Abstract
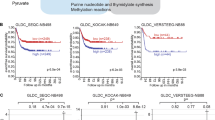
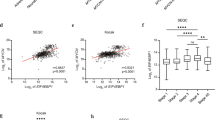
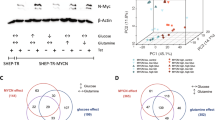
Cancer cells exhibit dysregulation of critical genes including those involved in lipid biosynthesis, with subsequent defects in metabolism. Here, we show that ELOngation of Very Long chain fatty acids protein 4 (ELOVL4), a rate-limiting enzyme in the biosynthesis of very-long polyunsaturated fatty acids (n-3, ≥28 C), is expressed and transcriptionally repressed by the oncogene MYCN in neuroblastoma cells. In keeping, ELOVL4 positively regulates neuronal differentiation and lipids droplets accumulation in neuroblastoma cells. At the molecular level we found that MYCN binds to the promoter of ELOVL4 in close proximity to the histone deacetylases HDAC1, HDAC2, and the transcription factor Sp1 that can cooperate in the repression of ELOVL4 expression. Accordingly, in vitro differentiation results in an increase of fatty acid with 34 carbons with 6 double bonds (FA34:6); and when MYCN is silenced, FA34:6 metabolite is increased compared with the scrambled. In addition, analysis of large neuroblastoma datasets revealed that ELOVL4 expression is highly expressed in localized clinical stages 1 and 2, and low in high-risk stages 3 and 4. More importantly, high expression of ELOVL4 stratifies a subsets of neuroblastoma patients with good prognosis. Indeed, ELOVL4 expression is a marker of better overall clinical survival also in MYCN not amplified patients and in those with neuroblastoma-associated mutations. In summary, our findings indicate that MYCN, by repressing the expression of ELOVL4 and lipid metabolism, contributes to the progression of neuroblastoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
20 January 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41388-021-02164-y
References
Snaebjornsson MT, Janaki-Raman S, Schulze A. Greasing the wheels of the cancer machine: the role of lipid metabolism in cancer. Cell Metab. 2020;31:62–76.
Buck MD, O’Sullivan D, Klein Geltink RI, Curtis JD, Chang CH, Sanin DE, et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell. 2016;166:63–76.
Sulciner ML, Gartung A, Gilligan MM, Serhan CN, Panigrahy D. Targeting lipid mediators in cancer biology. Cancer Metastasis Rev. 2018;37:557–72.
DeBose-Boyd RA, Ye J. SREBPs in lipid metabolism, insulin signaling, and beyond. Trends Biochem Sci. 2018;43:358–68.
Rohrig F, Schulze A. The multifaceted roles of fatty acid synthesis in cancer. Nat Rev Cancer. 2016;16:732–49.
Snaebjornsson MT, Schulze A. Tumours use a metabolic twist to make lipids. Nature. 2019;566:333–4.
Riscal R, Skuli N, Simon MC. Even cancer cells watch their cholesterol! Mol Cell. 2019;76:220–31.
Dickerman BA, Garcia-Albeniz X, Logan RW, Denaxas S, Hernan MA. Avoidable flaws in observational analyses: an application to statins and cancer. Nat Med. 2019;25:1601–6.
McGregor GH, Campbell AD, Fey SK, Tumanov S, Sumpton D, Blanco GR, et al. Targeting the metabolic response to statin-mediated oxidative stress produces a synergistic antitumor response. Cancer Res. 2020;80:175–88.
Yao L, Han C, Song K, Zhang J, Lim K, Wu T. Omega-3 polyunsaturated fatty acids upregulate 15-PGDH expression in cholangiocarcinoma cells by inhibiting miR-26a/b expression. Cancer Res. 2015;75:1388–98.
Bhattacharjee S, Jun B, Belayev L, Heap J, Kautzmann MA, Obenaus A, et al. Elovanoids are a novel class of homeostatic lipid mediators that protect neural cell integrity upon injury. Sci Adv. 2017;3:e1700735.
Jun B, Mukherjee PK, Asatryan A, Kautzmann MA, Heap J, Gordon WC, et al. Elovanoids are novel cell-specific lipid mediators necessary for neuroprotective signaling for photoreceptor cell integrity. Sci Rep. 2017;7:5279.
Agbaga MP, Brush RS, Mandal MN, Henry K, Elliott MH, Anderson RE. Role of Stargardt-3 macular dystrophy protein (ELOVL4) in the biosynthesis of very long chain fatty acids. Proc Natl Acad Sci USA. 2008;105:12843–8.
Zhang K, Kniazeva M, Han M, Li W, Yu Z, Yang Z, et al. A 5-bp deletion in ELOVL4 is associated with two related forms of autosomal dominant macular dystrophy. Nat Genet. 2001;27:89–93.
Bazan NG. Docosanoids and elovanoids from omega-3 fatty acids are pro-homeostatic modulators of inflammatory responses, cell damage and neuroprotection. Mol Asp Med. 2018;64:18–33.
