Abstract
It is well documented that hypoxia activates the hypoxia-inducible factor 1-alpha (HIF1α)/vascular endothelial growth factor A (VEGFA) axis to promote angiogenesis in breast cancer. However, it is unclear how this axis is negatively regulated. In this study, we demonstrated that miR-153 directly inhibits expression of HIF1α by binding to the 3′UTR of HIF1A mRNA, as well as suppresses tube formation of primary human umbilical vein endothelial cells (HUVECs) and breast cancer angiogenesis by decreasing the secretion of VEGFA. Importantly, expression of miR-153 was induced by hypoxia-stimulated ER stress, which activates IRE1α and its downstream transcription factor X-box binding protein 1 (XBP1). X-box binding protein 1 directly binds to the promoter of the miR-153 host gene PTPRN and activates transcription. These results indicate that hypoxia induces miR-153 to fine tune the HIF1α/VEGFA axis in breast cancer angiogenesis and miR-153 could be used for breast cancer anti-angiogenesis therapy.
Similar content being viewed by others
Introduction
Angiogenesis is a physiological process that form new vessels from pre-existing ones [1, 2], which is critical for solid tumor growth, invasion, and metastasis [1, 3]. Vascular endothelial growth factor A (VEGFA) is one of the most important proangiogenic factors secreted by tumor cells [4, 5]. The expression of VEGFA is directly induced by hypoxia-inducible factor 1-alpha (HIF1α) at the transcriptional level under hypoxia [6, 7]. Bevacizumab, a monoclonal antibody against VEGFA, became the first commercially available anti-angiogenesis drug in multiple solid tumors, although it was prohibited for use for treating metastatic breast cancer by the US FDA in 2011.
MicroRNAs (miRNAs) are small RNA molecules that silences gene expression through binding to the 3′-UTR of gene’s mRNA [8,9,10]. A number of studies suggest that miR-153 inhibits tumor growth and metastasis by targeting the snail family transcription repressor 1 (SNAI1), the zinc finger E-box binding homeobox 2 (ZEB2) [11], the metadherin (MTDH) [12], the ADAM metallopeptidase domain 19 (ADAM19) [13], the AKT serine/threonine kinase 1 (AKT1) [14], the HECT domain E3 ubiquitin protein ligase 3 (HECTD3) [15], and the BCL2 family apoptosis regulator (MCL-1) [16]. We recently reported that miR-153 was induced by mifepristone and inhibited the breast cancer stem cells by downregulating the kruppel like factor 5 (KLF5) [17]. However, miR-153 was reported to be oncogenic in prostate cancer by targeting the phosphatase and tensin homolog (PTEN) [18] and in colorectal cancer by targeting the forkhead box O3a (FOXO3a) [19]. Therefore, the functions and action mechanisms of miR-153 in cancers are not completely clear. In this study, we demonstrated that miR-153 inhibits breast cancer angiogenesis by targeting HIF1α.
Expression of miR-153 is downregulated in lung cancer [14, 20], glioblastoma [21, 22], and oral cancer [11]. Breast cancer patients with higher expression levels of miR-153 had a significantly better 5-year survival rate [15]. Loss of miR-153 expression is associated with promoter methylation [23]. Epigenetic regulators, including 4-phenylbutyric acid and 5-aza-2′-deoxycytidine, were reported to induce miR-153 expression [21,22,23]. In addition, miR-153 is induced by mifepristone [17] and glucose [24]. However, the mechanisms of miR-153 induction by these factors are unknown. Interestingly, mifepristone and glucose also induce endoplasmic reticulum (ER) stress [25, 26]. Here, we found that miR-153 was induced by hypoxia through ER stress.
Hypoxia, as stress, causes unfolded or misfolded proteins accumulation within the ER because of the energy depletion and the changes in the redox environment [27, 28]. Unfolded or misfolded proteins further activate three ER transmembrane stress sensors, including the activating transcription factor 6 (ATF6), the pancreatic ER kinase (PKR)-like ER kinase (PERK), and the inositol-requiring enzyme 1 (IRE1), and determine the cell fate according to the duration and intensity of ER stress [29,30,31]. It has been reported that ER stress regulates miRNA expression [32, 33].
