Abstract
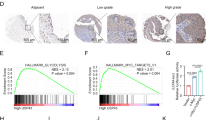
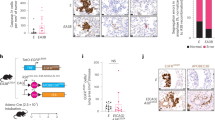
Rapid progression is the major cause of the poor prognosis of hepatocellular carcinoma (HCC); however, the underlying mechanism remained unclear. Here, we found Calpain-2 (CAPN2), a well-established protease that accelerates tumor progression in several malignancies, is overexpressed in HCC and acts as an independent predictor for poor outcomes. Furthermore, CAPN2 promoted the proliferation and invasion of HCC, and showed a positive correlation with the levels of invasion-related markers. Mechanistically, a novel CAPN2-SRC positive regulatory loop was identified upstream of β-catenin to prevent its ubiquitination and degradation, and subsequently promoted HCC progression: CAPN2 could proteolyze PTP1B to form a truncation of approximately 42 kDa with increased phosphatase activity, resulting in reduced SRC Y530 phosphorylation and increased SRC kinase activity; meanwhile, CAPN2 itself was a bone fide substrate of SRC that was primarily phosphorylated at Y625 by SRC and exhibited increased proteolysis activity upon phosphorylation. Interestingly, the CAPN2-SRC loop could not only restrain most of cytoplasmic β-catenin degradation by inhibiting GSK3β pathway, but also prevented TRIM33-induced nuclear β-catenin degradation even in β-catenin-mutant cells. Present study identified a CAPN2-SRC positive loop responsible for intracellular β-catenin accumulation and signaling activation, and targeting CAPN2 protease activity might be a promising approach for preventing HCC progression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Zhu XD, Sun HC. Emerging agents and regimens for hepatocellular carcinoma. J Hematol Oncol. 2019;12:110.
Finn RS, Zhu AX. Evolution of systemic therapy for hepatocellular carcinoma. Hepatology. 2021;73:150–7.
Lee TK, Guan XY, Ma S. Cancer stem cells in hepatocellular carcinoma - from origin to clinical implications. Nat Rev Gastroenterol Hepatol. 2022;19:26–44.
Ahn JC, Teng PC, Chen PJ, Posadas E, Tseng HS, Lu S, et al. Detection of circulating tumor cells and their implications as a biomarker for diagnosis, prognostication, and therapeutic monitoring in hepatocellular carcinoma. Hepatology. 2021;73:422–36.
Kong FH, Ye QF, Miao XY, Liu X, Huang SQ, Xiong L, et al. Current status of sorafenib nanoparticle delivery systems in the treatment of hepatocellular carcinoma. Theranostics. 2021;11:5464–90.
Ye QH, Zhu WW, Zhang JB, Qin Y, Lu M, Lin GL, et al. GOLM1 modulates EGFR/RTK cell-surface recycling to drive hepatocellular carcinoma metastasis. Cancer Cell. 2016;30:444–58.
Baudry M, Bi X. Calpain-1 and Calpain-2: the Yin and Yang of synaptic plasticity and neurodegeneration. Trends Neurosci. 2016;39:235–45.
Xie XQ, Wang MJ, Li Y, Lei LP, Wang N, Lv ZY, et al. miR-124 intensified oxaliplatin-based chemotherapy by targeting CAPN2 in colorectal cancer. Mol Ther Oncolytics. 2020;17:320–31.
Xu F, Gu J, Lu C, Mao W, Wang L, Zhu QL, et al. Calpain-2 enhances non-small cell lung cancer progression and chemoresistance to paclitaxel via EGFR-pAKT pathway. Int J Biol Sci. 2019;15:127–37.
Storr SJ, Lee KW, Woolston CM, Safuan S, Green AR, Macmillan RD, et al. Calpain system protein expression in basal-like and triple-negative invasive breast cancer. Ann Oncol. 2012;23:2289–96.
Lee WJ, Shin CH, Ji H, Jeong SD, Park MS, Won HH, et al. hnRNPK-regulated LINC00263 promotes malignant phenotypes through miR-147a/CAPN2. Cell Death Dis. 2021;12:290.
Zhang Y, Zhu X, Qiao X, Gu X, Xue JQ, Han YS, et al. LIPH promotes metastasis by enriching stem-like cells in triple-negative breast cancer. J Cell Mol Med. 2020;24:9125–34.
