Abstract
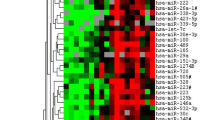
MicroRNAs (miRs) are small non-coding RNAs that can have large impacts on oncogenic pathways. Possible functions of dysregulated miRs have not been studied in neurofibromatosis type 1 (NF1) plexiform neurofibromas (PNFs). In PNFs, Schwann cells (SCs) have biallelic NF1 mutations necessary for tumorigenesis. We analyzed a miR microarray comparing with normal and PNF SCs and identified differences in miR expression, and we validated in mouse PNFs versus normal mouse SCs by qRT-PCR. Among these, miR-155 was a top overexpressed miR, and its expression was regulated by RAS/MAPK signaling. Overexpression of miR-155 increased mature Nf1−/− mouse SC proliferation. In SC precursors, which model tumor-initiating cells, pharmacological and genetic inhibition of miR-155 decreased PNF-derived sphere numbers in vitro, and we identified Maf as a miR-155 target. In vivo, global deletion of miR-155 significantly decreased tumor number and volume, increasing mouse survival. Fluorescent nanoparticles entered PNFs, suggesting that an anti-miR might have therapeutic potential. However, treatment of established PNFs using anti-miR-155 peptide nucleic acid-loaded nanoparticles marginally decreased tumor numbers and did not reduce tumor growth. These results suggest that miR-155 plays a functional role in PNF growth and/or SC proliferation, and that targeting neurofibroma miRs is feasible, and might provide novel therapeutic opportunities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Boyd KP, Korf BR, Theos A. Neurofibromatosis type 1. J Am Acad Dermatol. 2009;61:1–14.
Varan A, Sen H, Aydin B, Yalcin B, Kutluk T, Akyuz C. Neurofibromatosis type 1 and malignancy in childhood. Clin Genet. 2016;89:341–5.
Prada CE, Jousma E, Rizvi TA, Wu J, Dunn RS, Mayes DA, et al. Neurofibroma-associated macrophages play roles in tumor growth and response to pharmacological inhibition. Acta Neuropathol. 2013;125:159–68.
Rice FL, Houk G, Wymer JP, Gosline SJC, Guinney J, Wu J, et al. The evolution and multi-molecular properties of NF1 cutaneous neurofibromas originating from C-fiber sensory endings and terminal Schwann cells at normal sites of sensory terminations in the skin. PloS ONE. 2019;14:e0216527.
Zhu Y, Ghosh P, Charnay P, Burns DK, Parada LF. Neurofibromas in NF1: schwann cell origin and role of tumor environment. Science. 2002;296:920–2.
Wu J, Williams JP, Rizvi TA, Kordich JJ, Witte D, Meijer D, et al. Plexiform and dermal neurofibromas and pigmentation are caused by Nf1 loss in desert hedgehog-expressing cells. Cancer Cell. 2008;13:105–16.
Chen Z, Mo J, Brosseau J-P, Shipman T, Wang Y, Liao C-P, et al. Spatiotemporal loss of NF1 in schwann cell lineage leads to different types of cutaneous neurofibroma susceptible to modification by the hippo pathway. Cancer Discov. 2019;9:114–29.
Radomska KJ, Coulpier F, Gresset A, Schmitt A, Debbiche A, Lemoine S, et al. Cellular origin, tumor progression, and pathogenic mechanisms of cutaneous neurofibromas revealed by mice with Nf1 knockout in boundary cap cells. Cancer Discov. 2019;9:130–47.
Zheng H, Chang L, Patel N, Yang J, Lowe L, Burns DK, et al. Induction of abnormal proliferation by nonmyelinating schwann cells triggers neurofibroma formation. Cancer Cell. 2008;13:117–28.
McCormick F. Ras signaling and NF1. Curr Opin Genet Dev. 1995;5:51–5.
Le LQ, Parada LF. Tumor microenvironment and neurofibromatosis type I: connecting the GAPs. Oncogene. 2007;26:4609–16.
Ratner N, Miller SJ. A RASopathy gene commonly mutated in cancer: the neurofibromatosis type 1 tumour suppressor. Nat Rev Cancer. 2015;15:290–301.
Kraniak JM, Sun D, Mattingly RR, Reiners JJ Jr, Tainsky MA. The role of neurofibromin in N-Ras mediated AP-1 regulation in malignant peripheral nerve sheath tumors. Mol Cell Biochem. 2010;344:267–76.
Wu J, Dombi E, Jousma E, Scott Dunn R, Lindquist D, Schnell BM, et al. Preclincial testing of sorafenib and RAD001 in the Nf1(flox/flox);DhhCre mouse model of plexiform neurofibroma using magnetic resonance imaging. Pediatr Blood Cancer. 2012;58:173–80.
Jessen WJ, Miller SJ, Jousma E, Wu J, Rizvi TA, Brundage ME, et al. MEK inhibition exhibits efficacy in human and mouse neurofibromatosis tumors. J Clin Invest. 2013;123:340–7.