In this study, we demonstrated that miR-153 downregulated HIF1α expression by binding to its mRNA 3′-UTR and inhibited breast cancer angiogenesis by inactivating the HIF1α/VEGFA axis. More importantly, we found that hypoxia induced miR-153 expression in breast cancer cells through the ER stress/IRE1α/X-box binding protein 1 (XBP1) pathway. Therefore, we conclude that miR-153 inhibits breast cancer angiogenesis and hypoxia induces miR-153 to fine tune the HIF1α/VEGFA-stimulated angiogenesis. These discoveries help us better understand the regulation of hypoxia-induced angiogenesis and provide a novel therapeutic strategy for breast cancer.
Results
miR-153 inhibits hypoxia-induced HIF1α expression in breast cancer cells
There is a consensus miR-153-binding site at the HIF1A-3′-UTR (Fig. 1a). To test whether miR-153 directly targets HIF1α, we cloned wild-type (WT) and mutant sequences of the miR-153-binding site into the pMIR-REPORT system and performed dual-luciferase report assays. The miR-153 mimics significantly inhibited the luciferase activity of WT pMIR-HIF1A-3′-UTR but not that of mutant in the HEK293T cell line (Fig. 1b). Consistently, the miR-153 mimics dramatically inhibited hypoxia-induced HIF1α protein expression in the MDA-MB-231 and HCC1937 breast cancer cell lines, and the MCF10A immortalized breast epithelial cell line (Fig. 1c). Furthermore, the miR-153 inhibitor increased hypoxia-induced HIF1α protein expression in these cell lines (Fig. 1d).
miR-153 inhibits the HIF1α protein expression. a The predicted miR-153 binding sequence and the mutant sequence at the 3′-UTR of HIF1A mRNA. b miR-153 suppressed the luciferase activity of the HIF1A-3′-UTR but not the HIF1A-3′-UTR-Mutant. HEK293T cells were transfected with the miR-153 mimics (50 nM) and pMIR-HIF1A-3′-UTR or pMIR-HIF1A-3′-UTR-Mutant (800 ng/ml) together with pCMV-Renilla control (8 ng/ml). Following transfection for 48 h, the cell lysate were collected for a dual-luciferase report assay. *P < 0.05 and **P < 0.01, t-test. c miR-153 mimics (50 nM) inhibited the increase in hypoxia-induced HIF1α protein expression in the MDA-MB-231, HCC1937, and MCF10A cell lines. The HIF1α protein level was detected by using western blot analysis. d The inhibitor of miR-153 (50 nM) increased expression of HIF1α induced by hypoxia in the MDA-MB-231, HCC1937, and MCF10A cell lines. The HIF1α protein level was detected by using western blot analysis
To further understand how miR-153 affects HIF1A mRNA expression, we detected the half-life of HIF1A mRNA after manipulating miR-153 expression. After actinomycin D treatment, the miR-153 mimics shortened the half-life of HIF1A mRNA (Fig. S1A), whereas the miR-153 inhibitor prolonged the half-life of HIF1A mRNA (Fig. S1B). These results suggest that miR-153 downregulated HIF1α expression through promoting HIF1A mRNA degradation.
miR-153 inhibits expression of HIF1α target gene VEGFA in breast cancer cells under hypoxia
As miR-153 targets HIF1α, it should inhibit expression of HIF1α target genes. We transfected breast cancer cell lines, MDA-MB-231 and HCC1937, with the miR-153 mimics and exposed the cells to hypoxia (Fig. 2a). Under hypoxia, transcription of HIF1A was suppressed, but transcription of the HIF1α typical downstream target genes, including VEGFA and GLUT1, was significantly induced in both cell lines (Fig. 2b). As predicted, miR-153 significantly decreased the mRNA levels of HIF1A, VEGFA, and GLUT1 under hypoxia (Fig. 2b). Then, the conditional media (CM) were harvested to detect the protein level of VEGFA with ELISA (Fig. 2c). In agreement with the mRNA levels, miR-153 almost completely blocked hypoxia-induced VEGFA protein expression increase in both breast cancer cell lines.
