Abstract
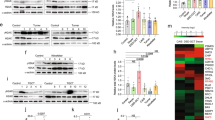
Individuals with a PTEN germline mutation receive the molecular diagnosis of PTEN hamartoma tumor syndrome (PHTS). PHTS displays a complex spectrum of clinical phenotypes including harmartomas, predisposition to cancers, and autism spectrum disorder (ASD). Clear-cut genotype-phenotype correlations are yet to be established due to insufficient information on the PTEN function being impacted by mutations. To fill this knowledge gap, we compared functional impacts of two selected missense PTEN mutant alleles, G132D and M134R, each respectively being associated with distinct clinical phenotype, ASD or thyroid cancer without ASD using gene-edited human induced pluripotent stem cells (hiPSCs). In homozygous hiPSCs, PTEN expression was severely reduced by M134R mutation due to shortened protein half-life. G132D suppressed PTEN expression to a lesser extent than Μ134R mutation without altering protein half-life. When challenged with γ-irradiation, G132D heterozygous cells exited radiation-induced G2 arrest earlier than wildtype and M134R heterozygous hiPSCs despite the similar DNA damage levels as the latter two. Immunoblotting analyses suggested that γ-irradiation induced apoptosis in G132D heterozygous cells to lesser degrees than in the hiPSCs of other genotypes. These data suggest that ASD-associated G132D allele promotes genome instability by premature cell cycle reentry with incomplete DNA repair.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The authors will share any data relating to this work upon reasonable request and execution of data sharing agreement.
References
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science. 1997;275:1943–7.
Steck PA, Pershouse MA, Jasser SA, Yung WKA, Lin H, Ligon AH, et al. Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat Genet. 1997;15:356–62.
Nelen MR, Padberg GW, Peeters EAJ, Lin AY, Helm Bvanden, Frants RR, et al. Localization of the gene for Cowden disease to chromosome 10q22–23. Nat Genet. 1996;13:114–6.
Liaw D, Marsh DJ, Li J, Dahia PLM, Wang SI, Zheng Z, et al. Germline mutations of the PTEN gene in Cowden disease, an inherited breast and thyroid cancer syndrome. Nat Genet. 1997;16:64–67.
Yehia L, Eng C. PTEN hamartoma tumor syndrome. In: Adam MP, Everman DB, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ, et al., editors. GeneReviews®. Seattle, WA: University of Washington; 1993 http://www.ncbi.nlm.nih.gov/books/NBK1488/. Accessed 22 Dec 2022.
Di Cristofano A, Pesco B, Cordon-Cardo C, Pandolfi PP. Pten is essential for embryonic development and tumour suppression. Nat. Genet. 1998;19. https://doi.org/10.1038/1235.
Schwerd T, Khaled AV, Schürmann M, Chen H, Händel N, Reis A, et al. A recessive form of extreme macrocephaly and mild intellectual disability complements the spectrum of PTEN hamartoma tumour syndrome. Eur J Hum Genet. 2016;24:889–94.
Mighell TL, Thacker S, Fombonne E, Eng C, O’Roak BJ. An integrated deep-mutational-scanning approach provides clinical insights on PTEN genotype-phenotype relationships. Am J Hum Genet. 2020;106:818–29.
Yehia L, Ngeow J, Eng C. PTEN-opathies: from biological insights to evidence-based precision medicine. J Clin Investig. 2019;129:452–64.
Sarn N, Thacker S, Lee H, Eng C. Germline nuclear-predominant Pten murine model exhibits impaired social and perseverative behavior, microglial activation, and increased oxytocinergic activity. Mol Autism. 2021;12:41.
Skelton PD, Stan RV, Luikart BW. The role of PTEN in neurodevelopment. CXP. 2019;5:60–71.
Tilot AK, Bebek G, Niazi F, Altemus JB, Romigh T, Frazier TW, et al. Neural transcriptome of constitutional Pten dysfunction in mice and its relevance to human idiopathic autism spectrum disorder. Mol Psychiatry. 2016;21:118–25.
Shen WH, Balajee AS, Wang J, Wu H, Eng C, Pandolfi PP, et al. Essential role for nuclear PTEN in maintaining chromosomal integrity. Cell. 2007;128:157–70.
Chen ZH, Zhu M, Yang J, Liang H, He J, He S, et al. PTEN interacts with histone H1 and controls chromatin condensation. Cell Rep. 2014;8:2003–14.
