemoattractant recognized by Tsr in the nanomolar concentration range [40,41,42]. These results indicate that hormone taxis might be common among enteric bacteria and not limited to pathogens, but the scope of this behavior remained unclear.
The goal of this study was therefore to systematically investigate the extent to which E. coli—as a model commensal bacterium with a well-studied chemotaxis system—can exhibit taxis toward different catecholamines, thyroid hormones and polyamines. Our results demonstrate that E. coli responds, using both low and high abundance receptors, to a wide range of the physiologically important compounds that are known to accumulate in the gut lumen, including dopamine, NE, 3,4-dihydroxymandelic acid (DHMA), melatonin and spermidine, but not to their metabolic precursors. Notably, we show that the same compounds also affect growth of E. coli, in an apparent correlation with its chemotactic preferences. We therefore propose that the observed chemotactic responses may be used by commensal and pathogenic E. coli to locate optimal growth niches in the gut lumen, but also to avoid the antimicrobial activity of the mucous layer of the gut.
Results
Chemotactic response of the wild type E. coli towards gut compounds
To analyze the intracellular response of the chemotaxis pathway of E. coli towards gut compounds, we used a previously described assay based on Förster (fluorescence) Resonance Energy Transfer (FRET). Here, the phosphorylation-dependent interaction between CheY fused to yellow fluorescent protein (CheY-YFP) and its phosphatase CheZ fused to cyan fluorescent protein (CheZ-CFP) is monitored as a readout of the pathway activity [43, 44] (Fig. S1). The response was tested by stimulating cells expressing the FRET pair with serial dilutions of chemicals and measuring the subsequent change in FRET ratio (i.e., the ratio of the YFP to CFP fluorescence emission). As previously shown [43, 45, 46], stimulation with an attractant results in a rapid decrease in the FRET ratio, reflecting a decrease in the kinase activity. Stimulation with a repellent has an opposite effect, i.e., it increases the FRET ratio. Because continuous stimulation elicits adaptive changes in receptor methylation that gradually offset the effects of either attractant or repellent, the FRET ratio typically transiently overshoots upon removal of the chemoeffector (Fig. S1) [43, 44].
We started our analysis with the biosynthetic pathway for catecholamine hormones (Fig. S2), the most widely studied compounds in molecular endocrinology. In the wild-type E. coli cells, the two major neurotransmitters of the catecholamine pathway, dopamine and NE, elicited biphasic responses (Fig. 1a–d). Dopamine was sensed as a repellent at concentrations below 1 mM, with the FRET ratio increasing upon addition of dopamine, and then decreasing upon its removal, which is opposite to the effect of canonical attractant α-amethyl-d,l-aspartate (MeAsp) (Fig. 1a). In contrast, at 10 mM dopamine signaled as attractant (Fig. 1a, b).
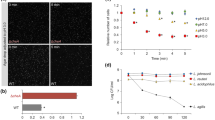
FRET-based analysis of the chemotaxis pathway response to catecholamines. a, c, e Examples of FRET measurements of responses to dopamine (a), norepinephrine (NE) (c), and 3,4-dihydroxymandelic acid (DHMA) (e). The ratio of YFP/CFP fluorescence reflects the efficiency of FRET and thus activity of the chemotaxis pathway. Buffer-adapted cells were stimulated with step-like addition and subsequent removal of compounds, indicated by downward and upward arrows, respectively. Saturating stimulation with 1 mM α-methyl-DL-aspartate (MeAsp) was used as a positive control. b, d, f Dose-response curves of wild-type cells (filled circles), Tar-only cells (open squares) or Tsr-only cells (open diamonds) to dopamine (b), NE (d), and DHMA (f). Each point represents the mean FRET-measured values of the kinase activity, normalized to the activity in buffer, from at least three independent experiments, with error bars indicating the standard error of the mean. Values above one correspond to a repellent response, while values below one correspond to an attractant response
The response to NE was generally less pronounced, and it had an inverse pattern compared to the dopamine response. NE behaved as a weak attractant at low concentrations, but it produced a repellent response above 1 mM (Fig. 1c, d). The attractant response to low concentrations of NE is overall consistent with previous microfluidic measurements, although in those studies the chemotaxis towards low levels of NE was stronger and comparable to the chemotaxis towards amino acid attractants [39, 40]. This response was shown to rely on the conversion of NE into DHMA, which could be induced by the NE-mediated signaling [40,41,42]. Indeed, we observed that the responses of the wild-type E. coli cells to NE and DHMA were similar, with DHMA eliciting a weak attractant response up to 50 µM but a strong repellent response at higher concentrations (Fig. 1e, f), but in contrast to those previous studies no further enhancement of response could be observed upon growing E. coli cultures in presence of NE (not shown). No effects on the chemotaxis pathway activity could be observed for l-tyrosine, consistent with previous work [28], or for L-3,4-dihydroxyphenylanine (L-DOPA) (Fig. S3A). Notably, although L-DOPA and l-tyrosine can be detected in the GI tract, these precursors are rapidly converted into dopamine in the gut lumen [8, 47,48,49] and therefore unlikely to form stable gradients. Finally, although epinephrine elicited an apparent repellent response (not shown), interpretation of these data was complicated by a strong autofluorescence of epinephrine that interfered with FRET measurements.
We next tested two major thyroid hormones, serotonin and melatonin. E. coli shows no chemotactic response to l-tryptophan, the precursor of thyroid hormones [28], and no obvious chemotaxis was observed for serotonin (Fig. S3B). However, the wild-type E. coli cells exhibited strong dose-dependent repellent response to melatonin at concentrations above 0.1 mM (Fig. 2a, b).
Pathway response to melatonin. a Example of a FRET measurement response in the wild-type cells, performed as in Fig. 1. b Dose–response curves from indicated strains. Error bars correspond to the standard error of the mean for three independent experiments
Finally, of the two tested polyamines, E. coli displayed no significant reaction to putrescine (Fig. S3B) but showed a strong repellent response towards spermidine in the millimolar concentration range (Fig. 3a, b).
