Abstract
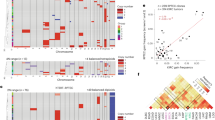
Oncogenesis is considered to result from chromosomal instability, in addition to oncogene and tumor-suppressor alterations. Intermediate to aneuploidy and chromosomal instability, genome doubling is a frequent event in tumor development but the mechanisms driving tetraploidization and its impact remain unexplored. Cell fusion, one of the pathways to tetraploidy, is a physiological process involved in mesenchymal cell differentiation. Besides simple genome doubling, cell fusion results in the merging of two different genomes that can be destabilized upon proliferation. By testing whether cell fusion is involved in mesenchymal oncogenesis, we provide evidence that it induces genomic instability and mediates tumor initiation. After a latency period, the tumor emerges with the cells most suited for its development. Furthermore, hybrid tumor genomes were stabilized after this selection process and were very close to those of human pleomorphic mesenchymal tumors. Thus genome restructuring triggered by cell fusion may account for the chromosomal instability involved in oncogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hahn WC, Counter CM, Lundberg AS, Beijersbergen RL, Brooks MW, Weinberg RA. Creation of human tumour cells with defined genetic elements. Nature. 1999;400:464–8.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Zack TI, Schumacher SE, Carter SL, Cherniack AD, Saksena G, Tabak B, et al. Pan-cancer patterns of somatic copy-number alteration. Nat Genet. 2013;45:1134.
Davoli T, Xu AW, Mengwasser KE, Sack LM, Yoon JC, Park PJ, et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell. 2013;155:948–62.
Davoli T, de Lange T. The causes and consequences of polyploidy in normal development and cancer. Annu Rev Cell Dev Biol. 2011;27:585–610.
Bielski CM, Zehir A, Penson AV, Donoghue MTA, Chatila W, Armenia J, et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat Genet. 2018;50:1189–95.
Margolis RL. Tetraploidy and tumor development. Cancer Cell. 2005;8:353–4.
Ganem NJ, Storchova Z, Pellman D. Tetraploidy, aneuploidy and cancer. Curr Opin Genet Dev. 2007;17:157–62.
Kwon M, Godinho SA, Chandhok NS, Ganem NJ, Azioune A, Thery M, et al. Mechanisms to suppress multipolar divisions in cancer cells with extra centrosomes. Genes Dev. 2008;22:2189–203.
Vitale I, Galluzzi L, Castedo M, Kroemer G. Mitotic catastrophe: a mechanism for avoiding genomic instability. Nat Rev Mol Cell Biol. 2011;12:385–92.
Vitale I, Galluzzi L, Senovilla L, Criollo A, Jemaà M, Castedo M, et al. Illicit survival of cancer cells during polyploidization and depolyploidization. Cell Death Differ. 2011;18:1403–13.
Fujiwara T, Bandi M, Nitta M, Ivanova EV, Bronson RT, Pellman D. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature. 2005;437:1043–7.
Santaguida S, Amon A. Short- and long-term effects of chromosome mis-segregation and aneuploidy. Nat Rev Mol Cell Biol. 2015;16:473–85.
Davoli T, Uno H, Wooten EC, Elledge SJ. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science. 2017;355:eaaf8399.
Turajlic S, Xu H, Litchfield K, Rowan A, Chambers T, Lopez JI, et al. Tracking cancer evolution reveals constrained routes to metastases: TRACERx renal. Cell. 2018;173:581–594.e12.
Bakhoum SF, Ngo B, Laughney AM, Cavallo J-A, Murphy CJ, Ly P, et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature. 2018;553:467–72.
Davoli T, Denchi EL, de Lange T. Persistent telomere damage induces bypass of mitosis and tetraploidy. Cell. 2010;141:81–93.
Targa A, Rancati G. Cancer: a CINful evolution. Curr Opin Cell Biol. 2018;52:136–44.
Duelli D, Lazebnik Y. Cell fusion: a hidden enemy? Cancer Cell. 2003;3:445–8.
Duelli D, Lazebnik Y. Cell-to-cell fusion as a link between viruses and cancer. Nat Rev Cancer. 2007;7:968–76.
Duelli DM, Padilla-Nash HM, Berman D, Murphy KM, Ried T, Lazebnik Y. A virus causes cancer by inducing massive chromosomal instability through cell fusion. Curr Biol. 2007;17:431–7.
