Abstract
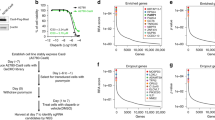
Inhibitors of the mitotic kinase PLK1 yield objective responses in a subset of refractory cancers. However, PLK1 overexpression in cancer does not correlate with drug sensitivity, and the clinical development of PLK1 inhibitors has been hampered by the lack of patient selection marker. Using a high-throughput chemical screen, we discovered that cells deficient for the tumor suppressor ARID1A are highly sensitive to PLK1 inhibition. Interestingly this sensitivity was unrelated to canonical functions of PLK1 in mediating G2/M cell cycle transition. Instead, a whole-genome CRISPR screen revealed PLK1 inhibitor sensitivity in ARID1A deficient cells to be dependent on the mitochondrial translation machinery. We find that ARID1A knock-out (KO) cells have an unusual mitochondrial phenotype with aberrant biogenesis, increased oxygen consumption/expression of oxidative phosphorylation genes, but without increased ATP production. Using expansion microscopy and biochemical fractionation, we see that a subset of PLK1 localizes to the mitochondria in interphase cells. Inhibition of PLK1 in ARID1A KO cells further uncouples oxygen consumption from ATP production, with subsequent membrane depolarization and apoptosis. Knockdown of specific subunits of the mitochondrial ribosome reverses PLK1-inhibitor induced apoptosis in ARID1A deficient cells, confirming specificity of the phenotype. Together, these findings highlight a novel interphase role for PLK1 in maintaining mitochondrial fitness under metabolic stress, and a strategy for therapeutic use of PLK1 inhibitors. To translate these findings, we describe a quantitative microscopy assay for assessment of ARID1A protein loss, which could offer a novel patient selection strategy for the clinical development of PLK1 inhibitors in cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The RNA seq and CRISPR screen data from this publication have been deposited to the Gene Expression Omnibus database [58] (https://www.ncbi.nlm.nih.gov/geo/) and assigned the identifier GSE193942. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE [59] partner repository with the dataset identifier PXD031110.
Change history
06 May 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41388-022-02338-2
References
Bruinsma W, Raaijmakers JA, Medema RH. Switching Polo-like kinase-1 on and off in time and space. Trends Biochem Sci. 2012;37:534–42.
Liu Z, Sun Q, Wang X. PLK1, a potential target for cancer therapy. Transl Oncol. 2017;10:22–32.
Yim H. Current clinical trials with polo-like kinase 1 inhibitors in solid tumors. Anticancer Drugs. 2013;24:999–1006.
Gutteridge RE, Ndiaye MA, Liu X, Ahmad N. Plk1 inhibitors in cancer therapy: from laboratory to clinics. Mol Cancer Ther. 2016;15:1427–35.
Liu X. Targeting polo-like kinases: a promising therapeutic approach for cancer treatment. Transl Oncol. 2015;8:185–95.
Elsayed I, Wang X. PLK1 inhibition in cancer therapy: potentials and challenges. Future Med Chem. 2019;11:1383–6.
Kadoch C, Hargreaves DC, Hodges C, Elias L, Ho L, Ranish J, et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat Genet. 2013;45:592–601.
Shen J, Peng Y, Wei L, Zhang W, Yang L, Lan L, et al. ARID1A deficiency impairs the DNA damage checkpoint and sensitizes cells to PARP inhibitors. Cancer Disco. 2015;5:752–67.
Dykhuizen EC, Hargreaves DC, Miller EL, Cui K, Korshunov A, Kool M, et al. BAF complexes facilitate decatenation of DNA by topoisomerase IIalpha. Nature. 2013;497:624–7.
Yamamoto S, Tsuda H, Takano M, Tamai S, Matsubara O. Loss of ARID1A protein expression occurs as an early event in ovarian clear-cell carcinoma development and frequently coexists with PIK3CA mutations. Mod Pathol. 2012;25:615–24.
Ayhan A, Mao T-L, Seckin T, Wu C-H, Guan B, Ogawa H, et al. Loss of ARID1A expression is an early molecular event in tumor progression from ovarian endometriotic cyst to clear cell and endometrioid carcinoma. Int J Gynecol Cancer. 2012;22:1310–5.
Kadoch C, Crabtree GR. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci Adv. 2015;1:e1500447.
Savas S, Skardasi G. The SWI/SNF complex subunit genes: their functions, variations, and links to risk and survival outcomes in human cancers. Crit Rev Oncol Hematol. 2018;123:114–31.
