Abstract
Parasites exist in every ecosystem and can have large influence on food web structure and function, yet, we know little about parasites’ effect on food web dynamics. Here we investigate the role of microbial parasitism (viruses of bacteria, phytoplankton and cyanobacteria, and parasitic chytrids on cyanobacteria) on the dynamics of trophic pathways and food web functioning during a cyanobacteria bloom, using linear inverse food web modeling parameterized with a 2-month long data set (biomasses, infection parameters, etc.). We show the importance of grazing on heterotrophic bacteria (the microbial pathway: DOC → bacteria → consumer) and how consumers depended on bacteria during peak-cyanobacteria bloom, which abundance was partly driven by the viral activity. As bacteria become the main energy pathway to the consumers, the system takes a more web-like structure through increased omnivory, and may thereby facilitate the system’s persistence to the cyanobacteria outbreak. We also showed how the killing of cyanobacteria host cells by chytrids had important impact on the food web dynamics by facilitating grazing on the cyanobacteria, and by offering alternative pathways to the consumers. This seemed to increase the system’s ability to return to a mix of trophic pathways, which theoretically increases the stability of the system.
Similar content being viewed by others
Introduction
Parasites exist in all ecosystems, where they have the potential to influence food web structure and properties [1,2,3]. Since Marcogliese & Cone’s [4] plea for researchers to include parasites in food web and ecological studies, a multitude of studies have shown how parasites can alter biochemical cycles, influence species richness, change productivity, increase trophic chain length and number of links, and cause changes in the topology of the trophic network and functioning of the ecosystem (e.g., [1, 2, 5]). The way parasites tend to structure food webs has also been suggested as a possible link to more stable systems [6], but as the author suggests, this need further verification by including parasites in network analyses. Most of the lessons done on parasite’s effect on food webs are based on comparisons of food web networks with and without parasites, while investigating the actual changes in food webs under different degree of infection has rarely been done. Hence, one question remaining unclear is how parasitism may affect the dynamics of food webs, particularly in terms of energy flows [7]. Parasites are likely to affect food web dynamics and the way energy flows in the system in a variety of ways. They directly influence the variation in the host population over time, with consequent indirect effects on interacting species in the system [2]. They are also often themselves subject to predation of free living stages, or via contaminant predation [8, 9]. Parasites may also indirectly modulate the flow of energy, by modifying the behavior or morphology of their host, increasing their susceptibility to predation [10, 11]. Empirical data and models of parasites’ influence on energy flows and dynamics over time are virtually non-existing [7].
Cyanobacteria, one of the most widespread plankton in aquatic systems, are subject to a variety of parasites such as fungi, protists, bacteria, and viruses [12]. When blooming, cyanobacteria have the potential to decrease phytoplankton diversity by outcompeting other species, often resulting in monospecific blooms [13]. Disease and parasites play an important role in hampering their dominance allowing for a more diverse system, both in terms of species and energy flows. The classical view of cyanobacteria as a trophic dead end, however, is largely debated (e.g., [14]). Although grazing rates on cyanobacteria is often low and inefficient [15], many zooplankters possess the capacity to accommodate to bloom situations and their potential toxicity. Some larger grazers (e.g., copepods and cladocerans) can break down large filamentous cyanobacteria, circumventing the size constraint [16, 17]. Increased frequency, duration, and intensity of blooms are expected to select for better adapted zooplankton [18,19,20]. Despite their ubiquity, the fate of cyanobacteria production and to what degree their production is used in a system is still questioned [21].
Herbivory, is only one of the potential trophic pathways for the cyanobacterial production to reach higher trophic levels. First, the extensive excretion by cyanobacteria stimulates bacterial activity and the microbial loop, offering alternative flows of energy via bacteria to protozoans and zooplankton [22, 23]. Viral infections also have an important influence on bacteria, phytoplankton, and cyanobacteria [24]. Their lytic infections result in the lysis of the host cell, resulting in additional release of dissolved organic matter and detritus, and with consequent reduced biomass of infected host populations [25, 26]. Second, dead organic matter produced by cyanobacteria is a potential important food source for detritivores. Finally, the less studied trophic pathways, the parasitism with free living stages and/or modification of behavior, or morphology increasing the host vulnerability to predation [10, 11]. Parasitic fungal chytrids are common parasites of larger phytoplankton such as colonial and filamentous cyanobacteria [27, 28]. An infection results in the death of the infected cell as they release edible zoospores [29], and can also lead to a mechanical fractionation making the remaining filament parts more vulnerable to predation [30]. Thus, the chytrid parasitism can be an important trophic link from inedible algae, such as filamentous cyanobacteria, to the zooplankton community [31, 32].
In this study we investigated the trophic dynamics in a microbial pelagic community during the course of a cyanobacteria bloom, and how parasitism affects flow-dynamics and overall functioning of the system. By using a food web modeling approach, calibrated with an extensive data set following a cyanobacteria bloom during 2 months with detailed data on chytrid parasitism on cyanobacteria, zooplankton community, heterotrophic bacteria, and phytoplankton (including cyanobacteria) and viruses infecting them both, we directly estimated the carbon flows in the system. We focused on the herbivory, bacterivory, detritivory, and chytridivory (i.e., feeding on chytrid zoospores) trophic pathways (Fig. 1), by specifically addressing three questions: (i) what is the fate of the filamentous cyanobacteria production, (ii), how do the consumers in the system sustain their energetic demands during a bloom, and (iii) what role does parasitism (viruses and chytrids) have on partitioning of energy and overall functioning of the system? Our results suggest that the bacterivory pathway may have a significant role by offering an alternative trophic pathway to the system during stress, experienced throughout the peak bloom, and that the viral activity (infecting heterotrophic bacteria mainly) is likely to influence its dynamic. We also show that neither the zooplankton community nor the parasites of cyanobacteria were likely to control the cyanobacteria bloom. More importantly though, the parasites had an essential role by facilitating grazing on the cyanobacteria, and by offering alternative energy pathways to the consumers. This had seemingly important consequences on the system’s ability to return to a mix of trophic pathways, which theoretically should increase the stability of the system.