Ferrari G, Cusella-De Angelis G, Coletta M, Paolucci E, Stornaiuolo A, Cossu G, et al. Muscle regeneration by bone marrow-derived myogenic progenitors. Science. 1998;279:1528–30.
Terada N, Hamazaki T, Oka M, Hoki M, Mastalerz DM, Nakano Y, et al. Bone marrow cells adopt the phenotype of other cells by spontaneous cell fusion. Nature. 2002;416:542–5.
Weimann JM, Johansson CB, Trejo A, Blau HM. Stable reprogrammed heterokaryons form spontaneously in Purkinje neurons after bone marrow transplant. Nat Cell Biol. 2003;5:959–66.
Rizvi AZ, Swain JR, Davies PS, Bailey AS, Decker AD, Willenbring H, et al. Bone marrow-derived cells fuse with normal and transformed intestinal stem cells. Proc Natl Acad Sci USA. 2006;103:6321–5.
Alvarez-Dolado M, Pardal R, Garcia-Verdugo JM, Fike JR, Lee HO, Pfeffer K, et al. Fusion of bone-marrow-derived cells with Purkinje neurons, cardiomyocytes and hepatocytes. Nature. 2003;425:968–73.
Camargo FD, Green R, Capetanaki Y, Jackson KA, Goodell MA, Capetenaki Y. Single hematopoietic stem cells generate skeletal muscle through myeloid intermediates. Nat Med. 2003;9:1520–7.
Johansson CB, Youssef S, Koleckar K, Holbrook C, Doyonnas R, Corbel SY, et al. Extensive fusion of haematopoietic cells with Purkinje neurons in response to chronic inflammation. Nat Cell Biol. 2008;10:575–83.
Nygren JM, Liuba K, Breitbach M, Stott S, Thorén L, Roell W, et al. Myeloid and lymphoid contribution to non-haematopoietic lineages through irradiation-induced heterotypic cell fusion. Nat Cell Biol. 2008;10:584–92.
Zhou X, Merchak K, Lee W, Grande JP, Cascalho M, Platt JL. Cell Fusion Connects Oncogenesis with Tumor Evolution. Am J Pathol. 2015;185:2049–60.
Aichel O. “Über Zellverschmelzung mit qualitativ abnormer chromosomenverteilung als ursache der geschwulstbildung” [About cell fusion with qualitatively abnormal. chromosome distribution as cause for tumor formation.]. In: Vorträge und aufsätze über entvickelungsmechanik der organismen. Leipzig: Wilhelm Engelmann; 1911. p. 92–111.
Pawelek JM, Chakraborty AK. Fusion of tumour cells with bone marrow-derived cells: a unifying explanation for metastasis. Nat Rev Cancer. 2008;8:377–86.
Pawelek JM. Tumour-cell fusion as a source of myeloid traits in cancer. Lancet Oncol. 2005;6:988–93.
Powell AE, Anderson EC, Davies PS, Silk AD, Pelz C, Impey S, et al. Fusion between Intestinal epithelial cells and macrophages in a cancer context results in nuclear reprogramming. Cancer Res. 2011;71:1497–505.
Gast CE, Silk AD, Zarour L, Riegler L, Burkhart JG, Gustafson KT, et al. Cell fusion potentiates tumor heterogeneity and reveals circulating hybrid cells that correlate with stage and survival. Sci Adv. 2018;4:eaat7828.
Chakraborty A, Lazova R, Davies S, Bäckvall H, Ponten F, Brash D, et al. Donor DNA in a renal cell carcinoma metastasis from a bone marrow transplant recipient. Bone Marrow Transplant. 2004;34:183–6.
Clawson GA, Kimchi E, Patrick SD, Xin P, Harouaka R, Zheng S, et al. Circulating tumor cells in melanoma patients. PLoS ONE. 2012;7:e41052.
Clawson GA. Cancer. Fusion for moving. Science. 2013;342:699–700.
Clawson GA, Matters GL, Xin P, Imamura-Kawasawa Y, Du Z, Thiboutot DM, et al. Macrophage-tumor cell fusions from peripheral blood of melanoma patients. PLoS ONE. 2015;10:e0134320.
Clawson GA, Matters GL, Xin P, McGovern C, Wafula E, dePamphilis C, et al. ‘Stealth dissemination’ of macrophage-tumor cell fusions cultured from blood of patients with pancreatic ductal adenocarcinoma. PLoS ONE. 2017;12:e0184451.
