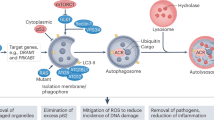
Abstract
Cancer stem cells (CSCs), which are distinct subpopulations of tumor cells, have a substantially higher tumor-initiating capacity and are closely related to poor clinical outcomes. Damage to organelles can trigger CSC pool exhaustion; however, the underlying mechanisms are poorly understood. ZER6 is a zinc-finger protein with two isoforms possessing different amino termini: p52-ZER6 and p71-ZER6. Since their discovery, almost no study reported on their biological and pathological functions. Herein, we found that p52-ZER6 was crucial for CSC population maintenance; p52-ZER6-knocking down almost abolished the tumor initiation capability. Through transcriptomic analyses together with in vitro and in vivo studies, we identified insulin like growth factor 1 receptor (IGF1R) as the transcriptional target of p52-ZER6 that mediated p52-ZER6 regulation of CSC by promoting pro-survival mitophagy. Moreover, this regulation of mitophagy-mediated CSC population maintenance is specific to p52-ZER6, as p71-ZER6 failed to exert the same effect, most possibly due to the presence of the HUB1 domain at its N-terminus. These results provide a new perspective on the regulatory pathway of pro-survival mitophagy in tumor cells and the molecular mechanism underlying p52-ZER6 oncogenic activity, suggesting that targeting p52-ZER6/IGF1R axis to induce CSC pool exhaustion may be a promising anti-tumor therapeutic strategy.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Data availability
All data supporting the findings of this study can be freely accessed by any researcher for non-commercial purposes upon reasonable request.
References
Lee TK, Guan XY, Ma S. Cancer stem cells in hepatocellular carcinoma - from origin to clinical implications. Nat Rev Gastroenterol Hepatol. 2022;19:26–44.
Liu C, Liu L, Chen X, Cheng J, Zhang H, Shen J, et al. Sox9 regulates self-renewal and tumorigenicity by promoting symmetrical cell division of cancer stem cells in hepatocellular carcinoma. Hepatology. 2016;64:117–29.
Hermann PC, Bhaskar S, Cioffi M, Heeschen C. Cancer stem cells in solid tumors. Semin Cancer Biol. 2010;20:77–84.
Vasquez EG, Nasreddin N, Valbuena GN, Mulholland EJ, Belnoue-Davis HL, Eggington HR, et al. Dynamic and adaptive cancer stem cell population admixture in colorectal neoplasia. Cell Stem Cell. 2022;29:1213–28.
Gao Y, Zhang Z, Li K, Gong L, Yang Q, Huang X, et al. Linc-DYNC2H1-4 promotes EMT and CSC phenotypes by acting as a sponge of miR-145 in pancreatic cancer cells. Cell Death Dis. 2017;8:2924.
Huang M, Zhang D, Wu JY, Xing K, Yeo E, Li C, et al. Wnt-mediated endothelial transformation into mesenchymal stem cell-like cells induces chemoresistance in glioblastoma. Sci Transl Med. 2020;12:1.
Duan H, Liu Y, Gao Z, Huang W. Recent advances in drug delivery systems for targeting cancer stem cells. Acta Pharm Sin B. 2021;11:55–70.
Katajisto P, Dohla J, Chaffer CL, Pentinmikko N, Marjanovic N, Iqbal S, et al. Asymmetric apportioning of aged mitochondria between daughter cells is required for stemness. Science. 2015;348:340–3.
Chen Q, Weng K, Lin M, Jiang M, Fang Y, Chung SSW, et al. SOX9 modulates the transformation of gastric stem cells through biased symmetric cell division. Gastroenterology. 2023;1:3.
Raggi C, Taddei ML, Sacco E, Navari N, Correnti M, Piombanti B, et al. Mitochondrial oxidative metabolism contributes to a cancer stem cell phenotype in cholangiocarcinoma. J Hepatol. 2021;74:1373–85.
Steinbichler TB, Dudas J, Skvortsov S, Ganswindt U, Riechelmann H, Skvortsova II. Therapy resistance mediated by cancer stem cells. Semin Cancer Biol. 2018;53:156–67.
Conroy AT, Sharma M, Holtz AE, Wu C, Sun Z, Weigel RJ. A novel zinc finger transcription factor with two isoforms that are differentially repressed by estrogen receptor-alpha. J Biol Chem. 2002;277:9326–34.
Stabach PR, Thiyagarajan MM, Weigel RJ. Expression of ZER6 in ERalpha-positive breast cancer. J Surg Res. 2005;126:86–91.