Simplified representation of the four trophic pathways considered in this study during a Dolichospermum macrosporum bloom in lake Aydat: (1) chytridivory (i.e., feeding on chytrid zoospores), (2) herbivory, (3) bacterivory (or microbial loop), and (4) detritivory. These pathways were viewed and analyzed from two perspectives, either as (a) a ratio of the total cyanobacteria production, addressing the questions “what is the fate of the filamentous cyanobacteria production?”, or (b) a ratio of the total ingestion by consumers, addressing the question “how do the consumers in the system sustain their energetic demands during a bloom?” (see Trophic pathways in Methods section). a The pathway cyanobacteria to DET to zooplankton was not considered, but only the proportion of cyanobacteria production becoming detritus. For a detailed flow-diagram of the complete system see Supplementary Fig. A1
Methods
Food web modeling approach
We studied the pelagic community during a bloom of the filamentous cyanobacteria Dolichospermum macrosporum (Anabaena macrospora), infected by parasitic Chytridiomycota (chytrids), during 2 months in Lake Aydat, a small eutrophic lake located in the French Massif Central. For a food web perspective, we used linear inverse food web modeling to simulate the carbon flows (mgC m−2 d−1) between trophic compartments in the system (Supplementary Appendix A). Carbon was used as a proxy for energy flows. The advantage of using carbon as a unit is that biomasses and biological rates are often measured in this unit, and since carbon account of ~50% of dry mass in most species [33]. A linear inverse model is a steady-state model, constructed by applying linear equations (mass balance equations), and linear inequality equations (constraint, in form of minimum or/and maximum range) to define and constrain the possible value for each flow in the system [34, 35]. Totally 17 compartments were defined for lake Aydat’s pelagic system: 12 living (picophytoplankton, nanophytoplankton, and microphytoplankton, the latter consisting >91% of the cyanobacteria D. macrosporum during bloom condition, sporangia, and zoospore stages of the chytrids, heterotrophic bacteria, free living virus, heterotrophic nanoflagellates, ciliates, rotifers, cladocerans, copepods), two non-living (dissolved organic matter (DOC), detritus), and three external (gross primary production (GPP), respiration, loss) compartments, interconnected with 83 flows (Supplementary Figure A1). Detritus is here defined as the upper size threshold of dissolved organic matter (>0.7 µm), and loss as the carbon lost out of the system via sinking or predation by organisms at higher tropic levels not included in the model. While the mesozooplankton was represented by three compartments (rotifers, cladocera, copepods), species specific data was used when calculating the diet constraints for the respective compartments (Supplementary Appendix A). GPP by the phytoplankton compartments was the only external incoming flow. The microphytoplankton compartment will hereafter be referred to as the cyanobacteria compartment.
The development of the LIM was originally done in order to reconstruct complete sets of food web flows from incomplete observations. It is especially used in the study of microbial food webs as the direct estimation of processes is more problematic for smaller than larger animals. A model is said to be inverse when applied to estimate unknown parameters from the system’s state, apart from forward models that describe state dynamics using known parameters and relations [34]. The linear inverse model is based on a set of linear equations and inequalities that define a space of solutions that a Monte Carlo Markov Chain (MCMC) algorithm will allow to describe. The result is a set of solutions, each solution giving one value for each flow. Thanks to this MCMC-sampling procedure, at each time step, a mean value and confidence interval can be calculated for each flow, allowing an estimate of the flow uncertainty. Most constraints are based on relationships between flows, so estimating biomasses of each compartment is not necessary, but is valuable when parameterizing inequalities used to constrain the flows. Here, the LIM was necessary because of the high degree of functional diversity considered. To parameterize the model, data from an extensive field study in Lake Aydat were used, in combination with literature information (Supplementary Appendix B).
Sampling was done from the 9th of September to the 27th of October 2010, corresponding to the seasonal bloom of D. macrosporum, with a sampling resolution of 3 days. Abundances, biomasses, and several infection parameters in regard to chytrid infection on cyanobacteria, and viral infections on bacteria were measured for each sampling date. Viral infection of phytoplankton and cyanobacteria were made at nine sampling occasions. Grazing of cyanobacteria was assumed dependent on the filament size (Supplementary Appendix A). Also, the free living zoospores are valuable food to zooplankton [27, 31]. The transfer of phytoplankton biomass to higher trophic levels via the zoospores has been referred to as the “mycoloop” [27]. Field measurements of several of these processes were included in the model (Supplementary Appendix AB). Meanwhile, viruses are commonly infecting both phytoplankton and bacteria, where a lytic infection (in comparison to the lysogenic infection) leads to the lysis and death of the infected cell with consequent DOC and detritus releases [24]. In regard to viral infections, in this study we considered lytic infections of bacteria and all phytoplankton compartments based on field measurements of infection rates, and grazing on viruses by heterotrophic nanoflagellates using abundances of free living viruses [36]. Further, net primary production, which was used to deduce the GPP in the model, was estimated using a vertically generalized productivity model ([37]; Supplementary Appendix C). Totally 17 equations and 105 inequality equations were defined in the model, of which 34 were upper limit diet constraints (Supplementary Table A2). In total, 73 out of the 83 flows were directly constrained with an inequality equation, while the remaining 10 flows were derived from the model.
To reflect the temporal dynamics in the (steady-state) food web model, the model was adjusted for each sampling date using date-specific abundance, biomass, and infection parameters (resulting in 17 time steps). Since the 17 time steps had the same underling equations (Supplementary Table A2), the measured biomasses and flow rates included in the model equations made the model return a unique solution for each time step. The LIM package in R (version 3.1.1, www.r-project.org) was used to construct and analyze the model (Supplementary Appendix A). We used the MCMC-sampling procedure to obtain probability distributions based on 200,000 iterations for each flow [38]. Model performances were visually assessed using biplots of individual flows and convergence plots.
Trophic pathways
Ratio between flow values have previously been used to distinguish and identify food web typology and dominant trophic pathways [39, 40]. We re-defined some of the ratios proposed by Sakka et al. [40], in order to address our two first research questions. The ratios were calculated from the 83 flows for all 17 time steps, where all 200,000 LIM-solutions were used. First, to analyze the fate of the cyanobacterial production we calculated feeding ratios in relation to the cyanobacteria production (Fig. 1a):
“Cyanobacterivory” as the consumption rate of cyanobacteria by any consumer divided by the available cyanobacteria net-production.
“Bacterivory” as the consumption rate of bacteria (sustained on DOC exudated only by cyanobacteria), by any consumer divided by the available cyanobacteria net-production.
“Chytridivory” as the consumption rate of chytrids zoospores by any consumer, divided by the available cyanobacteria net-production.
“Detritus production” as the production rate of detritus by cyanobacteria divided by the available cyanobacteria net-production. This is not a feeding ratio directly, but simply the proportion of cyanobacteria production turning into detritus. Of this proportion, at most 25% was utilized by detritivores. Due to the model construction, the exact proportion is not possible to derive.
To address how consumers sustain their energetic demands during a bloom, we calculated feeding ratios in relation to the total feeding rate of the consumers in the system (Fig. 2b):
“Herbivory pathway” as the consumption rate of phytoplankton by any consumer divided by total consumption rate by consumer.
