Abstract
Background
An adverse maternal environment (AME) predisposes progeny towards cognitive impairment in humans and mice. Cognitive impairment associates with hippocampal dysfunction. An important regulator of hippocampal function is the hippocampal serotonergic system. Dysregulation of hippocampal serotonin receptor 2c (HTR2c) expression is linked with cognitive impairment. HTR2c contains multiple mRNA variants and isoforms that are epigenetically regulated including DNA methylation, histone modifications, and small nucleolar RNA MBII-52. We tested the hypotheses that AME increases HTR2c variant expression and alters epigenetic modifications along the HTR2c gene locus.
Methods
We create an AME through maternal Western diet and prenatal environmental stress in the mouse. We analyzed hippocampal HTR2c and variants’ expression, DNA methylation and histone modifications along the gene locus, and MBII-52 levels in postnatal day 21 offspring.
Results
AME significantly increased the expressions of total HTR2c and full-length variants (V201 and V202) concurrently with an altered epigenetic profile along the HTR2c gene locus in male offspring hippocampi. Moreover, increased full-length variants’ expression in AME males was in line with increased MBII-52 levels.
Conclusions
AME affects male offspring hippocampal expression of HTR2c and full-length variants via epigenetic mechanisms. Altered hippocampal HTR2c expression may contribute to cognitive impairment seen in adult males in this model.
Impact
-
The key message of our article is that an adverse maternal environment increases expression of total HTR2c mRNA and protein, alters proportions of HTR2c mRNA variants, and impacts HTR2c epigenetic modifications in male offspring hippocampi relative to controls.
-
Our findings add to the literature by providing the first report of altered HTR2c mRNA variant expression in association with altered epigenetic modifications in the hippocampus of offspring mice exposed to an adverse maternal environment.
-
Our findings suggest that an adverse maternal environment affects the expression of genes previously determined to regulate cognitive function through an epigenetic mechanism in a sex-specific manner.
Similar content being viewed by others
Introduction
Adverse maternal environment (AME) and the consumption of a Western diet (WD) in early life predispose progeny toward poor health outcomes and increase the risk of long-term neuropsychiatric morbidities in humans1,2 and animal models.3,4,5,6 In our mouse model of AME and postweaning WD, adult male progeny demonstrate impaired learning and memory function.7 Furthermore, AME male offspring demonstrate reduced hippocampal neurogenesis and brain-derived neurotrophic factor (BDNF) protein levels.8 However, the pathogenesis underlying these findings remain unclear. A relevant pathogenesis to consider is the impact of AME upon hippocampal serotonergic system gene expression. Previous studies link perturbations in serotonergic system functions to multiple pathological disease states defined by memory loss,9,10,11,12 yet, whether AME specifically impacts serotonergic system gene expression remains unknown.
Serotonin (5-hydroxytryptamine, 5-HT) serves as a neurotransmitter for multiple cerebral functions including learning and memory.13,14,15 Serotonin acts through multiple receptors classified as serotonin receptor (HTR) 1 to HTR7. Several serotonin receptors’ subtypes have been further identified based on their respective affinity to agonist and cellular signaling pathways leading to a total of 14 recognized HTRs.16,17 The hippocampus expresses the majority HTRs.18,19,20 Previous studies demonstrate that altered hippocampal expression of HTRs contributes to the pathogenesis of neuropsychiatric disorders-related and age-related cognitive decline.21,22,23
The HTR2 subfamily requires specific attention because of the many studies linking antagonism of this receptor subfamily to improved cognition in various animal models.24 Moreover, the particularly dense expression of the HTR2 subfamily characterizes the hippocampus.25,26 The HTR2 subfamily includes HTR2a, HTR2b, and HTR2c subtypes. For example, reducing HTR2c activity through either a pharmacological approach (blockade with the HTR2c antagonist) or a transgenic approach (HTR2c knockout) improves reversal learning in rats and mice.27,28 Administration of a selective HTR2c antagonist prior to environmental stress repairs defects in hippocampal spatial memory.21 In addition, reducing HTR2c activity increases BDNF protein expression and neuron numbers in the hippocampus in other animal models.29,30
Most species demonstrate significant conservation of the HTR2c gene, which produces multiple mRNA variants and different isoforms due to alternative splicing. The mouse HTR2c gene contains 6 exons. Exon V consists of two parts: Va and Vb, and the latter is subject to alternative splicing.31,32 Inclusion of exon Vb in the pre-mRNA transcript produces a functional full-length version of HTR2c (HTR2c-Fl), whereas skipping of exon Vb of the HTR2c pre-mRNA alters the amino acid reading frame and leads to the production of a truncated protein (HTR2c-Tr). This isoform has no canonical receptor function and yet shows robust expression levels throughout the brain.33,34 HTR2c-Tr interacts with functional HTR2c-Fl and decreases the latter’s availability.33
Regulation of HTR2c expression and alternative splicing involves epigenetic mechanisms such as DNA methylation, histone covalent modifications, and noncoding RNAs.35,36,37 DNA hypermethylation at the promoter region decreases gene expression.35 While DNA methylation in the body of the gene regulates exon alternative splicing.38 Covalent histone modifications including acetylation (ac) and methylation also affect gene transcription. In general, histone H3 lysine (K) 9 acetylation (ac), H3K14ac, and H3K4 trimethylation (me3) lead to gene activation while H3K36me3 marks actively transcribed regions.39,40 In contrast, H3K9me3 and H3K27me3 typically lead to gene silencing.39 All of these histone covalent modifications display vulnerability to perinatal insults.41,42
Noncoding RNAs also participate in the epigenetic regulation of genes and alternative splicing of pre-mRNAs.43 Small nucleolar (sno)-RNA molecules primarily guide chemical modifications of other RNAs. Each snoRNA molecule acts as a guide for only one (or two) individual modifications in a target RNA.44 The brain-specific small nucleolar (sno)-RNA molecules SNORD115 (MBII-52) contain a recognition sequence for the HTR2c pre-RNA and regulate alternative splicing of HTR2c exon Vb.31,32,37,45 MBII-52 promotes the inclusion of exon Vb and thereby the production of a HTR2c-Fl.32
Despite this previous work and the impact of AME on neurodevelopmental issues such as memory and learning as well as neurogenesis, the effects of AME on HTR2c expression and its epigenetic profile in the hippocampus remain unknown. We therefore hypothesized that AME would increase HTR2c variant expression in the hippocampus due to the impaired learning and memory function previously demonstrated in this model. We further hypothesized that AME would alter hippocampal epigenetic characteristics along dysregulated HTR2c gene locus.
