Abstract
Mid-summer peaks in the abundance of Thaumarchaeota and nitrite concentration observed on the Georgia, USA, coast could result from in situ activity or advection of populations from another source. We collected data on the distribution of Thaumarchaeota, ammonia-oxidizing betaproteobacteria (AOB), Nitrospina, environmental variables and rates of ammonia oxidation during six cruises in the South Atlantic Bight (SAB) from April to November 2014. These data were used to examine seasonality of nitrification in offshore waters and to test the hypothesis that the bloom was localized to inshore waters. The abundance of Thaumarchaeota marker genes (16S rRNA and amoA) increased at inshore and nearshore stations starting in July and peaked in August at >107 copies L−1. The bloom did not extend onto the mid-shelf, where Thaumarchaeota genes ranged from 103 to 105 copies L−1. Ammonia oxidation rates (AO) were highest at inshore stations during summer (to 840 nmol L−1 d−1) and were always at the limit of detection at mid-shelf stations. Nitrite concentrations were correlated with AO (R = 0.94) and were never elevated at mid-shelf stations. Gene sequences from samples collected at mid-shelf stations generated using Archaea 16S rRNA primers were dominated by Euryarchaeota; sequences from inshore and nearshore stations were dominated by Thaumarchaeota. Thaumarchaeota were also abundant at depth at the shelf-break; however, this population was phylogenetically distinct from the inshore/nearshore population. Our analysis shows that the bloom is confined to inshore waters during summer and suggests that Thaumarchaeota distributions in the SAB are controlled primarily by photoinhibition and secondarily by water temperature.
Similar content being viewed by others
Introduction
Studies of seasonal variation in bacterioplankton metatranscriptomes [1, 2] revealed that Thaumarchaeota were abundant in Georgia coastal waters during mid-summer. Thaumarchaeota 16S rRNA genes (rrs) and Archaea ammonia monooxygenase genes (amoA) were detected at >107 copies L−1 in August during quarterly sampling from the ferry dock at Marsh Landing, Sapelo Island, Georgia [3, 4]. These peaks coincided with a mid-summer peak in nitrite concentration [4, 5]. Nitrite is an intermediate in the geochemical process of nitrification, produced by the oxidation of ammonia and removed by oxidation to nitrate [6, 7]. Members of the Archaea (Thaumarchaeota, ammonia-oxidizing archaea or AOA) and Bacteria (Beta- and Gamma-proteobacteria) are known to oxidize ammonia, whereas nitrite is oxidized by several groups of Bacteria [8]. Recently, members of the genus Nitrospira have been identified that can perform both steps of nitrification (complete ammonia oxidation (AO) or comammox; [9, 10]); however, marker genes from organisms capable of comammox have not been detected in the marine environment.
Because our previous studies focused on one location in a dynamic coastal environment, they could not distinguish between in situ growth of the Thaumarchaeota population there vs. local population dynamics coupled to broader scale circulation patterns that could include advection or mixing from a population outside the immediate study area. The Marsh Landing sampling site is at the mouth of the Duplin River, a tidal creek draining ~ 1200 ha of salt marsh inshore of Sapelo Island. The station is located adjacent to, and exchanges water with, Doboy Sound and thence the South Atlantic Bight (SAB) on every tidal cycle. The SAB is characterized by a broad, shallow (<50 m) continental shelf that extends ~ 1000 km along the southeast coast of the United States, from approximately West Palm Beach FL to Cape Hatteras NC, and 130 km across the shelf at its apex at Savannah, GA, USA. Water properties in the nearshore zone of the SAB are influenced by river flow and tidal exchange with extensive salt marshes and estuaries. Nearshore (z < 15 m) flow in the SAB trends from north to south, while shelf-break and slope waters (z > 50 m) are influenced by the Gulf Stream, flowing south to north [11]. The water column over the continental shelf is well-mixed by tidal currents (semidiurnal, 2–3 m tidal amplitude at the apex of the SAB), but is stratified over deeper water at the shelf-break, with a thermocline between 50 and 100 m [11].
We extended our survey of Thaumarchaeota populations to continental shelf and shelf-break waters of the SAB to evaluate the areal extent of the bloom and to detect source populations, if any, that might influence population dynamics at our primary study site. We sampled at approximately monthly intervals from April to November, 2014, to determine whether the seasonal dynamics of Thaumarchaeota populations in the SAB reflected, or drove, the dynamics we recorded at Marsh Landing. Although covariance of nitrite concentrations and Thaumarchaeota abundance at Marsh Landing strongly suggested that this population oxidizes ammonia, Hollibaugh et al. [4] did not measure AO rates. Thus, we measured AO using 15N-labeled ammonium, with the goal of assessing the magnitude of rates and determining whether rates covaried with the distribution of Thaumarchaeota. In addition to recording Thaumarchaeota abundance, we also determined the abundance of amoA genes from ammonia-oxidizing betaproteobacteria (AOB) to evaluate their distribution relative to AOA and AO. Finally, we measured the abundance of 16S rRNA genes (rrs) from Nitrospina, an important member of the guild of marine nitrite-oxidizing bacteria (NOB; [12,13,14,15]), to evaluate coupling between ammonia- and nitrite oxidation in these waters.
Methods
Sampling Site
Samples were collected on six cruises of the R/V Savannah across the SAB in April, May, July, August, September, and November of 2014 (See Supplemental Methods for details). Stations occupied on each cruise are shown in Fig. 1. Surface and bottom water samples were collected at all continental shelf stations. We collected samples at depths of 10, 75 or 100, 200, and 350 or 400 m at shelf-break stations. Samples were collected using 12 L Niskin bottles mounted on a rosette frame equipped with a Sea-Bird SBE 25 CTD. Profiles of environmental variables: conductivity (C, S/m), temperature (T, oC), salinity (S, PSU), dissolved oxygen (DO, mL L−1, SBE43), relative fluorescence (WetLabs WetStar fluorometer, calibrated to extracted chlorophyll, μg L−1), photosynthetically active radiation (PAR, μmoles of photons m−2 s−1, Li-Cor 2π sensor, 400–700 nmol L−1) and turbidity assessed as optical backscatter (OBS, Wetlabs BB, 600 nmol L−1, arbitrary units) were taken during hydrocasts by sensors integrated into the CTD package. Note that the OBS sensor saturated at some of the turbid inshore stations (OBS > 18.5). DNA was collected by filtering ~ 1 L of water from each station and depth sampled through 0.22 µm pore size Durapore filters (EMD Millipore). The filters were placed in Whirl-Pak bags, 2 mL of lysis buffer (0.75 mol L−1 sucrose, 40 mmol L−1 EDTA, 50 mmol L−1 Tris; pH 8.3) was added, and the filters were frozen on dry ice and stored at −80 °C until they were processed as described previously [16].