“Microbial pathway” as the consumption rate of bacteria and viruses by any consumer divided by total consumption rate by consumer. This pathway represents the transfer of DOC to higher trophic levels through the “microbial loop”.
“Chytrid pathway” as the consumption rate of zoosporic chytrids by any consumer divided by total consumption rate by consumer. This pathway represents the transfer of phytoplankton biomass to higher trophic levels through the “mycoloop”.
“Detritus pathway” as the consumption rate of detritus by any consumer divided by total consumption rate by consumer.
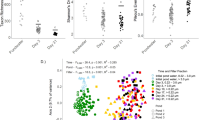
Abundance and biomasses of the major compartments considered in the food web model. a The abundance of cyanobacteria Dolichospermum macrosporum (PH3) during the 2 months sampled, and the prevalence of infected cyanobacteria cells by parasitic chytrids. The gray-shaded areas shows the bloom period, defined as cyanobacteria abundances >10 × 106 cells per liter and constitute >80% of phytoplankton biomass [18]. The dark gray area indicates the peak-bloom period. b The biomass of cyanobacteria D. macrosporum (PH3) and smaller phytoplankton (PH1 and PH2), c biomass of bacteria and viruses, and frequency of infected bacteria cells (FIC), eukaryotes (assumed to consist of PH1 and PH2 mainly, FIE) and D. macrosporum (FID), and d biomass of cladocera, copepod, and rotifers. For detailed biomasses see Supplementary Table A1
The detailed equations can be found in Table S1. The resulting ratios (expressed as a fraction) were plotted as the 90% credibility interval (the centered interval comprising 90% of the 200,000 iterations), against time. The minimum and maximum range of mean ratio values for the 17 time steps are reported in the text, if nothing else is stated.
In order to detect shifts in the mean ratios representing the trophic pathways during the 2-month period, we used a sequential t test approach designed to detect statistically significant shifts in mean level and magnitude of fluctuations in time series (STARS, [41]). The program identifies outliers for the periods identified, and then reports a “corrected mean value”. The cut-off length parameter was set l = 5 in order to allow identification of potential shifts during the different bloom period (Fig. 2a). A decrease in the cut-off length makes it harder for the program to detect a shift, and therefore was the significance level-parameter increased to 0.50 to facilitate detection. Only shifts with a final p = 0.05 significant level were reported.
Ecological network indices
Ecological network analysis (ENA, [42]) was used to compute several indices to characterize the structure and functioning of the system.
Total system throughput (T..) is calculated as the sum of all flows in the system, an indicator of system activity [43].
System omnivory index (SOI) is calculated as the weighted mean of the omnivory indices of each consumer compartment, using the logarithm of their consumption as weights, and where omnivory is the variance of the trophic level of its preys. Measures the omnivory in a system, and the distribution of feeding interactions among trophic levels [44].
Finn cycling index (FCI) is the ratio between flows generated by cycling over the Total system throughput [45].
The NetIndices package in R was used for the calculations, based on all 200,000 solutions of the 83 flows for the 17 time steps. The indices were presented as anomalies, which were calculated as the difference at each time step from the mean, plotted against time.
Results
Four periods defining the bloom development are in the following text referred to as the “pre-bloom”, “bloom”, “peak-bloom”, and “post-bloom” period (Fig. 2a). The 2 months sampling period was characterized by high cyanobacteria biomasses (Fig. 2b). The bloom increased gradually until the peak period (max. c.a. 8700 mgC m−2), and then decreased drastically. At abundances around 11 × 106 cells per liter (day 21), the first chytrid-infected cyanobacteria cells were detected. The infection increased rapidly and peaked 9 days after the cyanobacteria. Meanwhile, bacterial and viral biomasses varied in parallel (Fig. 2c). Biomasses were lowest during the “pre-peak period”, and then increased gradually until the end of the sampling period except for a sudden decrease in association with the collapse of the bloom. Cladocera dominated the mesozoopankton biomass (Fig. 2d), with two peaks during the sampled period. Their first increase was observed during the cyanobacteria “bloom period”, prior to the “peak-bloom”. The second increase occurred after the peak bloom. Both rotifer and copepod biomasses were comparably low and with slight increases during the “post-bloom period” (Fig. 2d). Worth noting is that the species composition for the three zooplankton compartments, used to calculate the feeding flows, varied during the sampled period (Supplementary Figure A2).
System productivity
In order to study the trophic dynamics during the course of the cyanobacterial bloom, we considered four major trophic pathways: herbivory, bacterivory, detrivory, and chytridivory (Fig. 1). The total production of these sources (i.e., phytoplankton, bacteria, detritus, and chytridic zoospores, respectively) ranged between 994 and 3743 mgC m−2 d−1. Their relative importance varied greatly (Fig. 3). Phytoplankton stood for the major production in the system (56–88% of total production), of which 75–99% were produced by the cyanobacteria compartment (i.e., 46–82% of the total production). Bacteria were the second most important source (5–26%), followed by detritus (4–10%). The relative importance of zoospore production was very low, maximum 2%.
The production in the system. a Total “available” production, which refers to the part of the production available to the secondary producers after respiration and dissolved organic matter release. Technically this includes the biomass accumulation. b The relative importance in production of the major primary sources considered: primary production by phytoplankton, secondary production by bacteria and viruses, and chytrids, and the overall contribution to the detritus (DET) pool (here defined as the detritus produces minus the part lost due to sinking). The dotted green line shows the relative importance of Cyanobacteria production. The gray-shaded areas indicates the bloom and peak-bloom periods, see Fig. 2a for details
The fate of cyanobacteria production
The major part of the cyanobacteria produced in the system became detritus (59–90%; Fig. 4b). Of this, >75% were lost through sinking, and the rest were assumed to stay in the system available to, e.g., detritivores or dissolution to DOC. Consequently, at most 33–56% (e.g., 1−(0.59×0.75)) of the cyanobacteria production was available to biological processes (other processes considered in the model except for loss out of the system).
