Abstract
Forkhead box C1 (FOXC1) is required for neural crest and ocular development, and mutations in FOXC1 lead to inherited Axenfeld–Rieger syndrome. Here, we find that FOXC1 and paired box 6 (PAX6) are co-expressed in the human limbus and central corneal epithelium. Deficiency of FOXC1 and alternation in epithelial features occur in patients with corneal ulcers. FOXC1 governs the fate of the corneal epithelium by directly binding to lineage-specific open promoters or enhancers marked by H3K4me2. FOXC1 depletion not only activates the keratinization pathway and reprograms corneal epithelial cells into skin-like epithelial cells, but also disrupts the collagen metabolic process and interferon signaling pathways. Loss of interferon regulatory factor 1 and PAX6 induced by FOXC1 dysfunction is linked to the corneal ulcer. Collectively, our results reveal a FOXC1-mediated regulatory network responsible for corneal epithelial homeostasis and provide a potential therapeutic target for corneal ulcer.
Similar content being viewed by others
Introduction
Corneal integrity and transparency are essential for normal vision. The outermost surface of the cornea is covered with a non-keratinized stratified squamous epithelium consisting of multilayer corneal epithelial cells (CECs) on the avascular stroma. The self-renewal and regeneration of the corneal epithelium are sustained by a population of limbal stem cells (LSCs), which are segregated in the basal layer of the limbal zone of the peripheral cornea.1,2 LSCs continuously generate committed transient amplifying cells that migrate toward the central cornea and differentiate into mature CECs.3,4 However, various insults including chronic inflammation, infection, Stevens–Johnson syndrome, and genetic mutations lead to LSC dysfunction, causing corneal blindness characterized by opacification, neovascularization, and conjunctivalization of the cornea.5,6
Cell fate determination and identity are governed by lineage-specific transcription factors (TFs) and epigenetic modifications. Key TFs not only determine organ development, but also define tissue specificity during adulthood. The master regulator paired box 6 (PAX6) plays a vital role in the developing visual system, and is required for corneal epithelial identity and homeostasis.7,8,9 Loss of PAX6 induces the corneal epithelium to acquire an opaque skin-epithelium phenotype, which leads to visual disorder.8
Forkhead Box C1 (FOXC1) is a key TF of ocular development.10 FOXC1 mutations in human patients cause ocular anterior segment defects, and pathological corneal neovascularization can be observed in patients with Axenfeld–Rieger syndrome harboring FOXC1 mutations or copy-number alterations.10,11 Previous findings indicated that Foxc1 maintains corneal transparency and avascularity in the neural crest-derived corneal stromal and endothelial cells.11 Neural crest-specific Foxc1 null mutation in mice can result in disorganized corneal stroma with excessive growth of blood vessels and failure of corneal endothelial formation.11 However, the role of FOXC1 in the corneal epithelial homeostasis and pathogenesis has not been reported.
In the present study, we identify FOXC1 as a novel core regulator that underlies the lineage commitment of human corneal epithelium. FOXC1 regulates the corneal epithelial lineage-specific gene expression and interferon signaling pathways by occupying open cis-regulatory elements that are marked by histone H3 lysine 4 dimethylation (H3K4me2). FOXC1-depleted LSCs show the activation of an epidermis-specific gene program and the repression of IRF1 and PAX6, which are pathological characteristics of the human corneal ulcer. These findings indicate that the FOXC1 regulatory network is required for corneal epithelial homeostasis, and FOXC1 may serve as a potential target for corneal ulcer treatment.
Results
Co-expression of FOXC1 and PAX6 in the corneal epithelium
To determine whether FOXC1 functions in the corneal epithelium, we first detected its expression pattern during corneal development. We found that Foxc1 was evidently expressed in the ocular surface structure and neural crest-derived cells during the early developmental stage in mice (Fig. 1a). The corneal epithelium showed Foxc1 expression from the embryonic developmental stage to adulthood in mice (Fig. 1a). Foxc1 was detected in the corneal stroma at embryonic day 15.5 (E15.5). In contrast, Pax6 was expressed in the outmost layer of the mouse ocular surface at E12.5. Pax6 was also expressed in the corneal epithelium but not in the corneal stroma during development and adulthood (Fig. 1a).
a Immunofluorescence staining of Foxc1 and Pax6 during mouse corneal development (E12.5, E15.5, E18.5, postnatal day 18.5, and postnatal 5 weeks). Scale bars, 100 μm. Blue = DAPI, Green = Foxc1/Pax6. b Immunofluorescence staining of FOXC1 and PAX6 in the human cornea at E13 and E22 weeks. Scale bars, 100 μm. c Immunofluorescence staining of FOXC1 and PAX6 in the limbus and corneal epithelium of adult human. Scale bar, 100 μm
As expected, FOXC1 and PAX6 co-expressed robustly in the human corneal epithelium of 13-week and 22-week-old embryos (Fig. 1b). Their expressions were also maintained in the adult limbus and central corneal epithelium (Fig. 1c). These observations implied that FOXC1 may play an important role in corneal epithelial development.