Dombi E, Baldwin A, Marcus LJ, Fisher MJ, Weiss B, Kim A, et al. Activity of selumetinib in neurofibromatosis type 1-related plexiform neurofibromas. N. Engl J Med. 2016;375:2550–60.
Gross AM, Wolters PL, Dombi E, Baldwin A, Whitcomb P, Fisher MJ, et al. Selumetinib in children with inoperable plexiform neurofibromas. N. Engl J Med. 2020;382:1430–42.
Jousma E, Rizvi TA, Wu J, Janhofer D, Dombi E, Dunn RS, et al. Preclinical assessments of the MEK inhibitor PD-0325901 in a mouse model of neurofibromatosis type 1. Pediatr Blood Cancer. 2015;62:1709–16.
Baulcombe D. DNA events. An RNA microcosm. Science 2002;297:2002–3.
Gordon JE, Wong JJ, Rasko JE. MicroRNAs in myeloid malignancies. Br J Haematol. 2013;162:162–76.
Berdasco M, Esteller M. Aberrant epigenetic landscape in cancer: how cellular identity goes awry. Dev Cell. 2010;19:698–711.
Vasilatou D, Papageorgiou S, Pappa V, Papageorgiou E, Dervenoulas J. The role of microRNAs in normal and malignant hematopoiesis. Eur J Haematol. 2010;84:1–16.
O’Connell RM, Rao DS, Chaudhuri AA, Baltimore D. Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol. 2010;10:111–22.
Babar IA, Cheng CJ, Booth CJ, Liang X, Weidhaas JB, Saltzman WM, et al. Nanoparticle-based therapy in an in vivo microRNA-155 (miR-155)-dependent mouse model of lymphoma. Proc Natl Acad Sci USA. 2012;109:E1695–704.
Jiang S, Zhang LF, Zhang HW, Hu S, Lu MH, Liang S, et al. A novel miR-155/miR-143 cascade controls glycolysis by regulating hexokinase 2 in breast cancer cells. EMBO J. 2012;31:1985–98.
Rodriguez A, Vigorito E, Clare S, Warren MV, Couttet P, Soond DR, et al. Requirement of bic/microRNA-155 for normal immune function. Science 2007;316:608–11.
Thai TH, Calado DP, Casola S, Ansel KM, Xiao C, Xue Y, et al. Regulation of the germinal center response by microRNA-155. Science 2007;316:604–8.
Fabani MM, Abreu-Goodger C, Williams D, Lyons PA, Torres AG, Smith KG, et al. Efficient inhibition of miR-155 function in vivo by peptide nucleic acids. Nucleic Acids Res. 2010;38:4466–75.
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M, et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 2005;438:685–9.
Ma L, Reinhardt F, Pan E, Soutschek J, Bhat B, Marcusson EG, et al. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat Biotechnol. 2010;28:341–7.
Zhang Y, Roccaro AM, Rombaoa C, Flores L, Obad S, Fernandes SM, et al. LNA-mediated anti-miR-155 silencing in low-grade B-cell lymphomas. Blood 2012;120:1678–86.
Subramanian S, Thayanithy V, West RB, Lee CH, Beck AH, Zhu S, et al. Genome-wide transcriptome analyses reveal p53 inactivation mediated loss of miR-34a expression in malignant peripheral nerve sheath tumours. J Pathol. 2010;220:58–70.
Chai G, Liu N, Ma J, Li H, Oblinger JL, Prahalad AK, et al. MicroRNA-10b regulates tumorigenesis in neurofibromatosis type 1. Cancer Sci. 2010;101:1997–2004.
Presneau N, Eskandarpour M, Shemais T, Henderson S, Halai D, Tirabosco R, et al. MicroRNA profiling of peripheral nerve sheath tumours identifies miR-29c as a tumour suppressor gene involved in tumour progression. Br J Cancer. 2013;108:964–72.
Gong M, Ma J, Li M, Zhou M, Hock JM, Yu X. MicroRNA-204 critically regulates carcinogenesis in malignant peripheral nerve sheath tumors. Neuro Oncol. 2012;14:1007–17.
Miller SJ, Rangwala F, Williams J, Ackerman P, Kong S, Jegga AG, et al. Large-scale molecular comparison of human schwann cells to malignant peripheral nerve sheath tumor cell lines and tissues. Cancer Res. 2006;66:2584–91.
Higgs G, Slack F. The multiple roles of microRNA-155 in oncogenesis. J Clin Bioinformatics. 2013;3:17.
Martin EC, Krebs AE, Burks HE, Elliott S, Baddoo M, Collins-Burow BM, et al. miR-155 induced transcriptome changes in the MCF-7 breast cancer cell line leads to enhanced mitogen activated protein kinase signaling. Genes Cancer. 2014;5:353–64.
Trotta R, Chen L, Costinean S, Josyula S, Mundy-Bosse BL, Ciarlariello D, et al. Overexpression of miR-155 causes expansion, arrest in terminal differentiation and functional activation of mouse natural killer cells. Blood 2013;121:3126–34.