Groszer M, Erickson R, Scripture-Adams DD, Dougherty JD, Le Belle J, Zack JA, et al. PTEN negatively regulates neural stem cell self-renewal by modulating G0-G1 cell cycle entry. Proc Natl Acad Sci USA. 2006;103:111–6.
McEllin B, Camacho CV, Mukherjee B, Hahm B, Tomimatsu N, Bachoo RM, et al. PTEN loss compromises homologous recombination repair in astrocytes: implications for GBM therapy with temozolomide or PARP inhibitors. Cancer Res. 2010;70:5457–64.
Ming M, Feng L, Shea CR, Soltani K, Zhao B, Han W, et al. PTEN positively regulates UVB-induced DNA damage repair. Cancer Res. 2011;71:5287–95.
Puc J, Keniry M, Li HS, Pandita TK, Choudhury AD, Memeo L, et al. Lack of PTEN sequesters CHK1 and initiates genetic instability. Cancer Cell. 2005;7:193–204.
Mighell TL, Evans-Dutson S, O’Roak BJ. A saturation mutagenesis approach to understanding PTEN lipid phosphatase activity and genotype-phenotype relationships. Am J Hum Genet. 2018;102:943–55.
He X, Ni Y, Wang Y, Romigh T, Eng C. Naturally occurring germline and tumor-associated mutations within the ATP-binding motifs of PTEN lead to oxidative damage of DNA associated with decreased nuclear p53. Hum Mol Genet. 2011;20:80–89.
He X, Arrotta N, Radhakrishnan D, Wang Y, Romigh T, Eng C. Cowden syndrome-related mutations in PTEN associate with enhanced proteasome activity. Cancer Res. 2013;73:3029–40.
Bax M, McKenna J, Do-Ha D, Stevens CH, Higginbottom S, Balez R, et al. The ubiquitin proteasome system is a key regulator of pluripotent stem cell survival and motor neuron differentiation. Cells. 2019;8:E581.
Hartwell LH, Weinert TA. Checkpoints: controls that ensure the order of cell cycle events. Science. 1989;246:629–34.
Kastan MB, Bartek J. Cell-cycle checkpoints and cancer. Nature. 2004;432:316–23.
Hitomi M, Yang K, Stacey AW, Stacey DW. Phosphorylation of cyclin D1 regulated by ATM or ATR controls cell cycle progression. Mol Cell Biol. 2008;28:5478–93.
Firsanov DV, Solovjeva LV, Svetlova MP. H2AX phosphorylation at the sites of DNA double-strand breaks in cultivated mammalian cells and tissues. Clin Epigenet. 2011;2:283–97.
Cruz C, Castroviejo-Bermejo M, Gutiérrez-Enríquez S, Llop-Guevara A, Ibrahim YH, Gris-Oliver A, et al. RAD51 foci as a functional biomarker of homologous recombination repair and PARP inhibitor resistance in germline BRCA-mutated breast cancer. Ann Oncol. 2018;29:1203–10.
Mariotti LG, Pirovano G, Savage KI, Ghita M, Ottolenghi A, Prise KM, et al. Use of the γ-H2AX assay to investigate DNA repair dynamics following multiple radiation exposures. PLoS ONE. 2013;8:e79541.
Yang J, Yu Y, Hamrick HE, Duerksen-Hughes PJ. ATM, ATR and DNA-PK: initiators of the cellular genotoxic stress responses. Carcinogenesis. 2003;24:1571–80.
So S, Davis AJ, Chen DJ. Autophosphorylation at serine 1981 stabilizes ATM at DNA damage sites. J Cell Biol. 2009;187:977–90.
Hirose Y, Katayama M, Mirzoeva OK, Berger MS, Pieper RO. Akt activation suppresses Chk2-mediated, methylating agent–induced G2 arrest and protects from temozolomide-induced mitotic catastrophe and cellular senescence. Cancer Res. 2005;65:4861–9.
Keane NA, Glavey SV, Krawczyk J, O’Dwyer M. AKT as a therapeutic target in multiple myeloma. Expert Opin Ther Targets. 2014;18:897–915.
Jackson MP, Hewitt EW. Cellular proteostasis: degradation of misfolded proteins by lysosomes. Essays Biochem. 2016;60:173–80.