Location of South Atlantic Bight stations sampled for this study. The area shown in light gray in each panel is above sea level, the darker gray is below sea level (depth indicated by isobaths) and the blue line separates the mainland from salt marsh and barrier islands. Sampling dates are: 2–6 April, 15–18 May, 15–19 July, 4–7 August, 23–30 September, and 3–7 November
Approximately 100 mL of the filtrate was frozen at −20 °C for analysis of nitrate, nitrite, ammonium, and urea. Nitrate plus nitrite (NOx) was determined by reducing nitrate (NO3−) to nitrite (NO2−) with cadmium [17]. Nitrite was determined following Strickland and Parsons [18]. Nitrate was calculated as the difference between NOx and NO2− with negative values, reflecting loss of NO2− during cadmium reduction [18], assumed to be 0. Ammonium was measured using a fluorometric method described in Holmes et al. [19]. Urea was measured following Mulvenna and Savidge [20] and Revilla et al. [21]. Samples (~ 1 L) for chlorophyll analysis were collected on 47 mm diameter Whatman GF/C glass fiber filters. Chlorophyll a was extracted from the filters into 90% acetone and measured spectrophotometrically as described in Strickland and Parsons [18].
Quantitative PCR
Quantitative PCR (qPCR) was performed as described previously [16] using the primers and probes given in Table S1. Note that Archaea amoA was quantified using the Wuchter et al. [22] primer set, which is biased against amoA genes from the deep-water clade of AOA [16, 23] that we encountered at shelf-break stations. Gene abundance (copies L−1) was calculated as described in Kalanetra et al. [24]. Additional details are provided in Supplemental Materials.
Phylogenetic analysis
DNA was collected on the July cruise from surface water samples taken at inshore (SAPES, DOBES, and ALTES), nearshore (C1), and mid-shelf (C7) samples; and from samples taken at 10 and 200 m at a shelf-break station (C17). Archaea rrs (~ 900 bp) and amoA (~ 635 bp) fragments were amplified from these samples using primers listed in Table S1. Amplicons were then cloned and sequenced as described previously [16, 24].
Operational taxonomic units (OTUs) were defined with identity cutoffs of 99% for Thaumarchaeota rrs, 98% for Archaea amoA and 97% for Euryarchaeota rrs and were determined from the similarity matrix for the final, edited nucleotide alignments. Phylogenetic trees of Archaea amoA, Thaumarchaeota rrs and Euryarchaeota rrs were constructed using MR BAYES [25] implemented in Geneious 10.1.3 [26].
AO rates
AO rates were measured in samples contained in 50 mL polypropylene tubes by adding 15N-labeled ammonium (>99 atom-percent 15NH4Cl; Cambridge Isotope Laboratories, Tewksbury, MA, USA) to a final concentration of 50 nM. Tubes were wrapped in aluminum foil and placed in a flowing surface seawater bath on deck. This bath was up to 15 °C warmer than in situ temperatures at depth at shelf-break stations, but was the same as bottom water temperatures at all other stations (Data set S1). Incubations were terminated after 24 h by freezing the whole tube at −20 °C. Control tubes were frozen immediately after adding 15NH4Cl. The 15NOx produced was converted to 15N2O using the “denitrifier method” [27] then processed as described previously [28,29,30]. 15N2O concentrations and δ15N were used to calculate AO rates with modified equations described previously [30,31,32]. Examination of the 15N data from the August cruise suggested a methodological error, leading us to discard it from further consideration.
Data archives
Nucleotide sequences representing OTUs have been submitted to NCBI Genbank under accession numbers KX697614-KX697730. The environmental data included in Data set S1, including metadata, have been archived with BCO-DMO under project “GCE-LTER,” identifier “SAB NITRO CRUISES 2014.”
Data analysis
The stations sampled were aggregated into four regions for subsequent analysis: inshore, nearshore, mid-shelf, and shelf-break; based on their locations and environmental characteristics. Inshore stations were between the barrier island complex and the mainland and have characteristics similar to the Marsh Landing station. Nearshore stations are outside the barrier island complex to depths of ~ 15 m. Stations at the edge of the shelf-break (depth ≥ 50 m) can be influenced by periodic intrusions of slope water; thus, they are considered separately from mid-shelf stations in this study (Fig. 1). Mid-shelf stations are between those defined as nearshore and shelf-break (depths between ~ 15–50 m, see Data set S1).
PAR attenuation coefficients were calculated from plots of ln(PAR) vs. depth following Lin et al. [33] and Liu et al. [34]. The OBS values reported are averages of 100 scans of unbinned output from the sensor package on the downcast, generally covering <2 m of the upper water column. Data were curated manually to ensure that plots of ln(PAR) vs. depth were linear and that OBS was constant through the depth interval analyzed from a given profile.
Chlorophyll a concentrations were estimated from relative fluorescence measured by the fluorometer in the CTD sensor package, calibrated against extracted chlorophyll. We found a significant correlation between the relative fluorescence signal and extracted chlorophyll a concentration (n = 280, R = 0.91, p < 0.05; data not shown).
Model II, major axis linear regressions [35] calculated using R [36] were used to evaluate relationships between pairs of variables. Confidence intervals for the slopes and intercepts were computed from 999 bootstrap permutations. Pearson’s product moment correlations were used to calculate correlation coefficients (R) between pairs of variables. Variables were log-transformed as needed prior to analysis.
Redundancy analysis (RDA) was performed with 999 Monte-Carlo permutations using the software package CANOCO v4.5 [37] to determine the relationship between the distribution of Thaumarchaeota marker genes and environmental variables. One analysis (RDA-1) examined the distribution of Thaumarchaeota marker genes in surface waters (z < 50 m) on all cruises. A second (RDA-2) focused on inshore stations where seasonal variation in the abundance of Thaumarchaeota marker genes was greatest in the SAB. A third (RDA-3) analyzed the onshore-offshore pattern of spatial variation across the SAB. As the abundance of Thaumarchaeota marker genes was uniformly low throughout the study area in April, May, and November, we restricted RDA-3 to data collected during the July, August, and September cruises, as they represented the strongest spatial gradients of Thaumarchaeota abundance in the study area during the period of observation.
Results
Environmental conditions in SAB continental shelf waters
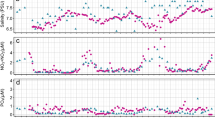
The water column was well-mixed at inshore and continental shelf stations during the months sampled (Data set S1). Temperature, salinity, DO, and relative fluorescence in surface waters changed seasonally, while OBS and the PAR attenuation coefficient did not (Fig. 2, Data set S1). Temperature increased from April (16 °C) to August (30 °C), and then decreased in fall (to 19 °C). Temperature was uniform across inshore, nearshore, and mid-shelf stations (Fig. 2a). Freshwater inflow from the Altamaha River lowered salinity at inshore stations, especially during April (to 23 PSU, Fig. 2b). The concentration of DO ranged from 3 to 7 mL L−1 between April and November and did not vary across the study area (Fig. 2c). Chlorophyll a concentration calculated from relative fluorescence, did not show a strong seasonal cycle but decreased from inshore stations to the shelf-break (Fig. 2d and Data set S1). The PAR attenuation coefficient and OBS covaried (Pearson’s R = 0.91, p < 0.001, data not shown), were highest at inshore stations, and did not change seasonally (Fig. 2e, f).