The results from the food web models in form of trophic pathways and some associated raw data. a The total cyanobacteria production in the system, which was used as a denominator to calculate the feeding ratios addressing the question “what is the fate of the cyanobacteria production”. b The proportion of cyanobacteria production becoming detritus (detritus production, the model was constructed so that >75% of this detritus is lost out of the system through sinking, hence, only a smaller fraction of this is available to consumers), and c the relative importance of alternative trophic pathways reaching the consumers: “Bacterivory”, “Cyanobacterivory”, and “Chytridivory”. d The average size of the cyanobacteria filament, the average biomass of zooplankton feeding cyanobacteria, and the frequency of infected cyanobacteria filaments (%). These are all raw data of which some were used to parameterize the model. e The total ingestion by the consumers in the system, which was used as a denominator to calculate the trophic pathways addressing the question “how do the consumers in the system sustain their energetic demands during a bloom”. f The relative importance of the alternative trophic pathways reaching the consumers: “Microbial”, “Herbivory”, “Chytridivory” and “Detritus” pathways. g The relative importance of grazing on cyanobacteria or smaller phytoplankton in relation to the total phytoplankton grazing (i.e., herbivory). h Bacterial production as estimated from the model, and the bacterial mortality due to viral lysis (%) which is part of the raw data used to parameterize the model. The microbial pathway is shown in shaded blue, which units can be seen in (f). The dotted lines in the ratio plots indicates the weighted mean value and the significant shifts identified using the STARS software (see Methods section for details). The gray-shaded areas indicates the bloom and peak-bloom periods, see Fig. 2a for details
The dominant trophic pathways from cyanobacteria showed significant changes during the period sampled (STARS results Fig. 4c; Table S2). Prior to the bloom, grazing on cyanobacteria was very low (1%, corrected mean (CM) from STARS analyses), while bacterivory was the major trophic pathway (CM: 8%). As the bloom started to build up the grazing on cyanobacteria increased until the collapse of the bloom when grazing on both cyanobacteria and bacteria decreased significantly (CM: 2%, 4%). During the following post-bloom period the cyanobacterivory increased again, and the three trophic pathways (i.e. cyanobacterivory, bacterivory, and chytridivory) were similarly important (CM: 6%, 4%, 3%). The most noteworthy difference was that between the pre-bloom and post-bloom period for cyanobacterivory. While cyanobacterivory was at its lowest levels during the pre-bloom period (max. 1.6%), it peaked during the post-bloom period (max. 7.3%), despite the fact that both periods showed low biomasses of cyanobacteria (Fig. 2b).
The increase in cyanobacterivory during the post-bloom period showed strong association with the increase of zooplankton biomass feeding on cyanobacteria (Spearman’s ρ = 0.86, p«0.001), which in turn showed association with the filament size of the cyanobacteria (Spearman’s ρ = −0.49, p = 0.048). The filament size, in turn, decreased as the frequency of chytrid-infected cyanobacteria cells increased (Fig. 4d).
How consumers sustain their energetic demands during bloom
The total ingestion by all consumers in the system showed little variation during the sampled period (Fig. 4e). The relative importance of the different trophic pathways, however, changed significantly (Fig. 4f; Table S2). During the pre-bloom period, a mix of trophic pathways supported the consumers (CM: herbivory 34%, bacterivory 39%, and detritivory pathway 28%). During the peak-bloom period the system shifted dramatically by becoming dependent on the bacterial pathway (CM: 61%). Once the biomass-peak of cyanobacteria decreased, the system shifted back to a mix of trophic pathways (CM: herbivory 31%, bacterivory 36%, and detritivory 28%) where grazing on chytrids zoospores (CM: 7%) contributed partly.
The bacterivory pathway correlated strongly with the bacterial production calculated from field measurements (Spearman’s ρ = 0.81, p«0.001). This is not surprising since major part of the bacterial production is assumed to be grazed by consumers. What is more interesting, however, is the negative correlation between bacterial production and the mortality of bacteria due to viral lysis (Spearman’s ρ = −0.57, p = 0.019; Fig. 4h).
When looking at the herbivory pathway only, it becomes evident that the herbivory consumers are strongly relying on smaller phytoplankton (CM: 81%) before the cyanobacterial bloom reached its maximum (Fig. 4g). After this shift, the herbivory consumers grazed equally on smaller phytoplankton (CM: 51%) and cyanobacteria (CM: 49%). The shift in herbivory pathway showed association to the filament size of the cyanobacteria (Fig. 4d).
Food web functioning
The indices tended to divide into two groups depending on their response pattern. The first group showed a significant increase in their index values during the peak bloom, including the T.. and the SOI (Fig. 5). The second group showed a significant increase during the post-bloom period mainly. This was the case for the FCI and the DivF (Fig. 5). The FCI was also significantly higher during peak bloom, although the post-peak period was significantly higher (Fig. 5).
Anomaly in the Ecological Network Indices (ENA) for Total System Throughput (T..), System Omnivory Index (SOI), Finn Cycling Index (FCI), and the Diversity of Flows (DivF) during the 2-month period modeled. Anomalies are calculated as the value at a particular date minus the average value for the whole period. The asterisks indicate periods significantly different from all other periods (i.e., bloom, pre-bloom, post-bloom, and peak-bloom periods) (Supplementary Appendix D)
Discussion
Cyanobacteria blooms are a worldwide problem, increasingly present in both marine and freshwater systems [47, 48]. These blooms are highly influenced by bottom up factors such as light [49], temperature [47], or water stability [50]. In return the important cyanobacterial biomass reported during blooms impact the entire trophic food web. Here, we choose to focus on the biotic part of such events. We used food web modeling to analyze and understand the trophic dynamics in a microbial pelagic system during the course of a cyanobacteria bloom, with a particular interest in parasite influences. Parasites are part of any natural system [2, 3], still, we lack understanding on how parasites affects food web dynamics [7]. To our knowledge, this is the first food web modeling study to follow the entire dynamics of a cyanobacteria bloom, and analyze the potential impact of parasites on food web dynamics during such disturbance.
The fate of cyanobacteria production
The cyanobacteria bloom in Lake Aydat in 2010 was not controlled by the zooplankton, as indicated by the low grazing comprising in average 3% of the produced biomass (Fig. 4c). Despite the sparse grazing, cyanobacteria contributed up to 13% of the consumer’s energetic demand (during post-bloom period), and >41% of the cyanobacteria produced were either directly or indirectly utilized by the system. In fact, most of the cyanobacteria produced reached the higher trophic levels via bacteria or grazing on detritus, and to a smaller extent via consumption of parasitic zoospores. This supports the growing pieces of evidence that cyanobacteria are not strictly trophic dead ends [51]. Cyanobacteria are typically considered an unsuitable food source for herbivores and omnivores. Many species, D. macrosporum included, are colony or filament building, giving them a natural size protection from grazing [15]. Although grazed, cyanobacteria are inefficient transferred to zooplankton due to their weak nutritional value [52] and their potential toxicity [18], Dolichospermum genus included [53]. Although this genus is known to be capable to produce both microcystins and anatoxin-a [54], recent investigation in lake Aydat showed no correlation between D. macrosporum and both toxins [55]. Unless nontoxic, D. macrosporum is nonetheless hardly edible and presents a poor food quality for zooplankton, the energetic path via the microbial loop can therefore play an important role by increasing the nutritional value of the cyanobacteria-derived biomass [56]. This “trophic upgrading” is also proposed for chytrids [28], since their zoospores contain high energy and fatty acid content [27], suggesting their value as food supplement. Altogether, although direct grazing on cyanobacteria is weak, the cyanobacteria biomass takes alternative energy paths to support the higher trophic levels in the system, of which some (bacterivory and chytridivory) may have important food quality improving effects.