Loss of FOXC1 and PAX6 is associated with human corneal ulcer
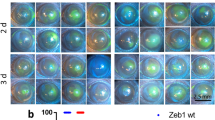
To study the potential relationship between FOXC1 and corneal pathogenesis, we examined 21 biopsy samples from patients with corneal ulcer. Remarkably, partial regions of the diseased corneal tissues from nine patients showed conspicuous absence of FOXC1, PAX6, and the corneal epithelial markers (KRT3 and KRT12). Instead, the keratinized epithelium-specific KRT1 or KRT10 appeared in these lesion areas (Fig. 2). These observations demonstrated the disruption of epithelial identity and homeostasis in corneal ulcer, and suggested that FOXC1 deficiency was probably associated with disorder of the corneal epithelium.
Gene expression program of CEC differentiation identifies FOXC1 as a key regulator
We then amplified human LSCs ex vivo and identified them by LSC markers KRT19, TP63, and PAX6 (Fig. 3a).12,13 Our primary LSCs showed a high proliferative potential, as evidenced by robust expression of cell proliferative marker KI67 (Fig. 3a). An air-lifting culture system was previously developed as a superior three-dimensional (3D) differentiation model of stratified epithelia.14,15 In this 3D model, LSCs were seeded into inserts of transwell plates and expanded in the culture medium. After the LSCs reached confluency, the medium in the upper chambers was removed to expose the cells to the air, and a small volume of medium in the lower chamber contacted with the cells to provide nutrition, which mirrors the condition under which CECs differentiate in vivo. Following induction in an air-lifting culture system, we obtained multilayer differentiated corneal epithelial sheets (dCESs) (Fig. 3b). Expression of PAX6 and CEC marker KRT124,16 (but not KRT19) in the dCESs indicated their terminal differentiation (Fig. 3b). In parallel with the in vivo-expression pattern, FOXC1 was also robustly expressed both in LSCs and in dCESs (Fig. 3a, b).
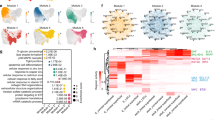
a Immunofluorescence staining of the indicated genes in primary LSCs. Scale bars, 50 μm. b Hematoxylin and eosin staining (upper left) in the dCESs induced through air-lifting culture. Immunofluorescence staining of the indicated genes in the dCESs induced through air-lifting culture. Scale bar, 100 μm. c Volcano plot showing gene expression levels in LSCs versus those in the dCESs. d Heatmap showing normalized transcripts-per-million reads (TPM) values of the downregulated TFs in the dCESs. e GO biological process (BP) analysis of genes expressed at higher levels in the dCESs than in LSCs. f GSEA showing the canonical pathways enriched in the preferentially expressed gene set in the dCESs. g Heatmap showing normalized TPM values of the top 25 (fold change) upregulated TFs with high expression levels in the dCESs. h GSSM of the top 25 (fold change) upregulated TFs with high expression levels in the dCESs. The cut-off value (0.75) is indicated by the dashed line
To explore the gene regulation program of corneal epithelial differentiation, we performed differential gene expression analysis for RNA-Seq data between LSCs and dCESs. The dCESs withdrew from cell-cycle progression, as evidenced by loss of KI67 (Fig. 3c). In addition, we obtained a set of TFs that were downregulated upon differentiation, including TP63, MYC, and CEBPD,17 which were well demonstrated to promote LSC proliferation (Fig. 3d).
In contrast, we identified 2011 genes that were associated with CEC differentiation, such as the corneal epithelial markers KRT3,18 KRT12, ALDH3A1,19 and CLU20 (Fig. 3c). Gene ontology (GO) analysis showed that genes preferentially expressed in dCESs are linked to several biological processes, such as extracellular structure/matrix organization, cell–cell adhesion, glycoprotein metabolic process, and collagen metabolic process (Fig. 3e). Gene set enrichment analysis (GSEA) revealed that multiple canonical pathways, including activation of matrix metalloproteinases, O-linked glycosylation of mucins, extracellular matrix glycoproteins, tight junction interactions, cell adhesion molecules cams, extracellular matrix organization, and keratan sulfate keratin metabolism, were significantly upregulated during CEC differentiation (Fig. 3f).
We also identified a cohort of TFs that were highly expressed and upregulated in dCESs, including FOXC1, PAX6, and the well-known pro-differentiation regulators KLF4 and KLF521 (Fig. 3g). The parallel expression pattern observed for FOXC1 and PAX6, both ex vivo and in vivo, implied that FOXC1 may play a key role in the corneal epithelium. In addition, to identify the core regulators of CEC differentiation, we selected the top 25 upregulated TFs by fold changes and computed their functional similarities by GO-terms Semantic Similarity Measures (GSSM, Fig. 3h), a tool for semantic comparisons of GO annotations.22 The high functional similarity score of FOXC1 implied that it was a hub regulator of these 25 candidates.