Pathway response to spermidine. a Example of FRET measurement of the chemotaxis pathway response in the wild-type cells, performed as in Fig. 1. b Dose–response curves from indicated strains. Error bars correspond to the standard error of the mean of three independent experiments
Responses mediated by Tar and Tsr
To further characterize the roles of the two most abundant E. coli chemoreceptors—Tar and Tsr—FRET measurements were also performed in strains expressing only one of them at levels similar to the net endogenous chemoreceptor expression in the wild type. We observed that both Tar and Tsr could mediate—sometimes opposite—responses to most of the tested hormones, indicating that chemotaxis of wild-type E. coli results from interplay between Tar- and Tsr-mediated responses. The Tsr-mediated response apparently makes a larger contribution to the behavior of the wild-type cells, consistent with Tsr being the most abundant receptor under our growth conditions [50, 51]. Specifically, Tar mediated an attractant response to dopamine (Fig. 1b and Fig. S4A), whereas the Tsr-only strain showed the same biphasic trend as the wild type, switching from repellent to attractant (Fig. 1b and Fig. S4B). For NE, Tar showed an attractant response, whereas Tsr sensed NE as a repellent over the entire concentration range (Fig. 1d and Fig. S4C,D). For DHMA, both Tar and Tsr mediated repellent responses (Fig. 1f and Fig. S4E, F). These results suggest that the NE sensing at high concentrations could not be solely explained by its conversion into DHMA, since Tar responses to NE and DHMA were clearly different. Both Tar and Tsr mediated a repellent response to melatonin, similar to the one observed for the wild-type strain (Fig. 2b and Fig. S4G, H).
In contrast to the observed responses to hormones, the wild type sensing of spermidine could clearly not be accounted by a combination of Tar- and Tsr-mediated responses (Fig. 3b). These were weak and biphasic, with attractant responses at high concentration where the wild type showed a stronger repellent response, indicating that spermidine sensing in the wild type E. coli must be mediated by one of the minor receptors.
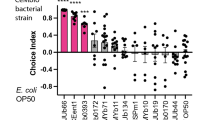
Microfluidic assay of chemotaxis
To verify the chemotactic responses to the gut compounds measured by FRET, we additionally analyzed the chemotactic behavior of E. coli in microfluidic channels. Here we used a recently described assay that allows measurements of the average motion of a bacterial population in linear chemical gradients, characteristic for chemotaxis [52, 53]. The observed chemotactic behavior (Fig. 4 and Fig. S5) was consistent with FRET measurements. The wild-type strain showed a statistically significant repellent taxis to all tested hormones in the 0–1 mM gradient, similar to the dominant response observed in this concentration range by FRET, except for epinephrine, which attractant taxis that is consistent with a previous report [39] was observed (Fig. 4). Chemotaxis of E. coli cells expressing only Tar or Tsr was also generally consistent with the FRET data, although in several cases the responses were variable and may require further verification (Fig. S5). Notably, only the wild type showed strong repellent reaction to spermidine, whereas the Tar- and Tsr-only strains showed either attractant or no response. Although the observed chemotactic drift away or towards the gut compounds was overall weaker than the drift in gradients of MeAsp [52, 53], it was similar to chemotaxis observed toward the metabolized strong attractant aspartate (Fig. 4). This suggests that degradation or uptake of attractants can markedly weaken the response that can be measured using this microfluidic assay, which may not be surprising given the relatively high density of bacteria within the channel in these experiments.
Chemotactic response in gradients of gut compounds. Chemotactic drift (defined as vch/αv0, see Methods) was measured in gradients of dopamine (Dop), NE, DHMA (DH), epinephrine (Epi), melatonin (Mel) or spermidine (S) established in a microfluidic device (Inset; measurement area is indicated by an orange rectangle). Zero chemotactic drift corresponds to non-responding cells and one to direct swimming up (negative values) or down (positive values) the gradient. For the negative control (∅), drift of the wild-type cells was measured in buffer in the absence of gradients. For the positive control, the drift was measured in a gradient of 0–1 mM aspartate (Asp). As a reference (not shown), the chemotactic drift of the wild-type cells in a gradient of non-metabolized attractant MeAsp had an average value of −0.1 [52]. Error bars indicate the standard error of the mean. A one-tailed student t-test was performed to assess the significance of the response being different from 0 (**P ≤ 0.05; *P ≤ 0.1)
Mechanism of the spermidine response
As mentioned above, our data indicated that the strong repellent response to spermidine observed in the wild type is likely to be mediated by one of the low-abundance receptors, Tap or Trg. As neither Trg nor Tap can function as the only receptor in E. coli [54], we tested chemotaxis to spermidine with trg and tap deletion strains (Fig. 5a). Whereas ∆tap cells responded to spermidine like the wild type, the response of ∆trg cells was comparable to the one observed for the Tar- and Tsr-only strains (Fig. S6A). This clearly suggests that the repellent response to spermidine is mediated by Trg. To further confirm the involvement of Trg, wild-type cells were adapted to ribose (a Trg-specific attractant) before stimulation with spermidine (Fig. S6B). Adaptation to ribose indeed abolished the repellent response to spermidine, implying that the interaction of the periplasmic ribose-binding protein (RBP) with the sensory domain of Trg, which is known to mediate response to ribose [55, 56], interferes with spermidine sensing.