Even if the average grazing rates on cyanobacteria was low, the dynamics in how cyanobacteria were utilized by the system changed during the course of the bloom. By the end of the bloom, grazing of cyanobacteria was five times higher than during the pre-bloom period (Fig. 4c), despite similar abundances (Fig. 2a). Grazing by zooplankton increased in response to lower filament size of the cyanobacteria (Fig. 4c,d), which was part of our model assumptions (i.e., filament size thresholds for zooplankton species). Importantly though, increased zooplankton biomasses (Fig. 4d) also contributed to the increased estimated grazing rates. This suggests a functional response in the zooplankton community to one or several changes observed during the post-bloom period, suggestively to increased availability of food in form of smaller, more available cyanobacteria, and zoosporic chytrids. Several factors can affect the filaments size [57], and splitting up of filaments can be caused by decreased temperature, nutrient depletion, cutting up of filaments by some zooplankton grazers, or due to the lyse of one cell within the filament [17, 58,59,60]. Sigee et al, [61] reported the role of fungal parasitism in the fractionation of filaments. Moreover, Gerphagnon et al, [30] showed that Rhizosiphon crassum, the unique chytrid species infecting D. macrosporum vegetative cells in our system, was able to such mechanistic fractionation. Our results support such observations and indicate that parasitic chytrids can influence the food web dynamics by making the cyanobacteria more available to the consumers in the system.
How consumers sustain their energetic demands during bloom
Throughout most parts of the sampled period the consumers in the system relied on a mix of trophic pathways (Fig. 4f). During the peak bloom, however, the consumers shifted to become largely dependent on the microbial pathway. Since the discovery of the microbial loop, its significance for the functioning of fresh and marine pelagic systems has been repeatedly demonstrated [62, 63]. Important players driving the microbial loop are the viruses [24, 64]. Bacteriophages are capable to control their host populations [65, 66], and their abundances are often strongly correlated [24, 64], as was the case in this study (Fig. 2c). We also observed a significant negative relationship between bacterial production and mortality of bacteria due to viral lysis (Fig. 4h). Hence, viral activity which influences bacterial production may have been an important factor for the overall trophic dynamics, particularly during the peak bloom. Further, the simultaneous increase in T… and SOI during peak bloom (Fig. 5) indicates that as the system activity increased due to the immense cyanobacteria production, the system became more omnivorous due to the increased bacterivory, changing from a more chain-like to web-like structure. Since omnivores are capable to switch between preys they are less sensitive to fluctuations of one of the prey populations, and web-like networks are therefore often associated to ecosystems able to absorb and recover from perturbations [44]. Similarly, Fagan [67] showed that increased degree of omnivory has stabilizing effects on community dynamics, and more omnivorous systems recover quicker from perturbation. Indeed, the sudden shift to the bacterivory pathway leading to a more omnivorous food web was a response to the change in phytoplankton community, which during peak bloom was almost exclusively composed of cyanobacteria (>96%). Here we show how the system shifted to a more web-like structure during increased perturbation, potentially driven by the viral activity. Evidently, bacteria and viruses play a vital role by offering alternative energy pathways which may help the system to persist during high perturbation, as during a cyanobacteria bloom.
Parasites effect on food web functioning
In this study, the carbon flows directly associated to virus and chytrid parasites were very low (Fig. 4c,f). This is in contrast to previous modeling studies showing high grazing rates on chytrid zoospores supporting up to 38% and 50% of the microzooplankton diet [68, 69]. This difference can partly be explained by the low infection rates of cyanobacteria during our study (<8% infected cells) compared to previous reported rates [28]. However, the parasitic chytrids had important effects on the food web dynamic and functioning. Chytrids made the system less dependent on a few trophic pathways. Both by modifying the carbon flow between cyanobacteria and their grazers, and via the additional chytrid pathway (Fig. 4c). Simultaneously, the DivF and the recycling in the system (FCI) increased during the post-bloom period (Fig. 5). Stability, in terms of a system’s ability to resist changes during perturbation [43], has been related to the possible choices of energy pathways in a system [70, 71]. Hence, DivF can be used as an indicator of stability [43, 72]. Also cycling in a system can act as a buffer during perturbation and increase the ability of the system to resist changes, increasing its resistance [73]. This results is an interesting paradox, while the chytrids kill their host, they bring stability to the system. This draws parallels to the stabilizing effect of the “Killing the winner” mechanism, which refers to a density-related control of a dominant bacteria population by their viruses or their parasites, thereby liberating niches for unaffected populations and give chance to more species to develop and coexist, and thereby increasing the diversity and stability of the microbial system [74, 75]. Our results suggest a similar mechanism beget by the chytrids, by effecting the dominant cyanobacteria population the system became more diverse in trophic pathways and stability increased.
In order to test our observations related to viral and chytrid parasitism, we applied structural equation modeling (SEM) to our simulated results (Fig. 6; Supplementary Appendix E). We assumed that viral and chytrid activity can directly affect food web structure (DivF or SOI), but also indirectly by modifying the dominant trophic pathways (i.e., detritivory, bacterivory, herbivory, cyanobacterivory). These are also the causal pathways that we will focus our following discussion around. First of all, the suggested strong impact of viruses on the microbial pathway (standardized coefficient: −0.48/−0.46, respectively for SOI and DivF) was likely due to the decreased viral infection rates during peak bloom, with consequent increased bacterial production and consumption (Fig. 4f,h). The SEM suggested that the viruses affected the bacterial grazing rates by protozoans, and as a consequence, viruses effect on protozoan grazing rates on smaller phytoplankton is likely to be reverse (0.31/0.30). These processes seemed to have important indirect effects on the level of omnivory in the system (−0.46 × 0.70 = −0.32, and 0.30 × −0.40 = −0.12, Fig. 6a), and as we argued above, may be essential for the system to sustain during peak-bloom perturbations, when the stability of the system had decreased (Fig. 5). Further, the SEM results confirmed the positive effect by chytrids on the cyanobacteria grazing rate (0.27). The increased cyanobacterivory and the additional chytrid pathway that made the system less dependent on a few trophic pathways (therefore the negative effect on the other trophic pathways; Fig. 6b), turned out as important direct (0.26) and indirect effects (particularly via the herbivory pathway, −0.23 × 0.59 = −0.14) on the DivF in the SEM model, and hence, the stability of the system. Expectedly though, parasites cannot explain all the dynamics observed in the system as indicated by the remaining large error terms (Fig. 6; Supplementary Table E1). The SEM model has also a weakness by not accounting for potential time lags between prey and consumer, or host and parasite populations’ growth responses, which suggests that the causality of the model should be interpreted with care. We suggest that these types of causal relationships between parasite activity and their effect on the food web, could with advantage be tested in experimental setups or bioassays in future studies. However, it is still intriguing that two exogenous variables representing parasite activity can explain up to 69% of the variation for some food web variables in a complex system. Through the process of killing their hosts, viruses and chytrids have the potential to alter the food web dynamics and structure, helping the system to withstand perturbations during cyanobacteria outbreaks, and bring stability to the system.