Methods
Animals
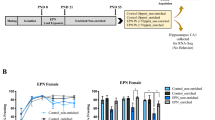
All experiments were conducted according to the Public Health Services Policy on Human Care and Use of Laboratory Animals and all procedures were approved by the Medical College of Wisconsin Institutional Animal Care and Use Committee.46 The mouse model of AME used in this study has been previously described.7 Briefly, AME was induced in mice by exposing six-week-old C57/Bl6 female mice randomly to either a control diet (CD, 10% fat without cholesterol and sucrose, #D14020502, Research Diet Inc., New Brunswick, NJ) or a Western diet (WD, 40% fat, comprised of increased saturated fat, cholesterol, and sucrose, #D12079B, Research Diet Inc.) for 5 weeks prior to pregnancy and throughout lactation. Dams in the CD group experienced a normal environment throughout pregnancy and are designated as Control (Con). Dams fed a WD experienced a ‘stressed’ environment the last third of pregnancy. The combination of chronic WD and gestational stress is designated as Adverse Maternal Environment (AME). The stressed environment consisted of daily random environmental changes as well as a static change in the maternal environment consisting of 1/3 of the standard amount of bedding from embryonic day (E)13 to E19. The acute random environmental changes included altered light cycles on three nonconsecutive days, three cage changes during the daytime on E15, and the short-term introduction of a novel object in the cage for a day. The period of prenatal stress was limited to E13–E19 to minimize newborn mortality, avoid interference with implantation and still target a period of rapid development and environmental vulnerability. Dams were delivered spontaneously, and litters were culled to six pups. At postnatal day 21 (P21), pups from both Con and AME groups were anesthetized and killed. Brains were quickly removed, hippocampi (HP) were dissected, flash-frozen in liquid nitrogen, and stored at −80 °C for molecular studies. For immunohistochemistry studies, animals were individually fixed via intra-cardiac perfusion with ice-cold 0.9% normal saline (VWR, Radnor, PA), followed by ice-cold 4% paraformaldehyde (Electron Microscopy Sciences, Hatfield, PA) for 5 min each for a total volume of 10–15 ml fixative. Whole brains were removed and postfixed at 4 °C overnight. Brains were then transferred to 70% ethanol, paraffin-embedded, and sectioned coronally at 4μm per section. A total of 40 pregnant female mice were used in the study with N = 6 litters/group for each experiment below.
RNA isolation and real-time RT-PCR
Total hippocampal RNA extraction and cDNA syntheses were performed as previously described.47 mRNA levels of genes interested were calculated relative to hypoxanthine phosphoribosyltransferase 1 (HPRT1, Mm.PT.39a.22214828, Integrated DNA Technologies) which was used as an internal control. mRNA levels of following genes were determined in this study: HTR1a (Mm00434106_s1, ThermoFisher Scientific), HTR1b (Mm00439377_s1), HTR2a (Mm00555764_m1), HTR2c (Mm00434127_m1), HTR3a (Mm.PT.58.30882666, Integrated DNA Technologies), HTR4 (Mm00434129_m1), HTR5a (Mm00434132_m1), HTR6 (Mm.PT.58.7237075), HTR7 (Mm00434133_m1) and MBII-52. Alternative splicing of mouse HTR2c gene generates 7 mRNA variants designated as V201-207 (Ensembl: ENSMUSG00000041380). V201 and V202 contain exon Vb, which encode HTR2c-Fl protein. V202 contains a retained intron designated as exon II′ and represents the longest transcript. V203 and V207 lack exon Vb and encode truncated isoforms. No specific assays can be designed for variant V201 and the assay for total HTR2c spans exon 3-4 junction which is common to all coding transcripts V201-203 and V207, we, therefore, calculated the relative amount of V201 transcript by subtracting the sum of 2(−ΔCt (variant transcript – HPRT1)) of V202, V203, and V207 from the 2(−ΔCt (total HTR2c – HPRT1)). Primer and probe sequences for HTR2c variants, HTR2c-Fl, and MBII-52 (acc# AF357427.1) are listed in Table 1. HTR2c-Tr transcript levels were calculated by the sum of V203 and V207 levels.