Horton JD, Shah NA, Warrington JA, Anderson NN, Park SW, Brown MS, et al. Combined analysis of oligonucleotide microarray data from transgenic and knockout mice identifies direct SREBP target genes. Proc Natl Acad Sci USA. 2003;100:12027–32.
Nohturfft A, Zhang SC. Coordination of lipid metabolism in membrane biogenesis. Annu Rev Cell Dev Biol. 2009;25:539–66.
Mizutani Y, Sun H, Ohno Y, Sassa T, Wakashima T, Obara M, et al. Cooperative synthesis of ultra long-chain fatty acid and ceramide during keratinocyte differentiation. PLoS One. 2013;8:e67317.
Thiam AR, Beller M. The why, when and how of lipid droplet diversity. J Cell Sci. 2017;130:315–24.
Teo W, Caprariello AV, Morgan ML, Luchicchi A, Schenk GJ, Joseph JT, et al. Nile Red fluorescence spectroscopy reports early physicochemical changes in myelin with high sensitivity. Proc Natl Acad Sci USA. 2021;118:e2016897118.
Agostini M, Romeo F, Inoue S, Niklison-Chirou MV, Elia AJ, Dinsdale D, et al. Metabolic reprogramming during neuronal differentiation. Cell Death Differ. 2016;23:1502–14.
Petroni A, Blasevich M, La Spada P, Papini N, Galli C. Arachidonic acid synthesis and lipid metabolism in retinoic acid-differentiated neuroblastoma cells. J Lipid Mediat Cell Signal. 1996;14:39–44.
Maris JM, Hogarty MD, Bagatell R, Cohn SL. Neuroblastoma. Lancet. 2007;369:2106–20.
Piacentini M, Annicchiarico-Petruzzelli M, Oliverio S, Piredda L, Biedler JL, Melino E. Phenotype-specific “tissue” transglutaminase regulation in human neuroblastoma cells in response to retinoic acid: correlation with cell death by apoptosis. Int J Cancer. 1992;52:271–8.
Ross RA, Spengler BA, Biedler JL. Coordinate morphological and biochemical interconversion of human neuroblastoma cells. J Natl Cancer Inst. 1983;71:741–7.
Lutz W, Stohr M, Schurmann J, Wenzel A, Lohr A, Schwab M. Conditional expression of N-myc in human neuroblastoma cells increases expression of alpha-prothymosin and ornithine decarboxylase and accelerates progression into S-phase early after mitogenic stimulation of quiescent cells. Oncogene. 1996;13:803–12.
Iraci N, Diolaiti D, Papa A, Porro A, Valli E, Gherardi S, et al. A SP1/MIZ1/MYCN repression complex recruits HDAC1 at the TRKA and p75NTR promoters and affects neuroblastoma malignancy by inhibiting the cell response to NGF. Cancer Res. 2011;71:404–12.
Lin YC, Lin JH, Chou CW, Chang YF, Yeh SH, Chen CC. Statins increase p21 through inhibition of histone deacetylase activity and release of promoter-associated HDAC1/2. Cancer Res. 2008;68:2375–83.
Depuydt P, Koster J, Boeva V, Hocking TD, Speleman F, Schleiermacher G, et al. Meta-mining of copy number profiles of high-risk neuroblastoma tumors. Sci Data. 2018;5:180240.
Matthay KK, Maris JM, Schleiermacher G, Nakagawara A, Mackall CL, Diller L, et al. Neuroblastoma. Nat Rev Dis Prim. 2016;2:16078.
Thiele CJ. Biology of pediatric peripheral neuroectodermal tumors. Cancer Metastasis Rev. 1991;10:311–9.
Pieraccioli M, Nicolai S, Pitolli C, Agostini M, Antonov A, Malewicz M, et al. ZNF281 inhibits neuronal differentiation and is a prognostic marker for neuroblastoma. Proc Natl Acad Sci USA. 2018;115:7356–61.
Zirath H, Frenzel A, Oliynyk G, Segerstrom L, Westermark UK, Larsson K, et al. MYC inhibition induces metabolic changes leading to accumulation of lipid droplets in tumor cells. Proc Natl Acad Sci USA. 2013;110:10258–63.
Ruiz-Perez MV, Sainero-Alcolado L, Oliynyk G, Matuschek I, Balboni N, Ubhayasekera S, et al. Inhibition of fatty acid synthesis induces differentiation and reduces tumor burden in childhood neuroblastoma. iScience. 2021;24:102128.
Amelio I, Bertolo R, Bove P, Candi E, Chiocchi M, Cipriani C, et al. Cancer predictive studies. Biol Direct. 2020;15:18.
Han Y, Ye X, Wang C, Liu Y, Zhang S, Feng W, et al. Integration of molecular features with clinical information for predicting outcomes for neuroblastoma patients. Biol Direct. 2019;14:16.