Dose-response curves of the pathway response to spermidine. Responses were measured by FRET as in Fig. 1 for a ∆trg cells (red crosses) and ∆tap cells (black crosses) and b ∆potD (open triangles), ∆potA (open diamonds), and ∆potD/potD+ (filled inverted triangles) cells (see Figure S6). Differences between Δtrg and Δtap responses in a and between ΔpotD and ΔpotA or ∆potD/potD+ responses b are significant according to student t-test performed with responses to 10 mM spermidine (P ≤ 0.01)
Signaling via periplasmic binding proteins (BPs) is common not only to ribose but to all Trg- or Tap-specific chemoattractants for which the sensing mechanisms have been established. These periplasmic BPs are components of the ATP-binding cassette (ABC) transporters, but they also interact with the low-abundance receptors and regulate their activity upon binding to ligands. We thus hypothesized that the preferential E. coli ABC transporter for spermidine, PotABCD [24, 57], might be involved in the Trg-mediated response. In the PotABCD transporter complex, PotD is the periplasmic BP, PotA is the membrane-associated ATPase, and PotB and PotC are two membrane-spanning components of the transmembrane channel [24, 58]. Notably, the crystal structure of the PotD protein complex with spermidine is very similar to E. coli d-Glucose/d-Galactose-binding protein (GBP) [59], another interaction partner of Trg.
Indeed, in the strain deleted for potD, the specific repellent response to spermidine was abolished, similar to the effect of trg deletion (Fig. 5b and Fig. S6C). Complementation with ectopically expressed potD restored this response (Fig. S6D). In contrast, a potA strain deleted for the membrane-associated ATPase behaved similarly to the wild type (Fig. 5b). Hence, our data suggest that the interaction of the periplasmic BP PotD with Trg—and not spermidine uptake—is required for the Trg-mediated repellent response to spermidine.
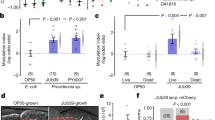
Effects of the gut compounds on E. coli growth
To better understand the physiological relevance of the observed chemotactic responses, we analyzed the effects of tested compounds on growth of a planktonic E. coli culture. Indeed, several compounds affected different stages of E. coli growth (Fig. 6a, Fig. S7). To capture these effects with a single number, we calculated the area below the curve as a measure of time-averaged OD (Fig. 6b). Similar differences between cultures were observed when comparing their maximal OD (Fig. S8). We observed that L-DOPA, epinephrine and, particularly, dopamine enhanced E. coli growth, in agreement with previous reports [11,12,13]. As oxidation of dopamine results in the development of black color during incubation, which could lower the precision of the optical density measurements despite baseline subtraction, we directly confirmed the growth-promoting effect of dopamine by quantifying the number of viable cells in the culture (Fig. S9). Although the exact mechanism of this growth stimulation by dopamine remains to be investigated, it might be related to the dopamine-mediated enhancement of iron uptake [11, 12].
Effect of the gut compounds on E. coli growth. a Examples of growth curves of E. coli MG1655 grown at 37 °C in TB containing dopamine, DHMA, melatonin or spermidine at indicated final concentrations (black line), and of control culture grown in TB (red line). Optical density of culture was measured at 600 nm as described in Methods. b The time-averaged OD, calculated as the area below the growth curves and divided by the duration of the experiment. For each experiment, the time-averaged OD was normalized to the time-averaged OD of the control culture in TB. Gray bars represent the catecholamine group; black bars the thyroid group; and white bars the polyamine group. A one-tailed student t-test was performed to assess the significance of the difference from the control (**P ≤ 0.01; *P ≤ 0.05). Each bar represents the mean of at least three independent experiments, with error bars indicating standard deviation
In contrast, DHMA and melatonin were growth-inhibitory, and the strongest growth inhibition was observed for spermidine (Fig. 6a, b and Fig. S8). The effect of spermidine was only observed above 1 mM, which matches the concentration range of chemorepellent response. In summary, the chemotactic preferences of E. coli were seemingly consistent with the effects of the tested compounds on E. coli growth and with their expected stability in the GI tract (Table S1).
Discussion
While most of the microbial endocrinology studies have focused on the effects of bacterial species on the concentrations of hormones in the GI tract, less attention has been paid to how the microbes themselves recognize these compounds [4,5,6,7, 60]. In this study, we used E. coli as the model enteric bacterium to investigate chemotactic responses to a range of compounds that are present in the human gut. Remarkably, we observed chemotactic responses to compounds that are known to accumulate in the gut—with the notable exception of serotonin—namely dopamine, NE, epinephrine, DHMA, melatonin and spermidine [8, 61,62,63,64]. Although accurate concentration measurements across the gut are complicated by the heterogeneity of the gut environment and the strong dependence on food content and health state of the host [65,66,67], these compounds can reach micro- to millimolar levels across the gut lumen [8, 10, 68]. Even higher concentrations are expected to be present in the vicinity of the mucous layer where these compounds are secreted, consistent with the observed concentration range of the chemotactic response. In contrast, no response could be detected for their precursors with similar chemical structure (e.g., l-tyrosine, l-tryptophan, l-DOPA, and putrescine), which are primarily derived from food sources and rapidly turned over in the GI tract [10, 48]. They are therefore unlikely to form long-lived gradients that can be used for orientation in the intestine. Our results thus provide further evidence for the hypothesis that bacteria can specifically utilize host signals to detect their GI location [14, 39,40,41,42].
For all hormones analyzed, chemotaxis in the wild type appears to be the result of interplay between responses mediated by the two high-abundance E. coli chemoreceptors, Tar and Tsr. For the most part, the Tsr-mediated response makes a larger contribution, in agreement with the higher expression of Tsr under our growth conditions. Such interplay between Tar- and Tsr-mediated responses is similar to the previously characterized tactic behavior in gradients of pH and temperature [69, 70], but contrasts with the responses to conventional chemoattractants that specifically bind to the periplasmic sensory domains of the receptors. Considering the similar range of hormone concentrations that are detected by Tar and Tsr, it is likely that all tested hormones are sensed indirectly, by perturbing some aspect of cell physiology. Although the exact molecular mechanisms of this sensing remains to be elucidated, it nevertheless appears to be specific, since chemically closely related precursor compounds elicit no response.