Protein isolation and immunoblot
Hippocampal tissue proteins isolation and immunoblots were performed as previously described.47 Antibodies against HTR1b (ab13896, Abcam), HTR2c (LS-C368171, Lifespan BioSciences Inc), HTR5a (LS-C356466, Lifespan BioSciences,) and HTR7 (LS-C358892, Lifespan BioSciences) at 1:50 dilution were used to determine protein abundance and Vinculin (#13901, Cell Signaling) at 1:10,000 dilution was used as a loading control.
Immunohistochemistry (IHC)
IHC was used to localize HTR2c expression in the hippocampus and performed using a Leica Bond Rx automated staining platform at the Histology Core Lab in the Department of Pathology Medical College of Wisconsin. Briefly, hippocampal coronal sections at 4 µm from Con and AME brains were deparaffinized, rehydrated, and subject to antigen retrieval treatment. Sections were then blocked with Protein Block (DAKO, Carpinteria, CA, Cat #x090930-2) for 30 min at room temperature (RT) and followed by incubation with rabbit anti-HTR2c (LS-C386171, Lifespan Biosciences) 1:5000 for 60 min at RT. After washing in TBST twice, sections were then exposed to donkey anti-rabbit 1:750 for 45 min at RT. Followed by washing in TBST twice, signals were developed with DAB solution (DAKO, Carpinteria, CA) and counterstained with hematoxylin. Sections were washed and mounted with CytosealTM 60 (ThermoFisher Scientific) mounting media. Images were captured by NanoZoomer Digital pathology slide scanner (Hamamatsu Photonics, Japan).
DNA isolation and pyrosequencing
Hippocampal DNA was extracted from P21 old mice using DNeasy® Blood & Tissue Kit (Qiagen, #69504). DNA was subjected to sodium bisulfite treatment using Epitech fast DNA bisulfite kit (Qiagen, #59824) as per the manufacturer’s protocol to determine site-specific CpG methylation. DNA methylation of the validation-set samples was determined through PCR amplification with biotinylated primers (Intergrated DNA Technologies, Coralville, IA). Primers were designed based on the genomic sequence obtained from Ensembl Genome Browser (Ensembl: ENSMUST00000096299.8) using PyroMark Assay Design Software version 2.0. Amplified products were confirmed with agarose gel electrophoresis. Percent of methylation was quantified by PyroMark Q48 Autoprep Pyrosequencer (Qiagen, Valencia, CA). Six primer sets (Table 2) were used to examine the methylation status of 5 CpG sites in the promoter region spanning from −1 to −240bp nucleotide position from transcription start site which is set as +1, 4 CpG sites around exon II′ region, 5 CpG sites in exon Va and 5 CpG sites in exon Vb regions, respectively.
Chromatin isolation and chromatin immunoprecipitation (ChIP) assay
Hippocampal chromatin was isolated from P21 old mice using tryChIPTM chromatin shearing tissue kit (#520237, Covaris Inc) and focused-ultrasonicator M220 (Covaris Inc.). ChIP assays with antibodies against histone H3 lysine (K) 4 trimethylation (H3K4me3, #9751, Cell Signaling Technology), H3K9me3 (#13969, Cell Signaling Technology), H3K27me3 (#9733, Cell Signaling Technology), H3K36me3 (#4909, Cell Signaling Technology), H3K9 acetylation (H3K9ac, #9649, Cell Signaling Technology) and H3K14ac (#7627, Cell Signaling Technology,) were performed using SimpleChIP® Plus Enzymatic Chromatin IP kit (#9005, Cell Signaling Technology). Real-time PCR was used to quantitate the amount of immunoprecipitation DNA from HTR2c promoter, exon II′, exon Vb, 3′ UTR, and an intergenic region was used as an internal control.48,49 Primers and probes used in the ChIP assay are listed in Table 1. Two control experiments were performed simultaneously with our ChIP experiments. First, we performed a “mock” ChIP that included input but did not utilize antibody. Second, we performed a ChIP that utilized an anti-rabbit secondary antibody as a negative control. Two percent of input was used as a loading control.
Statistics
GraphPad Prism 6 (GraphPad Software, San Diego, CA) was used to perform all analyses. All data presented are expressed as mean ± SD. ANOVA (Fisher’s protected least significant difference) and Mann–Whitney test were used to determine statistical significance between the control and AME groups in gene expression, DNA methylation, and histone modifications. Pearson’s r was used to measure HTR2c and BDNF correlation. Significance was set as p < 0.05.
Results
AME increases HTR2c expression in male hippocampus
We first examined the effect of AME on the mRNA levels of nine HTR genes including HTR1a, HTR1b, HTR2a, HTR2c, HTR3a, HTR4, HTR5a, HTR6, and HTR7 in postnatal day 21 (P21) offspring hippocampi. In male offspring hippocampi, we found that AME significantly increased HTR2c mRNA levels but decreased HTR5a and HTR7 mRNA levels compared to the controls. In female offspring hippocampi, we found that AME significantly increased HTR1b mRNA levels compared to the controls (Fig. 1a). We next determined the effect of AME on protein levels of HTR1b, HTR2c, HTR5a, and HTR7 in P21 offspring hippocampi. In male offspring hippocampi, we found that AME significantly increased HTR2c protein levels (Fig. 1b) without affecting the protein levels of HTR1b (Fig. 1c), HTR5a (Supplementary Fig. S1A) and HTR7 (Supplementary Fig. S1B) compared to the controls. In female offspring hippocampi, however, we found that AME significantly decreased HTR1b (Fig. 1c) and HTR2c protein levels (Fig. 1b) compared to the controls. Given that AME had consistent effects upon HTR2c mRNA and protein levels in P21 male hippocampi, we then focused on the effects of AME on HTR2c for subsequent molecular studies.
