Abstract
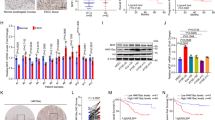
Enhancer of zeste homolog 2 (EZH2) and SET domain bifurcated 1 (SETDB1, also known as ESET) are oncogenic methyltransferases implicated in a number of human cancers. These enzymes typically function as epigenetic repressors of target genes by methylating histone H3 K27 and H3-K9 residues, respectively. Here, we show that EZH2 and SETDB1 are essential to proliferation in 3 SCC cell lines, HSC-5, FaDu, and Cal33. Additionally, we find both of these proteins highly expressed in an aggressive stem-like SCC sub-population. Depletion of either EZH2 or SETDB1 disrupts these stem-like cells and their associated phenotypes of spheroid formation, invasion, and tumor growth. We show that SETDB1 regulates this SCC stem cell phenotype through cooperation with ΔNp63α, an oncogenic isoform of the p53-related transcription factor p63. Furthermore, EZH2 is upstream of both SETDB1 and ΔNp63α, activating these targets via repression of the tumor suppressor RUNX3. We show that targeting this pathway with inhibitors of EZH2 results in activation of RUNX3 and repression of both SETDB1 and ΔNp63α, antagonizing the SCC cancer stem cell phenotype. This work highlights a novel pathway that drives an aggressive cancer stem cell phenotype and demonstrates a means of pharmacological intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq and ChIP-seq data are available from the Gene Expression Omnibus database (accession no. GSE202789) Methods for CRISPR activation, CRISPR depletion, domain-focused CRISPR screening, cDNA overexpression, EPZ-6438 and GSK126 treatment, GFP depletion assays, tumor xenografts, ChIP-seq, and RNA-seq are described in the Supplementary Information. Antibodies and reagents are also listed there, as well as RT-qPCR primer sequences and sgRNA sequences.
References
Yan W, Wistuba II, Emmert-Buck MR, Erickson HS. Squamous cell carcinoma—similarities and differences among anatomical sites. Am J Cancer Res. 2011;1:275–300.
Paver EC, Currie AM, Gupta R, Dahlstrom JE. Human papilloma virus related squamous cell carcinomas of the head and neck: diagnosis, clinical implications and detection of HPV. Pathology. 2020;52:179–91.
Liu-Smith F, Jia J, Zheng Y. UV-induced molecular signaling differences in melanoma and non-melanoma skin cancer. Adv Exp Med Biol. 2017;996:27–40.
Kikuchi K, Inoue H, Miyazaki Y, Ide F, Kojima M, Kusama K. Epstein-Barr virus (EBV)-associated epithelial and non-epithelial lesions of the oral cavity. Jpn Dent Sci Rev. 2017;53:95–109.
Johnson N. Tobacco use and oral cancer: a global perspective. J Dent Educ. 2001;65:328–39.
Crosbie EJ, Einstein MH, Franceschi S, Kitchener HC. Human papillomavirus and cervical cancer. Lancet. 2013;382:889–99.
Bacciu A, Mercante G, Ingegnoli A, Ferri T, Muzzetto P, Leandro G, et al. Effects of gastroesophageal reflux disease in laryngeal carcinoma. Clin Otolaryngol Allied Sci. 2004;29:545–8.
Lee SH. Chemotherapy for lung cancer in the era of personalized medicine. Tuberc Respir Dis. 2019;82:179–89.
Deng HY, Wang WP, Wang YC, Hu WP, Ni PZ, Lin YD, et al. Neoadjuvant chemoradiotherapy or chemotherapy? A comprehensive systematic review and meta-analysis of the options for neoadjuvant therapy for treating oesophageal cancer. Eur J Cardiothorac Surg. 2017;51:421–31.
Tagliamento M, Genova C, Rijavec E, Rossi G, Biello F, Dal Bello MG, et al. Afatinib and Erlotinib in the treatment of squamous-cell lung cancer. Expert Opin Pharmacother. 2018;19:2055–62.
Kaidar-Person O, Gil Z, Billan S. Precision medicine in head and neck cancer. Drug Resist Updat. 2018;40:13–6.
Oshimori N. Cancer stem cells and their niche in the progression of squamous cell carcinoma. Cancer Sci. 2020;111:3985–92.
Dragu DL, Necula LG, Bleotu C, Diaconu CC, Chivu-Economescu M. Therapies targeting cancer stem cells: Current trends and future challenges. World J Stem Cells. 2015;7:1185–201.
Fisher ML, Balinth S, Hwangbo Y, Wu C, Ballon C, Wilkinson JE, et al. BRD4 regulates transcription factor ∆Np63α to drive a cancer stem cell phenotype in squamous cell carcinomas. Cancer Res. 2021;81:6246–58.
Mills AA, Zheng B, Wang XJ, Vogel H, Roop DR, Bradley A. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature. 1999;398:708–13.
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dötsch V, et al. p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell. 1998;2:305–16.