Furthermore, there seemed to be a correlation between the chemotactic preferences and effects of individual compounds on E. coli growth (Table S1). Although this relation was less clear for hormones that elicited biphasic responses—dopamine and NE—high concentrations of dopamine led to a strong attractant response that was consistent with the growth-promoting effect of dopamine. NE had no significant effect on growth, but since NE may play a role in the activation of the immune response [6, 71], its sensing may be important for avoidance of the immune system and thus for bacterial survival [6, 13, 72].
For the compounds that elicited a repellent response at high concentrations, namely DHMA, melatonin and spermidine, there was a pronounced correlation with growth inhibition. Although low concentrations of DHMA were previously shown to elicit a highly sensitive attractant response [40,41,42], at high concentrations of DHMA that affect growth, the behavior of E. coli in gradients is apparently dominated by the repellent response. The chemotactic response to melatonin and the observed growth inhibition are consistent with recently proposed effects of this hormone on the microbiome [17, 73]. In the context of the correlation observed between chemotaxis and growth effects, it is interesting that serotonin affects neither growth nor chemotaxis of E. coli.
Of the two tested polyamines, putrescine and spermidine, only the latter elicited a chemotactic response. This might be consistent with putrescine being rapidly taken up or converted to spermidine or spermine in the small intestine [10]. In contrast, levels of spermidine in the GI tract are expected to be high [10, 74]. The only compound that elicited a straight attractant response in the wild-type cells was epinephrine, consistent with a previous study [39] and correlating again with its growth-promoting effect for E. coli.
Although the mechanism of hormone sensing by E. coli chemoreceptors remains to be investigated, we could show that spermidine is specifically sensed as a repellent by the low abundance receptor Trg. So far, Trg has been only implicated in attractant responses to ribose, glucose and galactose, mediated by the interactions of its sensory domain with periplasmic binding proteins, RBP (for ribose) and GBP (for glucose and galactose). Our results suggest that the response to spermidine similarly involves the periplasmic binding protein PotD, which is part of the spermidine uptake system PotABCD. To our knowledge, this is the first example of a repellent response mediated by a minor chemoreceptor. It is also the first clear example of a repellent response involving a periplasmic binding protein. Although nickel-binding protein had been implicated initially in the repellent response to nickel mediated by Tar [75], this finding was subsequently disproved [76].
In general, considering the role of chemotaxis in the host–microbe interactions may provide a deeper understanding of the behavior of enteric bacteria within the host. Because bacterial chemotaxis is primarily a single-cell behavior, chemotaxis may be highly important for the survival and proliferation of Proteobacteria in the GI tract [38], even though they represent only minor constituents of the normal gut microbiota. The apparent correlation found between chemotactic preferences, growth effects and the nature of tested gut compounds (i.e., food-derived or secreted) suggest that bacteria are exposed to gradients of hormones and other host-derived compounds in the mammalian gut. Consequently, E. coli, and most likely other motile enteric bacteria, seem to have evolved specific tactic responses as a way not only to avoid harmful (or to locate beneficial) levels of these compounds, but also to orient themselves in the gut. In this context, repellent responses observed at high concentrations of compounds that are secreted from the gut epithelium may enable bacteria to limit immediate contact with the mucous layer of the GI tract, which is known to contain high levels of antimicrobial proteins and immunoglobulin A (IgA) [1, 77]. Furthermore, the interplay between these repellent responses and attractant responses mediated by low concentrations of NE and DHMA, as observed previously [14, 39,40,41,42] and in our work, could explain chemotactic accumulation of E. coli at a certain distance from the mucosal surface [78]. This area might represent a specific growth niche in the intestine [79], where bacteria can benefit from rapidly diffusing nutrients released by the epithelium without being harmed by the mucosal antimicrobials. Intestinal inflammation has been previously shown to lead to an enhance release of nutrients [38] and electron acceptors [36, 80] along with a reduced hormone secretion [81,82,83]. This might shift the accumulation pattern of EHEC or Salmonella, which possess chemotaxis systems that are nearly identical to that of commensal E. coli, towards the mucosal surface, possibly promoting proliferation of Proteobacteria and infection [84].
Methods
Bacterial strains and plasmids
All strains and plasmids used in this work are listed in Table S2. Strains used for the microfluidics and FRET measurements were derived from E. coli RP437 [85], the wild-type strain for chemotaxis. FRET strains were transformed with a plasmid expressing CheY-YFP and CheZ-CFP pair from a bi-cistronic construct pVS88 [43, 44], and where indicated with a plasmid expressing either Tar (pVS1092) or Tsr (pVS160). Strain VS104 [Δ(cheY cheZ)] carrying pVS88 was used as the wild-type for FRET. The ∆potD and ∆potA deletions were introduced by phage P1vir transduction from the respective mutants from the Keio collection [86]. KmR cassettes were eliminated via FLP recombination [87]. For potD complementation (potD+), ∆potD was transformed with the constructed plasmid derivative of pKG116 containing potD (pJL02). For growth experiments, MG1655 was used as E. coli wild type.
Reagents
l-Tyrosine (≥98% purity), L-Dopa-(phenyl-d3) (L-3,4-dihydroxyphenylalanine, 98% purity), Dopamine hydrochloride, (−)-Norepinephrine,(−)-Epinephrine (≥98% purity), DL-3,4-Dihydroxymandelic acid (DHMA, 98% purity), Serotonin hydrochloride, Melatonin powder (≥98% purity), Putrescine dihydrochloride, Spermidine (≥99% purity), α-methyl-DL-aspartate (MeAsp), were obtained from Sigma-Aldrich (Taufkirchen, Germany), and l-serine from Acros Organics—Thermo Fisher Scientific (Nidderau, Germany).