Seasonal and spatial variability of: a temperature (°C), b salinity (PSU), c dissolved oxygen concentration (mL L−1), d chlorophyll from relative fluorescence (µg L−1), e PAR extinction coefficient (m−1), and f the optical backscatter signal (OBS, relative units) in surface waters (<50 m) of the South Atlantic Bight. The data were aggregated into inshore (solid line, X), nearshore (long dash, triangle), mid-shelf (medium dash, circle), and shelf regions (short dash, square) based on location, water column depth, and optical properties. The graph displays mean and standard error of all measurements made in samples taken in a given region on each cruise. The full data set is presented in Data set S1
Concentrations of NH4+, NO2−, and NO3− were highest at inshore stations and were much lower or undetectable in surface waters further offshore. Ammonium concentration at inshore stations averaged 2.1 μmol L−1 (range 0.2-6.2), whereas at other stations it was <0.8 μmol L−1 (Fig. 3a, Data set S1). Concentrations of NO2− on the continental shelf were low (mean < 0.3 μmol L−1, range 0–0.3), but were higher at inshore stations from July to September (mean > 0.9 μmol L−1; Fig. 3b, Data set S1). The concentration of NO3− was highest on average at inshore stations (mean 1.61 μmol L−1, range 0.1–8.5); Fig. 3c, Data set S1). The average concentration of urea was also highest at inshore stations (mean 0.3 μmol L−1, range 0.07−0.64) and ranged from 0 to 1.0 µmol L−1 in surface waters at all stations during all cruises (Fig. 3d, Data set S1).
Seasonal and spatial variability of the concentrations (μmol L−1) of a ammonium, b nitrite, c nitrate, and d urea; symbols as in Fig. 2
Seasonal distributions of nitrifying organisms
The abundance of Thaumarchaeota marker genes was similar in surface and bottom water samples taken at stations on the continental shelf (Data set S1). Abundances of Thaumarchaeota rrs and Archaea amoA covaried during all cruises (r2 = 0.83, p < 0.0001; Fig. S1a). Thaumarchaeota rrs and Archaea amoA genes were most abundant in inshore waters (105–108 copies L−1) and decreased to the mid-shelf (103–105 copies L−1). The abundances of both genes in surface waters of shelf-break stations were similar to those at mid-shelf stations except during April and November, when Thaumarchaeota abundances in surface samples from shelf-break stations were >105 copies L−1 (Fig. 4a, b; Data set S1).
Seasonal and spatial distributions of: a Archaea amoA (106 copies L−1), b Thaumarchaeota rrs (106 copies L−1), c) Betaproteobacteria amoA (103 copies L−1), and d Nitrospina sp. rrs genes (106 copies L−1), symbols as in Fig. 2
Seasonal patterns of Thaumarchaeota abundance differed among inshore, nearshore, mid-shelf, and shelf-break stations (Fig. 4a, b). Thaumarchaeota populations at inshore and nearshore stations increased >100-fold from April to August or September, then decreased in November (Fig. 4a, b; Data set S1). Thaumarchaeota abundances at mid-shelf stations varied ≤10-fold over the period of observation and did not show a strong seasonal signal (Fig. 4a, b; Data set S1). The abundance of Thaumarchaeota rrs and AOA amoA in surface waters at shelf-break stations decreased 100-fold from April to July, then increased in November (Fig. 4a, b; Data set S1).
The abundances of Betaproteobacteria amoA and Nitrospina sp. rrs genes ranged from 102 to 105 copies L−1 and 104 to 106 copies L−1, respectively; decreasing from inshore to shelf-break surface waters (Fig. 4c, d). They did not show a strong seasonal pattern of variation in abundance (<1 order of magnitude; Fig. 4c, d, Data set S1). The abundance of Betaproteobacteria amoA was only 0.1–10% of Archaea amoA at inshore stations during summer (Data set S1). Nitrospina rrs gene abundance was significantly, but weakly correlated with Thaumarchaeota rrs gene abundance across all samples collected (r2 = 0.1, slope = 0.004, p < 0.0001; Fig. S1b).
Vertical distribution of Thaumarchaeota at shelf-break stations
The water column was stratified at shelf-break stations, with the deepest surface mixed layer (~ 100 m) encountered during November (Fig. S2). Relative fluorescence displayed a pronounced subsurface maximum at ~ 100 m (Fig. S2). DO decreased below the surface mixed layer to <3 mL L−1 (Fig. S2). Nitrite concentrations were low in surface waters ( ≤ 0.2 μmol L−1) but peaked at ~ 100 m, coinciding with subsurface maxima in the abundance of Thaumarchaeota marker genes, AO rates and chlorophyll fluorescence (Fig. S2, Data set S1).
AO rates in the SAB
AO rates were greatest at inshore stations, similar to the distribution of Archaea amoA genes (Table 1). The maximum AO rate, 840 nmol L−1 d−1, was measured in bottom water at inshore station ALTES during September (Data set S1). AO rates at inshore stations were lowest in November (2.7 ± 1.1 nmol L−1 d−1; Table 1 and Data set S1). AO rates were extremely low (≤0.3 nmol L−1 d−1) at all nearshore and mid-shelf stations, except in September at nearshore stations A1 (surface: 104 nmol L−1 d−1, bottom: 132 nmol L−1 d−1) and C1 (bottom: 4.9 nmol L−1 d−1), which were comparable to AO rates measured at adjacent inshore stations (Data set S1). Maximum AO rates at shelf-break stations were detected at depth and coincided with maxima in Thaumarchaeota abundance (Data set S1).
Nitrite concentration correlated strongly with AO rates in the SAB, either considering all samples or only inshore samples (Table S2). AO rates covaried with abundances of Thaumarchaeota rrs, Betaproteobacteria amoA and [NH4+] in all samples (Table S2). AO rates were not significantly correlated with Thaumarchaeota amoA abundance because the primers we used underestimate Thaumarchaeota amoA abundance in the deep-water populations of Thaumarchaeota we found at the shelf-break.
Phylogenetic composition of Archaea populations in the SAB
We retrieved a total of 253 Archaea amoA gene sequences in seven libraries of cloned amplicons from water samples collected on the July cruise. Each of these libraries contained from 8 to 27 OTUs, defined at 98% nucleotide identity, with a total of 89 OTUs recovered (Table S3). Twenty OTUs were shared among all stations.