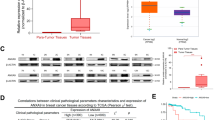
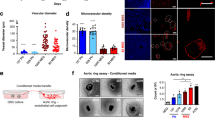
miR-153 inactivates the HIF1α/VEGFA axis in breast cancer. a Schematic illustration of the sample treatment, collection, and detection. MDA-MB-231 and HCC1937 cell lines were transfected with miR-153 mimics (50 nM) for 48 h and were then exposed to 1% O2 for 24 h. The cells were collected for real-time PCR to detect the HIF1A, GLUT1, and VEGFA mRNA levels. The conditional media (CM) were harvested for the enzyme-linked immunosorbent assay (ELISA) to detect the VEGFA protein levels. b The miR-153 mimics decreased the mRNA levels of HIF1A and inhibited the transcription of GLUT1 and VEGFA under hypoxia. 'N' means normoxia, and 'H' means hypoxia. **P < 0.01, t-test. c The miR-153 mimics inhibited the secreted VEGFA protein levels from the hypoxia-treated breast cancer cells. *P < 0.05, **P < 0.01, t-test. d There is a negative correlation trend between miR-153 and VEGFA mRNA in 732 breast cancer patients from the TCGA database. e There is a negative correlation trend between miR-153 and VEGFA mRNA in 18 clinical breast cancer samples
To test whether miR-153 inhibits VEGFA expression in breast cancer patients, we first analyzed data from the TCGA database and found that there was a negative correlation trend between miR-153 and VEGFA mRNA (r = −0.082, P = 0.026, Fig. 2d) in 732 breast cancer patients from TCGA database. We further collected 18 clinical breast cancer samples and detected the expression levels of VEGFA mRNA and miR-153 using the real-time polymerase chain reaction (real-time PCR). Consistently, we also found a negative correlation trend between them (r = -0.2417, P = 0.0179, Fig. 2e).
miR-153 suppresses the proliferation, migration, and tube formation of primary HUVECs by decreasing the breast cancer secreted VEGFA levels
As miR-153 inhibits expression of VEGFA in breast cancer cell lines under hypoxia, it may suppress angiogenesis of human umbilical vein endothelial cells (HUVECs) in a paracrine manner in vitro (Fig. 3a). To test this, we first obtained primary HUVECs and validated CD31 expression using flow cytometry and immunofluorescence staining (Fig. 3b). Then, we collected the CM for incubation with the primary HUVECs and performed DNA synthesis, wound healing, and tube formation assays in vitro (Figs. 3c-h, Fig. S2A-F). As a result, CM derived from either MDA-MB-231 or HCC1937 cells under hypoxia significantly promoted the proliferation, migration, and tube formation of primary HUVECs compared with CM under normoxia. miR-153 significantly inhibited the hypoxia-induced pro-angiogenesis function in both cancer cell lines. The addition of the exogenous VEGFA (20 ng/ml) into CM rescued the miR-153 caused inhibitory effect on the primary HUVECs (Figs. 3c-h, Fig. S2A-F). These results suggest that miR-153 inhibits hypoxia-induced angiogenesis in vitro by downregulating VEGFA protein expression in breast cancer cells.