Lazova R, Laberge GS, Duvall E, Spoelstra N, Klump V, Sznol M, et al. A melanoma brain metastasis with a donor-patient hybrid genome following bone marrow transplantation: first evidence for fusion in human cancer. PLoS ONE. 2013;8:e66731.
Kurgyis Z, Kemény LV, Buknicz T, Groma G, Oláh J, Jakab Á, et al. Melanoma-derived BRAF(V600E) mutation in peritumoral stromal cells: implications for in vivo cell fusion. Int J Mol Sci. 2016;17:E980.
LaBerge GS, Duvall E, Grasmick Z, Haedicke K, Pawelek J. A melanoma lymph node metastasis with a donor-patient hybrid genome following bone marrow transplantation: a second case of leucocyte-tumor cell hybridization in cancer metastasis. PLoS ONE. 2017;12:e0168581.
Luo F, Liu T, Wang J, Li J, Ma P, Ding H, et al. Bone marrow mesenchymal stem cells participate in prostate carcinogenesis and promote growth of prostate cancer by cell fusion in vivo. Oncotarget. 2016;7:30924–34.
Li G, Kikuchi K, Radka M, Abraham J, Rubin BP, Keller C. IL-4 receptor blockade abrogates satellite cell: Rhabdomyosarcoma fusion and prevents tumor establishment. STEM Cells. 2013;31:2304–12.
Mukhopadhyay KD, Bandyopadhyay A, Chang T-TA, Elkahloun AG, Cornell JE, Yang J, et al. Isolation and characterization of a metastatic hybrid cell line generated by ER negative and ER positive breast cancer cells in mouse bone marrow. PLoS ONE. 2011;6:e20473.
Kemény LV, Kurgyis Z, Buknicz T, Groma G, Jakab Á, Zänker K, et al. Melanoma cells can adopt the phenotype of stromal fibroblasts and macrophages by spontaneous cell fusion in vitro. Int J Mol Sci. 2016;17:E826.
Yano T, Tanaka M, Fukuda N, Ueda T, Nagase H. Loss of mutant mitochondrial DNA harboring the MELAS A3243G mutation in human cybrid cells after cell-cell fusion with normal tissue-derived fibroblast cells. Int J Mol Med. 2010;25:153–8.
Durrieu-Gaillard S, Dumay-Odelot H, Boldina G, Tourasse NJ, Allard D, André F, et al. Regulation of RNA polymerase III transcription during transformation of human IMR90 fibroblasts with defined genetic elements. Cell Cycle. 2018;17:605–15.
Chibon F, Mairal A, Fréneaux P, Terrier P, Coindre JM, Sastre X, et al. The RB1 gene is the target of chromosome 13 deletions in malignant fibrous histiocytoma. Cancer Res. 2000;60:6339–45.
Pérot G, Chibon F, Montero A, Lagarde P, de Thé H, Terrier P, et al. Constant p53 pathway inactivation in a large series of soft tissue sarcomas with complex genetics. Am J Pathol. 2010;177:2080–90.
Gibault L, Pérot G, Chibon F, Bonnin S, Lagarde P, Terrier P, et al. New insights in sarcoma oncogenesis: a comprehensive analysis of a large series of 160 soft tissue sarcomas with complex genomics. J Pathol. 2011;223:64–71.
Cancer Genome Atlas Research Network. Comprehensive and integrated genomic characterization of adult soft tissue sarcomas. Cell. 2017;171:950–965.e28.
Fletcher CDM, Bridge JA, Hogendoorm PCW. World Health Organization classification of tumours: pathology and genetics of tumours of soft tissue and bone. Lyon: IARC Press; 2012.
Kirsch DG, Dinulescu DM, Miller JB, Grimm J, Santiago PM, Young NP, et al. A spatially and temporally restricted mouse model of soft tissue sarcoma. Nat Med. 2007;13:992–7.
Dodd RD, Mito JK, Kirsch DG. Animal models of soft-tissue sarcoma. Dis Models Mech. 2010;3:557–66.
Lazebnik Y. The shock of being united and symphiliosis. Another lesson from plants? Cell Cycle. 2014;13:2323–9.
Ogle BM, Cascalho M, Platt JL. Biological implications of cell fusion. Nat Rev Mol Cell Biol. 2005;6:567–75.