Huang C, Wu S, Ji H, Yan X, Xie Y, Murai S, et al. Identification of XBP1-u as a novel regulator of the MDM2/p53 axis using an shRNA library. Sci Adv. 2017;3:1701383.
Huang C, Wu S, Li W, Herkilini A, Miyagishi M, Zhao H, et al. Zinc-finger protein p52-ZER6 accelerates colorectal cancer cell proliferation and tumour progression through promoting p53 ubiquitination. EBioMedicine. 2019;48:248–63.
Pei S, Minhajuddin M, Adane B, Khan N, Stevens BM, Mack SC, et al. AMPK/FIS1-mediated mitophagy is required for self-renewal of human AML stem cells. Cell Stem Cell. 2018;23:86–100.
Huang T, Xu T, Wang Y, Zhou Y, Yu D, Wang Z, et al. Cannabidiol inhibits human glioma by induction of lethal mitophagy through activating TRPV4. Autophagy. 2021;17:3592–606.
Liu K, Lee J, Kim JY, Wang L, Tian Y, Chan ST, et al. Mitophagy controls the activities of tumor suppressor p53 to regulate hepatic cancer stem cells. Mol Cell. 2017;68:281–92.
Fan S, Price T, Huang W, Plue M, Warren J, Sundaramoorthy P, et al. PINK1-dependent mitophagy regulates the migration and homing of multiple myeloma cells via the MOB1B-mediated Hippo-YAP/TAZ pathway. Adv Sci. 2020;7:1900860.
Gan ZY, Callegari S, Cobbold SA, Cotton TR, Mlodzianoski MJ, Schubert AF, et al. Activation mechanism of PINK1. Nature. 2022;602:328–35.
Yin K, Lee J, Liu Z, Kim H, Martin DR, Wu D, et al. Mitophagy protein PINK1 suppresses colon tumor growth by metabolic reprogramming via p53 activation and reducing acetyl-CoA production. Cell Death Differ. 2021;28:2421–35.
Chen Q, Lei JH, Bao J, Wang H, Hao W, Li L, et al. BRCA1 deficiency impairs mitophagy and promotes inflammasome activation and mammary tumor metastasis. Adv Sci. 2020;7:1903616.
Youle RJ, Narendra DP. Mechanisms of mitophagy. Nat Rev Mol Cell Biol. 2011;12:9–14.
Luo J, Gong L, Yang Y, Zhang Y, Liu Q, Bai L, et al. Enhanced mitophagy driven by ADAR1-GLI1 editing supports the self-renewal of cancer stem cells in hepatocellular carcinoma. Hepatology. 2023;12:4.
You B, Xia T, Gu M, Zhang Z, Zhang Q, Shen J, et al. AMPK-mTOR-mediated activation of autophagy promotes formation of dormant polyploid giant cancer cells. Cancer Res. 2022;82:846–58.
Yan Y, Zhang D, Zhou P, Li B, Huang SY. HDOCK: a web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 2017;45:365–73.
Yan Y, Tao H, He J, Huang SY. The HDOCK server for integrated protein-protein docking. Nat Protoc. 2020;15:1829–52.
Ludikhuize MC, Meerlo M, Gallego MP, Xanthakis D, Burgaya Julia M, Nguyen NTB, et al. Mitochondria define intestinal stem cell differentiation downstream of a FOXO/Notch axis. Cell Metab. 2020;32:889–900.
Hoang DM, Pham PT, Bach TQ, Ngo ATL, Nguyen QT, Phan TTK, et al. Stem cell-based therapy for human diseases. Signal Transduct Target Ther. 2022;7:272.
Xu L, Liu X, Peng F, Zhang W, Zheng L, Ding Y, et al. Protein quality control through endoplasmic reticulum-associated degradation maintains haematopoietic stem cell identity and niche interactions. Nat Cell Biol. 2020;22:1162–9.
Ghaffari S. Lysosomal regulation of metabolism in quiescent hematopoietic stem cells: more than just autophagy. Cell Stem Cell. 2021;28:374–7.
Picard M, Shirihai OS. Mitochondrial signal transduction. Cell Metab. 2022;34:1620–53.
Dohla J, Kuuluvainen E, Gebert N, Amaral A, Englund JI, Gopalakrishnan S, et al. Metabolic determination of cell fate through selective inheritance of mitochondria. Nat Cell Biol. 2022;24:148–54.
Han JX, Luo LL, Wang YC, Miyagishi M, Kasim V, Wu SR. SGLT2 inhibitor empagliflozin promotes revascularization in diabetic mouse hindlimb ischemia by inhibiting ferroptosis. Acta Pharmacol Sin. 2022;1:22.