Summary diagram of the effects of the parasite activity (virus and chytrids) on the dynamics of food web dynamics and food web structure, from structural equation modeling (SEM). SEM allows for testing the causality of a path model. We focused only on the parasites as endogenous, explanatory variables to the dynamics and structure of the system, even if we acknowledge that other factors are important. We assumed that viral and chytrid activity can directly affect food web structure, either in form of (a) diversity of flows (DivF), or (b) System Omnviory Index (SOI), but also indirectly by modifying the food web dynamics, in form of the dominant trophic pathways (i.e., detritivory, bacterivory, herbivory, cyanobacterivory). The thickness of the arrows is scaled to standardized coefficients from path analyses and illustrates the relative strength. Red arrows indicate positive effects, and blue negative effects. The effects of parasite activity on food web structure are split between direct effects and indirect effects via changes in trophic pathway dynamics. The indirect effects are calculated by the product of the coefficients along the path. The path analyses were ran using the SEM package in R, after assuring all assumptions were met (Supplementary Appendix E). The standardized coefficients for the error term (indicated by the small error into each endogenous variable box) are a, herbivory 0.82, microbial 0.79, detritus 0.97, cyanobacteria 0.78, and SOI 0.31, and b herbivory 0.82, microbial 0.77, detritus 0.97, cyanobacteria 0.77, and DivF 0.47
To conclude, we show how the microbial pathway is of major importance during peak-cyanobacteria bloom, partly driven by the viral activity. By shifting to the bacterivory pathway, the system took a more web-like structure through increased omnivory, which may have increased the resistance of the system during the cyanobacteria outbreak. We also show that neither the zooplankton community nor the parasites of cyanobacteria were likely to control the cyanobacteria bloom. Importantly though, by killing their host, the parasites had an essential role by facilitating grazing on the cyanobacteria, and by offering alternative energy pathways to the consumers. This had important consequences on the system’s ability to return to a mix of trophic pathways, which theoretically increase the stability of the system. Our results are in line with previous works showing how parasites can add stability to food webs by modifying the food web structure (reviewed in [6]), and that parasites are a key to a healthy ecosystem [76]. Future contributions would benefit from including qualitative information about the food web, such as food sources quality, and how the influence of infection on the genetic structure of the host population [77] can affect food web dynamics and functioning.
References
Dunne JA, Lafferty KD, Dobson AP, Hechinger RF, Kuris AM, Martinez ND, et al. Parasites affect food web structure primarily through increased diversity and complexity. PLoS Biol. 2013;11:e1001579.
Hatcher MJ, Dick JTA, Dunn AM. Diverse effects of parasites in ecosystems: linking interdependent processes. Front Ecol Environ. 2012;10:186–94.
Lafferty KD, Dobson AP, Kuris AM. Parasites dominate food web links. Proc Natl Acad Sci. 2006;103:11211–6.
Marcogliese DJ, Cone DK. Food webs: a plea for parasites. Trends Ecol Evol. 1997;12:320–5.
Kuris AM, Hechinger RF, Shaw JC, Whitney KL, Aguirre-Macedo L, Boch CA, et al. Ecosystem energetic implications of parasite and free-living biomass in three estuaries. Nature. 2008;454:515–8.
Poulin R. Network analysis shining light on parasite ecology and diversity. Trends Parasitol. 2010;26:492–8.
Lafferty KD, Allesina S, Arim M, Briggs CJ, De Leo G, Dobson AP, et al. Parasites in food webs: the ultimate missing links. Ecol Lett. 2008;11:533–46.
Johnson PTJ, Dobson A, Lafferty KD, Marcogliese DJ, Memmott J, Orlofske SA, et al. When parasites become prey: ecological and epidemiological significance of eating parasites. Trends Ecol Evol. 2010;25:362–71.
Thieltges DW, Amundsen P-A, Hechinger RF, Johnson PTJ, Lafferty KD, Mouritsen KN, et al. Parasites as prey in aquatic food webs: implications for predator infection and parasite transmission. Oikos. 2013;122:1473–82.
Lafferty KD, Morris AK. Altered behavior of parasitized killifish increases susceptibility to predation by bird final hosts. Ecology. 1996;77:1390–7.
Thomas F, Poulin R. Manipulation of a mollusk by a trophically transmitted parasite: convergent evolution or phylogenetic inheritance? Parasitology. 1998;116:431–6.
Gerphagnon M, MacArthur DJ, Latour D, Gachon CMM, Van Ogtrop F, Gleason FH, et al. Microbial players involved in the decline of filamentous and colonial cyanobacterial blooms with a focus on fungal parasitism. Environ Mircrobiol. 2015;17:2573–87.
Wynne TT, Stumpf RP. Spatial and temporal patterns in the seasonal distribution of toxic cyanobacteria in western Lake Erie from 2002-2014. Toxins. 2015;7:1649–63.
Wilson AE, Sarnelle O, Tillmanns AR. Effects of cyanobacterial toxicity and morphology on the population growth of freshwater zooplankton: meta-analyses of laboratory experiments. Limnol Oceanogr. 2006;51:1915–24.
De Bernardi R, Giussani G. Are blue-green algae a suitable food for zooplankton? An overview. Hydrobiologia. 1990;200:29–41.
Kâ S, Mendoza-Vera J, Bouvy M, Champalbert G, N’Gom-Kâ R, Pagano M. Can tropical freshwater zooplankton graze efficiently on cyanobacteria? Hydrobiologia. 2012;679:119–38.
Bouvy M, Pagano M, Troussellier M. Effects of a cyanobacterial bloom (Cylindrospermopsis raciborskii) on bacteria and zooplankton communities in Ingazeira reservoir (northeast Brazil). Aquat Microb Ecol. 2001;25:215–27.