FRET assay
The FRET measurements were performed as previously described [43, 46]. Briefly, E. coli cells were grown in tryptone broth (TB) media (1% tryptone, 0.5% NaCl), supplemented with the respective antibiotics (100 mg/mL ampicillin; 17 mg /mL chloramphenicol) and inducers (Table S2) at 34 °C and 275 r.p.m. Cells were harvested at OD600 0.6. by centrifugation (4000 × g for 5 min), washed with tethering buffer (10 mM KPO4, 0.1 mM EDTA, 1 μM methionine, 10 mM lactic acid, pH 7), and stored at 4 °C for 30 min. The sample was attached to a polylysine-coated coverslip, placed in a flow chamber under constant flow (300 µl/min) of tethering buffer using a syringe pump (Harvard Apparatus, Massachusetts, United States), which was used for stimulation with compounds of interest (Fig. S1). Measurements were performed on an upright fluorescence microscope (custom-modified Zeiss Axiovert 200 microscope, Carl Zeiss Microscopy GmbH, Jena, Germany) equipped with a PCI-6034 counting board connected to a computer with custom written LabView7 software (National Instruments). CFP fluorescence was excited at 436/20 nm through a 455 nm dichroic mirror by a 75 W Xenon lamp. To detect CFP and YFP emissions, 480/40 nm band pass and 520 nm long pass emission filters were used, respectively. Fluorescence of a monolayer of 300–500 cells was continuously recorded in the cyan and yellow channels using photon counters with a 1.0 s integration time. FRET response was measured as the change in the ratio of YFP/CFP due to energy transfer and normalized to the response of buffer-adapted cells to saturating stimulation with known strong attractant, either α-methyl-d,l-aspartate (MeAsp) or l-serine.
Microfluidics assay
The measurement of the chemotactic drift was performed as previously described [52]. Briefly, E. coli RP437 cells grown to mid-exponential growth phase were harvested by centrifugation (4000 × g for 5 min), washed with chemotaxis buffer (10 mM KPO4, 0.1 mM EDTA, 67 mM NaCl, pH 7) and stored at 4 °C for 30 min to inhibit protein synthesis. The sample was placed in silicone elastomer (SYLGARD 184, 1:10 crosslinker to base ratio, Dow Corning, USA) hand-made chemotaxis chamber, consisting of two reservoirs linked via a small channel (length L = 2 mm, width w = 1 mm). One chamber contains suspended bacteria and the other contains suspended bacteria mixed with the target compound at the indicated concentration. After 30 min, a linear gradient of chemoattractant is formed in the channel to which the cells respond. The response of each strain to the gradient was recorded in the middle of the channel using video-microscopy (×10 magnification under phase contrast illumination, Mikrotron 4CXP camera running at 100 frames per seconds for 100 s, with a 717 × 717 μm2–512 × 512 px2 –field of view, focal plane halfway through the 50 μm sample depth). A high-throughput computer analysis of the films yielded the average chemotactic drift velocity of the population vch, the population-averaged swimming speed of the cells v0 and the fraction of swimming cells, α, which enables to estimate the chemotactic drift vch/αv0, where zero corresponds to non-responding cells and one to a population where all cells swim directly down the gradient.
Growth experiments
E. coli cells from an overnight culture (37 °C and 200 r.p.m. in TB) were diluted until OD600 0.05 in a total volume of 110 µl in a 96-well plate (Greiner Bio-One, Frickenhausen, Germany). OD600 was measured using a plate reader (Tecan Infinite M1000, Tecan Deutschland GmbH, Crailsheim, Germany) for 14 h at 37 °C and 180 r.p.m. TB medium was supplemented with the analyzed compounds at the indicated concentrations. For the dopamine experiments, an additional baseline subtraction was performed, with TB without cells but supplemented with dopamine as a negative control. Growth was analyzed by calculating the area under the curve divided by the duration for each individual experiment (14 h), giving the time-averaged OD. The average of all experiments was then calculated and normalized to the control. Statistical analysis was performed to assess the difference to the control with a one-tailed student t-test. P-values lower than 0.05 were considered statistically significant.
Colony forming unit (CFU) assay
The CFU assay was performed by diluting an overnight culture 1:100 in a total volume of 10 ml of TB containing dopamine at indicated concentrations and growing it at 37 °C for 6 h. Cells were serially diluted 10−6 and 10−7 and 100 µl of the samples were plated on LB plates in triplicates and incubated at 37 °C over night. The number of colonies in each plate was counted and the CFU/ml (number of colonies/volume of inoculation×dilution) was calculated.
References
Hooper LV, Macpherson AJ. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat Rev Immunol. 2010;10:159–69.
Backhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI. Host-bacterial mutualism in the human intestine. Science. 2005;307:1915–20.
Sommer F, Backhed F. The gut microbiota–masters of host development and physiology. Nat Rev Microbiol. 2013;11:227–38.
Lyte M. Microbial endocrinology in the pathogenesis of infectious disease. Microbiol Spectr. 2016;4:VMBF-0021-2015.
Neuman H, Debelius JW, Knight R, Koren O. Microbial endocrinology: the interplay between the microbiota and the endocrine system. FEMS Microbiol Rev. 2015;39:509–21.
Pacheco AR, Sperandio V. Inter-kingdom signaling: chemical language between bacteria and host. Curr Opin Microbiol. 2009;12:192–8.
Rhee SH, Pothoulakis C, Mayer EA. Principles and clinical implications of the brain-gut-enteric microbiota axis. Nat Rev Gastroenterol Hepatol. 2009;6:306–14.
Eisenhofer G, Aneman A, Friberg P, Hooper D, Fandriks L, Lonroth H, et al. Substantial production of dopamine in the human gastrointestinal tract. J Clin Endocrinol Metab. 1997;82:3864–71.