Martincorena I, Roshan A, Gerstung M, Ellis P, Van Loo P, McLaren S, et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science. 2015;348:880–6.
Martincorena I, Fowler JC, Wabik A, Lawson ARJ, Abascal F, Hall MWJ, et al. Somatic mutant clones colonize the human esophagus with age. Science. 2018;362:911–7.
Johnson RT, Rao PN. Mammalian cell fusion: induction of premature chromosome condensation in interphase nuclei. Nature. 1970;226:717–22.
Rao PN, Johnson RT. Mammalian cell fusion: studies on the regulation of DNA synthesis and mitosis. Nature. 1970;225:159–64.
Vignery A. Macrophage fusion. J Exp Med. 2005;202:337–40.
Chen EH, Olson EN. Unveiling the mechanisms of cell-cell fusion. Science. 2005;308:369–73.
Dittmar T, Zänker KS. Tissue regeneration in the chronically inflamed tumor environment: implications for cell fusion driven tumor progression and therapy resistant tumor hybrid cells. Int J Mol Sci. 2015;16:30362–81.
Lu X, Kang Y. Efficient acquisition of dual metastasis organotropism to bone and lung through stable spontaneous fusion between MDA-MB-231 variants. Proc Natl Acad Sci USA. 2009;106:9385–90.
Ding J, Jin W, Chen C, Shao Z, Wu J. Tumor associated macrophage x cancer cell hybrids may acquire cancer stem cell properties in breast cancer. PLoS ONE. 2012;7:e41942.
Giroux V, Rustgi AK. Metaplasia: tissue injury adaptation and a precursor to the dysplasia-cancer sequence. Nat Rev Cancer. 2017;17:594–604.
Elinav E, Nowarski R, Thaiss CA, Hu B, Jin C, Flavell RA. Inflammation-induced cancer: crosstalk between tumours, immune cells and microorganisms. Nat Rev Cancer. 2013;13:759–71.
Van Mater D, Xu E, Reddy A, Añó L, Sachdeva M, Huang W, et al. Injury promotes sarcoma development in a genetically and temporally restricted manner. JCI Insight. 2018;3:123687.
Walsh S, Nygren J, Pontén A, Jovinge S. Myogenic reprogramming of bone marrow derived cells in a W41Dmdmdx deficient mouse model. PLoS ONE. 2011;6:e27500.
Kovacs G. Premature chromosome condensation: evidence for in vivo cell fusion in human malignant tumours. Int J Cancer. 1985;36:637–41.
Kovacs G, Georgii A. Spontaneous cell fusion in human malignancies: possible mechanism leading to heterogeneity. Lancet. 1985;1:350.
Pérot G, Derré J, Coindre J-M, Tirode F, Lucchesi C, Mariani O, et al. Strong smooth muscle differentiation is dependent on myocardin geneamplification in most human retroperitoneal leiomyosarcomas. Cancer Res. 2009;69:2269–78.
Commo F, Guinney J, Ferté C, Bot B, Lefebvre C, Soria J-C, et al. rCGH: a comprehensive array-based genomic profile platform for precision medicine. Bioinformatics. 2016;32:1402–4.
Acknowledgements
This work was supported by the Fondation ARC pour la Recherche Contre le Cancer (to T.L.) and the Fondation Recherche Médicale (to L.L. and L.D.). We acknowledge personnel of CREFRE US006 for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Delespaul, L., Merle, C., Lesluyes, T. et al. Fusion-mediated chromosomal instability promotes aneuploidy patterns that resemble human tumors. Oncogene 38, 6083–6094 (2019). https://doi.org/10.1038/s41388-019-0859-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0859-6
This article is cited by
-
Cell fusion upregulates PD-L1 expression for evasion from immunosurveillance
Cancer Gene Therapy (2024)
-
Single-cell landscape of undifferentiated pleomorphic sarcoma
Oncogene (2024)
-
Acquisition of cancer stem cell capacities after spontaneous cell fusion
BMC Cancer (2021)
-
Cell fusion enhances energy metabolism of mesenchymal tumor hybrid cells to sustain their proliferation and invasion
BMC Cancer (2021)
-
Comparison of hybrid clones derived from human breast epithelial cells and three different cancer cell lines regarding in vitro cancer stem/ initiating cell properties
BMC Cancer (2020)