Abstract
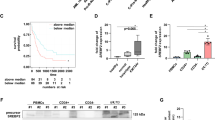
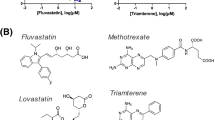
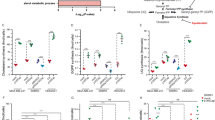
TET2 (ten-eleven-translocation) protein is a Fe(II)- and α-ketoglutarate-dependent dioxygenase that catalyzes DNA demethylation to regulate gene expression. While TET2 gene is frequently mutated in hematological cancer, its enzymatic activity is also compromised in various solid tumors. Whether TET2 deficiency creates vulnerability for cancer cells has not been studied. Here we reported that TET2 deficiency is associated with the change of lipid metabolism processes in acute myeloid leukemia (AML) patient. We demonstrate that statins, the inhibitors of β-Hydroxy β-methylglutaryl-CoA (HMG-CoA) reductase and commonly used cholesterol-lowering medicines, significantly sensitize TET2 deficient tumor cells to apoptosis. TET2 directly regulates the expression of HMG-CoA synthase (HMGCS1) by catalyzing demethylation on its promoter region, and conversely TET2 deficiency leads to significant down-regulation of HMGCS1 expression and the mevalonate pathway. Consistently, overexpression of HMGCS1 in TET2-deficient cells rescues statin-induced apoptosis. We further reveal that decrease of geranylgeranyl diphosphate (GGPP), an intermediate metabolite in the mevalonate pathway, is responsible for statin-induced apoptosis. GGPP shortage abolishes normal membrane localization and function of multiple small GTPases, leading to cell dysfunction. Collectively, our study reveals a vulnerability in TET2 deficient tumor and a potential therapeutic strategy using an already approved safe medicine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Baylin SB, Ohm JE. Epigenetic gene silencing in cancer - a mechanism for early oncogenic pathway addiction? Nat Rev Cancer. 2006;6:107–16.
Kazanets A, Shorstova T, Hilmi K, Marques M, Witcher M. Epigenetic silencing of tumor suppressor genes: paradigms, puzzles, and potential. Biochim Biophys Acta. 2016;1865:275–88.
He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q, et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science. 2011;333:1303–7.
Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Masse A, et al. Mutation in TET2 in myeloid cancers. N. Engl J Med. 2009;360:2289–301.
Tefferi A, Pardanani A, Lim KH, Abdel-Wahab O, Lasho TL, Patel J, et al. TET2 mutations and their clinical correlates in polycythemia vera, essential thrombocythemia and myelofibrosis. Leukemia. 2009;23:905–11.
Xu YP, Lv L, Liu Y, Smith MD, Li WC, Tan XM, et al. Tumor suppressor TET2 promotes cancer immunity and immunotherapy efficacy. J Clin Invest. 2019;129:4316–31.
Yang H, Liu Y, Bai F, Zhang JY, Ma SH, Liu J, et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene. 2013;32:663–9.
Jin SG, Jiang Y, Qiu R, Rauch TA, Wang Y, Schackert G, et al. 5-Hydroxymethylcytosine is strongly depleted in human cancers but its levels do not correlate with IDH1 mutations. Cancer Res. 2011;71:7360–5.
Kudo Y, Tateishi K, Yamamoto K, Yamamoto S, Asaoka Y, Ijichi H, et al. Loss of 5-hydroxymethylcytosine is accompanied with malignant cellular transformation. Cancer Sci. 2012;103:670–6.
Jawert F, Hasseus B, Kjeller G, Magnusson B, Sand L, Larsson L. Loss of 5-hydroxymethylcytosine and TET2 in oral squamous cell carcinoma. Anticancer Res. 2013;33:4325–8.
Xu W, Yang H, Liu Y, Yang Y, Wang P, Kim SH, et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell. 2011;19:17–30.
Laukka T, Mariani CJ, Ihantola T, Cao JZ, Hokkanen J, Kaelin WG Jr, et al. Fumarate and succinate regulate expression of hypoxia-inducible genes via TET enzymes. J Biol Chem. 2016;291:4256–65.
Killian JK, Kim SY, Miettinen M, Smith C, Merino M, Tsokos M, et al. Succinate dehydrogenase mutation underlies global epigenomic divergence in gastrointestinal stromal tumor. Cancer Disco. 2013;3:648–57.
Wu D, Hu D, Chen H, Shi G, Fetahu IS, Wu F, et al. Glucose-regulated phosphorylation of TET2 by AMPK reveals a pathway linking diabetes to cancer. Nature 2018;559:637–41.
Zhang X, Yang J, Shi D, Cao Z. TET2 suppresses nasopharyngeal carcinoma progression by inhibiting glycolysis metabolism. Cancer Cell Int. 2020;20:363.
Chen L, Ma MY, Sun M, Jiang LY, Zhao XT, Fang XX, et al. Endogenous sterol intermediates of the mevalonate pathway regulate HMGCR degradation and SREBP-2 processing. J Lipid Res. 2019;60:1765–75.
Goldstein JL, Brown MS. Regulation of the mevalonate pathway. Nature. 1990;343:425–30.
Waller DD, Park J, Tsantrizos YS. Inhibition of farnesyl pyrophosphate (FPP) and/or geranylgeranyl pyrophosphate (GGPP) biosynthesis and its implication in the treatment of cancers. Crit Rev Biochem Mol Biol. 2019;54:41–60.