HTR2c protein levels inversely correlates with BDNF protein levels in male hippocampus
Previous studies in other models reveal that HTR2c activity negatively affects hippocampal BDNF protein levels in rats and mice.29,30 Moreover, AME decreases male offspring hippocampal BDNF protein levels in the model that is the focus of this manuscript.8 We, therefore, determined the correlation of HTR2c and BDNF protein levels in P21 hippocampi from this current study. We found that HTR2c protein levels were inversely correlated with BDNF protein levels in male offspring hippocampi (Fig. 1c, **p < 0.01) but not in female offspring hippocampi (Fig. 1d). Using IHC, we found that apparent increased HTR2c expression in males was not confined to a particular subfield in the hippocampus (Fig. 2).
AME increases mRNA levels of HTR2c-Fl variants in male hippocampus
Regulation of HTR2c expression transcriptionally depends in part upon the regulation of mRNA variants. We subsequently determined the effect of AME on the expression of HTR2c mRNA variants (Fig. 3a–f).
a A schematic of mouse HTR2c gene. White boxes represent noncoding exons while black boxes represent coding exons. Exon II′ and Vb region are subject to alternative splicing. b A schematic representation of HTR2c mRNA variants. Horizontal lines above exons represent the locations of primers for the variants. The horizontal line above exons III/VI of V201 represents the location of the primers for total HTR2c transcripts while the horizontal line above exons Va/b of V201 represents the location of the primers for HTR2c-Fl transcripts. c HTR2c variant mRNA levels. d HTR2c-Fl mRNA levels. e HTR2c-Tr mRNA levels. f HTR2c-Fl/Tr ratio. Data were presented as mean ± SD. N = 6 litters/group. *p < 0.05, **p < 0.01.
AME significantly increased mRNA levels of HTR2c variants V201 and V202 in P21 male offspring hippocampi compared to the controls (Fig. 3c). AME also significantly increased HTR2c-Fl mRNA levels and HTR2c-Fl/Tr ratio in P21 male hippocampi compared to the controls (Fig. 3d–f). However, AME did not affect HTR2c variant expressions and HTR2c-Fl/Tr ratio in P21 female hippocampi compared to the controls (Fig. 3c–f).
AME alters epigenetic profile of HTR2c gene in male hippocampus
Epigenetic modifications such as DNA methylation and histone covalent modifications modulate alternative splicing of mRNA.50 We subsequently examined the effect of AME on DNA methylation and histone modifications along the HTR2c gene locus, as well as the expression of noncoding RNA MBII-52.
AME did not affect CpG methylation in the HTR2c gene proximal promoter region in DNA from P21 offspring hippocampi in both sexes (Fig. 4a). In contrast, AME significantly increased CpG methylation at upstream CpG site and 2 CpG sites within exon II′ (Fig. 4b), as well as 2 CpG sites in exon Vb region (Fig. 4g) but not in exon Va region in DNA from P21 male hippocampi (Fig. 4f). AME significantly increased CpG methylation at 1 CpG site in exon Va region in DNA from P21 female hippocampi.
Data were presented as mean ± SD. a A schematic of HTR2c promoter region. Vertical lines represent CpG sites examined. Numbers below the vertical lines represent the location of each CpG site relative to the transcription start site that is set as +1. b Percent of CpG methylation in HTR2c promoter. c A schematic of HTR2c V202 exon II′ area. Vertical lines represent CpG sites, 87 bp represents the location of CG1 site relative to the start of exon II′. d Percent of CpG methylation around exon II′ region. e HTR2c exon V sequence. Five CpGs (boxed) in exon Va (top) region and 5 CpGs (boxed) in exon Vb (bottom) region were examined, respectively. f Percent of CpG methylation in exon Va region. g Percent of CpG methylation in exon Vb region. N = 6 litters/group. *p < 0.05.
AME also increased densities of active histone marks along HTR2c gene locus in P21 male offspring hippocampal chromatin. AME significantly increased the densities of active histone marks H3K9ac, H3K14ac, and H3K36me3 in the promoter region of HTR2c in P21 male offspring hippocampal chromatin compared to the controls (Fig. 5a). Furthermore, AME significantly increased the densities of H3K9ac at the V202 exon II′ region (Fig. 5b) and H3K14ac at exon Vb in P21 male offspring hippocampal chromatin compared to the controls (Fig. 5c, d). AME did not affect the densities of H3K4me3, H3K9me3, and H3K27me3 along HTR2c locus in P21 male offspring hippocampal chromatin (Fig. 5b-e). Moreover, AME did not affect any histone marks (H3K9ac, H3K14ac, and H3K36me3) examined in P21 female hippocampal chromatin in contrast to the findings in male chromatin (Supplementary Fig. S2A–D).