The diversity and richness of Archaea amoA sequences were highest at inshore stations DOBES and ALTES, and at 200 m at the shelf-break station (Table S3). Most of the Archaea amoA gene sequences from inshore (SAPES, 91%; DOBES, 78%; ALTES, 81%) and nearshore (C1, 71%) stations fell into the ‘coastal water/sediment’ clade [38], with most sequences from SAPES (71%) and C1 (42%) closely related to ‘Candidatus Nitrosopumilus maritimus’ strain SCM1 (>98% identity; Fig. S3). Eighty-six percent of the sequences from mid-shelf Station C7 and 61% of the sequences from 10 m at the shelf-break fell into the WCA (Water column clade A, a shallow water, marine ecotype; [38]); along with 29% of the sequences retrieved from Station C1 and 22% of the sequences retrieved from DOBES. Eighty nine percent of the OTUs recovered from 200 m at Station C17 were unique to that depth and 18 of 27 OTUs, ~ 65% of the sequences recovered from that library, fell into WCB (Water Column clade B, [38]), a deep water, marine ecotype (Fig. S3). Three OTUs recovered from 10 m at Station C17 and 1 OTU from Station C7 also fell into WCB. The remaining OTUs recovered from 200 m at Station C17 fell into WCA (Fig. S3), along with three OTUs from 10 m at Station C17. Two OTUs from Station ALTES clustered with the soil clade.
Approximately 80% of the Archaea rrs sequences retrieved from 200 m at shelf-break station C17 were from Thaumarchaeota and 90% of these, and both Thaumarchaeota sequences recovered from 10 m at this station, grouped with reference sequences from deep water (Fig. S4). Only two of the sequences recovered from the shelf-break clustered with the open ocean, surface water isolate ‘Ca. Nitrosopelagicus brevis’ strain CN25 [39] at >93% identity (Fig. S4). Most of the Thaumarchaeota rrs sequences retrieved from inshore or nearshore stations were closely related (>98% identity) to ‘Ca. N. maritimus’ strain SCM1 and to newly isolated Thaumarchaeota strains from Puget Sound (‘Ca. Nitrosopumilus sp.’ strains PS0 and HCA1; [40]) and the northern Adriatic Sea (‘Ca. Nitrosopumilus piranensis’ strain D3C and ‘Ca. Nitrosopumilus adriaticus’ strain NF5; [41]; Fig. S4).
Clone libraries of Archaea rrs genes were dominated by Euryarchaeota (67.6% of all sequences retrieved, Table S3), especially in samples from offshore stations (93%). Most of these grouped with Marine Group II Archaea ([42, 43]; Fig. S5). Thaumarchaeota rrs sequences retrieved from inshore stations were most similar to members of the Nitrosopumilaceae, including sequences retrieved previously from Marsh Landing [3], whereas rrs sequences retrieved from shelf-break samples were most similar to sequences from other open ocean and deep-water samples. This split in phylogenetic composition of populations of Thaumarchaeota rrs genes parallels that of the amoA sequences we retrieved from the same samples.
Relationship of Thaumarchaeota distributions to environmental factors
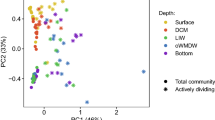
The first two axes of RDA-1, which evaluated the relationship between abundances of Thaumarchaeota marker genes and environmental variables in surface water samples (z < 50 m) taken during all cruises, explained 67.2% of total variation, of which 66.7% was explained by Axis 1 (Table S4). The biplot of RDA-1 (Fig. 5a) shows that OBS, temperature, Bacteria rrs gene abundance, and DO (in order of decreasing importance) contributed significantly (p < 0.05) to explaining the variance in Thaumarchaeota gene abundances (Fig. 5a and Table S4). OBS explained 54% of the variance in Thaumarchaeota gene abundances in this analysis, while temperature explained 7% (Table S4). Temperature explained 33% of a total of 63% of the seasonal variation in Thaumarchaeota gene abundance at inshore stations explained by the first two axes of RDA-2. OBS explained an additional 12% of the variance in this analysis (Fig. 5b, Table S4). OBS explained most of the spatial variation (the onshore-offshore gradient) of Thaumarchaeota gene abundances during the summer (69% of a total of 80.1% of the variance explained by the first two axes of RDA-3; Fig. 5c, Table S4). Although the PAR attenuation coefficient (kz) covaried with OBS (Pearson’s R = 0.89, p < 0.001), it did not contribute significantly to any of the RDA analyses (p > 0.1; Table S4). Bacterial rrs gene abundance, temperature, DO, urea concentration, and relative fluorescence also contributed significantly to explaining seasonal variation in Thaumarchaeota abundance at inshore stations in RDA-2 and to explaining spatial variation in the intensity of the bloom across the SAB in RDA-3 (Fig. 5b, c; Table S4).
Redundancy analysis (RDA) ordination plots of Thaumarchaeota rrs and Archaea amoA genes vs. environmental factors for a surface water (<50 m) samples of the SAB on all cruises, b in nearshore waters, and c for surface waters at all stations on the July, August, and September cruises. The length and angle of arrows show the contribution of each variable to the RDA axes (also reported in Table S4). RF, relative fluorescence (chlorophyll a); T, temperature; S, salinity; DO, dissolved oxygen concentration; B-rrs, Bacteria 16S rRNA gene abundance; T-rrs, Thaumarchaeota 16S rRNA gene abundance; amoA, Archaea amoA gene abundance
Discussion
Thaumarchaea populations are partitioned by habitat in the SAB
Clone libraries from inshore and nearshore samples shared few Thaumarchaeota OTUs with samples from 75 or 200 m at the shelf-break, and phylotypes from each region were closely related to reference sequences from sites with similar oceanographic characteristics (Figs. S3, S4). Phylogenetic analyses showed that inshore and nearshore populations contained the same ecotypes, likely as a result of mixing between these water masses. We encountered amoA sequences that are most closely related to the low-salinity-adapted ‘Ca. Nitrosoarchaeum limnia’ strain SFB1 [44] at inshore stations ALTES and DOBES, which are more strongly influenced by freshwater input from the Altamaha River than SAPES or the nearshore stations.
Euryarchaeota dominated libraries of Archaea rrs genes, accounting for 67% of all Archaea rrs sequences retrieved from our samples (Table S3). This is similar to the distribution of Archaea in the Northeastern South China Sea [45]. Most of these sequences were affiliated with the Marine Group II and III Archaea (Figure S5), especially in samples from mid-shelf and shelf-break stations. We recovered relatively fewer Euryarchaeota rrs sequences from 200 m at shelf-break station C17 compared with the 10 m sample from this station, consistent with the global distribution of MGII Archaea, which are reported to decrease in abundance with depth [14, 15, 42, 43, 46, 47]. The marked difference in the distribution of Euryarchaeota and Thaumarchaeota populations in our samples suggests very different responses of these groups to environmental factors, especially photoinhibition, that may be controlling their distributions in the SAB.