Miller SJ, Jessen WJ, Mehta T, Hardiman A, Sites E, Kaiser S, et al. Integrative genomic analyses of neurofibromatosis tumours identify SOX9 as a biomarker and survival gene. EMBO Mol Med. 2009;1:236–48.
Hall A, Choi K, Liu W, Rose J, Zhao C, Yu Y, et al. RUNX represses Pmp22 to drive neurofibromagenesis. Sci Adv. 2019;5:eaau8389.
Kluiver J, Poppema S, de Jong D, Blokzijl T, Harms G, Jacobs S, et al. BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J Pathol. 2005;207:243–9.
Xiao C, Calado DP, Galler G, Thai T-H, Patterson HC, Wang J, et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 2007;131:146–59.
Zhou Y, Wang X, Liu Z, Huang X, Li X, Cheng K, et al. Prognostic role of microRNA-155 expression in gliomas: a meta-analysis. Clin Neurol Neurosurg. 2019;176:103–9.
Sherman LS, Ratner N. Immunocytochemical assay for Ras activity. Methods Enzymol. 2001;333:348–55.
Yin Q, Wang X, McBride J, Fewell C, Flemington E. B-cell receptor activation induces BIC/miR-155 expression through a conserved AP-1 element. J Biol Chem. 2008;283:2654–62.
Brundage ME, Tandon P, Eaves DW, Williams JP, Miller SJ, Hennigan RH, et al. MAF mediates crosstalk between Ras-MAPK and mTOR signaling in NF1. Oncogene 2014;33:5626–36.
Zhang C, Guo ZM. Multiple functions of Maf in the regulation of cellular development and differentiation. Diabetes Metab Res Rev. 2015;31:773–8.
Liu M, Tong Z, Ding C, Luo F, Wu S, Wu C, et al. Transcription factor c-Maf is a checkpoint that programs macrophages in lung cancer. J Clin Invest. 2020;130:2081–96.
Su W, Hopkins S, Nesser NK, Sopher B, Silvestroni A, Ammanuel S, et al. The p53 transcription factor modulates microglia behavior through microRNA-dependent regulation of c-Maf. J Immunol. 2014;192:358–66.
Kim M, Wende H, Walcher J, Kuhnemund J, Cheret C, Kempa S, et al. Maf links Neuregulin1 signaling to cholesterol synthesis in myelinating Schwann cells. Genes Dev. 2018;32:645–57.
Ganguly S, Chaubey B, Tripathi S, Upadhyay A, Neti PV, Howell RW, et al. Pharmacokinetic analysis of polyamide nucleic-acid-cell penetrating peptide conjugates targeted against HIV-1 transactivation response element. Oligonucleotides 2008;18:277–86.
O’Connell RM, Taganov KD, Boldin MP, Cheng G, Baltimore D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc Natl Acad Sci USA. 2007;104:1604–9.
Banerjee A, Schambach F, DeJong CS, Hammond SM, Reiner SL. Micro-RNA-155 inhibits IFN-gamma signaling in CD4+ T cells. Eur J Immunol. 2010;40:225–31.
Solomon J, Warren K, Dombi E, Patronas N, Widemann B. Automated detection and volume measurement of plexiform neurofibromas in neurofibromatosis 1 using magnetic resonance imaging. Comput Med Imaging Graph. 2004;28:257–65.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS. Human MicroRNA targets. PLoS Biol. 2004;2:e363.
Wu J, Keng VW, Patmore DM, Kendall JJ, Patel AV, Jousma E, et al. Insertional mutagenesis identifies a STAT3/Arid1b/beta-catenin pathway driving neurofibroma initiation. Cell Rep. 2016;14:1979–90.
Woodrow KA, Cu Y, Booth CJ, Saucier-Sawyer JK, Wood MJ, Saltzman WM. Intravaginal gene silencing using biodegradable polymer nanoparticles densely loaded with small-interfering RNA. Nat Mater. 2009;8:526–33.
Cheng CJ, Saltzman WM. Enhanced siRNA delivery into cells by exploiting the synergy between targeting ligands and cell-penetrating peptides. Biomaterials. 2011;32:6194–203.
Acknowledgements
We thank Drs. Mark Saltzman and Christopher J. Cheng (Yale School of Engineering and Applied Science) for providing fluorescence-conjugated nanoparticles to fulfill part of the in vivo experiments. This work was supported by NIH R01 NS097233 to J.W. NIH R0 NS28840 and R37 NS096356 to N.R.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict ofinterest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Na, Y., Hall, A., Choi, K. et al. MicroRNA-155 contributes to plexiform neurofibroma growth downstream of MEK. Oncogene 40, 951–963 (2021). https://doi.org/10.1038/s41388-020-01581-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01581-9
This article is cited by
-
The role and regulation of Maf proteins in cancer
Biomarker Research (2023)