Han YT, Ye XF, Cheng J, Zhang SY, Feng WX, Han Z, et al. Integrative analysis based on survival associated co-expression gene modules for predicting Neuroblastoma patients’ survival time. Biol Direct. 2019;14:4. https://doi.org/10.1186/s13062-018-0229-2.
Sumsion GR, Bradshaw MS, Beales JT, Ford E, Caryotakis GRG, Garrett DJ, et al. Diverse approaches to predicting drug-induced liver injury using gene-expression profiles. Biol Direct. 2020;15:1. https://doi.org/10.1186/s13062-019-0257-6.
Mihaylov I, Kandula M, Krachunov M, Vassilev D. A novel framework for horizontal and vertical data integration in cancer studies with application to survival time prediction models. Biol Direct. 2019;14:22. https://doi.org/10.1186/s13062-019-0249-6.
Carroll PA, Diolaiti D, McFerrin L, Gu H, Djukovic D, Du J, et al. Deregulated Myc requires MondoA/Mlx for metabolic reprogramming and tumorigenesis. Cancer Cell. 2015;27:271–85.
Amelio I, Bertolo R, Bove P, Buonomo OC, Candi E, Chiocchi M, et al. Liquid biopsies and cancer omics. Cell Death Discov. 2020;6:131.
Harris ZN, Dhungel E, Mosior M, Ahn TH. Massive metagenomic data analysis using abundance-based machine learning. Biol Direct. 2019;14:12.
Liu L, Wang G, Wang L, Yu C, Li M, Song S, et al. Computational identification and characterization of glioma candidate biomarkers through multi-omics integrative profiling. Biol Direct. 2020;15:10.
Chen JC, Tyler AD. Systematic evaluation of supervised machine learning for sample origin prediction using metagenomic sequencing data. Biol Direct. 2020;15:29.
Chierici M, Francescatto M, Bussola N, Jurman G, Furlanello C. Predictability of drug-induced liver injury by machine learning. Biol Direct. 2020;15:3.
Bellomaria A, Barbato G, Melino G, Paci M, Melino S. Recognition mechanism of p63 by the E3 ligase Itch Novel strategy in the study and inhibition of this interaction. Cell Cycle. 2012;11:3638–48.
Ciocci M, Iorio E, Carotenuto F, Khashoggi HA, Nanni F, Melino S. H2S-releasing nanoemulsions: a new formulation to inhibit tumor cells proliferation and improve tissue repair. Oncotarget. 2016;7:84338–58.
Nicolai S, Pieraccioli M, Smirnov A, Pitolli C, Anemona L, Mauriello A, et al. ZNF281/Zfp281 is a target of miR-1 and counteracts muscle differentiation. Mol Oncol. 2020;14:294–308.
Lamastra FR, De Angelis R, Antonucci A, Salvatori D, Prosposito P, Casalboni M, et al. Polymer composite random lasers based on diatom frustules as scatterers. Rsc Adv. 2014;4:61809–16.
Pallucca R, Visconti S, Camoni L, Cesareni G, Melino S, Panni S, et al. Specificity of epsilon and Non-epsilon Isoforms of Arabidopsis 14-3-3 Proteins Towards the H+-ATPase and Other Targets. Plos ONE. 2014;9:e90764. https://doi.org/10.1371/journal.pone.0090764.
Acknowledgements
We would like to thank Dott. Butera Alessio for technical assistance and Professor Manfred Schwab for SHEP Tet21N cell line.
Funding
This research was funded by Associazione Italiana per la Ricerca contro il Cancro (AIRC) to GM (IG#20473; 2018–2022), Ministry of Health and MAECI Italy-China Science and Technology Cooperation (#PGR00961) to GM. Work has been also supported by Regione Lazio through LazioInnova Progetto Gruppo di Ricerca n 85-2017-14986.
Author information
Authors and Affiliations
Contributions
FR, MA, NGB, GR, and GM designed the experiments. MA, GR, and GM supervised the project. FR and JC performed the biochemical experiments. BJ and JC performed LC-MS/MS. FR performed the bioinformatic analysis. FR, MA, GR, and GM analyzed the data. MA, GR, and GM wrote the manuscript and all the authors commented and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Rugolo, F., Bazan, N.G., Calandria, J. et al. The expression of ELOVL4, repressed by MYCN, defines neuroblastoma patients with good outcome. Oncogene 40, 5741–5751 (2021). https://doi.org/10.1038/s41388-021-01959-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01959-3
This article is cited by
-
Are lipid droplets the picnic basket of brain tumours?
Cell Death Discovery (2024)
-
Identification of Key lncRNAs Associated with Immune Infiltration and Prognosis in Gastric Cancer
Biochemical Genetics (2024)
-
A comprehensive review of the family of very-long-chain fatty acid elongases: structure, function, and implications in physiology and pathology
European Journal of Medical Research (2023)
-
EIF4EBP1 is transcriptionally upregulated by MYCN and associates with poor prognosis in neuroblastoma
Cell Death Discovery (2022)
-
Targeting lipid metabolism in cancer: neuroblastoma
Cancer and Metastasis Reviews (2022)