Seasonality of Thaumarchaeota in the SAB
The data collected during the present study establish that a mid-summer peak in Thaumarchaeota abundance occurs at all of the inshore and nearshore stations of the SAB we sampled (Fig. 4). Thaumarchaeota never became abundant at mid-shelf stations (Fig. 4), and the deep-water population at the shelf-break, which was present throughout the study (Fig. S2), was phylogenetically distinct from the inshore population (Figs S3 and S4), ruling it out as a potential source of the inshore population. These observations are consistent with data from our long-term monitoring at Marsh Landing and suggest that observations made at Marsh Landing represent processes in the broader inshore and nearshore area of the SAB. We conclude that the seasonal bloom of Thaumarchaeota we documented at Marsh Landing is the result of local growth, triggered by some combination of environmental variables, rather than the consequence of fortuitous sampling of a population that developed elsewhere.
Effect of light on Thaumarchaeota distributions
The mid-summer peak in Thaumarchaeota abundance at nearshore and inshore stations in the SAB contrasts with the mid-winter maxima of Thaumarchaeota abundance reported for coastal waters at high latitudes, such as the North Sea [22, 48, 49] and polar oceans [50, 51]. Light has been suggested to inhibit Thaumarchaeota growth [40, 52], though the question of whether light affects Thaumarchaeota directly or via photochemical production of reactive oxygen species [53] remains to be resolved [54]. The summer disappearance of Thaumarchaeota from the euphotic zone at high latitudes could thus be a response to the strong seasonal increase in irradiance there. Thaumarchaeota never became abundant at mid-shelf stations of the SAB, despite the similarity of seasonal patterns of water temperature and DO between inshore and mid-shelf stations (Fig. 2d). Light penetration is considerably greater at mid-shelf stations compared with inshore and nearshore stations (Fig. 2f), where turbidity owing to sediment resuspended by strong tidal mixing (Fig. 2f, S6) and CDOM supplied by river discharge [55] result in high attenuation coefficients (Fig. 2e). OBS emerged as a significant (positive) environmental factor in our RDA analysis of Thaumarchaeota distribution data. Greater light penetration in the offshore water column could thus have inhibited Thaumarchaeota growth at mid-shelf stations of the SAB, though other factors, such as decreasing fluxes of regenerated ammonia along the gradient from inshore stations to the shelf-break, may also have influenced growth of Thaumarchaeota. Support for this hypothesis comes from the significant, if minor, contribution of the abundance of Bacteria rrs to distributions of Thaumarchaeota revealed by the RDA analyses.
Mid-winter maxima of Thaumarchaeota abundance have also been reported from studies at lower latitudes [56]. These maxima may be the result of enhanced vertical mixing during winter, as suggested by Mincer et al. [15]. Kalanetra et al. [24] also suggested that mixing might cause the mid-winter peak in Thaumarchaeota abundance in coastal waters off the West Antarctic Peninsula; however, subsequent work in this area [57] indicated that this population was actively growing. Similar seasonal changes in vertical mixing (Fig. S2) may explain the elevated concentrations of Thaumarchaeota we observed in surface waters at this station in April and November.
Effect of water temperature
RDA analysis suggests that water temperature influences the seasonal abundance and activity of Thaumarchaeota at inshore stations (Fig. 5b). Thaumarchaeota isolates from Puget Sound have been shown to have high Q10 values [40] with maximum growth rates at >25 °C. Analyses of the effects of incubation temperature on nitrite production and accumulation made using a nitrifying enrichment culture raised from a Marsh Landing inoculum [5] revealed similar temperature dependencies for activity of this population, and further, that temperatures above 40 °C completely inhibit AO by these populations. The mid-summer nitrite peak at inshore stations seen in this study can be explained as a consequence of the uncoupling of ammonia- and nitrite oxidation. Comparison of data sets from other coastal waters [5, 58] suggests that this is a common pattern for nitrification in temperate estuarine waters.
Competition with phytoplankton
Phytoplankton have been suggested to compete with Thaumarchaeota for NH4+ in the North Sea [22] and to inhibit the growth of Thaumarchaeota during the spring and summer in Antarctic coastal waters [51]. Martens-Habbena et al. [59] and Urakawa et al. [60] have demonstrated that phytoplankton are unlikely to be able to compete directly with Thaumarchaeota for ammonia, and chlorophyll concentration did not emerge as a major explanatory variable in our RDA analyses. Positive correlations between Thaumarchaeota and phytoplankton populations have been noted in a few studies [16, 61].
Role of ammonia availability
Ammonia availability has also been suggested to control the distribution of Thaumarchaeota [59, 60, 62]. Ammonium concentrations measured in the SAB ranged from the limit of analytical detection at mid-shelf and shelf-break stations to 6.4 µmol L−1 in inshore waters (Data set S1). Ammonium concentrations at the limit of analytical detection (~ 50 nmol L−1) may not be low enough to limit Thaumarchaeota, which are reported to have thresholds for ammonium uptake that are ≤ 10 nmol L−1 [59]. AO rates in our samples were not correlated strongly with ammonia concentration (Table S2), and ammonium concentration did not contribute significantly to the Thaumarchaeota-environment relationship in our RDA analyses (Table S4). Although ammonium availability does not appear to be the key factor determining the distribution of Thaumarchaeota abundance in the SAB, the summer bloom of Thaumarchaeota at inshore stations coincided with relatively high ammonium concentrations (mean 2.1 μmol L−1, range 0.2–6.2), whereas ammonium concentrations were consistently low at mid-shelf stations (Fig. 3a, Data set S1).
The significant, but small, contribution of urea to Thaumarchaeota-environment relationships (p = 0.006, 8% of the variance explained in RDA-2; Fig. 5b, Table S4) suggests that seasonal variability in urea concentration may contribute to the timing of the inshore bloom. Urea can serve as an alternate source of reduced nitrogen for AOA [63,64,65], potentially having a minor role in nitrification in the SAB.
Role of copper limitation
The availability of free copper (Cu2+) has also been suggested to limit growth of Thaumarchaeota in coastal environments [66]. We addressed this question directly in a separate study [67]. We found that Cu2+ concentrations at six inshore sites, including Marsh Landing, were consistently at sub-fmol L−1 levels as a result of complexation by two ligand classes, well below the concentrations reported to limit the growth the Thaumarchaeota [68], yet a mid-summer bloom of Thaumarchaeota was apparent at all of the stations sampled [67].
Shelf-break populations
The highest AO rates at shelf-break stations coincided with peak concentrations of Thaumarchaeota abundance, nitrite and chlorophyll (Figure S2, Data set S1). The AO rate at the bottom of the euphotic zone at the shelf-break was much lower than AO rates measured at inshore stations during summer, although AO rates measured were likely higher than in situ because of the elevated incubation temperature. Thaumarchaeota ecotypes exhibit different growth rates and cell-specific AO rates [40, 59, 69,70,71]. Smith et al. [72] found that AO rates in Monterey Bay surface waters correlated with the abundance of AOA from the WCA clade, whereas deep-water samples, dominated by AOA from the WCB clade, consistently had low AO rates. We found two distinct Thaumarchaeota populations at inshore vs. shelf-break samples from the SAB (Figs. S3, S4). The former was composed mainly of organisms with amoA sequences most similar to “Ca. N. maritimus” strain SCM1, whereas the latter contained a mix of AOA from the WCA and WCB clades (Fig. S3). Thus, greater cell-specific AO rates of “Ca. N. maritimus” strain SCM1 [59] compared with open ocean strains [59, 71] could contribute to higher AO rates in inshore waters.