miR-153 suppresses the proliferation, migration, and tube formation of primary HUVECs in a paracrine manner. a Schematic illustration of the sample treatment and collection. The breast cancer cells were transfected with miR-153 mimics (50 nM) for 48 h and were then exposed to 1% O2 for 24 h. The conditional medium (CM) was harvested for culturing the primary HUVECs. b CD31 expression of primary HUVECs was detected using flow cytometry and immunofluorescence assays. The representative images for immunofluorescence are shown. c The DNA synthesis of primary HUVECs was inhibited by CM collected from hypoxia-treated HCC1937 cells transfected with miR-153 mimics. The exogenous VEGFA (20 ng/ml) could rescue the phenotype. DNA synthesis was detected by using the Click-iTTM EdU Alexa Fluor® 647 Imaging Kit. Representative images are shown. d The quantitative results of c. **P < 0.01, t-test. e The migration of primary HUVECs was inhibited by CM collected from hypoxia-treated HCC1937 cells transfected with miR-153 mimics in the wound-healing assay. The exogenous VEGFA (20 ng/ml) could rescue the phenotype. Representative images are shown. f The quantitative results of e. *P < 0.05 and **P < 0.01, t-test. g The tube formation of primary HUVECs was inhibited by CM collected from hypoxia-treated HCC1937 cells transfected with miR-153 mimics. The exogenous VEGFA (20 ng/ml) rescued the phenotype. Representative images are shown. Scale bar, 50 μm. h The quantitative results of g. **P < 0.01, t-test
miR-153 inhibits breast tumor growth and angiogenesis in vivo
To test whether miR-153 suppresses angiogenesis and tumor growth in vivo, we orthotopically injected MDA-MB-231-Luc cells into the fat pad of BALB/c nude mice (Fig. S3). When the tumor size reached 50 mm3, the mice were given miR-153 or control agomir (1 nmol) by subcutaneous injection around the tumors. miR-153 significantly inhibited the tumor growth based on the tumor volume and the tumor weight (Fig. S3A and D). The bioluminescence images using the IVIS system also showed that the average intensity of the miR-153 group was less than that of the control group in the MDA-MB-231-Luc xenograft tumor model (Fig. S3B). The mRNA levels of HIF1A and VEGFA in xenografts of the miR-153 group were lower than those of the control group (Fig. S3E). We also demonstrated that miR-153 significantly decreased the microvessel numbers in the xenograft tumors, as assessed by CD31 immunohistochemistry (Fig. S3F-G).
To investigate whether miR-153 regulates tumor growth and angiogenesis through suppressing HIF1α expression, we ectopically overexpressed HIF1α in MDA-MB-231 cells (Fig. 4a) and orthotopically injected MDA-MB-231-pCDH-HIF1α cells into the fat pad of BALB/c nude mice. Interestingly, when HIF1α was ectopically overexpressed in the tumors, miR-153 failed to suppress MDA-MB-231 xenograft tumor growth (Figs. 4b-d). Meanwhile, restoring HIF1α expression reversed the phenotype of miR-153 in tumor angiogenesis (Figs. 4e, f), and the inhibition effect of miR-153 on VEGFA mRNA could be rescued (Fig. 4g). Taken together, these results suggest that miR-153 inhibits the tumor growth and angiogenesis in vivo by inhibiting the HIF1α/VEGFA axis, at least in part.
miR-153 inhibits the tumor angiogenesis by downregulating the HIF1α/VEGFA axis. a Overexpression of HIF1α in the MDA-MB-231 cell lines, as detected by western blot. b–d The miR-153 agomir suppressed the growth of the MDA-MB-231-pCDH-vector tumors but not that of the MDA-MB-231-pCDH-HIF1α tumors in a nude mouse xenograft model. Tumor growth was measured every 3 days. All tumors were collected and weighed on the last day of the experiment. Data were represented as the mean ± s.d. of six mice for each group (12 tumors). *P < 0.05, **P < 0.01, t-test. e, f miR-153 decreased the number of microvessels, as measured by the CD31 immunohistochemical staining, in the MDA-MB-231-pCDH-vector xenograft tumors, but not in the MDA-MB-231-pCDH-HIF1α xenograft tumors. Represented images are shown. Scale bar, 100 μm. **P < 0.01, t-test. g miR-153 decreased the expression levels of HIF1A and VEGFA at the mRNA level in the MDA-MB-231-pCDH-vector xenograft tumors, but not in the MDA-MB-231-pCDH-HIF1α xenograft tumors, as detected using RT-qPCR. *P < 0.05, **P < 0.01, t-test
Hypoxia induces miR-153 expression via triggering ER stress
Considering miR-153 inhibits hypoxia-induced HIF1α and angiogenesis, we wondered whether miR-153 is regulated by hypoxia. As shown in Fig. 5a, hypoxia significantly increased the miR-153 levels in both MDA-MB-231 and HCC1937 cells. Because all miR-153 inducers, including mifepristone, intermittent high glucose, and hypoxia, could trigger ER stress, we examined whether hypoxia-induced ER stress in MDA-MB-231 and HCC1937 cells. As expected, the BIP, ATF6, IRE1α, and PERK protein expression levels were induced by hypoxia including 1% O2 (Fig. 5a) and CoCl2 (Fig. S4A). Similar results were observed in MCF10A cells (Fig. S4A).