Data were presented as mean ± SD. a A schematic of HTR2c V202 genomic. White boxes represent noncoding exons while black boxes represent coding exons. b Histone code at the promoter region. c Histone code in HTR2c V202 exon II′ region. d Histone code in exon Vb region. e Histone code in 3′ UTR region. N = 6 litters/group. *p < 0.05.
Lastly, AME significantly increased MBII-52 expression in the hippocampus compared to the controls, coincidentally with upregulated expression of full-length variants V201 and V202 in males but not in females (Fig. 6).
Discussion
Our study demonstrates that AME programs an increase in the expressions of total HTR2c and full-length variants in the hippocampus of P21 male offspring. Our study further demonstrates that these changes occur concurrently with changes in DNA methylation, histone modifications, and small nucleolar RNA levels that are typically associated with increases in mRNA levels and changes in mRNA variant proportions. The relevance of these findings lies in the context that they occur in a model of AME-induced postnatal impaired learning, memory, and hippocampal neurogenesis in male offspring considering the key role HTR2c plays in these processes. To our knowledge, this is the first time a pre-perinatal event has been demonstrated to impact postnatal HTR2c variant expression and associated epigenetic modifications in offspring hippocampus.
AME impairs nonspatial and spatial learning and memory in adult male mice fed with WD in our model.7 Our finding of AME-induced upregulation of HTR2c expression selectively in male offspring hippocampi in this study suggests that HTR2c may play a role in cognitive functions in this model. Previous studies in non-pre-perinatal models similarly demonstrate that cognitive function impairment associates with dysregulated HTR2c expression.21,27 For instance, HTR2c antagonism improves cognitive flexibility in various animal models.21,24,27 Specifically, reducing the activity of the HTR2c either by a pharmacological approach (blockade with the HTR2c antagonist SB242084) or a transgenic (HTR2c knockout) approach improves reversal learning in rats and mice.27,28 Additionally, administration of selective HTR2c antagonist prior to environmental stress repairs defects in hippocampal long-term potentiation and spatial memory caused by a combination of environmental stressors in mice.21 More importantly, reducing HTR2c activity with antagonist increases BDNF protein expression and neuron numbers in the hippocampus of rats and mice exposed to a chronic unpredictable mild stress condition.29,30,51 Furthermore, HTR2c antagonist SB242084 stimulated progenitor cell proliferation in the dentate gyrus in rats.51 In our study, we found that HTR2c expression inversely correlated with BDNF protein levels in male hippocampi. Together with our previously reported AME-induced reduction of BDNF protein levels concurrently with impaired neurogenesis in male hippocampi,8 we speculate that upregulated HTR2c expression may play a role in BDNF downregulation and subsequently impair neurogenesis in the male hippocampus, which in turn contribute to learning and memory deficits later in life in our model.7 The inconsistent results of HTR1b and HTR2c between mRNA and protein levels in AME female offspring hippocampi in this study indicate that translational regulations may also be involved, at least in the female hippocampi.
Our findings of AME-induced increase in the expression of HTR2c variants V201 and V202 imply that upregulated V201 and V202 contribute to increased total HTR2c expression in male hippocampi in our model. Changes in mRNA levels generally implicate the presence of active epigenetic mechanisms such as DNA methylation and histone covalent modifications. Specific to HTR2c, previous investigations find that DNA hypermethylation at the promoter region represses HTR2c expression.35 In the present study, however, we found that AME did not significantly affect DNA methylation status in HTR2c promoter in either sex. We found instead that AME significantly increased CpG methylation in exon II′ region and specific sites within the exon Vb region. These increases occur with the context of increased mRNA levels of variants V201 and V202 that encode HTR2c-Fl protein in males, suggesting that DNA methylation in exon II′ and Vb region may facilitate their inclusions. The results of increased CpG methylation in HTR2c exons II′ and Vb are similar to what has been previously reported for the rat IGF-1 gene.38 For the IGF-1 gene, controlled cortical impact (CCI) upregulates IGF-1B (the exon 5-containing variant) expression and increases CpG methylation around exon 5 region in rat hippocampi. Together, these findings are in accord with other studies that support an important role for exonal DNA methylation playing in the regulation of alternative splicing.52,53 Moreover, these findings also support the role of DNA methylation in marking a genomic sequence as an exon or intron.54 Another potential contribution of DNA methylation to HTR2c V201 and V202 mRNA alternative splicing may lie in the association between DNA methylation and the specific histone modifications outlined below, as supported by evidence that a dynamic interaction exists between DNA methylation and the histone code.55
Histone modifications appear to play a key role in the process of alternative splicing and subsequent mRNA variant expression.38,56,57 For example, our group previously demonstrated increased occupancy of the activating histone marks (H3K9ac, H3K36me3, and H3K4me3) in IGF-1 exon 5 region are concurrent with increased IGF-1B expression induced by CCI in rat hippocampi.38 Similarly, Chen et al. showed that H3K14ac induces an exon skipping event in human CD4+ T cells.56,58 We found in the present study that AME significantly increased H3K14ac in exon Vb region in male hippocampi. We speculate that increased H3K14ac may promote exon Vb inclusion in male hippocampi in our model. Collectively, these observations support the concept of crosstalk between histone modification and DNA methylation in the context of exon–intron architecture and splicing.59
AME-induced increased H3K14ac accumulation occurred in regions other than HTR2c exon Vb in male hippocampi in the current study. AME also increased H3K14ac accumulation in the proximal promoter region in male hippocampi. Additionally, AME increased densities of the other two activating marks H3K9ac and H3K36me in the promoter region in male hippocampi as well. Interestingly, AME increased density of H3K9ac rather than H3K14ac in exon II′ region which is unique to V202 suggesting that H3K9ac may get involved in exon II′ inclusion in males. Although H3K36me3 is often associated with actively transcribed regions, it also appears at promoter regions.38 For instance, our group previously demonstrated that increased H3K36me accumulation not only occurs in IGF-1 exon 5 but also in the IGF-1 promoter region. Both of which occurred concurrently with upregulated IGF-1 expression induced by CCI in rat hippocampi.38 We speculate that region-specific histone modifications play a role in HTR2c alternative splicing and full-length variant transcriptional upregulation in male hippocampi in the current model.