Tolstorukov MY, Sansam CG, Lu P, Koellhoffer EC, Helming KC, Alver BH, et al. Swi/Snf chromatin remodeling/tumor suppressor complex establishes nucleosome occupancy at target promoters. Proc Natl Acad Sci USA. 2013;110:10165–70.
Williamson CT, Miller R, Pemberton HN, Jones SE, Campbell J, Konde A, et al. ATR inhibitors as a synthetic lethal therapy for tumours deficient in ARID1A. Nat Commun. 2016;7:13837.
Trizzino M, Barbieri E, Petracovici A, Wu S, Welsh SA, Owens TA, et al. The tumor suppressor ARID1A controls global transcription via pausing of RNA polymerase II. Cell Rep. 2018;23:3933–45.
Winston F, Carlson M. Yeast SNF/SWI transcriptional activators and the SPT/SIN chromatin connection. Trends Genet. 1992;8:387–91.
Valente AJ, Maddalena LA, Robb EL, Moradi F, Stuart JA. A simple ImageJ macro tool for analyzing mitochondrial network morphology in mammalian cell culture. Acta Histochem. 2017;119:315–26.
Li W, Köster J, Xu H, Chen C-H, Xiao T, Liu JS, et al. Quality control, modeling, and visualization of CRISPR screens with MAGeCK-VISPR. Genome Biol. 2015;16:281.
Hoppe MM, Jaynes P, Wardyn JD, Upadhyayula SS, Tan TZ, Lie S, et al. Quantitative imaging of RAD51 expression as a marker of platinum resistance in ovarian cancer. EMBO Mol Med. 2021;13:e13366.
Wu JN, Roberts CW. ARID1A mutations in cancer: another epigenetic tumor suppressor? Cancer Disco. 2013;3:35–43.
Ashizawa M, Saito M, Min AKT, Ujiie D, Saito K, Sato T, et al. Prognostic role of ARID1A negative expression in gastric cancer. Sci Rep. 2019;9:6769.
Miller RE, Brough R, Bajrami I, Williamson CT, McDade S, Campbell J, et al. Synthetic lethal targeting of ARID1A-mutant ovarian clear cell tumors with dasatinib. Mol Cancer Ther. 2016;15:1472–84.
Gorlick R, Kolb EA, Keir ST, Maris JM, Reynolds CP, Kang MH, et al. Initial testing (stage 1) of the polo-like kinase inhibitor volasertib (BI 6727), by the Pediatric Preclinical Testing Program. Pediatr Blood Cancer. 2014;61:158–64.
Rudolph D, Steegmaier M, Hoffmann M, Grauert M, Baum A, Quant J, et al. BI 6727, a Polo-like kinase inhibitor with improved pharmacokinetic profile and broad antitumor activity. Clin Cancer Res. 2009;15:3094–102.
Ciccia A, Elledge SJ. The DNA damage response: making it safe to play with knives. Mol Cell. 2010;40:179–204.
van Vugt MA, Bras A, Medema RH. Polo-like kinase-1 controls recovery from a G2 DNA damage-induced arrest in mammalian cells. Mol Cell. 2004;15:799–811.
Wu C, Lyu J, Yang EJ, Liu Y, Zhang B, Shim JS. Targeting AURKA-CDC25C axis to induce synthetic lethality in ARID1A-deficient colorectal cancer cells. Nat Commun. 2018;9:3212.
Doench JG, Fusi N, Sullender M, Hegde M, Vaimberg EW, Donovan KF, et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol. 2016;34:184–91.
Lissanu Deribe Y, Sun Y, Terranova C, Khan F, Martinez-Ledesma J, Gay J, et al. Mutations in the SWI/SNF complex induce a targetable dependence on oxidative phosphorylation in lung cancer. Nat Med. 2018;24:1047–57.
Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, et al. Sequence and organization of the human mitochondrial genome. Nature. 1981;290:457–65.
Ogiwara H, Takahashi K, Sasaki M, Kuroda T, Yoshida H, Watanabe R, et al. Targeting the vulnerability of glutathione metabolism in ARID1A-deficient cancers. Cancer Cell. 2019;35:177–90 e178.
Liu C, Tate T, Batourina E, Truschel ST, Potter S, Adam M, et al. Pparg promotes differentiation and regulates mitochondrial gene expression in bladder epithelial cells. Nat Commun. 2019;10:4589.