Ger KA, Hansson LA, Lürling M. Understanding cyanobacteria-zooplankton interactions in a more eutrophic world. Freshw Biol. 2014;59:1783–98.
Hairston NG Jr, Holtmeier CL, Lampert W, Weider LJ, Post DM, Fischer JM, et al. Natural selection for grazer resistance to toxic cyanobacteria: evolution of phenotypic plasticity? Evolution. 2001;55:2203–14.
Sarnelle O, Wilson AE. Local adaptation of Daphnia pulicaria to toxic cyanobacteria. Limnol Oceanogr. 2005;50:1565–70.
Lesutienė J, Bukaveckas PA, Gasiūnaitė ZR, Pilkaitytė R, Razinkovas-Baziukas A. Tracing the isotopic signal of a cyanobacteria bloom through the food web of a Baltic Sea costal lagoon. Estuar Coast Shelf S. 2014;138:47–56.
Hessen DO, Andersen T, Lyche A. Carbon metabolism in a humic lake: pool sizes and cycling through zooplankton. Limnol Oceanogr. 1990;35:84–99.
Wylie JL, Currie DJ. The relative importance of bacteria and algae as food sources for crustacean zooplankton. Limnol Oceanogr. 1991;36:708–28.
Sime-Ngando T. Environmental bacteriophages: viruses of microbes in aquatic ecosystems. Front Microbiol. 2014;5:355.
Hewson I, O’Neil JM, Dennison WC. Virus like particles associated with Lyngbya majuscule (Cyanophyta; Oscillatoriacea) bloom decline in Moreton Bay, Australia. Aquat Microb Ecol. 2001;25:207–13.
Tucker S, Pollard P. Identification of cyanophage Ma-LBP and infection of the cyanobacterium Microcystis aeruginosa from an Australian subtropical lake by the virus. Appl Environ Microbiol. 2005;71:629–35.
Kagami M, de Bruin A, Ibelings B, Van Donk E. Parasitic chytrids: their effects on phytoplankton communities and food-web dynamics. Hydrobiologia. 2007;578:113–29.
Sime-Ngando T. Phytoplankton chytridiomycosis: fungal parasites of phytoplankton and their imprints on the food web dynamics. Front Microbiol. 2012;3:1–13.
Ibelings BW, de Bruin A, Kagami M, Rijkeboer M, Brehm M, Van Donk E. Host parasite interactions between freshwater phytoplankton and chytrid fungi (chytridiomycota). J Phycol. 2004;40:437–53.
Gerphagnon M, Latour D, Colombet J, Sime-Ngando T. Fungal parasitism: life cycle, dynamics and impact on cyanobacterial blooms. PLoS ONE. 2013;8:e60894.
Gleason FH, Kagami M, Marano AV, Sime-Ngando T. Fungal zoospores are valuable food resources in aquatic ecosystems. Inoculum Newsl Mycol Soc Am. 2009;60:1–3.
Kagami M, Miki T, Takimoto G. Mycoloop: chytrids in aquatic food webs. Front Microbiol. 2014;5:166.
Belgrano A. Aquatic food-webs’ ecology: old and new challenges. In: Belgrano A, Scharler UM, Dunne J, Ulanowicz RE, editors. Aquatic food webs: an ecosystem approach. New York: Oxford University Press; 2005. p. 9.
Niquil N, Saint-Béat B, Johnson GA, Soetaert K, van Oevelen D, Bacher C. et al. Inverse modeling in modern ecology and application to coastal ecosystems. In: Wolanski E, McLusky DS, editors. Treatise on Estuarine and Coastal Science. Waltham: Academic Press; 2012. Vol. 9.
van Oevelen D, Van den Meersche K, Meysman FJR, Soetaert K, Middelburg JJ, Vézina AF. Quantifying food web flows using linear inverse models. Ecosystems. 2010;13:32–45.
Bettarel Y, Sime-Ngando T, Bouvy M, Arfi R, Ambard C. Low consumption of virus-sized particles by heterotrophic nanoflagellates in two lakes of the French Massif Central. Aquat Microb Ecol. 2005;39:205–9.
Behrenfeld MJ, Falkowski PG. Photosynthetic rates derived from satellite-based chlorophyll concentration. Limnol Oceanogr. 1997;42:1–20.
van den Meersche K, Soetaert K, Van Oevelen D. xsample(): An R Function for Sampling Linear Inverse Problems. J Stat Softw. 2009;30:1–15.
Legendre L, Rassoulzadegan F. Plankton and nutrient dynamics in marine waters. Ophelia. 1995;41:153–72.
Sakka Hlaili A, Niquil N, Legendre L. Planktonic food webs revisited: reanalysis of results from the linear inverse approach. Prog Oceanogr. 2014;120:216–29.
Radionov SN. A sequential algorithm for testing climate regime shifts. Geophys Res Lett. 2004;31:L09204.
Ulanowicz RE. Growth and development: ecosystems phenomenology. New York: Springer; 1986.
Rutledge RW, Basorre BL, Mulholland RJ. Ecological stability: an information theory viewpoint. J Theor Biol. 1976;57:355.
Libralato S. System Omnivory Index. Ecol Indic. 2008;4:3472–7.
Finn JT. Flow analysis of models of the Hubbard Brook ecosystem. Ecology. 1980;61:562–71.
Ulanowicz RE, Norden JS. Symmetrical overhead in flow networks. Int J Syst Sci. 1990;21:429–37.
Pearl HW, Huisman J. Blooms like it hot. Science. 2008;320:57–58.
Taranu ZE, Gregory-Eaves I, Leavitt PR, Bunting L, Buchaca T, Catalan J, et al. Acceleration of cyanobacterial dominance in north temperate-subarctic lakes during the Anthropocene. Ecol Lett. 2015;18:375–84.
Havens KE, Phlips EJ, Cichra MF, Li BL. Light availability as a possible regulator of cyanobacteria species composition in a shallow subtropical lake. Freshw Biol. 1998;39:547–56.
Pearl HW. A comparison of cyanobacterial bloom dynamics in freshwater, estuarine and marine environments. Phycologia. 1996;35:25–35.
Perga M-E, Domaizon I, Guillard J, Hamelet V, Anneville O. Are cyanobacteria blooms trophic dead ends? Oecologia. 2013;172:551–62.
von Elert EV, Martin-Creuzburg D, Le Coz JR. Absence of sterols constrains carbon transfer between cyanobacteria and a freshwater herbivore (Daphnia galeata). Proc Biol Sci. 2003;270:1209–14.