Although we found an increase in DNA methylation at one CpG site in exon Va region in AME females, there were no differences in the densities of the histone marks examined that were in line with unaltered mRNA levels of HTR2c total and its variants. DNA methylation alone often is not enough to impact mRNA levels.60,61 For example, Gutierrez-Arcelus et al. demonstrated that DNA methylation alone does not significantly drive allele-specific expression in the cells from the human umbilical cord.60 In addition, DNA methylation alone is inconsequential for transcription in A. thaliana plant.61 Taken together, we speculate that DNA methylation in exon Va region alone fails to impact HTR2c transcription in AME females in our model.
Noncoding RNA small nucleolar RNA SNORD115 also regulates alternative splicing of HTR2c exon Vb (MBII-52).31,32,37 Brain-specific MBII-52 harbors a phylogenetically conserved 18-nucleotide antisense element that perfectly complements a region of HTR2c primary transcript that undergoes post-transcriptional changes and subsequently impacts potency.62 MBII-52 directly promotes alternative exon Vb of HTR2c inclusion and favors the production of full-length receptor isoforms with higher potency.62 We similarly found that AME increased MBII-52 expression in male hippocampi concurrently with upregulated expression of total HTR2c and full-length variants, suggesting that increased MBII-52 expression may facilitate exon Vb inclusion and the production of HTR2c-Fl transcripts in AME male hippocampi in our model.
Our study has limitations like all studies. All molecular analyses performed in this study utilized the whole homogenized hippocampus, thus limiting our ability to understand the contribution of hippocampal subfields but allowing for a comprehensive hippocampal investigation.
In summary, we have identified that an adverse maternal environment increases total HTR2c and full-length variants’ expression in association with altered epigenetic profile along HTR2c gene locus in male hippocampi. These results advance the field by providing the first report of altered HTR2c mRNA variant expression in accord with altered epigenetic profiles in the hippocampus of mice exposed to an adverse maternal environment. A generalizable concept inherent to our studies is the potential relevance of epigenetic mechanisms in non-promoter regions of a gene.
References
Alastalo, H. et al. Early life stress and physical and psychosocial functioning in late adulthood. PLoS ONE 8, e69011 (2013).
Barker, D. J. et al. Type 2 (non-insulin-dependent) diabetes mellitus, hypertension and hyperlipidaemia (syndrome X): relation to reduced fetal growth. Diabetologia 36, 62–67 (1993).
Arcego, D. M. et al. Early life adversities or high fat diet intake reduce cognitive function and alter BDNF signaling in adult rats: interplay of these factors changes these effects. Int. J. Dev. Neurosci. 50, 16–25 (2016).
Tozuka, Y. et al. Maternal obesity impairs hippocampal BDNF production and spatial learning performance in young mouse offspring. Neurochem. Int. 57, 235–247 (2010).
Weaver, I. C. et al. Epigenetic programming by maternal behavior. Nat. Neurosci. 7, 847–854 (2004).
Cordner, Z. A. et al. Maternal high-fat diet results in cognitive impairment and hippocampal gene expression changes in rat offspring. Exp. Neurol. 318, 92–100 (2019).
Ke, X. et al. Adverse maternal environment and western diet impairs cognitive function and alters hippocampal glucocorticoid receptor promoter methylation in male mice. Physiol. Rep. 8, e14407 (2020).
Ke, X., Huang, Y., Fu, Q., Lane, R. H. & Majnik, A. Adverse maternal environment alters microRNA-10b-5p expression and its epigenetic profile concurrently with impaired hippocampal neurogenesis in male mouse hippocampus. Dev. Neurosci. 43, 95–105 (2021).
Cowen, P. & Sherwood, A. C. The role of serotonin in cognitive function: evidence from recent studies and implications for understanding depression. J. Psychopharmacol. 27, 575–583 (2013).
Jenkins, T. A., Nguyen, J. C., Polglaze, K. E. & Bertrand, P. P. Influence of tryptophan and serotonin on mood and cognition with a possible role of the gut-brain axis. Nutrients 8, 56 (2016).
Lai, M. K. et al. Loss of serotonin 5-Ht2a receptors in the postmortem temporal cortex correlates with rate of cognitive decline in Alzheimer’s disease. Psychopharmacology 179, 673–677 (2005).
Xu, Y. et al. Neurotransmitter receptors and cognitive dysfunction in Alzheimer’s disease and Parkinson’s disease. Prog. Neurobiol. 97, 1–13 (2012).
