Abstract
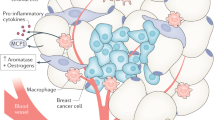
The International Agency for Research on Cancer determined that obesity is the primary preventable cause of breast cancer. The nuclear receptor peroxisome proliferator activated receptor γ (PPARγ) binds inflammatory mediators in obesity and its expression is reduced in human breast cancer. We created a new model to better understand how the obese microenvironment alters nuclear receptor function in breast cancer. The obesity related cancer phenotype was PPARγ dependent; deletion of PPARγ in mammary epithelium which is a tumor suppressor in lean mice unexpectedly increased tumor latency, reduced the luminal progenitor (LP) tumor cell fraction, and increased autophagic and senescent cells. Loss of PPARγ expression in mammary epithelium of obese mice increased expression of 2-aminoadipate semialdehyde synthase (AASS) which regulates lysine catabolism to acetoacetate. PPARγ-associated co-repressors and activators regulated AASS expression via a canonical response element. AASS expression was significantly reduced in human breast cancer, and AASS overexpression or acetoacetate treatment inhibited proliferation and induced autophagy and senescence in human breast cancer cell lines. Genetic or pharmacologic HDAC inhibition promoted autophagy and senescence in mammary tumor cells in vitro and in vivo. We concluded that lysine metabolism is a novel metabolic tumor suppressor pathway in breast cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Data are available on reasonable request from the authors.
References
American Cancer Society. Current year estimates for breast cancer. 2021. Cancer.org.
Tung NM, Garber JE. BRCA1/2 testing: therapeutic implications for breast cancer management. Br J Cancer. 2018;119:141–52.
Dawson SJ, Rueda OM, Aparicio S, Caldas C. A new genome driven integrated classification of breast cancer and its implications. EMBO J. 2013;32:617–28.
Bensinger SJ, Tontonoz P. Integration of metabolism and inflammation by lipid activated nuclear receptors. Nature. 2008;454:470–7.
Vona-Davis L, Rose DP. The obesity-inflammation-eicosanoid axis in breast cancer. J Mamm Gl Biol Neopl. 2013;18:291–307.
Youssef J, Badr M. Peroxisome proliferator activated receptors and cancer: challenges and opportunities. Br J Pharmacol. 2011;164:68–82.
Jiang WG, Douglas-Jones A, Mansel RE. Expression of peroxisome proliferator activated receptor γ (PPARγ) and the PPARγ coactivator, PGC-1, in human breast cancer correlates with clinical outcomes. Int J Cancer. 2003;106:752–7.
Kramer K, Wu J, Crowe DL. Tumor suppressor control of the cancer stem cell niche. Oncogene. 2016;35:4165–78.
Kushi LH, Doyle C, McCullough M, Rock CL, Demark-Wahnefried W, Bandera EV, et al. American Cancer Society guidelines on nutrition and physical activity for cancer prevention. CA Cancer J Clin. 2012;62:30–67.
Iyengar NM, Hudis CA, Dannenberg AJ. Obesity and inflammation: new insights into breast cancer development and progression. Am Soc Clin Oncol Educ Book. 2013;33:45–51.
Howe LR, Subbaramaiah K, Hudis CA, Dannenberg AJ. Molecular pathways: adipose inflammation as a mediator of obesity associated cancer. Clin Cancer Res. 2013;19:6074–83.
Ligibel JA, Strickler HD. Obesity and its impact on breast cancer: tumor incidence, recurrence, survival, and possible interventions. Am Soc Clin Oncol Educ Book. 2013;33:52–9.
Rose DP, Vona-Davis L. Biochemical and molecular mechanisms for the association between obesity, chronic inflammation, and breast cancer. Biofactors. 2014;40:1–12.
Ewertz M, Jensen MB, Gunnarsdottir KA, Hojris I, Jakobsen EH, Nielsen D, et al. Effect of obesity on prognosis after early stage breast cancer. J Clin Oncol. 2011;29:25–31.
von Drygalski A, Tran TB, Messer K, Pu M, Corringham S, Nelson C, et al. Obesity is an independent predictor of poor survival in metastatic breast cancer: retrospective analysis of a patient cohort whose treatment included high dose chemotherapy and autologous stem cell support. Int J Breast Cancer. 2011;2011:523276.