Hypoxia upregulates miR-153 expression by triggering ER stress protein IRE1α. a Hypoxia (1% O2) triggered ER stress and upregulated miR-153 expression in MDA-MB-231 and HCC1937 cell lines. The HIF1α, BIP, PERK, IRE1α, and ATF6 protein levels were detected by western blot analysis, and the miR-153 level was detected by the real-time polymerase chain reaction. *P < 0.05, **P < 0.01, t-test. b Tunicamycin (TM, 5 μg/ml) upregulated miR-153 expression in MDA-MB-231 and HCC1937 cell lines. The BIP, PERK, IRE1α, and ATF6 protein levels were detected by western blot analysis, and the miR-153 level was detected by RT-qPCR. *P < 0.05, **P < 0.01, t-test, compared with the value of the TM 0-h treatment. c Knockdown of IRE1α specifically blocked hypoxia-induced miR-153 expression in MDA-MB-231 and HCC1937 cell lines. The HIF1α, BIP, PERK, IRE1α, and ATF6 protein levels were detected by western blot analysis, and the miR-153 level was detected by the real-time polymerase chain reaction. *P < 0.05, **P < 0.01, t-test
Subsequently, we treated MDA-MB-231 and HCC1937 cells with a typical ER stress inducer, tunicamycin (TM), and detected the miR-153 levels using RT-qPCR. TM induced ER stress in both MDA-MB-231 and HCC1937 cells because the protein levels of BIP, PERK, IRE1α, and ATF6 were upregulated (Fig. 5b). Importantly, the miR-153 expression levels were induced by TM in both cell lines in a time-dependent manner (Fig. 5b). To illustrate the mechanism by which ER stress induces miR-153 expression under hypoxia, we knocked down PERK, IRE1α, and ATF6 and examined miR-153 expression. Knockdown of IRE1α, but not ATF6 or PERK, blocked hypoxia-induced miR-153 expression in MDA-MB-231 and HCC1937 cells (Fig. 5c and Fig. S4C). This is also true in MCF10A cells (Fig. S4B-C).
XBP1 transcription factor mediates hypoxia-induced miR-153 expression
Hypoxia induces miR-153 expression in an ER stress/IRE1α-dependent manner. X-box binding protein 1 is an important downstream transcription factor of IRE1α. When XBP1 was knocked down in MDA-MB-231 and HCC1937 cells, either hypoxia or TM failed to induce miR-153 expression (Figs. 6a, b). Similar results were observed in MCF10A (Fig. S5A-B).
X-box binding protein 1 induces miR-153 expression by binding to the PTPRN promoter. a Knockdown of XBP1 inhibited the upregulation of miR-153 induced by hypoxia in the MDA-MB-231 and HCC1937 cell lines. **P < 0.01, t-test, compared with the value of CONsi with the 1% O2 0-h treatment. b Knockdown of XBP1 inhibited the upregulation of miR-153 induced by TM in MDA-MB-231 and the HCC1937 cell lines. **P < 0.01, t-test, compared with the value of CONsi with the TM 0-h treatment. c Illustration of the miR-153 location in two host genes, PTPRN and PTPRN2. The XBP1 binding sites in the promoter region of these two host genes are indicated. d Hypoxia induces expression of miR-153 and PTPRN, but not PTPRN2, at the mRNA level in the MCF10A, MDA-MB-231, and HCC1937 cell lines, as detected by RT-qPCR. e Hypoxia induced PTPRN protein expression, as detected by western blot analysis. f Hypoxia-induced XBP1 bound to the promoter of PTPRN, as determined by the chromatin immunoprecipitation (ChIP) assay. g The PTPRN promoter containing the XBP1 binding motif was significantly activated by hypoxia in HEK293T cells, as determined by dual-luciferase assay. **P < 0.01, t-test. h The work model of this study
The miR-153 gene is located in genes encoding two major type 1 diabetes autoantigens, islet-associated protein PTPRN and PTPRN2 [24]. There are consensus XBP1-binding sites in their promoter regions (Fig. 6c). We tested whether hypoxia induces expression of PTPRN and PTPRN2 genes in breast cells and found that only PTPRN, but not PTPRN2, was induced at the transcriptional level in MCF10A, MDA-MB-231, and HCC1937 cells (Fig. 6d). Consistently, hypoxia increased the PTPRN protein expression levels in these cells (Fig. 6e). Finally, we demonstrated that XBP1 bound to the PTPRN promoter under hypoxia using the chromatin immunoprecipitation assay (Fig. 6f). Furthermore, hypoxia activated the PTPRN promoter with the XBP1-binding site but not the PTPRN promoter without the XBP1-binding site using the dual-luciferase assay (Fig. 6g). These findings suggest that XBP1 induces miR-153 expression by directly binding to the promoter of its host gene PTPRN in breast cancer cells under hypoxia.