Shen C, Yu Y, Li H, Liu MQ, Shen HL, Yang PY. Global profiling of proteolytically modified proteins in human metastatic hepatocellular carcinoma cell lines reveals CAPN2 centered network. Proteomics. 2012;12:1917–27.
Perugorria MJ, Olaizola P, Labiano I, Esparza-Baquer A, Marzioni M, Marin JJG, et al. Wnt-β-catenin signalling in liver development, health and disease. Nat Rev Gastroenterol Hepatol. 2019;16:121–36.
Liu Y, Chen H, Zheng P, Zheng YX, Luo Q, Xie GH, et al. ICG-001 suppresses growth of gastric cancer cells and reduces chemoresistance of cancer stem cell-like population. J Exp Clin Cancer Res. 2017;36:125.
Liu Q, He L, Li S, Li FY, Deng GZ, Huang XJ, et al. HOMER3 facilitates growth factor-mediated β-Catenin tyrosine phosphorylation and activation to promote metastasis in triple negative breast cancer. J Hematol Oncol. 2021;14:6.
Moro L, Simoneschi D, Kurz E, Arbini AA, Jang SW, Guaragnella N, et al. Epigenetic silencing of the ubiquitin ligase subunit FBXL7 impairs c-SRC degradation and promotes epithelial-to-mesenchymal transition and metastasis. Nat Cell Biol. 2020;22:1130–42.
Leung HW, Leung CON, Lau EY, Chung KPS, Mok EH, Lei MML, et al. EPHB2 activates β-catenin to enhance cancer stem cell properties and drive sorafenib resistance in hepatocellular carcinoma. Cancer Res. 2021;81:3229–40.
Pleiman CM, Hertz WM, Cambier JC. Activation of phosphatidylinositol-3’ kinase by Src-family kinase SH3 binding to the p85 subunit. Science. 1994;263:1609–12.
Ngan E, Stoletov K, Smith HW, Common J, Muller WJ, Lewis JD, et al. LPP is a Src substrate required for invadopodia formation and efficient breast cancer lung metastasis. Nat Commun. 2017;8:15059.
Liu L, Wang T, Yang X, Xu CX, Liao ZH, Wang XD, et al. MTNR1B loss promotes chordoma recurrence by abrogating melatonin-mediated β-catenin signaling repression. J Pineal Res. 2019;67:e12588.
Lin J, Song T, Li C, Mao W. GSK-3β in DNA repair, apoptosis, and resistance of chemotherapy, radiotherapy of cancer. Biochim Biophys Acta Mol Cell Res. 2020;1867:118659.
Song Y, Liu Y, Pan S, Xie S, Wang ZW, Zhu X. Role of the COP1 protein in cancer development and therapy. Semin Cancer Biol. 2020;67:43–52.
Lyle CL, Belghasem M, Chitalia VC. c-Cbl: an important regulator and a target in angiogenesis and tumorigenesis. Cells. 2019;8:498.
Xue J, Chen Y, Wu Y, Wang ZY, Zhou AD, Zhang SC, et al. Tumour suppressor TRIM33 targets nuclear β-catenin degradation. Nat Commun. 2015;6:6156.
Cortesio CL, Chan KT, Perrin BJ, Burton NO, Zhang S, Zhang ZY, et al. Calpain 2 and PTP1B function in a novel pathway with Src to regulate invadopodia dynamics and breast cancer cell invasion. J Cell Biol. 2008;180:957–71.
Blom N, Sicheritz-Pontén T, Gupta R, Gammeltoft S, Brunak S. Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics. 2004;4:1633–49.
Wang C, Xu H, Lin S, Deng WK, Zhou JQ, Zhang Y, et al. GPS 5.0: an update on the prediction of kinase-specific phosphorylation sites in proteins. Genomics Proteom Bioinforma. 2020;18:72–80.
Gao Q, Zhu H, Dong L, Shi WW, Chen R, Song ZJ, et al. Integrated proteogenomic characterization of HBV-related hepatocellular carcinoma. Cell. 2019;179:561–77.