Popov LD. Mitochondrial biogenesis: an update. J Cell Mol Med. 2020;24:4892–9.
Zorov DB, Filburn CR, Klotz LO, Zweier JL, Sollott SJ. Reactive oxygen species (ROS)-induced ROS release: a new phenomenon accompanying induction of the mitochondrial permeability transition in cardiac myocytes. J Exp Med. 2000;192:1001–14.
Tillberg PW, Chen F, Piatkevich KD, Zhao Y, Yu CC, English BP, et al. Protein-retention expansion microscopy of cells and tissues labeled using standard fluorescent proteins and antibodies. Nat Biotechnol. 2016;34:987–92.
Koc E, Cimen H, Kumcuoglu B, Abu N, Akpinar G, Haque M, et al. Identification and characterization of CHCHD1, AURKAIP1, and CRIF1 as new members of the mammalian mitochondrial ribosome. Front Physiol. 2013;4:1–15.
Mao TL, Ardighieri L, Ayhan A, Kuo KT, Wu CH, Wang TL, et al. Loss of ARID1A expression correlates with stages of tumor progression in uterine endometrioid carcinoma. Am J Surg Pathol. 2013;37:1342–8.
Ma X, Wang L, Huang, Li Y, Yang D, Li T, et al. Polo-like kinase 1 coordinates biosynthesis during cell cycle progression by directly activating pentose phosphate pathway. Nat Commun. 2017;8:1506.
Li Z, Li J, Bi P, Lu Y, Burcham G, Elzey BD, et al. Plk1 phosphorylation of PTEN causes a tumor-promoting metabolic state. Mol Cell Biol. 2014;34:3642–61.
Gutteridge RE, Singh CK, Ndiaye MA, Ahmad N. Targeted knockdown of polo-like kinase 1 alters metabolic regulation in melanoma. Cancer Lett. 2017;394:13–21.
Matsumoto T, Wang PY, Ma W, Sung HJ, Matoba S, Hwang PM. Polo-like kinases mediate cell survival in mitochondrial dysfunction. Proc Natl Acad Sci USA. 2009;106:14542–6.
Yao CH, Wang R, Wang Y, Kung CP, Weber JD, Patti GJ. Mitochondrial fusion supports increased oxidative phosphorylation during cell proliferation. Elife. 2019;8:1–19.
Liu W, Acin-Perez R, Geghman KD, Manfredi G, Lu B, Li C. Pink1 regulates the oxidative phosphorylation machinery via mitochondrial fission. Proc Natl Acad Sci USA. 2011;108:12920–4.
Nickerson JA, Wu Q, Imbalzano AN. Mammalian SWI/SNF enzymes and the epigenetics of tumor cell metabolic reprogramming. Front Oncol. 2017;7:49.
Richter U, Lahtinen T, Marttinen P, Suomi F, Battersby BJ. Quality control of mitochondrial protein synthesis is required for membrane integrity and cell fitness. J Cell Biol. 2015;211:373–89.
Škrtić M, Sriskanthadevan S, Jhas B, Gebbia M, Wang X, Wang Z, et al. Inhibition of mitochondrial translation as a therapeutic strategy for human acute myeloid leukemia. Cancer Cell. 2011;20:674–88.
Sharon D, Cathelin S, Mirali S, Trani JMD, Yanofsky DJ, Keon KA, et al. Inhibition of mitochondrial translation overcomes venetoclax resistance in AML through activation of the integrated stress response. Sci Transl Med. 2019;11:eaax2863.
Mennuni M, Filograna R, Felser A, Bonekamp NA, Giavalisco P, Lytovchenko O, et al. Metabolic resistance to the inhibition of mitochondrial transcription revealed by CRISPR-Cas9 screen. EMBO Rep. 2021;n/a:e53054.
Croucher PJ, Samuelsz E, Hassaine L, Ross B, Luebbermann M, Ridinger M, et al. Abstract 2019: oxidative phosphorylation (OXPHOS) dependency predicts response to the Polo-like kinase 1 (PLK1) inhibitor onvansertib in a phase 1b/2 of relapsed/refractory acute myeloid leukemia (R/R AML). Cancer Res. 2020;80:2019–19.
Helming KC, Wang X, Roberts CWM. Vulnerabilities of mutant SWI/SNF complexes in cancer. Cancer Cell. 2014;26:309–17.
Wang K, Kan J, Yuen ST, Shi ST, Chu KM, Law S, et al. Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer. Nat Genet. 2011;43:1219–23.