Aouizerate, B. et al. Updated overview of the putative role of the serotoninergic system in obsessive-compulsive disorder. Neuropsychiatr. Dis. Treat. 1, 231–243 (2005).
Buhot, M. C. Serotonin receptors in cognitive behaviors. Curr. Opin. Neurobiol. 7, 243–254 (1997).
Cools, R., Roberts, A. C. & Robbins, T. W. Serotoninergic regulation of emotional and behavioural control processes. Trends Cogn. Sci. 12, 31–40 (2008).
Nichols, D. E. & Nichols, C. D. Serotonin receptors. Chem. Rev. 108, 1614–1641 (2008).
Roth, B. L. Multiple serotonin receptors: clinical and experimental aspects. Ann. Clin. Psychiatry 6, 67–78 (1994).
Berumen, L. C., Rodriguez, A., Miledi, R. & Garcia-Alcocer, G. Serotonin receptors in hippocampus. Sci. World J. 2012, 823493 (2012).
Celada, P., Puig, M. V. & Artigas, F. Serotonin modulation of cortical neurons and networks. Front. Integr. Neurosci. 7, 25 (2013).
Meneses, A. Involvement of 5-Ht(2a/2b/2c) receptors on memory formation: simple agonism, antagonism, or inverse agonism? Cell. Mol. Neurobiol. 22, 675–688 (2002).
Busceti, C. L. et al. 5-Ht(2c) serotonin receptor blockade prevents tau protein hyperphosphorylation and corrects the defect in hippocampal synaptic plasticity caused by a combination of environmental stressors in mice. Pharmacol. Res. 99, 258–268 (2015).
Del’Guidice, T. et al. Stimulation of 5-Ht2c receptors improves cognitive deficits induced by human tryptophan hydroxylase 2 loss of function mutation. Neuropsychopharmacology 39, 1125–1134 (2014).
Meltzer, C. C. et al. Serotonin in aging, late-life depression, and Alzheimer’s disease: the emerging role of functional imaging. Neuropsychopharmacology 18, 407–430 (1998).
Svob Strac, D., Pivac, N. & Muck-Seler, D. The serotonergic system and cognitive function. Transl. Neurosci. 7, 35–49 (2016).
Fischette, C. T., Nock, B. & Renner, K. Effects of 5,7-dihydroxytryptamine on serotonin1 and serotonin2 receptors throughout the rat central nervous system using quantitative autoradiography. Brain Res. 421, 263–279 (1987).
Pazos, A., Probst, A. & Palacios, J. M. Serotonin receptors in the human brain–Iv. autoradiographic mapping of serotonin-2 receptors. Neuroscience 21, 123–139 (1987).
Boulougouris, V., Glennon, J. C. & Robbins, T. W. Dissociable effects of selective 5-Ht2a and 5-Ht2c receptor antagonists on serial spatial reversal learning in rats. Neuropsychopharmacology 33, 2007–2019 (2008).
Nilsson, S. R., Ripley, T. L., Somerville, E. M. & Clifton, P. G. Reduced activity at the 5-Ht(2c) receptor enhances reversal learning by decreasing the influence of previously non-rewarded associations. Psychopharmacology 224, 241–254 (2012).
Lu, Y., Ho, C. S., McIntyre, R. S., Wang, W. & Ho, R. C. Agomelatine-induced modulation of brain-derived neurotrophic factor (Bdnf) in the rat hippocampus. Life Sci. 210, 177–184 (2018).
Gumuslu, E. et al. The antidepressant agomelatine improves memory deterioration and upregulates Creb and Bdnf gene expression levels in unpredictable chronic mild stress (Ucms)-exposed mice. Drug Target Insights 8, 11–21 (2014).
Doe, C. M. et al. Loss of the imprinted Snorna Mbii-52 leads to increased 5htr2c pre-RNA editing and altered 5ht2cr-mediated behaviour. Hum. Mol. Genet. 18, 2140–2148 (2009).
Kishore, S. & Stamm, S. The Snorna Hbii-52 regulates alternative splicing of the serotonin receptor 2c. Science 311, 230–232 (2006).
Martin, C. B. et al. Rna splicing and editing modulation of 5-Ht(2c) receptor function: relevance to anxiety and aggression in Vgv mice. Mol. Psychiatry 18, 656–665 (2013).
Isles, A. R. Htr2c splice variants and 5ht2cr-mediated appetite. Trends Endocrinol. Metab. 28, 542–544 (2017).
Schachtschneider, K. M. et al. Impact of neonatal iron deficiency on hippocampal DNA methylation and gene transcription in a porcine biomedical model of cognitive development. BMC Genomics 17, 856 (2016).
Tang, B., Dean, B. & Thomas, E. A. Disease- and age-related changes in histone acetylation at gene promoters in psychiatric disorders. Transl. Psychiatry 1, e64 (2011).
Kishore, S. et al. The Snorna Mbii-52 (Snord 115) is processed into smaller RNAs and regulates alternative splicing. Hum. Mol. Genet. 19, 1153–1164 (2010).
Schober, M. E. et al. Traumatic brain injury increased Igf-1b mRNA and altered Igf-1 exon 5 and promoter region epigenetic characteristics in the rat pup hippocampus. J. Neurotrauma 29, 2075–2085 (2012).
Black, J. C., Van Rechem, C. & Whetstine, J. R. Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol. Cell 48, 491–507 (2012).