Kobayashi K. Role of catecholamine signaling in brain and nervous system functions: new insights from mouse molecular genetic study. J Investig Dermatol Symp Proc. 2001;6:115–21.
Milovic V. Polyamines in the gut lumen: bioavailability and biodistribution. Eur J Gastroenterol Hepatol. 2001;13:1021–5.
Freestone PP, Lyte M, Neal CP, Maggs AF, Haigh RD, Williams PH. The mammalian neuroendocrine hormone norepinephrine supplies iron for bacterial growth in the presence of transferrin or lactoferrin. J Bacteriol. 2000;182:6091–8.
Freestone PP, Haigh RD, Lyte M. Blockade of catecholamine-induced growth by adrenergic and dopaminergic receptor antagonists in Escherichia coli O157:H7, Salmonella enterica and Yersinia enterocolitica. BMC Microbiol. 2007;7:8.
Lyte M, Ernst S. Catecholamine induced growth of gram negative bacteria. Life Sci. 1992;50:203–12.
Sperandio V, Torres AG, Jarvis B, Nataro JP, Kaper JB. Bacteria-host communication: the language of hormones. Proc Natl Acad Sci USA. 2003;100:8951–6.
Cajochen C, Krauchi K, Wirz-Justice A. Role of melatonin in the regulation of human circadian rhythms and sleep. J Neuroendocrinol. 2003;15:432–7.
O’Mahony SM, Clarke G, Borre YE, Dinan TG, Cryan JF. Serotonin, tryptophan metabolism and the brain-gut-microbiome axis. Behav Brain Res. 2015;277:32–48.
Paulose JK, Wright JM, Patel AG, Cassone VM. Human gut bacteria are sensitive to melatonin and express endogenous circadian rhythmicity. PLoS ONE. 2016;11:e0146643.
Reigstad CS, Salmonson CE, Rainey JF 3rd, Szurszewski JH, Linden DR, Sonnenburg JL, et al. Gut microbes promote colonic serotonin production through an effect of short-chain fatty acids on enterochromaffin cells. FASEB J. 2015;29:1395–403.
Spohn SN, Mawe GM. Non-conventional features of peripheral serotonin signalling: the gut and beyond. Nat Rev Gastroenterol Hepatol. 2017;14:412–20.
Michael AJ. Polyamines in eukaryotes, bacteria, and archaea. J Biol Chem. 2016;291:14896–903.
Seiler N, Raul F. Polyamines and the intestinal tract. Crit Rev Clin Lab Sci. 2007;44:365–411.
Matsumoto M, Kakizoe K, Benno Y. Comparison of fecal microbiota and polyamine concentration in adult patients with intractable atopic dermatitis and healthy adults. Microbiol Immunol. 2007;51:37–46.
Kashiwagi K, Kobayashi H, Igarashi K. Apparently unidirectional polyamine transport by proton motive force in polyamine-deficient Escherichia coli. J Bacteriol. 1986;165:972–7.
Pistocchi R, Kashiwagi K, Miyamoto S, Nukui E, Sadakata Y, Kobayashi H, et al. Characteristics of the operon for a putrescine transport system that maps at 19 min on the Escherichia coli chromosome. J Biol Chem. 1993;268:146–52.
Hazelbauer GL. Bacterial chemotaxis: the early years of molecular studies. Annu Rev Microbiol. 2012;66:285–303.
Sourjik V, Armitage JP. Spatial organization in bacterial chemotaxis. EMBO J. 2010;29:2724–33.
Wadhams GH, Armitage JP. Making sense of it all: bacterial chemotaxis. Nat Rev Mol Cell Biol. 2004;5:1024–37.
Yang Y, M Pollard A, Hofler C, Poschet G, Wirtz M, Hell R, et al. Relation between chemotaxis and consumption of amino acids in bacteria. Mol Microbiol. 2015;96:1272–82.
Jani S, Seely AL, Peabody VG, Jayaraman A, Manson MD. Chemotaxis to self-generated AI-2 promotes biofilm formation in Escherichia coli. Microbiology. 2017;163:1778–90.
Laganenka L, Colin R, Sourjik V. Chemotaxis towards autoinducer 2 mediates autoaggregation in Escherichia coli. Nat Commun. 2016;7:12984.
Chandrashekhar K, Gangaiah D, Pina-Mimbela R, Kassem II, Jeon BH, Rajashekara G. Transducer like proteins of Campylobacter jejuni 81-176: role in chemotaxis and colonization of the chicken gastrointestinal tract. Front Cell Infect Microbiol. 2015;5:46.
Day CJ, King RM, Shewell LK, Tram G, Najnin T, Hartley-Tassell LE, et al. A direct-sensing galactose chemoreceptor recently evolved in invasive strains of Campylobacter jejuni. Nat Commun. 2016;7:13206.
Huang JY, Sweeney EG, Sigal M, Zhang HC, Remington SJ, Cantrell MA, et al. Chemodetection and destruction of host urea allows Helicobacter pylori to locate the epithelium. Cell Host Microbe. 2015;18:147–56.
Keilberg D, Ottemann KM. How Helicobacter pylori senses, targets and interacts with the gastric epithelium. Environ Microbiol. 2016;18:791–806.
Rahman H, King RM, Shewell LK, Semchenko EA, Hartley-Tassell LE, Wilson JC, et al. Characterisation of a multi-ligand binding chemoreceptor CcmL (Tlp3) of Campylobacter jejuni. PLoS Pathog. 2014;10:e1003822.
Rivera-Chavez F, Lopez CA, Zhang LF, Garcia-Pastor L, Chavez-Arroyo A, Lokken KL et al. Energy taxis toward host-derived nitrate supports a Salmonella pathogenicity Island 1-independent mechanism of invasion. MBio. 2016;7:mBio.00960-16.