Davis AA, Kaklamani VG. Metabolic syndrome and triple negative breast cancer: a new paradigm. Int J Breast Cancer. 2012;2012:809291.
van Keymeulen A, Rocha AS, Ousset M, Beck B, Bouvencourt G, Rock J, et al. Distinct stem cells contribute to mammary gland development and maintenance. Nature. 2011;479:189–95.
Shehata M, Teschendorff A, Sharp G, Novcic N, Russell IA, Avril S, et al. Phenotypic and functional characterization of the luminal cell hierarchy of the mammary gland. Breast Cancer Res. 2012;14:134–52.
Sauer SW, Opp S, Hoffmann GF, Koeller DM, Okun JG, Kolker S. Therapeutic modulation of cerebral L-lysine metabolism in a mouse model for glutaric aciduria type I. Brain. 2011;134:157–70.
Lavallard VJ, Meijer AJ, Codogno P, Gual P. Autophagy, signaling, and obesity. Pharmacol Res. 2012;66:513–25.
Jain K, Paranandi KS, Sridharan S, Basu A. Autophagy in breast cancer and its implications for therapy. Am J Cancer Res. 2013;3:251–65.
Maycotte P, Thorburn A. Targeting autophagy in breast cancer. World J Clin Oncol. 2014;5:224–40.
Chen S, Jiang YZ, Huang L, Zhou RJ, Yu KD, Liu Y, et al. The residual tumor autophagy marker LC3B serves as a prognostic marker in local advanced breast cancer after neoadjuvant chemotherapy. Clin Cancer Res. 2013;19:6853–62.
Gewirtz DA. The four faces of autophagy: implications for cancer therapy. Cancer Res. 2014;74:647–51.
Capparelli C, Guido C, Whitaker-Menezes D, Bonuccelli G, Balliet R, Pestell TG, et al. Autophagy and senescence in cancer associated fibroblasts metabolically supports tumor growth and metastasis via glycolysis and ketone production. Cell Cycle. 2012;11:2285–302.
Avena P, Anselmo W, Whitaker-Menezes D, Wang C, Pestell RG, Lamb RS, et al. Compartment specific activation of PPARγ governs breast cancer tumor growth via metabolic reprogramming and symbiosis. Cell Cycle. 2013;12:1360–70.
Capparelli C, Chiavarina B, Whitaker-Menezes D, Pestell TG, Pestell RG, Hulit J, et al. Cdk inhibitors (p16/p19/p21) induce senescence and autophagy in cancer associated fibroblasts fueling tumor growth via paracrine interactions without an increase in neo-angiogenesis. Cell Cycle. 2012;11:3599–610.
Brown NE, Jeselsohn R, Bihani T, Hu MG, Foltopoulou P, Kuperwasser C, et al. Cyclin D1 activity regulates autophagy and senescence in the mammary epithelium. Cancer Res. 2012;72:6477–89.
Bertucci F, Ng CKY, Patsouris A, Droin N, Piscuoglio S, Carbuccia N, et al. Genomic characterization of metastatic breast cancers. Nature. 2019;569:560–4.
van Geelen CT, Savas P, Teo ZL, Luen SJ, Weng CF, Ko YA, et al. Clinical indications of prospective genomic profiling of metastatic breast cancer patients. Breast Cancer Res. 2020;22:91.
Rovito D, Gionfriddo G, Barone I, Giordano C, Grande F, De Amicis F, et al. Ligand activated PPARγ downregulates CXCR4 gene expression through a novel identified PPAR response element and inhibits breast cancer progression. Oncotarget. 2016;7:65109–24.
Cui Z, Xie M, Wu Z, Shi Y. Relationship between histone deacetylase 3 (HDAC3) and breast cancer. Med Sci Monitor. 2018;24:2456–64.