FOXC1 occupies the H3K4me2-marked cis-regulatory elements
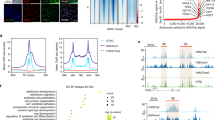
In general, TFs selectively bind to enhancers or promoters to control transcriptional programs. We mapped the genome-wide binding profile of FOXC1 in LSCs using chromatin immunoprecipitation sequencing (ChIP-Seq). ChIP-Seq data of H3K4me2, which is associated with both the transcription start sites (TSSs) and enhancers,23,24 was used to delineate the epigenetic feature of LSCs. The accessible chromatin was assessed by assay for transposase-accessible chromatin with sequencing (ATAC-Seq).25 These high-throughput sequencing data showed a high Pearson’s correlation coefficient between two biological replicates (Supplementary Fig. S1).
We found that FOXC1 principally bound to intronic and intergenic regions, and a small portion (~20%) of FOXC1 peaks was located at proximal promoters. In contrast, ~42% of H3K4me2 peaks were located at promoters and ~50% marked putative noncoding enhancers (Fig. 4a). ATAC and H3K4me2 signals were significantly enriched at the center of FOXC1 peaks (Fig. 4b). Genome-wide binding heatmaps showed that FOXC1 colocated with ATAC and H3K4me2 (Fig. 4c), suggesting that FOXC1 occupied open cis-regulatory elements. Interestingly, FOXC1 bound to its own promoter and multiple neighboring enhancers that were marked by H3K4me2 and ATAC (Fig. 4d), forming an autoregulatory loop. In addition, FOXC1, ATAC, and H3K4me2 were also significantly enriched at the TSSs (Fig. 4e).
a Barplot showing the distribution patterns of FOXC1 and ATAC peaks in LSCs. b Metaplot showing the average FOXC1, H3K4me2, and ATAC enrichment signals around the center of FOXC1 peaks. c Heatmaps showing the binding signals of FOXC1, ATAC, and H3K4me2 at the center of FOXC1 peaks. d Genome browser tracks for FOXC1, H3K4me2, and ATAC enrichment signals across the FOXC1 locus. e Metaplot showing the average FOXC1, H3K4me2, and ATAC enrichment signals around the center of TSSs. f Motifs enriched in ATAC and FOXC1 peaks. g Frequencies of PAX6, TP63, and KLF4 motifs around the centers of FOXC1 and ATAC peaks
Accessible chromatin regulatory elements can recruit TFs to control gene transcription programs. The function and identity of cells or tissues are generally dominated by cooperation between multiple lineage-specific TFs. Thus, we used the HOMER algorithm26 to search TF binding motifs in the FOXC1 and ATAC peaks. We found that both ATAC and FOXC1-bound regions were significantly coenriched for the motifs of key corneal epithelial regulators (TP63, PAX6, GRHL2,27,28 SOX9,29 EHF,30 KLF4, KLF5, and STAT3;31 Fig. 4f). These motifs were located at the center of the FOXC1 and ATAC peaks (Fig. 4g and Supplementary Fig. S2). Collectively, we delineated the chromatin landscapes of FOXC1 binding sites.
FOXC1 determines the corneal epithelial fate via activating PAX6 and repressing epidermal regulators
To explore the function and molecular mechanism of FOXC1 in the corneal epithelium, we performed short-hairpin RNA (shRNA)-mediated knockdown experiments. ZNF750 is known to initiate early- and late-differentiation programs in skin epithelial cells.32 We found that FOXC1 depletion significantly inhibited PAX6 expression and activated ZNF750 in LSCs (Supplementary Fig. S3a and S3b), suggesting that a cellular identity switch was triggered. About a third (107/194) of the downregulated genes in shFOXC1-treated LSCs were preferentially expressed in dCESs (Supplementary Fig. S3c), suggesting that FOXC1 plays a role in corneal epithelial differentiation. However, PAX6 depletion did not significantly affect expression of FOXC1 in LSCs (Fig. S3d). These results indicated that FOXC1 is the upstream regulator of PAX6. Thus, we speculated that FOXC1 dysfunction altered the lineage commitment of LSCs.
Next, FOXC1-deficient LSCs were induced to differentiate into stratified epithelium using the 3D air-lifting culture system. Although epithelial stratification was not affected when FOXC1 was knocked down, the morphology and arrangement of the dCESs were disordered (Fig. S4a). RNA-Seq analysis was used to determine the FOXC1-regulated gene network during differentiation. Hierarchical clustering showed a high degree of agreement between the two biological replicates (Supplementary Fig. S4b). Principal component analysis (PCA) indicated that gene expression of FOXC1-depleted dCESs was quite distinct from that of the control group (Fig. 5a). We identified 1404 downregulated genes and 1148 upregulated genes in shFOXC1-treated dCESs (Supplementary Fig. S4c).