Graff, J. & Tsai, L. H. Histone acetylation: molecular mnemonics on the chromatin. Nat. Rev. Neurosci. 14, 97–111 (2013).
Ke, X. et al. Intrauterine growth restriction affects hippocampal dual specificity phosphatase 5 gene expression and epigenetic characteristics. Physiol. Genomics 43, 1160–1169 (2011).
Ke, X. et al. Intrauterine growth retardation affects expression and epigenetic characteristics of the rat hippocampal glucocorticoid receptor gene. Physiol. Genomics 42, 177–189 (2010).
Frias-Lasserre, D. & Villagra, C. A. The importance of ncRNAs as epigenetic mechanisms in phenotypic variation and organic evolution. Front. Microbiol. 8, 2483 (2017).
Gjerde, D. T, Hoang, L. & Hornby, D. RNA Purification and Analysis: Sample Preparation, Extraction, Chromatography 25–26 (Wiley-VCH, 2009).
Cavaille, J. et al. Identification of brain-specific and imprinted small nucleolar RNA genes exhibiting an unusual genomic organization. Proc. Natl Acad. Sci. USA 97, 14311–14316 (2000).
American Physiological Society & World Medical Association General Assembly. Guiding principles for research involving animals and human beings. Am. J. Physiol. Cell Physiol. 282, 3 following instructions for authors (2002).
Cohen, S. et al. Adverse early life environment increases hippocampal microglia abundance in conjunction with decreased neural stem cells in juvenile mice. Int. J. Dev. Neurosci. 55, 56–65 (2016).
Fung, C. M. et al. Iugr prevents Igf-1 upregulation in juvenile male mice by perturbing postnatal Igf-1 chromatin remodeling. Pediatr. Res. 78, 14–23 (2015).
Lan, F. et al. Recognition of unmethylated histone H3 lysine 4 links Bhc80 to Lsd1-mediated gene repression. Nature 448, 718–722 (2007).
Flores, K. et al. Genome-wide association between DNA methylation and alternative splicing in an invertebrate. BMC Genomics 13, 480 (2012).
AlAhmed, S. & Herbert, J. Effect of agomelatine and its interaction with the daily corticosterone rhythm on progenitor cell proliferation in the dentate gyrus of the adult rat. Neuropharmacology 59, 375–379 (2010).
Lopez Soto, E. J. & Lipscombe, D. Cell-specific exon methylation and CTCF binding in neurons regulate calcium ion channel splicing and function. Elife 9, e54879 (2020).
Pant, D., Narayanan, S. P., Vijay, N. & Shukla, S. Hypoxia-induced changes in intragenic DNA methylation correlate with alternative splicing in breast cancer. J. Biosci. 45, 3 (2020).
Chodavarapu, R. K. et al. Relationship between nucleosome positioning and DNA methylation. Nature 466, 388–392 (2010).
Jin, B., Li, Y. & Robertson, K. D. DNA methylation: superior or subordinate in the epigenetic hierarchy? Genes Cancer 2, 607–617 (2011).
Chen, W., Feng, P., Ding, H. & Lin, H. Classifying included and excluded exons in exon skipping event using histone modifications. Front. Genet. 9, 433 (2018).
Kolasinska-Zwierz, P. et al. Differential chromatin marking of introns and expressed exons by H3k36me3. Nat. Genet. 41, 376–381 (2009).
Chen, W., Song, X. & Lin, H. Combinatorial pattern of histone modifications in exon skipping event. Front. Genet. 10, 122 (2019).
Schwartz, S. & Ast, G. Chromatin density and splicing destiny: on the cross-talk between chromatin structure and splicing. EMBO J. 29, 1629–1636 (2010).
Gutierrez-Arcelus, M. et al. Passive and active DNA methylation and the interplay with genetic variation in gene regulation. Elife 2, e00523 (2013).
Zhang, Y., Wendte, J. M., Ji, L. & Schmitz, R. J. Natural variation in DNA methylation homeostasis and the emergence of epialleles. Proc. Natl Acad. Sci. USA 117, 4874–4884 (2020).
Bratkovic, T., Modic, M., Camargo Ortega, G., Drukker, M. & Rogelj, B. Neuronal differentiation induces Snord115 expression and is accompanied by post-transcriptional changes of serotonin receptor 2c mRNA. Sci. Rep. 8, 5101 (2018).
Acknowledgements
Immunohistochemistry studies were carried out using the Clinical and Translational Research Core Lab in the Department of Pathology Medical College of Wisconsin.
Funding
Funding support for this study was provided by the Children’s Mercy Hospital and Department of Pediatrics, Medical College of Wisconsin.
Author information
Authors and Affiliations
Contributions
X.K., Y.H., Q.F., and A.M. performed experiments; X.K. analyzed data and prepared figures; X.K. and R.H.L. interpreted results of experiments; X.K. drafted the manuscript; R.H.L. and A.M. edited and revised the manuscript; R.H.L. approved the final version of the manuscript; R.H.L., X.K., and Y.H. conceived and designed research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent statement
Patient consent was not required for this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ke, X., Huang, Y., Fu, Q. et al. Adverse maternal environment affects hippocampal HTR2c variant expression and epigenetic characteristics in mouse offspring. Pediatr Res 92, 1299–1308 (2022). https://doi.org/10.1038/s41390-022-01962-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-022-01962-8