Ramadan WS, Talaat IM, Hachim MY, Lischka A, Gemoll T, El-Awady R. The impact of CBP expression in estrogen receptor positive breast cancer. Clin Epigenet. 2021;13:72.
Finn PF, Dice JF. Ketone bodies stimulate chaperone mediated autophagy. J Biol Chem. 2005;280:25864–70.
Rojas-Morales P, Tapia E, Pedraza-Chaverri J. β-hydroxybutyrate: a signaling metabolite in starvation response. Cell Signal. 2016;28:917–23.
Huang YH, Yang PM, Chuah QY, Lee YJ, Hsieh YF, Peng CW, et al. Autophagy promotes radiation induced senescence but inhibits bystander effects in human breast cancer cells. Autophagy. 2014;10:1212–28.
Sharma K, Goehe RW, Di X, Hicks MA, Torti SV, Torti FM, et al. A novel cytostatic form of autophagy in sensitization of non-small cell lung cancer cells to radiation by vitamin D analog EB1089. Autophagy. 2014;10:2346–61.
Wilson EN, Bristol ML, Di X, Maltese WA, Koterba K, Beckman MJ, et al. A switch between cytoprotective and cytotoxic autophagy in the radiosensitization of breast tumor cells by chloroquine and vitamin D. Horm Cancer. 2011;2:272–85.
Zhao Y, He J, Yang L, Luo Q, Liu Z. Histone deacetylase 3 modification of microRNA-31 promotes cell proliferation and aerobic glycolysis in breast cancer and is predictive of poor prognosis. J Breast Cancer. 2018;21:112–23.
Palmieri D, Lockman PR, Thomas FC, Hua E, Herring J, Hargrave E, et al. Vorinostat inhibits brain metastatic colonization in a model of triple negative breast cancer and induces DNA double strand breaks. Clin Cancer Res. 2009;15:6148–57.
De Cremoux P, Dalyai M, N’Dove O, Moutahir F, Rolland G, Chouchane-Mik O, et al. HDAC inhibition does not induce estrogen receptor in human triple negative breast cancer cell lines and patient derived xenografts. Breast Cancer Res Treat. 2015;149:81–9.
Ha K, Fiskus W, Choi DS, Bhaskara S, Cerchietti L, Devarai SG, et al. Histone deacetylase inhibitor treatment induces BRCAness and synergistic lethality with PARP inhibitor and cisplatin against human triple negative breast cancer cells. Oncotarget. 2014;5:5637–50.
Vasilatos SN, Katz TA, Oesterreich S, Wan Y, Davidson NE, Huang Y. Crosstalk between lysine specific demethylase 1 (LSD1) and histone deacetylases mediates antineoplastic efficacy of HDAC inhibitors in human breast cancer cells. Carcinogenesis. 2013;34:1196–207.
Tu Y, Hershman DL, Bhalla K, Fiskus W, Pellegrino CM, Andreopoulou E, et al. A phase I-II study of the histone deacetylase inhibitor vorinostat plus sequential weekly paclitaxel and doxorubicin-cyclophosphamide in locally advanced breast cancer. Breast Cancer Res Treat. 2014;146:145–52.
Wang J, Kim TH, Ahn MY, Lee J, Jung JH, Choi WS, et al. Sirtinol, a class III HDAC inhibitor, induces apoptotic and autophagic cell death in MCF7 human breast cancer cells. Int J Oncol. 2012;41:1101–9.
Acknowledgements
We thank Drs. Ke Ma, Balaji Ganesh, Hui Chen, and Feigen Seiler (University of Illinois Research Resources Center) for assistance with confocal microscopy, flow cytometry, mass spectrometry, and electron microscopy. This study was supported by Department of Defense Breast Cancer Research Program Award W81XWH-20-1-0029.
Author information
Authors and Affiliations
Contributions
JW and KK performed experiments and analyzed data. DLC supervised the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, J., Kramer, K. & Crowe, D.L. Lysine metabolism is a novel metabolic tumor suppressor pathway in breast cancer. Oncogene 42, 2402–2414 (2023). https://doi.org/10.1038/s41388-023-02766-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-023-02766-8