Buhaescu I, Izzedine H. Mevalonate pathway: a review of clinical and therapeutical implications. Clin Biochem. 2007;40:575–84.
Takai Y, Sasaki T, Matozaki T. Small GTP-binding proteins. Physiol Rev. 2001;81:153–208.
Wong WW, Dimitroulakos J, Minden MD, Penn LZ. HMG-CoA reductase inhibitors and the malignant cell: the statin family of drugs as triggers of tumor-specific apoptosis. Leukemia. 2002;16:508–19.
Farmer JA, Torre-Amione G. Comparative tolerability of the HMG-CoA reductase inhibitors. Drug Saf. 2000;23:197–213.
Davidson MH. Safety profiles for the HMG-CoA reductase inhibitors: treatment and trust. Drugs. 2001;61:197–206.
Illingworth DR, Tobert JA. HMG-CoA reductase inhibitors. Adv Protein Chem. 2001;56:77–114.
Bottorff M, Hansten P. Long-term safety of hepatic hydroxymethyl glutaryl coenzyme A reductase inhibitors: the role of metabolism-monograph for physicians. Arch Intern Med. 2000;160:2273–80.
Wang Y, Xiao M, Chen X, Chen L, Xu Y, Lv L, et al. WT1 recruits TET2 to regulate its target gene expression and suppress leukemia cell proliferation. Mol Cell. 2015;57:662–73.
Cheng M, Bhujwalla ZM, Glunde K. Targeting Phospholipid Metabolism in Cancer. Front Oncol. 2016;6:266.
Nguyen UT, Guo Z, Delon C, Wu Y, Deraeve C, Franzel B, et al. Analysis of the eukaryotic prenylome by isoprenoid affinity tagging. Nat Chem Biol. 2009;5:227–35.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell 2011;144:646–74.
Shen H, Laird PW. Interplay between the cancer genome and epigenome. Cell 2013;153:38–55.
Lee JV, Carrer A, Shah S, Snyder NW, Wei S, Venneti S, et al. Akt-dependent metabolic reprogramming regulates tumor cell histone acetylation. Cell Metab. 2014;20:306–19.
Wolf A, Agnihotri S, Munoz D, Guha A. Developmental profile and regulation of the glycolytic enzyme hexokinase 2 in normal brain and glioblastoma multiforme. Neurobiol Dis. 2011;44:84–91.
Chen M, Zhang J, Li N, Qian Z, Zhu M, Li Q, et al. Promoter hypermethylation mediated downregulation of FBP1 in human hepatocellular carcinoma and colon cancer. PLoS One. 2011;6:e25564.
Tefferi A, Lim KH, Abdel-Wahab O, Lasho TL, Patel J, Patnaik MM, et al. Detection of mutant TET2 in myeloid malignancies other than myeloproliferative neoplasms: CMML, MDS, MDS/MPN and AML. Leukemia. 2009;23:1343–5.
Abdel-Wahab O, Mullally A, Hedvat C, Garcia-Manero G, Patel J, Wadleigh M, et al. Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood. 2009;114:144–7.
Nibourel O, Kosmider O, Cheok M, Boissel N, Renneville A, Philippe N, et al. Incidence and prognostic value of TET2 alterations in de novo acute myeloid leukemia achieving complete remission. Blood. 2010;116:1132–5.
Bullinger L, Dohner K, Dohner H. Genomics of acute myeloid leukemia diagnosis and pathways. J Clin Oncol. 2017;35:934–46.
Istvan ES, Deisenhofer J. Structural mechanism for statin inhibition of HMG-CoA reductase. Science. 2001;292:1160–4.
Garcia-Bermudez J, Baudrier L, Bayraktar EC, Shen Y, La K, Guarecuco R, et al. Squalene accumulation in cholesterol auxotrophic lymphomas prevents oxidative cell death. Nature 2019;567:118–22.
Jaffe AB, Hall A. Rho GTPases: biochemistry and biology. Annu Rev Cell Dev Biol. 2005;21:247–69.
Acknowledgements
This work was supported by the National Key Research and Development Project of China (No. 2018YFA0800304 to H.-X.Y.), the NSFC grants (No. 82172595 to H.-X.Y.), and the Development Fund for Shanghai Talents (No. 2019109 to H.-X.Y.). H.-X.Y. is also supported by the Supporting Fund for Innovation and Entrepreneurship by Chongqing Returned Oversea Scholars.
Author information
Authors and Affiliations
Contributions
S-JS, YX, K-LG and H-XY conceived the general framework of this study. Y-JA contributed to the bioinformatics analysis. S-JS, Y-JA and H-XY designed the experiments. S-JS, Y-JA, K-LD, J-YZ, CZ and Y-PS performed experiments. S-JS, Y-JA, YX, K-LG and H-XY prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
KLG is a co-founder of Vivace Therapeutics. YX is a co-founder of Cullgen Inc. Other authors declare no competing financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, SJ., Ai, YJ., Duan, KL. et al. TET2 deficiency sensitizes tumor cells to statins by reducing HMGCS1 expression. Oncogene 41, 5385–5396 (2022). https://doi.org/10.1038/s41388-022-02531-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02531-3