Soares E, Zhou H. Master regulatory role of p63 in epidermal development and disease. Cell Mol Life Sci. 2018;75:1179–90.
Fisher ML, Balinth S, Mills AA. p63-related signaling at a glance. J Cell Sci. 2020;133:jcs228015.
Petitjean A, Ruptier C, Tribollet V, Hautefeuille A, Chardon F, Cavard C, et al. Properties of the six isoforms of p63: p53-like regulation in response to genotoxic stress and cross talk with DeltaNp73. Carcinogenesis. 2008;29:73–81.
Ayyanathan K, Lechner MS, Bell P, Maul GG, Schultz DC, Yamada Y, et al. Regulated recruitment of HP1 to a euchromatic gene induces mitotically heritable, epigenetic gene silencing: a mammalian cell culture model of gene variegation. Genes Dev. 2003;17:1855–69.
Karanth AV, Maniswami RR, Prashanth S, Govindaraj H, Padmavathy R, Jegatheesan SK, et al. Emerging role of SETDB1 as a therapeutic target. Expert Opin Ther Targets. 2017;21:319–31.
Regina C, Compagnone M, Peschiaroli A, Lena A, Annicchiarico-Petruzzelli M, Piro MC, et al. Setdb1, a novel interactor of ΔNp63, is involved in breast tumorigenesis. Oncotarget. 2016;7:28836–48.
Margueron R, Reinberg D. The Polycomb complex PRC2 and its mark in life. Nature. 2011;469:343–9.
Bachmann IM, Halvorsen OJ, Collett K, Stefansson IM, Straume O, Haukaas SA, et al. EZH2 expression is associated with high proliferation rate and aggressive tumor subgroups in cutaneous melanoma and cancers of the endometrium, prostate, and breast. J Clin Oncol. 2006;24:268–73.
Sneeringer CJ, Scott MP, Kuntz KW, Knutson SK, Pollock RM, Richon VM, et al. Coordinated activities of wild-type plus mutant EZH2 drive tumor-associated hypertrimethylation of lysine 27 on histone H3 (H3K27) in human B-cell lymphomas. Proc Natl Acad Sci USA. 2010;107:20980–5.
Zhao M, Hu X, Xu Y, Wu C, Chen J, Ren Y, et al. Targeting of EZH2 inhibits epithelial‑mesenchymal transition in head and neck squamous cell carcinoma via regulating the STAT3/VEGFR2 axis. Int J Oncol. 2019;55:1165–75.
Kim KH, Roberts CW. Targeting EZH2 in cancer. Nat Med. 2016;22:128–34.
Bae SC, Choi JK. Tumor suppressor activity of RUNX3. Oncogene. 2004;23:4336–40.
Xiao Z, Tian Y, Jia Y, Shen Q, Jiang W, Chen G, et al. RUNX3 inhibits the invasion and migration of esophageal squamous cell carcinoma by reversing the epithelial‑mesenchymal transition through TGF‑β/Smad signaling. Oncol Rep. 2020;43:1289–99.
Keyes WM, Pecoraro M, Aranda V, Vernersson-Lindahl E, Li W, Vogel H, et al. ΔNp63α is an oncogene that targets chromatin remodeler Lsh to drive skin stem cell proliferation and tumorigenesis. Cell Stem Cell. 2011;8:164–76.
Shi J, Wang E, Milazzo JP, Wang Z, Kinney JB, Vakoc CR. Discovery of cancer drug targets by CRISPR-Cas9 screening of protein domains. Nat Biotechnol. 2015;33:661–7.
Sniezek JC, Matheny KE, Westfall MD, Pietenpol JA. Dominant negative p63 isoform expression in head and neck squamous cell carcinoma. Laryngoscope. 2004;114:2063–72.
Somerville TDD, Xu Y, Miyabayashi K, Tiriac H, Cleary CR, Maia-Silva D, et al. TP63-mediated enhancer reprogramming drives the squamous subtype of pancreatic ductal adenocarcinoma. Cell Rep. 2018;25:1741–55.e7.
Bailey TL, Johnson J, Grant CE, Noble WS. The MEME Suite. Nucleic Acids Res. 2015;43:W39–49.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, et al. The human genome browser at UCSC. Genome Res. 2002;12:996–1006.
Romano RA, Birkaya B, Sinha S. Defining the regulatory elements in the proximal promoter of DeltaNp63 in keratinocytes: Potential roles for Sp1/Sp3, NF-Y, and p63. J Investig Dermatol. 2006;126:1469–79.
Antonini D, Rossi B, Han R, Minichiello A, Di Palma T, Corrado M, et al. An autoregulatory loop directs the tissue-specific expression of p63 through a long-range evolutionarily conserved enhancer. Mol Cell Biol. 2006;26:3308–18.
Tsusaka T, Shimura C, Shinkai Y. ATF7IP regulates SETDB1 nuclear localization and increases its ubiquitination. EMBO Rep. 2019;20:e48297.