Eldeeb MA, Thomas RA, Ragheb MA, Fallahi A, Fon EA. Mitochondrial quality control in health and in Parkinson’s disease. Physiol Rev. 2022;102:1721–55.
Wei M, Nurjanah U, Li J, Luo X, Hosea R, Li Y, et al. YY2-DRP1 axis regulates mitochondrial fission and determines cancer stem cell asymmetric division. Adv Sci. 2023;10:2207349.
Wallace DC. Mitochondria and cancer. Nat Rev Cancer. 2012;12:685–98.
Fang EF, Hou Y, Palikaras K, Adriaanse BA, Kerr JS, Yang B, et al. Mitophagy inhibits amyloid-beta and tau pathology and reverses cognitive deficits in models of Alzheimer’s disease. Nat Neurosci. 2019;22:401–12.
Bravo-San Pedro JM, Kroemer G, Galluzzi L. Autophagy and mitophagy in cardiovascular disease. Circ Res. 2017;120:1812–24.
D’Amico D, Mottis A, Potenza F, Sorrentino V, Li H, Romani M, et al. The RNA-binding protein PUM2 impairs mitochondrial dynamics and mitophagy during aging. Mol Cell. 2019;73:775–87.
Wu Q, Tian AL, Li B, Leduc M, Forveille S, Hamley P, et al. IGF1 receptor inhibition amplifies the effects of cancer drugs by autophagy and immune-dependent mechanisms. J Immunother Cancer. 2021;9:7.
Gong Y, Ma Y, Sinyuk M, Loganathan S, Thompson RC, Sarkaria JN, et al. Insulin-mediated signaling promotes proliferation and survival of glioblastoma through Akt activation. Neuro Oncol. 2016;18:48–57.
Stalnecker CA, Grover KR, Edwards AC, Coleman MF, Yang R, DeLiberty JM, et al. Concurrent inhibition of IGF1R and ERK increases pancreatic cancer sensitivity to autophagy inhibitors. Cancer Res. 2022;82:586–98.
Stan MN, Krieger C. Teprotumumab - a review of its adverse effects profile. J Clin Endocrinol Metab. 2023;45:77.
Dabin R, Wei C, Liang S, Ke C, Zhihan W, Ping Z. Astrocytic IGF-1 and IGF-1R orchestrate mitophagy in traumatic brain injury via exosomal miR-let-7e. Oxid Med Cell Longev. 2022;2022:3504279.
Visvader JE, Lindeman GJ. Cancer stem cells in solid tumours: accumulating evidence and unresolved questions. Nat Rev Cancer. 2008;8:755–68.
Tu Z, Karnoub AE. Mesenchymal stem/stromal cells in breast cancer development and management. Semin Cancer Biol. 2022;86:81–92.
Lin Z, Wu Y, Xu Y, Li G, Li Z, Liu T. Mesenchymal stem cell-derived exosomes in cancer therapy resistance: recent advances and therapeutic potential. Mol Cancer. 2022;21:179.
Melzer C, von der Ohe J, Lehnert H, Ungefroren H, Hass R. Cancer stem cell niche models and contribution by mesenchymal stroma/stem cells. Mol Cancer. 2017;16:28.
Paunovska K, Loughrey D, Dahlman JE. Drug delivery systems for RNA therapeutics. Nat Rev Genet. 2022;23:265–80.
Miyagishi M, Taira K. Strategies for generation of an siRNA expression library directed against the human genome. Oligonucleotides. 2003;13:325–33.
Acknowledgements
This study was financially supported by the National Natural Science Foundation of China (82372655, 82173029, 32070715 and 82203424); the Natural Science Foundation of Chongqing (CSTB2022NSCQ-MSX0611 and CSTB2022NSCQ-MSX0612).
Author information
Authors and Affiliations
Contributions
SW, VK and CH conceived and designed the project, analyzed and interpreted the experimental results, and wrote the manuscript; WL performed most of the experiments; analyzed and interpreted the experimental results, and wrote the manuscript; CH, LQ, YT, XZ, and LZ performed part of qRT-PCR and western blotting. HZ collected human clinical samples and performed clinical samples analysis. MM designed shRNA target sites and analyzed part of the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, W., Huang, C., Qiu, L. et al. p52-ZER6/IGF1R axis maintains cancer stem cell population to promote cancer progression by enhancing pro-survival mitophagy. Oncogene (2024). https://doi.org/10.1038/s41388-024-03058-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41388-024-03058-5