Jones S, Wang TL, Shih Ie M, Mao TL, Nakayama K, Roden R, et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science. 2010;330:228–31.
Bitler BG, Aird KM, Garipov A, Li H, Amatangelo M, Kossenkov AV, et al. Synthetic lethality by targeting EZH2 methyltransferase activity in ARID1A-mutated cancers. Nat Med. 2015;21:231–8.
Fukumoto T, Fatkhutdinov N, Zundell JA, Tcyganov EN, Nacarelli T, Karakashev S, et al. HDAC6 inhibition synergizes with Anti-PD-L1 therapy in ARID1A-inactivated ovarian cancer. Cancer Res. 2019;79:5482–9.
Bertolin G, Bulteau AL, Alves-Guerra MC, Burel A, Lavault MT, Gavard O, et al. Aurora kinase A localises to mitochondria to control organelle dynamics and energy production. Elife. 2018;7:1–28.
Joukov V, De Nicolo A. Aurora-PLK1 cascades as key signaling modules in the regulation of mitosis. Sci Signal. 2018;11:1–25.
Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30:207–10.
Perez-Riverol Y, Bai J, Bandla C, García-Seisdedos D, Hewapathirana S, Kamatchinathan S, et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 2022;50:D543–52.
Wardyn JD, Chan ASY, Jeyasekharan AD. A robust protocol for CRISPR-Cas9 gene editing in human suspension cell lines. Curr Protoc. 2021;1:e286.
Acknowledgements
We would like to thank Dr Chen Gao Bin and Prof. Li Shang at the Duke-NUS CRISPR Core Facility for their assistance with the CRISPR screen.
Funding
ADJ is supported by the Singapore Ministry of Health’s National Medical Research Council Transition Award (NMRC/TA/0052/2016). Work in ADJ’s and the Kappei laboratory is also funded by the Cancer Science Institute of Singapore, National University of Singapore, through the National Research Foundation Singapore and the Singapore Ministry of Education under its Research Centers of Excellence initiative. Part of this work was funded through a collaborative grant between ADJ and DSPT from the Singapore Ministry of Health’s National Medical Research Council (NMRC/CIRG/1400/2014). This research was also funded by the Singapore Ministry of Education under its Singapore Ministry of Education Academic Research Fund Tier 2 (MOE2018-T2-2-179) to Karen Crasta.
Author information
Authors and Affiliations
Contributions
Conceptualization: USS, PJ, ADJ. Data curation: USS, OA, AS, HY. Formal analysis: USS, SL, MMH, SJ, VK. Funding acquisition: ADJ. Investigation: USS, TSCN, AA, GKR, JDW, MYL, PCP, BWQT, LHu, RJ, KS, MB, YP, SL, LC, PMH. Methodology: USS, DK, SC, WLT, CF, ADJ. Project administration: ADJ. Resources: LHKL, ML, SP, KC, PT, DK, YWP, CF, DSPT, WLT, LHo, SC, ADJ. Software – OA, HY. Supervision: ADJ, PJ. Validation: USS, TSCN, AA, PJ, LHKL, SP, KC, PT, DK, YWP, CF, DSPT, WLT, AS, HY, OA. Visualization: USS, MMH, TSCN, PJ, ADJ. Writing – original draft – USS, PJ, ADJ. Writing – review and editing - LHKL, SP, KC, PT, DK, YWP, CF, DSPT, WLT, PJ, ADJ.
Corresponding author
Ethics declarations
Competing interests
ADJ: honoraria from Turbine Ltd, AstraZeneca, Antengene, Janssen and MSD, travel funding from Perkin Elmer, and research funding from Janssen and AstraZeneca. DSPT: honoraria from AstraZeneca, Roche, Bayer, MSD, Merck Serono, Tessa Therapeutics, Novartis, and Genmab and research funding from AstraZeneca, Bayer and Karyopharm. The other co-authors have no conflicts of interest to declare.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Srinivas, U.S., Tay, N.S.C., Jaynes, P. et al. PLK1 inhibition selectively induces apoptosis in ARID1A deficient cells through uncoupling of oxygen consumption from ATP production. Oncogene 41, 1986–2002 (2022). https://doi.org/10.1038/s41388-022-02219-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02219-8
This article is cited by
-
Treating ARID1A mutated cancers by harnessing synthetic lethality and DNA damage response
Journal of Biomedical Science (2022)