a PCA of the air-lifting-induced dCESs treated with scrambled shRNA and shFOXC1. b Venn diagram showing the overlap between genes preferentially expressed in the dCESs and those that were downregulated after FOXC1 knockdown in the dCESs. c Heatmap showing normalized TPM values of the selected differentially expressed genes in scrambled shRNA-treated and shFOXC1-treated dCESs. d Immunofluorescence staining of the indicated genes in scrambled shRNA- and shFOXC1-treated dCESs. Scale bar, 100 μm. e Pie chart showing the percentages of differentially expressed genes, with or without FOXC1 binding, that were induced by FOXC1 knockdown in the dCESs. f Genome browser tracks for FOXC1, H3K4me2, and ATAC enrichment signals across the PAX6 and CLU loci in LSCs. g GO BP analysis of genes upregulated in the FOXC1-depleted dCESs. h GSEA of keratinization pathway in the gene expression matrix of scrambled shRNA- versus shFOXC1-treated dCESs
FOXC1 knockdown in dCESs downregulated 468 differentiation-associated genes (Fig. 5b), including the corneal epithelium marker genes PAX6, KRT3, KRT12, CLU, MUC1, MUC4, and MUC1633 (Fig. 5c, d and Supplementary Fig. S4d). GSEA also revealed an overall compromise of the differentiation-associated gene program in FOXC1-depleted dCESs (Supplementary Fig. S4e). Of note, FOXC1 inhibition only induced 745 differentially expressed genes in LSCs, which was far less than that (2551) in dCESs. A total of 2180 genes were regulated by FOXC1 only during CEC differentiation (Supplementary Fig. S4f). These results suggested that FOXC1 plays a vital role in the CEC differentiation.
The chromatin binding profile showed that a large percentage (62.3%) of the differentially expressed genes induced by FOXC1 knockdown was directly bound by FOXC1 (Fig. 5e). For example, FOXC1 occupied ATAC-marked and H3K4me2-marked enhancer elements of its downstream genes PAX6, CLU, ST3GAL4, and WNT7B (Fig. 5f and Supplementary Fig. S5a). FOXC1 also bound to the activated promoters of WNT7B, ST3GAL4, MUC1, and IGFBP3, which were repressed upon FOXC1 depletion (Supplementary Fig. S5a, b).
Furthermore, we observed activation of the epidermis-specific genes ZNF750, TINCR, CALML5, SFN, IRF6, KRT1, and KRT108 when FOXC1 was knocked down (Fig. 5c, d and Supplementary Fig. S4d). ZNF750 and TINCR are known to promote epidermal differentiation by inducing CALML5, which interacts with SFN.34 IRF6 is required for skin epithelial differentiation and barrier function.35,36 GO analysis showed that FOXC1-deficient dCESs possessed epidermis function and identity, evidenced by the upregulation of the gene program controlling skin epithelium-specific biological processes, like epidermal cell differentiation, keratinization/cornification, skin development, peptide cross-linking, water homeostasis, and establishment of skin barrier (Fig. 5g). The keratinization pathway was also activated by FOXC1 depletion (Fig. 5h).
Taken together, these results indicated that dysfunction of FOXC1 disrupted the corneal epithelial identity, and induced the conversion of the corneal epithelium to keratinized epidermis. We identified FOXC1 as a key regulator of the corneal epithelial fate determination and homeostasis.
FOXC1 maintains corneal epithelial homeostasis via interferon signaling pathways
To further explore the role of FOXC1 in corneal epithelial homeostasis, we analyzed the FOXC1-regulated gene network in dCESs. We found that FOXC1 controlled the biological processes of eye development, epithelium migration, and multiple signaling pathways, like regulation of ERK1/ERK2 cascade, regulation of mitogen-activated protein kinase (MAPK) activity, and protein kinase B signaling (Fig. 6a). Importantly, FOXC1 deficiency disrupted CEC differentiation-associated protein glycosylation, the glycoprotein metabolic process, and collagen metabolic process (Figs. 6a and 3e). Previous study has demonstrated that interferon signaling can inhibit angiogenesis by regulating the expression of vascular endothelial growth factor A (VEGFA) and soluble VEGF receptor 1 in the human cornea.37 Remarkably, we found that molecules involving in interferon signaling pathways (IRF1, IFNAR1, IFNAR2, IFNGR1, IFNGR2, IRF9, STAT1, and STAT2) and genes response to interferon signaling were significantly repressed in FOXC1-compromised dCESs (Fig. 6a–c and Supplementary Fig. S6a, b). FOXC1 regulated their expression by directly binding to open promoters or distal enhancers (Fig. 6d, e and Supplementary Fig. S6c).