Welker M, Von Döhren H. Cyanobacterial peptides—nature’s own combinatorial biosynthesis. FEMS Microbiol Rev. 2006;30:530–63.
Rapala J, Sivonen K. Assessment of environmental conditions that favor hepatotoxic and neurotoxic Anabaena spp. strains cultured under light limitation at different temperatures. Microb Ecol. 1998;36:181–92.
Sabart M, Crenn K, Perrière F, Abila A, Leremboure M, Colombet J, et al. Co-occurrence of microcystin and anatoxin-a in the freshwater lake Aydat (France): analytical and molecular approaches during a three-year survey. Harmful Algae. 2015;48:12–20.
Desvilettes C, Bec A. Formation and transfer of fatty acids in aquatic microbial food webs - role of heterotrophic protists. In: Arts MT, Brett M, Kainz M, editors. Lipids in aquatic ecosystems. New York: Springer; 2009. p. 25–42. .
Alam MGM, Jahan N, Thalib L, Wei B, Maekawa T. Effects of environmental factors on the seasonally change of phytoplankton populations in a closed freshwater pond. Environ Int. 2001;27:363–71.
Chan F, Pace ML, Howarth RW, Marino RM. Bloom formation in heterocystic nitrogen-fixing cyanobacteria: The dependence on colony size and zooplankton grazing. Limnol Oceanogr. 2004;49:2171–8.
Yamamoto Y, Nakahara H. Seasonal variations in the morphology of bloom-forming cyanobacteria in a eutrophic pond. Limnology. 2009;10:185–93.
Yamamoto Y, Suzuki K. Distribution and algal-lysing activity of fruiting myxobacteria in lake Suwa. J Phycol. 1990;26:457–62.
Sigee DC, Selwyn A, Gallois P, Dean AP. Patterns of cell death in freshwater colonial cyanobacteria during the late summer bloom. Phycologia. 2007;46:284–92.
Azam F, Fenchel T, Field JG, Gray JS, Meyer-Reil LA, Thingstad F. The ecological role of water-column microbes in the sea. Mar Ecol Prog Ser. 1983;10:257–63.
Pomeroy LR. The ocean’s food web, a changing paradigm. BioScience. 1974;24:499–504.
Fuhrman JA. Marine viruses and their biogeochemical and ecological effects. Nature. 1999;399:541–8.
Fischer UR, Velimirov B. High control of bacterial production by viruses in a eutrophic oxbow lake. Aquat Microb Ecol. 2002;27:1–12.
Suttle CA. Marine viruses—major players in the global ecosystem. Nat Rev Microbiol. 2007;5:801–12.
Fagan WF. Omnivory as a stabilizing feature of natural communities. Am Nat. 1997;150:554–67.
Grami B, Rasconi S, Niquil N, Jobard M, Saint-Beat B, Sime-Ngando T. Functional effects of parasites on food web properties during the spring diatom bloom in lake Pavin: a linear inverse modeling analysis. PLoS One. 2011;6:e23273.
Rasconi S, Grami B, Niquil N, Jobard M, Sime-Ngando T. Parasitic chytrids sustain zooplankton growth during inedible algal bloom. Front Microbiol. 2014;5:229.
MacArthur R. Fluctuations of animal populations, and a measure of community stability. Ecology. 1955;36:533–6.
Odum EP. Fundamentals of ecology. Philadelphia: Saunders; 1953. p. 56.
Tecchio S, Chaalali A, Raoux A, Tous Rius A, Lequesne J, Girardin V, et al. Evaluating ecosystem-level anthropogenic impacts in a stressed transitional environment: The case of the Seine estuary. Ecol Indic. 2015;61:833–45.
Saint-Béat B, Baird D, Asmus H, Asmus R, Bacher C, Pacella SR, et al. Trophic networks: How do theories link ecosystem structure and functioning to stability properties? A review. Ecol Indic. 2015;52:458–71.
Thingstad TF. Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol Oceanogr. 2000;45:1320–8.
Thingstad TF, Lignell R. Theoretical models for the control of bacterial growth rate, abundance, diversity and carbon demand. Aquat Microb Ecol. 1997;13:19–27.
Hudson PJ, Dobson AP, Lafferty KD. Is a healthy ecosystem one that is rich in parasites? Trends Ecol Evol. 2006;21:381–5.
Gsell AS, de Senerpont Domis LN, Verhoeven KF, van Donk E, Ibelings BW. Chytrid epidemics may increase genetic diversity of a diatom spring-bloom. ISME J. 2013;7:2057–9.
Acknowledgements
We thank Serena Rasconi (Wasser luster Lunz, Austria) and Boutheina Grami (University of Monastir, Tunisia) for important help and discussion during the initial phase of the construction of the food web model, Pascal Claquin (Université du Caen Basse Normandie, France) for valuable help on the calculation of primary production, Urania Christaki (Université du Littoral Côte d’Opale, France) for expert knowledge about protozoan feeding, and Justine Lequesne and Valérie Girardin for statistical advice in regard to effect size and statistical testing. At last, we want to thank three anonymous reviewers improving this manuscript by their valuable comments. PB was supported by Région Basse Normandie. MH, MG and this work were supported by a grant from the French ANR Programme Blanc ROME (Rare and Overlooked Microbial Eukaryotes in aquatic ecosystems) coordinated by TSN.
Author contributions
MH, MG, PB, and NN conceived the study. MG and JC collected field data and samples. PB, MG, and MH set up the database for data used in the model. MH constructed the model and ran statistical analyses, with input and feedback from MG, JC, ST, NN, and TSN. NN led the modeling work. TSN led the overall project. MH wrote the first draft of the manuscript and all authors contributed significantly to data interpretation and revisions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Haraldsson, M., Gerphagnon, M., Bazin, P. et al. Microbial parasites make cyanobacteria blooms less of a trophic dead end than commonly assumed. ISME J 12, 1008–1020 (2018). https://doi.org/10.1038/s41396-018-0045-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-018-0045-9
This article is cited by
-
Lytic bacteriophage have diverse indirect effects in a synthetic cross-feeding community
The ISME Journal (2020)
-
Trophic position, elemental ratios and nitrogen transfer in a planktonic host–parasite–consumer food chain including a fungal parasite
Oecologia (2020)
-
The fungicide azoxystrobin promotes freshwater cyanobacterial dominance through altering competition
Microbiome (2019)
-
Fungi in aquatic ecosystems
Nature Reviews Microbiology (2019)
-
Response of the eukaryotic plankton community to the cyanobacterial biomass cycle over 6 years in two subtropical reservoirs
The ISME Journal (2019)