Caruso S, Calatayud AL, Pilet J, Le Bella T, Rekik S, Imbeaud S, et al. Analysis of liver cancer cell lines identifies agents with likely efficacy against hepatocellular carcinoma and markers of response. Gastroenterology. 2019;157:760–76.
Chitalia VC, Foy RL, Bachschmid MM, Zeng LL, Panchenko MV, Zhou MI, et al. Jade-1 inhibits Wnt signalling by ubiquitylating beta-catenin and mediates Wnt pathway inhibition by pVHL. Nat Cell Biol. 2008;10:1208–16.
Shao H, Chou J, Baty CJ, Burke NA, Watkins SC, Stolz DB, et al. Spatial localization of m-calpain to the plasma membrane by phosphoinositide biphosphate binding during epidermal growth factor receptor-mediated activation. Mol Cell Biol. 2006;26:5481–96.
Tai WT, Chen YL, Chu PY, Chen LJ, Hung MH, Shiau CW, et al. Protein tyrosine phosphatase 1B dephosphorylates PITX1 and regulates p120RasGAP in hepatocellular carcinoma. Hepatology. 2016;63:1528–43.
Bruix J, Sherman M. Management of hepatocellular carcinoma: an update. Hepatology. 2011;53:1020–2.
Jiang Y, Sun A, Zhao Y, Ying WT, Sun HC, Yang XR, et al. Proteomics identifies new therapeutic targets of early-stage hepatocellular carcinoma. Nature. 2019;567:257–61.
Sun YF, Xu Y, Yang XR, Guo W, Zhang X, Qiu SJ, et al. Circulating stem cell-like epithelial cell adhesion molecule-positive tumor cells indicate poor prognosis of hepatocellular carcinoma after curative resection. Hepatology. 2013;57:1458–68.
Dong ZR, Sun D, Yang YF, Zhou W, Wu R, Wang XW, et al. TMPRSS4 drives angiogenesis in hepatocellular carcinoma by promoting HB-EGF expression and proteolytic cleavage. Hepatology. 2020;72:923–39.
Hu B, Xu Y, Li YC, Huang JF, Cheng JW, Guo W, et al. CD13 promotes hepatocellular carcinogenesis and sorafenib resistance by activating HDAC5-LSD1-NF-κB oncogenic signaling. Clin Transl Med. 2020;10:e233.
Acknowledgements
We thank all the patient enrolled for their agreement for using their tissue samples. Moreover, we thank James P. Mahaffey, PhD, from Liwen Bianji (Edanz) (www.liwenbianji.cn/), for editing the English text of a draft of this manuscript.
Funding
LR-Q was sponsored by National Natural Science Foundation of China (81772774, 82072876); Program of scientific and technological innovation action plan, Shanghai Municipal Commission of science and technology (20XD1434200, 20Y11903500). GL was sponsored by National Natural Science Foundation of China (81772808). RN was sponsored by the Shanghai International Science and Technology Collaboration Program (18410721900), and the National Natural Science Foundation of China (81472672). MX-L was sponsored by Shanghai Sailing Program (20YF1407900); National Natural Science Foundation of China (82002618). WY-C was sponsored by National Natural Science Foundation of China (81800190). ZH was sponsored by Shanghai Natural Science Foundation, Shanghai Municipal Commission of science and technology (19ZR1410300).
Author information
Authors and Affiliations
Contributions
RQL, LG, NR, and XLM contributed to the study design. XLM, KYZ, YDC, YT, HZ, YCW, and WGT contributed to the acquisition, analysis, and interpretation of the data. XLM, SHX, NR, LG, and RQL helped to draft the manuscript. NR, LG, and RQL critically revised the manuscript. All authors read and approve the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Shanghai Cancer Center Research Ethics Committee and Zhongshan Hospital Research Ethics Committee, and all individuals provided informed consent for inclusion of their tissue in this study. Moreover, animal study was approved by the Animal Experimentation Ethics Committee of Shanghai Cancer Center, Fudan University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ma, XL., Zhu, KY., Chen, YD. et al. Identification of a novel Calpain-2-SRC feed-back loop as necessity for β-Catenin accumulation and signaling activation in hepatocellular carcinoma. Oncogene 41, 3554–3569 (2022). https://doi.org/10.1038/s41388-022-02367-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02367-x