Conclusions
Thaumarchaeota were never abundant on the SAB continental shelf, and the deep-water population found at the shelf-break was phylogenetically distinct from the inshore population, indicating that the summer bloom observed previously at an inshore site does not originate offshore. OBS (light penetration) explains most of the variation in the spatial distribution of Thaumarchaeota in the SAB, suggesting that factors related to light penetration control the potential for the development of Thaumarchaeota populations. Within the inshore zone of the SAB where light penetration is consistently low, water temperature appears to exert secondary control on the potential for bloom development. The weak correlation between Thaumarchaeota and Nitrospina distributions and the strong correlation between AO and nitrite concentration is consistent with the mid-summer accumulation of nitrite in inshore waters that appears to result from differential responses of ammonia- and nitrite-oxidizing organisms to seasonal variation in water temperature [73,74,75,76,77,78,79,80].
References
Gifford SM, Sharma S, Rinta-Kanto JM, Moran MA. Quantitative analysis of a deeply sequenced marine microbial metatranscriptome. ISME J. 2011;5:461–472.
Gifford SM, Sharma S, Booth M, Moran MA. Expression patterns reveal niche diversification in a marine microbial assemblage. ISME J. 2013;7:281–298.
Hollibaugh JT, Gifford S, Bano N, Sharma S, Moran MA. Metatranscriptomic analysis of ammonia-oxidizing organisms in an estuarine bacterioplankton assemblage. ISME J. 2011;5:866–878.
Hollibaugh JT, Gifford SM, Moran MA, Ross MJ, Sharma S, Tolar BB. Seasonal variation in the metratranscriptomes of a Thaumarchaeota population from SE USA coastal waters. ISME J. 2014;8:685–698.
Schaefer SC, Hollibaugh JT. Temperature decouples ammonium and nitrite oxidation in coastal waters. Environ Sci Technol. 2017;51:3157–3164.
Francis CA, Beman JM, Kuypers MMM. New processes and players in the nitrogen cycle: the microbial ecology of anaerobic and archaeal ammonia oxidation. ISME J. 2007;1:19–27.
Ward BB. Nitrification in the Ocean. In: Ward B, Arp DJ, Klotz MJ, (eds). Nitrification. Washington, DC, USA: ASM Press; 2011. pp. 325–345.
Daims H, Lucker S, Le Paslier D, Wagner M. Diversity, environmental genomics and ecophysiology of nitrite-oxidizing bacteria. In: Ward BB, Arp DJ, Klotz MG, (eds). Nitrification. Washington DC, USA: ASM Press; 2011. pp. 295–322.
Daims H, Lebedeva EV, Pjevac P, Han P, Herbold C, Albertsen M, et al. Complete nitrification by Nitrospira bacteria. Nature. 2015;528:504–509.
Van Kessel MaHJ, Speth DR, Albertsen M, Nielsen PH, Op Den Camp HJM, Kartal B, Jetten MSM, Lücker S. Complete nitrification by a single microorganism. Nature. 2015;528:555–559.
Atkinson LP, Menzel DW, Bush KA, (eds). Oceanography of the Southeastern US Continental Shelf. Washington DC, USA: AGU; 1985. pp. 156.
Watson SW, Waterbury JB. Characteristics of two marine nitrite oxidizing bacteria, Nitrospina gracilis nov. gen. nov. sp. and Nitrococcus mobilis nov. gen. nov. sp. Arch Mikrobiol. 1971;77:203–230.
Suzuki MT, Preston CM, Béjà O, de la Torre JR, Steward GF, DeLong EF. Phylogenetic screening of ribosomal RNA gene-containing clones in Bacterial Artificial Chromosome (BAC) libraries from different depths in Monterey Bay. Microb Ecol. 2004;48:473–488.
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard N-U, et al. Community genomics among stratified microbial assemblages in the ocean’s interior. Science. 2006;311:496–503.
Mincer TJ, Church MJ, Taylor LT, Preston C, Karl DM, DeLong EF. Quantitative distribution of presumptive archaeal and bacterial nitrifiers in Monterey Bay and the North Pacific Subtropical Gyre. Environ Microbiol. 2007;9:1162–1175.
Tolar BB, King GM, Hollibaugh JT. An analysis of Thaumarchaeota populations from the Northern Gulf of Mexico. Front Microbiol. 2013;4:72.
Jones MN. Nitrate reduction by shaking with cadmium: alternative to cadmium columns. Water Res. 1984;18:643–646.
Strickland JDH, Parsons TR. A Practical Handbook of Seawater Analysis. 2 edn. Ottawa: Fisheries Research Board of Canada; 1972. Second Editionvol. Bulletin167.
Holmes RM, Aminot A, Kerouel R, Hooker BA, Peterson BJ. A simple and precise method for measuring ammonium in marine and freshwater ecosystems. Can J Fish Aquat Sci. 1999;56:1801–1808.
Mulvenna PF, Savidge G. A modified manual method for the determination of urea in seawater using diacetylmonoxime reagent. Estuar Coast Shelf Sci. 1992;34:429–438.
Revilla M, Alexander J, Glibert PM. Urea analysis in coastal waters: comparison of enzymatic and direct methods. Limnol Oceanogr Methods. 2005;3:290–299.
Wuchter C, Abbas B, Coolen MJL, Herfort L, van Bleijswijk J, Timmers P, et al. Archaeal nitrification in the ocean. Proc Natl Acad Sci. 2006;103:12317–12322.
Beman JM, Popp BN, Francis CA. Molecular and biogeochemical evidence for ammonia oxidation by marine Crenarchaeota in the Gulf of California. ISME J. 2008;2:429–441.
Kalanetra KM, Bano N, Hollibaugh JT. Ammonia-oxidizingArchaea in the Arctic Ocean and Antarctic coastal waters. Environ Microbiol. 2009;11:2434–2445.
Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17:754–755.
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647–1649.
Sigman DM, Casciotti KL, Andreani M, Barford C, Galanter M, Bohlke JK. A bacterial method for the nitrogen isotopic analysis of nitrate in seawater and freshwater. Anal Chem. 2001;73:4145–4153.
Dore JE, Popp BN, Karl DM, Sansone FJ. A large source of atmospheric nitrous oxide from subtropical North Pacific surface waters. Nature. 1998;396:63–66.
Popp BN, Sansone FJ, Rust TM, Merritt DA. Determination of concentration and carbon isotopic composition of dissolved methane in sediments and nearshore waters. Anal Chem. 1995;67:405–411.