a GO BP analysis of the genes downregulated in FOXC1-depleted dCESs. b Downregulated interferon alpha/beta/gamma (IFN A/B/G) signaling pathway genes in FOXC1-depleted dCESs. Node size represents fold change. c GSEA of IFN A/B signaling pathway in the gene expression matrix of scrambled shRNA-treated versus shFOXC1-treated dCESs. d, e Genome browser tracks for FOXC1, H3K4me2, and ATAC signals in LSCs, and RNA-Seq signals in the dCESs around the IRF1, IFNAR1, IFNGR1, and IRF9 loci. f GSSM of the top 15 (fold change) downregulated TFs in FOXC1-depleted dCESs. The cutoff value (0.75) is indicated by the dashed line. These 15 TFs were also highly expressed in the scramble group. g Immunofluorescence staining of IRF1 in adult limbus-cornea. Scale bar, 100 μm. h Immunofluorescence staining of IRF1 in cornea ulcer tissues. Scale bar, 100 μm
Moreover, we used GSSM to quantify GO semantic similarity of the top 15 (fold change) downregulated TFs with high expression levels, and showed that interferon-induced IRF1 had the highest functional similarity with others (Fig. 6f), suggesting its hub role. IRF1 also showed a high GO semantic similarity score among the CEC differentiation-associated TFs (Fig. 3h). IRF1 expression was also inhibited in LSCs when FOXC1 was knocked down (Supplementary Fig. S3b). The IRF1 motif was enriched in the open chromatin regions in LSCs (Supplementary Fig. S6d). We observed robust expression of IRF1 in the limbus and central corneal epithelium (Fig. 6g). Likewise, loss of IRF1 occurred in the lesion regions of the corneal ulcer tissues (Fig. 6h). These observations indicated that FOXC1 maintains corneal epithelial homeostasis via interferon signaling pathways.
Discussion
Human corneal epithelium, as a barrier tissue of the eye, suffers from external insults, thus requiring continuous self-renewal and differentiation of resident adult stem cells. LSCs are essential for the maintenance of corneal epithelial homeostasis and visual acuity.38 Previous reports demonstrated that FOXC1 is required for ocular anterior segment development in mice and humans, and functional studies on this gene mainly focused on its antiangiogenic role in corneal stroma.10,11,39,40 Here, we identified FOXC1 as a core TF that maintains the corneal epithelial identity during adulthood, which suggested a broader role of FOXC1 in the ocular system.
Specifically, several signaling pathways (such as interferon signaling, ERK1/ERK2 cascade, MAPK, and protein kinase B signaling) were affected in FOXC1-depleted dCESs, which indicated that these signal pathways may be important for the corneal epithelial identity. In addition, FOXC1 also controlled protein glycosylation and collagen metabolism in dCESs. Previous findings indicated that FOXC1 in neural crest-derived corneal stroma cells maintains corneal transparency and avascularity by regulating proangiogenic matrix metalloproteinases and VEGF signaling.11 Thus, FOXC1 appears to govern distinct biological processes in different tissues.
The present study revealed the chromatin binding profile of FOXC1 in LSCs. The function of TFs in the nucleus depends on the active chromatin environment. Thus, we also mapped the chromatin accessibility and H3K4me2 landscapes. As expected, FOXC1 controlled the corneal epithelial gene network by directly binding to promoters or enhancers marked by ATAC and H3K4me2 signals. Notably, motifs of several regulators important to corneal epithelial self-renewal and differentiation were co-enriched at the centers of ATAC and FOXC1 peaks. The proximity of these TFs implied the potential cooperation between them. This observation supported the notion that cooperative binding of cell-type-specific TFs orchestrates lineage identity and homeostasis.26,41,42 Collectively, our present work provides novel insights into the FOXC1-mediated regulatory network and epigenetic features required for corneal epithelial homeostasis.
Interestingly, we found that FOXC1 activated gene expression of the whole interferon signaling pathways in dCESs. Interferon signaling robustly represses angiogenesis in human corneal stroma and tumors.37,43 Thus, we speculated that FOXC1 could inhibit neo-angiogenesis by regulating interferon signaling in the limbus and corneal epithelium. The anti-angiogenic function of FOXC1 may also be achieved in corneal stromal cells via interferon signaling. Previous report has revealed that the interferon response gene IRF1 is important for the corneal innate immune response to bacterial infection.37 Of note, the expression patterns of FOXC1, PAX6, and IRF1 were consistent in corneal epithelial lineage and pathological cornea. Loss of FOXC1, IRF1, and PAX6 was observed in some regions of the corneal ulcer tissues. These findings raised the possibility that disturbance of the FOXC1/interferon signaling axis causes disorder of the corneal innate immune defense, which results in pathological changes in the corneal epithelium. The role and regulatory mechanism of IRF1 in corneal epithelium fate and homeostasis need to be explored in the future.
Materials and methods
Clinical materials
Human embryonic cornea was obtained voluntarily from the donor in compliance with the Ethics Committee of The Third Affiliated Hospital of Guangzhou Medical University (2018006). All human pathological tissues were obtained with the approval of the Ethics Committee of Zhongshan Ophthalmic Center of Sun Yat-sen University (2020KYPZ115).
Culture of primary LSCs
Human limbus tissues isolated from donors were cut into small pieces and digested at 37 °C with 0.2% collagenase IV (Gibco) for 2 h and then with 0.25% trypsin-EDTA (Gibco) for 15 min. LSC expansion was achieved in Matrigel-coated dishes. The components of the LSC medium have been described previously.8
Air-lifting culture protocol
LSCs were plated in transwell culture inserts (Corning), and the medium was added to the upper and lower compartments. The cells were cultured for approximately 3–6 days until they reached confluency, after which the medium was removed from the inserts. LSCs were induced to differentiate for another 5–14 days to form dCESs through contact with the medium in the lower chamber. The medium used for air-lifting culture was the same as that used to culture LSCs.