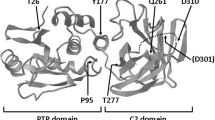
Smith IN, Thacker S, Jaini R, Eng C. Dynamics and structural stability effects of germline PTEN mutations associated with cancer versus autism phenotypes. J Biomol Struct Dyn. 2019;37:1766–82.
Kang SC, Jaini R, Hitomi M, Lee H, Sarn N, Thacker S, et al. Decreased nuclear Pten in neural stem cells contributes to deficits in neuronal maturation. Mol Psychiatry. 2020;11:43.
Tilot AK, Gaugler MK, Yu Q, Romigh T, Yu W, Miller RH, et al. Germline disruption of Pten localization causes enhanced sex-dependent social motivation and increased glial production. Hum Mol Genet. 2014;23:3212–27.
Papa A, Wan L, Bonora M, Salmena L, Song MS, Hobbs RM, et al. Cancer-associated PTEN mutants act in a dominant-negative manner to suppress PTEN protein function. Cell. 2014;157:595–610.
Yehia L, Seyfi M, Niestroj L-M, Padmanabhan R, Ni Y, Frazier TW, et al. Copy number variation and clinical outcomes in patients with germline PTEN mutations. JAMA Netw Open. 2020;3:e1920415.
Markkanen E, Meyer U, Dianov GL. DNA damage and repair in schizophrenia and autism: implications for cancer comorbidity and beyond. Int J Mol Sci. 2016;17:856.
Qing X, Zhang G, Wang Z-Q. DNA damage response in neurodevelopment and neuromaintenance. FEBS J. 2023;290:3300–10.
Wang M, Wei P-C, Lim CK, Gallina IS, Marshall S, Marchetto MC, et al. Increased Neural progenitor proliferation in a hiPSC model of autism induces replication stress-associated genome instability. Cell Stem Cell. 2020;26:221–233.e6.
Nelen MR, Kremer H, Konings IB, Schoute F, Essen AJ, van, Koch R, et al. Novel PTEN mutations in patients with Cowden disease: absence of clear genotype–phenotype correlations. Eur J Hum Genet. 1999;7:267–73.
Yehia L, Ni Y, Sadler T, Frazier TW, Eng C. Distinct metabolic profiles associated with autism spectrum disorder versus cancer in individuals with germline PTEN mutations. npj Genom Med. 2022;7:1–9.
Chen Y-H, Pruett-Miller SM. Improving single-cell cloning workflow for gene editing in human pluripotent stem cells. Stem Cell Res. 2018;31:186–92.
Hitomi M, Stacey DW. The checkpoint kinase ATM protects against stress-induced elevation of cyclin D1 and potential cell death in neurons. Cytometry A. 2010;77:524–33.
Hitomi M, Stacey DW. Cyclin D1 production in cycling cells depends on ras in a cell-cycle-specific manner. Curr Biol. 1999;9:1075–84.
Chumakova AP, Hitomi M, Sulman EP, Lathia JD. High-throughput automated single-cell imaging analysis reveals dynamics of glioblastoma stem cell population during state transition. Cytometry A. 2019;95:290–301.
Acknowledgements
We would like to thank Washington University Genome Engineering and hiPSC Core (GEiC) facility for preparing gene-edited human induced pluripotent stem cells used in this study. This study was supported, in part, by the Lisa Dean Moseley Foundation Grant (to CE and MH) and the Ambrose Monell Foundation (PTEN Switch grant to CE). CE is the Sondra J. and Stephen R. Hardis Endowed Chair of Cancer Genomic Medicine at the Cleveland Clinic, and an ACS Clinical Research Professor.
Author information
Authors and Affiliations
Contributions
CE was responsible for conception and design of the entire study, interpretation of data, and drafting and critical revision of the manuscript. MH was responsible for study design, experimental execution, data acquisition, data analysis, data interpretation, and drafting the manuscript. JV was responsible for designing the project and critical review and revision of the manuscript. SCK was responsible for data acquisition and analysis and reviewing the manuscript. All authors approved of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hitomi, M., Venegas, J., Kang, S.C. et al. Differential cell cycle checkpoint evasion by PTEN germline mutations associated with dichotomous phenotypes of cancer versus autism spectrum disorder. Oncogene 42, 3698–3707 (2023). https://doi.org/10.1038/s41388-023-02867-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-023-02867-4