Stecher B, Hapfelmeier S, Muller C, Kremer M, Stallmach T, Hardt WD. Flagella and chemotaxis are required for efficient induction of Salmonella enterica serovar Typhimurium colitis in streptomycin-pretreated mice. Infect Immun. 2004;72:4138–50.
Stecher B, Barthel M, Schlumberger MC, Haberli L, Rabsch W, Kremer M, et al. Motility allows S. Typhimurium to benefit from the mucosal defence. Cell Microbiol. 2008;10:1166–80.
Bansal T, Englert D, Lee J, Hegde M, Wood TK, Jayaraman A. Differential effects of epinephrine, norepinephrine, and indole on Escherichia coli O157:H7 chemotaxis, colonization, and gene expression. Infect Immun. 2007;75:4597–607.
Pasupuleti S, Sule N, Cohn WB, MacKenzie DS, Jayaraman A, Manson MD. Chemotaxis of Escherichia coli to norepinephrine (NE) requires conversion of NE to 3,4-dihydroxymandelic acid. J Bacteriol. 2014;196:3992–4000.
Sule N, Pasupuleti S, Kohli N, Menon R, Dangott LJ, Manson MD, et al. The norepinephrine metabolite 3,4-dihydroxymandelic acid is produced by the commensal microbiota and promotes chemotaxis and virulence gene expression in enterohemorrhagic Escherichia coli. Infect Immun. 2017;85:e00431–417.
Pasupuleti S, Sule N, Manson MD, Jayaraman A. Conversion of norepinephrine to 3,4-dihdroxymandelic acid in Escherichia coli requires the QseBC quorum-sensing system and the fear transcription factor. J Bacteriol. 2018;200:e00564–517.
Sourjik V, Berg HC. Receptor sensitivity in bacterial chemotaxis. Proc Natl Acad Sci USA. 2002;99:123–7.
Sourjik V, Berg HC. Functional interactions between receptors in bacterial chemotaxis. Nature. 2004;428:437–41.
Neumann S, Hansen CH, Wingreen NS, Sourjik V. Differences in signalling by directly and indirectly binding ligands in bacterial chemotaxis. EMBO J. 2010;29:3484–95.
Sourjik V, Vaknin A, Shimizu TS, Berg HC. In vivo measurement by FRET of pathway activity in bacterial chemotaxis. Methods Enzymol. 2007;423:365–91.
Schulz C, Eisenhofer G, Lehnert H. Principles of catecholamine biosynthesis, metabolism and release. Front Horm Res. 2004;31:1–25.
Bianchine JR, Calimlim LR, Morgan JP, Dujuvne CA, Lasagna L. Metabolism and absorption of L-3,4 dihydroxyphenylalanine in patients with Parkinson’s disease. Ann N Y Acad Sci. 1971;179:126–40.
Vieira-Coelho MA, Soares-Da-Silva P. Uptake and intracellular fate of L-DOPA in a human intestinal epithelial cell line: Caco-2. Am J Physiol. 1998;275:C104–112.
Kalinin Y, Neumann S, Sourjik V, Wu M. Responses of Escherichia coli bacteria to two opposing chemoattractant gradients depend on the chemoreceptor ratio. J Bacteriol. 2010;192:1796–800.
Salman H, Libchaber A. A concentration-dependent switch in the bacterial response to temperature. Nat Cell Biol. 2007;9:1098–100.
Colin R, Zhang R, Wilson LG. Fast, high-throughput measurement of collective behaviour in a bacterial population. J R Soc Interface. 2014;11:20140486.
Wilson LG, Martinez VA, Schwarz-Linek J, Tailleur J, Bryant G, Pusey PN, et al. Differential dynamic microscopy of bacterial motility. Phys Rev Lett. 2011;106:018101.
Feng X, Baumgartner JW, Hazelbauer GL. High- and low-abundance chemoreceptors in Escherichia coli: differential activities associated with closely related cytoplasmic domains. J Bacteriol. 1997;179:6714–20.
Ames GF. Bacterial periplasmic transport systems: structure, mechanism, and evolution. Annu Rev Biochem. 1986;55:397–425.
Binnie RA, Zhang H, Mowbray S, Hermodson MA. Functional mapping of the surface of Escherichia coli ribose-binding protein: mutations that affect chemotaxis and transport. Protein Sci. 1992;1:1642–51.
Furuchi T, Kashiwagi K, Kobayashi H, Igarashi K. Characteristics of the gene for a spermidine and putrescine transport system that maps at 15 min on the Escherichia coli chromosome. J Biol Chem. 1991;266:20928–33.
Kashiwagi K, Hosokawa N, Furuchi T, Kobayashi H, Sasakawa C, Yoshikawa M, et al. Isolation of polyamine transport-deficient mutants of Escherichia coli and cloning of the genes for polyamine transport proteins. J Biol Chem. 1990;265:20893–7.
Sugiyama S, Matsuo Y, Maenaka K, Vassylyev DG, Matsushima M, Kashiwagi K, et al. The 1.8-A X-ray structure of the Escherichia coli PotD protein complexed with spermidine and the mechanism of polyamine binding. Protein Sci. 1996;5:1984–90.
Dinan TG, Cryan JF. Regulation of the stress response by the gut microbiota: implications for psychoneuroendocrinology. Psychoneuroendocrinology. 2012;37:1369–78.
Haskel Y, Hanani M. Inhibition of gastrointestinal motility by MPTP via adrenergic and dopaminergic mechanisms. Dig Dis Sci. 1994;39:2364–7.
Pizzi C, Pignata S, Calderopoli R, D’Agostino L, Tritto G, D’Adamo G, et al. Cell kinetics and polyamine enzymes in the intestinal mucosa of rats with azoxymethane induced tumours. Int J Exp Pathol. 1994;75:305–11.
Straub RH, Wiest R, Strauch UG, Harle P, Scholmerich J. The role of the sympathetic nervous system in intestinal inflammation. Gut. 2006;55:1640–9.