Immunofluorescence staining
Immunofluorescence experiments were performed as previously described.8 Briefly, after fixation, dehydration, paraffin embedding, and de-paraffinization, the tissue sections were permeated and blocked with 0.3% Triton X-100 and 3% bovine serum albumin for 1 h. Then, the samples were incubated with primary antibodies overnight and with secondary antibodies for 1 h on the following day. Cell nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). The following antibodies were used: anti-FOXC1 (Bethyl Laboratories, A303-519A), anti-K19 (Thermo Scientific, MS-1902-P), anti-p63 (BioLegend, 619002), anti-Ki67 (CST, 9129S), anti-Cytokeratin 1 (Invitrogen, MA1-06312), anti-Cytokeratin 10 (Invitrogen, MA1-06319), anti-PAX6 (BioLegend, 901301), anti-Cytokeratin 12 (Abcam,ab124975), anti-Cytokeratin 3 (Abcam, ab68260), and anti-ZNF750 (Sigma, HPA023012).
RNA interference and quantitative real-time PCR (qPCR) analysis
Two shRNAs targeting FOXC1 were respectively cloned into the PLKO.1 lentivirus plasmid. LSCs were infected with lentivirus particles encoding shRNAs for 16 h and then selected with 2 μg/mL puromycin (Gibco) for 48 h. The RNeasy Mini Kit (Qiagen) was used to isolate total RNA from cells, and cDNAs were synthesized using the PrimeScript™ RT Master Mix Kit (Takara). qPCR was performed using the iTaq™ Universal SYBR® Green Supermix Kit (Bio–Rad). The following shRNAs were used:
shFOXC1-1: 5′-GAGCTTTCGTCTACGACTGTA-3′
shFOXC1-2: 5′-GTCACAGAGGATCGGCTTGAA-3′
shPAX6-1: 5′-CGTCCATCTTTGCTTGGGAAA-3′
shPAX6-2: 5′-AGTTTGAGAGAACCCATTATC-3′
Scramble: 5′-CAACAAGATGAAGAGCACCAA-3′
ChIP-Seq
Chromatin was fixed with 1% formaldehyde for 10 min and sheared in a sonification buffer (50 mM HEPES-NaOH, pH 7.5, 300 mM NaCl, 1 mM EDTA, 0.1%, Na-deoxycholate, 1% TritonX-100, and 0.1% SDS). Fragmented DNA was incubated with primary antibodies overnight at 4 °C and then with Protein A/G Dynabeads (1:1, Invitrogen) for 1 h. The beads were collected and washed sequentially with the following solutions: high-salt buffer (50 mM HEPES-NaOH, pH 7.5, 500 mM NaCl, 1 mM EDTA, 0.1%, Na-Deoxycholate, 1% TritonX-100, and 0.1% SDS), low-salt wash buffer (10 mM Tris-HCl, pH 8.0, 250 mM LiCl, 1 mM EDTA, 0.5% IGEPAL CA-630, and 0.5% Na-Deoxycholate), and TE buffer (10 mM Tri-HCl, pH 8.0, and 1 mM EDTA).
Elution of the immunoprecipitated protein/DNA complexes was performed using elution buffer (50 mM Tris-HCl, pH 8.0, 10 mM EDTA, and 1% SDS) at 65 °C for 4 h. Next, the protein/DNA complexes were incubated with proteinase K (Invitrogen) and RNase A (Invitrogen) for 1 h. De-crosslinked DNA was purified using the PCR Purification Kit (Qiagen), and then, the KAPA Hyper Prep Kit (Kapa Biosystems, KK8502) was used to construct DNA libraries for Illumina PE150 sequencing.
ATAC-Seq
In total, 50,000 cells were digested, collected, and lysed in ice-cold lysis buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 3 mM MgCl2, 0.5% IGEPAL CA-630, and 0.1% Tween-20) for 5 min. Then, Tn5 transposase reactions (Vazyme Biotech, TD501) were performed using the TruePrep DNA Library Prep Kit (Vazyme Biotech). After purification, the transposed DNA fragments were amplified and collected according to manufacturer’s instructions. Library DNA was sequenced with an Illumina HiSeq X10 instrument.
RNA-Seq data assay
STAR software44 was used to align the RNA-Seq reads to the human hg19 reference genome, and the RSEM tool45 was used to calculate the TPM values. Differentially expressed genes were identified using the DESeq2 R package,46 with a log2 fold-change value of ≥1 and a false-discovery rate of <0.05. GO biological process enrichment analysis was performed using the clusterprofiler R package47 with a p-value cut-off of 0.01 and a q-value cut-off of 0.05.