Bangsow C, Rubins N, Glusman G, Bernstein Y, Negreanu V, Goldenberg D, et al. The RUNX3 gene-sequence, structure and regulated expression. Gene. 2001;279:221–32.
Selvarajan V, Osato M, Nah GSS, Yan J, Chung TH, Voon DC, et al. RUNX3 is oncogenic in natural killer/T-cell lymphoma and is transcriptionally regulated by MYC. Leukemia. 2017;31:2219–27.
Otani S, Date Y, Ueno T, Ito T, Kajikawa S, Omori K, et al. Runx3 is required for oncogenic Myc upregulation in p53-deficient osteosarcoma. Oncogene. 2022;41:683–91.
Kudo Y, Tsunematsu T, Takata T. Oncogenic role of RUNX3 in head and neck cancer. J Cell Biochem. 2011;112:387–93.
Gao F, Huang C, Lin M, Wang Z, Shen J, Zhang H, et al. Frequent inactivation of RUNX3 by promoter hypermethylation and protein mislocalization in oral squamous cell carcinomas. J Cancer Res Clin Oncol. 2009;135:739–47.
Jili S, Eryong L, Lijuan L, Chao Z. RUNX3 inhibits laryngeal squamous cell carcinoma malignancy under the regulation of miR-148a-3p/DNMT1 axis. Cell Biochem Funct. 2016;34:597–605.
Sugiura H, Ishiguro H, Kuwabara Y, Kimura M, Mitsui A, Mori Y, et al. Decreased expression of RUNX3 is correlated with tumor progression and poor prognosis in patients with esophageal squamous cell carcinoma. Oncol Rep. 2008;19:713–9.
Kumar A, Singhal M, Chopra C, Srinivasan S, Surabhi RP, Kanumuri R, et al. Threonine 209 phosphorylation on RUNX3 by Pak1 is a molecular switch for its dualistic functions. Oncogene. 2016;35:4857–65.
Kanumuri R, Chelluboyina AK, Biswal J, Vignesh R, Pandian J, Venu A, et al. Small peptide inhibitor from the sequence of RUNX3 disrupts PAK1–RUNX3 interaction and abrogates its phosphorylation-dependent oncogenic function. Oncogene. 2021;40:5327–41.
Fujii S, Ito K, Ito Y, Ochiai A. Enhancer of zeste homologue 2 (EZH2) down-regulates RUNX3 by increasing histone H3 methylation. J Biol Chem. 2008;283:17324–32.
Lu B, Klingbeil O, Tarumoto Y, Somerville TDD, Huang YH, Wei Y, et al. A transcription factor addiction in leukemia imposed by the MLL promoter sequence. Cancer Cell. 2018;34:970–81.e8.
Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, Barcena C, et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature. 2015;517:583–8.
Levanon D, Groner Y. Structure and regulated expression of mammalian RUNX genes. Oncogene. 2004;23:4211–9.
Date Y, Ito K. Oncogenic RUNX3: a link between p53 deficiency and MYC dysregulation. Mol Cells. 2020;43:176–81.
Jiang L, Yu H, Ness S, Mao P, Guo F, Tang J, et al. Comprehensive analysis of co-mutations identifies cooperating mechanisms of tumorigenesis. Cancers. 2022;14:415.
Jian Z, Strait A, Jimeno A, Wang XJ. Cancer stem cells in squamous cell carcinoma. J Investig Dermatol. 2017;137:31–7.
Lin C, Li X, Zhang Y, Guo Y, Zhou J, Gao K, et al. The microRNA feedback regulation of p63 in cancer progression. Oncotarget. 2015;6:8434–53.
Acknowledgements
This work was supported by the Office of the Director, National Institutes of Health through award numbers 5P30CA045508 (Cancer Center Support Grant), CA247400 (to SB), CA225134 (to MLF), as well as R01CA190997 and R21OD018332 (to AAM). This project was also supported through the Cold Spring Harbor Laboratory and Northwell Health Affiliation.
Author information
Authors and Affiliations
Contributions
SB: Performed and designed experiments, paper writing. MLF: Performed and designed experiments. YH: Performed and designed experiments. CW: Performed and designed experiments. CB: Performed experiments. XS: Designed experiments. AAM: Experimental design and direction, paper writing, funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Balinth, S., Fisher, M.L., Hwangbo, Y. et al. EZH2 regulates a SETDB1/ΔNp63α axis via RUNX3 to drive a cancer stem cell phenotype in squamous cell carcinoma. Oncogene 41, 4130–4144 (2022). https://doi.org/10.1038/s41388-022-02417-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02417-4
This article is cited by
-
RUNX transcription factors: biological functions and implications in cancer
Clinical and Experimental Medicine (2024)
-
The functions of SET domain bifurcated histone lysine methyltransferase 1 (SETDB1) in biological process and disease
Epigenetics & Chromatin (2023)
-
p63: a crucial player in epithelial stemness regulation
Oncogene (2023)