Bubenik GA. Gastrointestinal melatonin: localization, function, and clinical relevance. Dig Dis Sci. 2002;47:2336–48.
Magro F, Vieira-Coelho MA, Fraga S, Serrao MP, Veloso FT, Ribeiro T, et al. Impaired synthesis or cellular storage of norepinephrine, dopamine, and 5-hydroxytryptamine in human inflammatory bowel disease. Dig Dis Sci. 2002;47:216–24.
Magro F, Fraga S, Ribeiro T, Soares-da-Silva P. Decreased availability of intestinal dopamine in transmural colitis may relate to inhibitory effects of interferon-gamma upon L-DOPA uptake. Acta Physiol Scand. 2004;180:379–86.
Sandrini S, Aldriwesh M, Alruways M, Freestone P. Microbial endocrinology: host-bacteria communication within the gut microbiome. J Endocrinol. 2015;225:R21–34.
Huether G. The contribution of extrapineal sites of melatonin synthesis to circulating melatonin levels in higher vertebrates. Experientia. 1993;49:665–70.
Paulick A, Jakovljevic V, Zhang S, Erickstad M, Groisman A, Meir Y, et al. Mechanism of bidirectional thermotaxis in Escherichia coli. eLife. 2017;6:e26607.
Yang Y, Sourjik V. Opposite responses by different chemoreceptors set a tunable preference point in Escherichia coli pH taxis. Mol Microbiol. 2012;86:1482–9.
Straub RH, Schaller T, Miller LE, von Horsten S, Jessop DS, Falk W, et al. Neuropeptide Y cotransmission with norepinephrine in the sympathetic nerve-macrophage interplay. J Neurochem. 2000;75:2464–71.
Lyte M. Microbial endocrinology and infectious disease in the 21st century. Trends Microbiol. 2004;12:14–20.
Voigt RM, Forsyth CB, Green SJ, Mutlu E, Engen P, Vitaterna MH, et al. Circadian disorganization alters intestinal microbiota. PLoS ONE. 2014;9:e97500.
Bartos F, Bartos D, Grettie DP, Campbell RA. Polyamine levels in normal human serum. Comparison of analytical methods. Biochem Biophys Res Commun. 1977;75:915–9.
de Pina K, Navarro C, McWalter L, Boxer DH, Price NC, Kelly SM, et al. Purification and characterization of the periplasmic nickel-binding protein NikA of Escherichia coli K12. Eur J Biochem. 1995;227:857–65.
Englert DL, Adase CA, Jayaraman A, Manson MD. Repellent taxis in response to nickel ion requires neither Ni2+ transport nor the periplasmic NikA binding protein. J Bacteriol. 2010;192:2633–7.
McGuckin MA, Linden SK, Sutton P, Florin TH. Mucin dynamics and enteric pathogens. Nat Rev Microbiol. 2011;9:265–78.
Allweiss B, Dostal J, Carey KE, Edwards TF, Freter R. The role of chemotaxis in the ecology of bacterial pathogens of mucosal surfaces. Nature. 1977;266:448–50.
Li H, Limenitakis JP, Fuhrer T, Geuking MB, Lawson MA, Wyss M, et al. The outer mucus layer hosts a distinct intestinal microbial niche. Nat Commun. 2015;6:8292.
Rivera-Chavez F, Winter SE, Lopez CA, Xavier MN, Winter MG, Nuccio SP, et al. Salmonella uses energy taxis to benefit from intestinal inflammation. PLoS Pathog. 2013;9:e1003267.
De Palma G, Collins SM, Bercik P, Verdu EF. The microbiota-gut-brain axis in gastrointestinal disorders: stressed bugs, stressed brain or both? J Physiol. 2014;592:2989–97.
Khan WI, Ghia JE. Gut hormones: emerging role in immune activation and inflammation. Clin Exp Immunol. 2010;161:19–27.
Mittal R, Debs LH, Patel AP, Nguyen D, Patel K, O’Connor G, et al. Neurotransmitters: the critical modulators regulating gut-brain axis. J Cell Physiol. 2017;232:2359–72.
Litvak Y, Byndloss MX, Tsolis RM, Baumler AJ. Dysbiotic Proteobacteria expansion: a microbial signature of epithelial dysfunction. Curr Opin Microbiol. 2017;39:1–6.
Parkinson JS, Houts SE. Isolation and behavior of Escherichia coli deletion mutants lacking chemotaxis functions. J Bacteriol. 1982;151:106–13.
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol. 2006;2:2006.
Cherepanov PP, Wackernagel W. Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene. 1995;158:9–14.
Acknowledgements
We thank Remy Colin for help with the microfluidics experiments, Seán Murray for help with analysis of the growth experiments, and Anja Paulick and Remy Colin for critically reading the manuscript. This work was supported by grants SO 421/11-1 from the Deutsche Forschungsgemeinschaft and 294761-MicRobE from the European Research Council.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lopes, J.G., Sourjik, V. Chemotaxis of Escherichia coli to major hormones and polyamines present in human gut. ISME J 12, 2736–2747 (2018). https://doi.org/10.1038/s41396-018-0227-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0227-5
This article is cited by
-
Chemotaxis and autoinducer-2 signalling mediate colonization and contribute to co-existence of Escherichia coli strains in the murine gut
Nature Microbiology (2023)
-
The ecological roles of bacterial chemotaxis
Nature Reviews Microbiology (2022)
-
Annotation of chemotaxis gene clusters and proteins involved in chemotaxis of Bacillus subtilis strain MB378 capable of biodecolorizing different dyes
Environmental Science and Pollution Research (2022)
-
Evaluation of the gut microbiome in association with biological signatures of inflammation in murine polytrauma and shock
Scientific Reports (2021)
-
Transporters of glucose and other carbohydrates in bacteria
Pflügers Archiv - European Journal of Physiology (2020)