ChIP-Seq and ATAC-Seq data assays
BWA software48 was used to map trimmed reads to the human hg19 reference genome, and the uniquely mapped reads were selected using the Picard MarkDuplicates tools (http://broadinstitute.github.io/picard/). For ChIP-seq data, we used MACS249 to call peaks with the following parameters: -f BAMPE -B –SPMR -q 0.001–call-summits–fix-bimodal–seed 11521 and–extsize 200. For ATAC peak calling, the following parameters were used: -f BAMPE -B –SPMR -q 0.001–call-summits–seed 11521–nomodel–shift -100–extsize 200. The HOMER mergePeaks command was used to obtain overlapping peaks between two biological replicates. Peak distributions were examined with the ChIPseeker package.50 Peak annotations were performed using the HOMER’s annotatePeaks.pl program with the default parameters. Peak summits were used to identify motifs using HOMER’s findMotifsGenome.pl program with the parameter: -size 200.
GSSM analysis
Bioconductor R package GOSemSim was used to compute GO semantic similarity between genes according to the published method.22 Functional similarity of genes was defined as the geometric mean of their semantic similarities in molecular function and cellular component of GO analysis.
Data availability
The data sets that support the findings of this study are available in this paper or the Supplementary Information. Next-generation high-throughput sequencing raw data are available from the corresponding author upon reasonable request.
References
Pellegrini, G. et al. Location and clonal analysis of stem cells and their differentiated progeny in the human ocular surface. J. Cell Biol. 145, 769–782 (1999).
Davanger, M. & Evensen, A. Role of the pericorneal papillary structure in renewal of corneal epithelium. Nature 229, 560–561 (1971).
Cotsarelis, G., Cheng, S. Z., Dong, G., Sun, T. T. & Lavker, R. M. Existence of slow-cycling limbal epithelial basal cells that can be preferentially stimulated to proliferate: implications on epithelial stem cells. Cell 57, 201–209 (1989).
Gonzalez, G., Sasamoto, Y., Ksander, B. R., Frank, M. H. & Frank, N. Y. Limbal stem cells: identity, developmental origin, and therapeutic potential. Wiley Interdiscip. Rev. 7, e303 (2018).
Nowell, C. S., Radtke, F. Corneal epithelial stem cells and their niche at a glance. J. Cell Sci. 130, 198119 (2017).
Osei-Bempong, C., Figueiredo, F. C. & Lako, M. The limbal epithelium of the eye—a review of limbal stem cell biology, disease and treatment. Bioessays 35, 211–219 (2013).
Shaham, O., Menuchin, Y., Farhy, C. & Ashery-Padan, R. Pax6: a multi-level regulator of ocular development. Prog. Retinal Eye Res. 31, 351–376 (2012).
Ouyang, H. et al. WNT7A and PAX6 define corneal epithelium homeostasis and pathogenesis. Nature 511, 358–361 (2014).
Gen, L. et al. Transcription factor PAX6 (Paired Box 6) controls limbal stem cell lineage in development and disease. J. Biol. Chem. 290, 20448 (2015).
Smith, R. S. et al. Haploinsufficiency of the transcription factors FOXC1 and FOXC2 results in aberrant ocular development. Hum. Mol. Genet. 9, 1021–1032 (2000).
Seo, S. et al. Forkhead box transcription factor FoxC1 preserves corneal transparency by regulating vascular growth. Proc. Natl Acad. Sci. USA 109, 2015–2020 (2012).
Ramos, T., Scott, D. & Ahmad, S. An update on ocular surface epithelial stem cells: cornea and conjunctiva. Stem Cells Int. 2015, 601731 (2015).
Senoo, M., Pinto, F., Crum, C. P. & McKeon, F. p63 is essential for the proliferative potential of stem cells in stratified epithelia. Cell 129, 523–536 (2007).
Liu, N. et al. Stem cell competition orchestrates skin homeostasis and ageing. Nature 568, 344–350 (2019).
Tan, E. K., He, H. & Tseng, S. C. G. Epidermal differentiation and loss of clonal growth potential of human limbal basal epithelial progenitor cells during intrastromal invasion. Invest. Ophthalmol. Vis. Sci. 52, 4534 (2011).
Kurpakus, M. A., Maniaci, M. T. & Esco, M. Expression of keratins K12, K4 and K14 during development of ocular surface epithelium. Curr. Eye Res. 13, 805–814 (1994).
Barbaro, V. et al. C/EBPδ regulates cell cycle and self-renewal of human limbal stem cells. J. Cell Biol. 177, 1037–1049 (2007).
Schermer, A., Galvin, S. & Sun, T. T. Differentiation-related expression of a major 64K corneal keratin in vivo and in culture suggests limbal location of corneal epithelial stem cells. J. Cell Biol. 103, 49–62 (1986).
Estey, T., Piatigorsky, J., Lassen, N. & Vasiliou, V. ALDH3A1: a corneal crystallin with diverse functions. Exp. Eye Res. 84, 3–12 (2007).
Nakamura, T., Nishida, K., Dota, A. & Kinoshita, S. Changes in conjunctival clusterin expression in severe ocular surface disease. Invest. Ophthalmol. Vis. Sci. 43, 1702–1707 (2002).