Beman JM, Popp BN, Alford SE. Quantification of ammonia oxidation rates and ammonia-oxidizing archaea and bacteria at high resolution in the Gulf of California and eastern tropical North Pacific Ocean. Limnol Oceanogr. 2012;57:711–726.
Beman JM, Chow C-E, King AL, Feng Y, Fuhrman JA, Andersson A, et al. Global declines in oceanic nitrification rates as a consequence of ocean acidification. Proc Natl Acad Sci. 2011;108:208–213.
Ward BB, Kilpatrick KA, Renger E, Eppley RW. Biological nitrogen cycling in the nitracline. Limnol Oceanogr. 1989;34:493–513.
Lin S, Zou T, Gao H, Guo X. The vertical attenuation of irradiance as a function of turbidity: a case of the Huanghai (Yellow) Sea in spring. Acta Oceanol Sin. 2009;28:66–75.
Liu Q, Lu X, Tolar B, Mou X, Hollibaugh J. Concentrations, turnover rates and fluxes of polyamines in coastal waters of the South Atlantic Bight. Biogeochem. 2015;123:1–17.
Legendre P, Legendre L. Numerical Ecology. Amsterdam: Elsevier Science BV; 1998. 2nd English edn.
R Development Core Team. R: A language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2015.
ter Braak CJF, Šmilauer P (2002). CANOCO Reference manual and CanoDraw for Windows User’s guide: Software for Canonical Community Ordination, v 4.5. Microcomputer Power: Ithica, USA.
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB. Ubiquity and diversity of ammonia-oxidizing Archaea in water columns and sediments of the ocean. PNAS. 2005;102:14683–14688.
Santoro AE, Dupont CL, Richter RA, Craig MT, Carini P, McIlvin MR, et al. Genomic and proteomic characterization of “Candidatus Nitrosopelagicus brevis”: An ammonia-oxidizing archaeon from the open ocean. Proc Natl Acad Sci. 2015;112:1173–1178.
Qin W, Amin SA, Martens-Habbena W, Walker CB, Urakawa H, Devol AH, et al. Marine ammonia-oxidizing archaeal isolates display obligate mixotrophy and wide ecotypic variation. Proc Natl Acad Sci. 2014;111:12504–12509.
Bayer B, Vojvoda J, Offre P, Alves RJE, Elisabeth NH, Garcia JAL, et al. Physiological and genomic characterization of two novel marine thaumarchaeal strains indicates niche differentiation. ISME J. 2016;10:1051–1063.
Fuhrman JA, Davis AA. Widespread Archaea and novel Bacteria from the deep sea as shown by 16S rRNA gene sequences. Mar Ecol Prog Ser. 1997;150:275–285.
Massana R, DeLong EF, Pedros-Alio C. A few cosmopolitan phylotypes dominate planktonic Archaeal assemblages in widely different oceanic provinces. Appl Environ Microbiol. 2000;66:1777–1787.
Blainey PC, Mosier A, Potanina A, Francis CA, Quake SR. Genome of a low-salinity ammonia-oxidizing Archaeon determined by single-cell and metagenomic analysis. PLoS ONE. 2011;6:e16626.
Liu H, Zhang CL, Yang C, Chen S, Cao Z, Zhang Z et al. Marine Group II dominates planktonic Archaea in water column of the Northeastern South China Sea. Front Microbiol. 2017;8:1098.
Herndl GJ, Reinthaler T, Teira E, van Aken H, Veth C, Pernthaler A, et al. Contribution of Archaea to total prokaryotic production in the deep Atlantic Ocean. Appl Environ Microbiol. 2005;71:2303–2309.
Orsi WD, Smith JM, Wilcox HM, Swalwell JE, Carini P, Worden AZ, et al. Ecophysiology of uncultivated marine euryarchaea is linked to particulate organic matter. ISME J. 2015;9:1747–1763.
Herfort L, Schouten S, Abbas B, Veldhuis MJW, Coolen MJL, Wuchter C, et al. Variations in spatial and temporal distribution of Archaea in the North Sea in relation to environmental variables. FEMS Microb Ecol. 2007;62:242–257.
Pitcher A, Wuchter C, Siedenberg K, Schouten S, Sinninghe-Damste JS. Crenarchaeol tracks winter blooms of ammonia-oxidizing Thaumarchaeota in the coastal North Sea. Limnol Oceanogr. 2011;56:2308–2318.
Church MJ, DeLong EF, Ducklow HW, Karner MB, Preston CM, Karl DM. Abundance and distribution of planktonic Archaea and Bacteria in the waters west of the Antarctic Peninsula. Limnol Oceanogr. 2003;48:1893–1902.
Murray AE, Preston CM, Massana R, Taylor LT, Blakis A, Wu K, et al. Seasonal and spatial variability of Bacterial and Archaeal assemblages in the coastal waters near Anvers Island, Antarctica. Appl Environ Microbiol. 1998;64:2585–2595.
Merbt SN, Stahl DA, Casamayor EO, Martí E, Nicol GW, Prosser JI. Differential photoinhibition of bacterial and archaeal ammonia oxidation. FEMS Microbiol Lett. 2012;327:41–46.
Tolar BB, Powers LC, Miller WL, Wallsgrove NJ, Popp BN, Hollibaugh JT. Ammonia oxidation in the ocean can be inhibited by nanomolar concentrations of hydrogen peroxide. Front Mar Sci. 2016a;3. https://doi.org/10.3389/fmars.2016.00237.
Hollibaugh JT. Oxygen and the activity and distribution of marine Thaumarchaeota. Environ Microbiol Rep. 2017;9:186–188.
Medeiros PM, Babcock-Adams L, Seidel M, Castelao RM, Di Iorio D, Hollibaugh JT et al. (2017). Export of terrigenous dissolved organic matter in a broad continental shelf. Limnol Oceanogr. In press. https://doi.org/10.1002/lno.10528.
Galand PE, Gutiérrez-Provecho C, Massana R, Gasol JM, Casamayor EO. Inter-annual recurrence of archaeal assemblages in the coastal NW Mediterranean Sea (Blanes Bay Microbial Observatory). Limnol Oceanogr. 2010;55:2117–2125.
Tolar BB, Ross MJ, Wallsgrove NJ, Liu Q, Aluwihare LI, Popp BN, et al. Contribution of ammonia oxidation to chemoautotrophy in Antarctic coastal waters. ISME J. 2016c;10:2605–2619.
Bristow LA, Sarode N, Cartee J, Caro-Quintero A, Thamdrup B, Stewart FJ. Biogeochemical and metagenomic analysis of nitrite accumulation in the Gulf of Mexico hypoxic zone. Limnol Oceanogr. 2015;60:1733–1750.
Martens-Habbena W, Berube PM, Urakawa H, de la Torre JR, Stahl DA. Ammonia oxidation kinetics determine niche separation of nitrifying Archaea and Bacteria. Nature. 2009;461:976–979.
Urakawa H, Martens-Habbena W, Huguet C, de la Torre JR, Ingalls AE, Devol AH, et al. Ammonia availability shapes the seasonal distribution and activity of archaeal and bacterial ammonia oxidizers in the Puget Sound Estuary. Limnol Oceanogr. 2014;59:1321–1335.
Wells LE, Cordray M, Bowerman S, Miller LA, Vincent WF, Deming JW. Archaea in particle-rich waters of the Beaufort Shelf and Franklin Bay, Canadian Arctic: clues to an allochthonous origin? Limnol Oceanogr. 2006;51:47–59.
Horak REA, Qin W, Schauer AJ, Armbrust EV, Ingalls AE, Moffett JW, et al. Ammonia oxidation kinetics and temperature sensitivity of a natural marine community dominated by Archaea. ISME J. 2013;7:2023–2033.
Alonso-Sáez L, Waller AS, Mende DR, Bakker K, Farnelid H, Yager PL, et al. Role for urea in nitrification by polar marineArchaea. Proc Natl Acad Sci. 2012;109:17989–17994.
Lu L, Han W, Zhang J, Wu Y, Wang B, Lin X, et al. Nitrification of archaeal ammonia oxidizers in acid soils is supported by hydrolysis of urea. ISME J. 2012;6:1978–1984.
Tolar BB, Wallsgrove NJ, Popp BN, Hollibaugh JT. Oxidation of urea-derived nitrogen by Thaumarchaeota-dominated marine nitrifying communities. Environ Microbiol. 2016b; In press. https://doi.org/10.1111/1462-2920.13457.
Jacquot JE, Horak REA, Amin SA, Devol AH, Ingalls AE, Armbrust EV, et al. Assessment of the potential for copper limitation of ammonia oxidation by Archaea in a dynamic estuary. Mar Chem. 2014;162:37–49.
Whitby H, Hollibaugh JT, van den Berg CMG. Chemical speciation of copper in a salt marsh estuary and bioavailability to Thaumarchaeota. Front Mar Sci. 2017;4:178.
Amin SA, Moffett JW, Martens-Habbena W, Jacquot JE, Han Y, Devol A, et al. Copper requirements of the ammonia-oxidizing archaeon Nitrosopumilus maritimus SCM1 and implications for nitrification in the marine environment. Limnol Oceanogr. 2013;58:2037–2045.
Lehtovirta-Morley LE, Stoecker K, Vilcinskas A, Prosser JI, Nicol GW. Cultivation of an obligate acidophilic ammonia oxidizer from a nitrifying acid soil. Proc Natl Acad Sci. 2011;108:15892–15897.
Santoro AE, Casciotti KL, Francis CA. Activity, abundance and diversity of nitrifying archaea and bacteria in the central California Current. Environ Microbiol. 2010;12:1989–2006.
Santoro AE, Casciotti KL. Enrichment and characterization of ammonia-oxidizing archaea from the open ocean: phylogeny, physiology and stable isotope fractionation. ISME J. 2011;5:1796–1808.
Smith JM, Casciotti KL, Chavez FP, Francis CA. Differential contributions of archaeal ammonia oxidizer ecotypes to nitrification in coastal surface waters. ISME J. 2014;8:1704–1714.
DeLong EF. Archaea in coastal marine environments. Proc Natl Acad Sci USA. 1992;89:5685–5689.
Hornek R, Pommerening-Roser A, Koops H-P, Farnleitner AH, Kreuzinger N, Kirschner A, et al. Primers containing universal bases reduce multiple amoA gene specific DGGE band patterns when analysing the diversity of beta-ammonia oxidizers in the environment. J Microbiol Methods. 2006;66:147–155.
Bano N, Hollibaugh JT. Diversity and distribution of DNA sequences with affinity to ammonia-oxidizing bacteria of the ß-subdivision of the Class Proteobacteria in the Arctic Ocean. Appl Environ Microbiol. 2000;66:1960–1969.
Rotthauwe JH, Witzel KP, Liesack W. The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Environ Microbiol. 1997;63:4704–4712.
Sanger F, Nicklen S, Coulson AR. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci. 1977;74:5463–5467.
Suzuki MT, Taylor LT, DeLong EF. Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5’-nuclease assays. Appl Environ Microbiol. 2000;66:4605–4614.
Zhang Z, Schwartz S, Wagner L, Miller W. A greedy algorithm for aligning DNA sequences. J Comput Biol. 2000;7:203–214.
Biers EJ, Sun S, Howard EC. Prokaryotic genomes and diversity in the surface ocean: interrogating the Global Ocean Sampling metagenome. Appl Environ Microbiol. 2009;75:2221–2229.
Acknowledgements
We thank the captain and crew of R/V Savannah and Drs. Daniela Di Iorio, Yeping Yuan, and Yuntao Wang for helping with sample collection, as well as Erica Malágon and Dr. Tommy Dornhoffer. We would especially like to thank Dr. Yeping Yuan for suggestions and assistance with creation of figures, and Dr. John Schalles from Creighton University, Omaha N.E. for providing the satellite image of the SAB shown in Figure S6. We thank anonymous reviewers for constructive comments that improved an earlier version of this manuscript. This research was funded by the US National Science Foundation (NSF OCE 13−35838 to J.T.H. and NSF OCE 12-37130 to B.N.P.). This is publication number 1059 from the University of Georgia Marine Institute and 10319 from SOEST.
Author contributions
Q.L., B.B.T., and J.T.H. collected samples and conducted rate measurements. Q.L., B.B.T., M.J.R., and C.M.S. performed qPCR and chlorophyll analyses. Q.L. and B.B.T. amplified and cloned amoA and 16S RNA genes, Q.L. and J.T.H. analyzed sequence data. C.M.S., J.B.C., and J.T.H. performed nutrient analyses. N.J.W. and B.N.P. performed 15N analyses. Q.L. and J.T.H. analyzed CTD data and compiled the ancillary data used in the RDA analysis. Q.L. conducted the RDA analysis. Q.L. and J.T.H. analyzed the data obtained and wrote the paper in collaboration with B.B.T. and B.N.P. All of the authors have had opportunities to review and comment on the draft MS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Liu, Q., Tolar, B.B., Ross, M.J. et al. Light and temperature control the seasonal distribution of thaumarchaeota in the South Atlantic bight. ISME J 12, 1473–1485 (2018). https://doi.org/10.1038/s41396-018-0066-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0066-4
This article is cited by
-
Exploring the interactions between algae and archaea
Marine Life Science & Technology (2024)
-
Phylotype resolved spatial variation and association patterns of planktonic Thaumarchaeota in eastern Chinese marginal seas
Marine Life Science & Technology (2023)
-
Diversity and salinity adaptations of ammonia oxidizing archaea in three estuaries of China
Applied Microbiology and Biotechnology (2023)
-
Pattern changes of microbial communities in urban river affected by anthropogenic activities and their environmental driving mechanisms
Environmental Sciences Europe (2022)
-
Transcriptional activity differentiates families of Marine Group II Euryarchaeota in the coastal ocean
ISME Communications (2021)