Klein, R. H. et al. Characterization of enhancers and the role of the transcription factor KLF7 in regulating corneal epithelial differentiation. J. Biol. Chem. 292, 18937–18950 (2017).
Han, Y. et al. Proteomic investigation of the interactome of FMNL1 in hematopoietic cells unveils a role in calcium-dependent membrane plasticity. J. Proteom. 78, 72–82 (2013).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Jacobs, J. et al. The transcription factor Grainy head primes epithelial enhancers for spatiotemporal activation by displacing nucleosomes. Nat. Genet. 50, 1011–1020 (2018).
Werth, M. et al. The transcription factor grainyhead-like 2 regulates the molecular composition of the epithelial apical junctional complex. Development 137, 3835–3845 (2010).
Menzel-Severing, J. et al. Transcription factor profiling identifies Sox9 as regulator of proliferation and differentiation in corneal epithelial stem/progenitor cells. Sci. Rep. 8, 10268 (2018).
Stephens, D. N. et al. The Ets transcription factor EHF as a regulator of cornea epithelial cell identity. J. Biol. Chem. 288, 34304–34324 (2013).
Nakamura, T. et al. LRIG1 inhibits STAT3-dependent inflammation to maintain corneal homeostasis. J. Clin. Invest. 124, 385–397 (2014).
Klein, R. H. & Andersen, B. Dynamic networking for epidermal differentiation. Dev. Cell 32, 661–662 (2015).
Hayashi, R. et al. Co-ordinated ocular development from human iPS cells and recovery of corneal function. Nature 531, 376–380 (2016).
Sun, B. K. et al. CALML5 is a ZNF750- and TINCR-induced protein that binds stratifin to regulate epidermal differentiation. Genes Dev. 29, 2225–2230 (2015).
Oberbeck, N. et al. The RIPK4–IRF6 signalling axis safeguards epidermal differentiation and barrier function. Nature 574, 249–253 (2019).
Richardson, R. J. et al. Irf6 is a key determinant of the keratinocyte proliferation-differentiation switch. Nat. Genet. 38, 1329–1334 (2006).
Kommineni, V. K., Nagineni, C. N., William, A., Detrick, B. & Hooks, J. J. IFN-gamma acts as anti-angiogenic cytokine in the human cornea by regulating the expression of VEGF-A and sVEGF-R1. Biochem. Biophys. Res Commun. 374, 479–484 (2008).
Pellegrini, G. & De Luca, M. Eyes on the prize: limbal stem cells and corneal restoration. Cell Stem Cell 15, 121–122 (2014).
Lehmann, O. J. et al. Chromosomal duplication involving the forkhead transcription factor gene FOXC1 causes iris hypoplasia and glaucoma. Am. J. Hum. Genet. 67, 1129–1135 (2000).
Bailey, J. N. et al. Genome-wide association analysis identifies TXNRD2, ATXN2 and FOXC1 as susceptibility loci for primary open-angle glaucoma. Nat. Genet. 48, 189–194 (2016).
Siersbaek, R. et al. Transcription factor cooperativity in early adipogenic hotspots and super-enhancers. Cell Rep. 7, 1443–1455 (2014).
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Kammertoens, T. et al. Tumour ischaemia by interferon-γ resembles physiological blood vessel regression. Nature 545, 98–102 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Chen, T. W. et al. ChIPseek, a web-based analysis tool for ChIP data. BMC Genomics 15, 539 (2014).
Acknowledgements
We thank Dr. Tong Chen and his colleagues at EHBIO Gene Technology (Beijing) for providing technical support for our bioinformatics analysis. This work was supported by The National Key Research and Development (R&D) Program of China (2016YFA0101700), the National Natural Science Foundation of China (31771626, 81622012), and the Guangdong Innovative and Entrepreneurial Research Team Program (2016ZT06S029).
Author information
Authors and Affiliations
Contributions
H.O. and J.Y. designed this project and provided guidance for writing. M.L. and L.Z. designed and performed the experiments. M.L. wrote the manuscript. M.L. and H.H. performed the bioinformatics analysis. L.L., J.L., S.G., J.Z., and B.W. contributed to the plasmid construction and cell culture. H.G., J.T., and Z.M. performed the immunofluorescence experiments. L.W., C.L., and Y.F. assisted in the experimental design and writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, M., Zhu, L., Liu, J. et al. Loss of FOXC1 contributes to the corneal epithelial fate switch and pathogenesis. Sig Transduct Target Ther 6, 5 (2021). https://doi.org/10.1038/s41392-020-00378-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-020-00378-2
This article is cited by
-
The single-cell transcriptomic atlas and RORA-mediated 3D epigenomic remodeling in driving corneal epithelial differentiation
Nature Communications (2024)
-
Novel maternal duplication of 6p22.3-p25.3 with subtelomeric 6p25.3 deletion: new clinical findings and genotype–phenotype correlations
Molecular Cytogenetics (2023)
-
Comprehensive 3D epigenomic maps define limbal stem/progenitor cell function and identity
Nature Communications (2022)