Discussion
Accumulating evidences show that miR-153 plays a suppressor role in tumors [22, 34,35,36,37] because miR-153 inhibits tumors by targeting several oncogenes associated with EMT (SNAI1 and ZEB2) [11, 38, 39], cancer stemness (KLF5 and nuclear factor, erythroid 2 like 2 (NRF2)) [17, 40], and survival (HECTD3 and MCL-1) [15, 16]. However, the functions and mechanisms of miR-153 in tumor angiogenesis have not been studied. We demonstrated that miR-153 suppresses breast cancer angiogenesis by targeting the HIF1α/VEGFA axis. Additionally, miR-153 is induced by hypoxia in an ER stress-IRE1α-XBP1-dependent manner. These findings revealed a novel function and mechanism of action of miR-153 as well as demonstrate a novel regulatory mechanism of miR-153.
It is well known that hypoxia stabilizes HIF1α at the posttranslational level [6, 7] and induces VEGFA expression to stimulate angiogenesis. Interestingly, hypoxia also switches on miR-153 expression by triggering the ER stress/IRE1α/XBP1 pathway to negatively regulate the HIF1α/VEGFA axis at the post-transcriptional level. This “brake mechanism” can avoid overactivation of the HIF1α/VEGFA axis under hypoxic physiological or pathological conditions. This discovery enriches our knowledge about the regulation of HIF1α at different levels.
Although the HIF1α protein level was upregulated under hypoxia (Figs. 1c, d), the HIF1A mRNA level was significantly downregulated in breast cancer cell lines (Fig. 2b). The HIF1α protein is no longer ubiquitinated and degraded under hypoxia. However, the mechanism by which the HIF1A mRNA levels are decreased by hypoxia is unknown. The miR-153 mimics reduced the mRNA levels of HIF1A and shortened its half-life (Fig. 2b and S1A), suggesting that the hypoxia-induced miR-153 may be partially attributed to the downregulation of the HIF1A mRNA level. Other HIF1α-negative regulators, such as miR-429, miR-519c, miR-199a, miR-20a, and miR-497, may also be involved [41,42,43,44,45,46].
In addition to hypoxia, mifepristone and high glucose also increase expression of miR-153 [17, 24]. Intriguingly, these stimuli also induce ER stress [25, 26]. As expected, the typical ER stress inducer, TM, also induced miR-153 expression in breast cell lines. Therefore, it is very possible that mifepristone and high glucose induce miR-153 through the ER stress/IRE1α/XBP1 pathway. We further showed that expression of miR-153 under hypoxia was directly mediated by activated transcription factor XBP1 by binding to the promoter of the miR-153 host gene PTPRN. Human miR-153 is located in the 19th intron of PTPRN and PTPRN2. However, only PTPRN was upregulated with miR-153 at the mRNA level by hypoxia in breast cell lines (Fig. 6d). By contrast, transcription of PTPRN2 was downregulated under hypoxia or could not be detected in MDA-MB-231 cells under normoxia (Fig. 6d). Therefore, miR-153 is mainly co-regulated with the PTPRN host gene in human breast epithelial cells. The role of PTPRN in hypoxia-driven cellular processes requires further investigation.
As a direct transcription factor of miR-153, XBP1 has been reported to enhance HIF1A transcriptional activity through the protein-protein interaction [47], as well as to induce expression of VEGFA [2, 48]. Knockdown of XBP1 suppresses the tumor growth of triple-negative breast cancer xenografts [47]. We found that miR-153 decreased expression of XBP1 at the transcription level (Fig. S5C). There is possible negative feedback inhibition between XBP1 and miR-153. X-box binding protein 1 induces transcription of miR-153, but miR-153 inhibits expression of XBP1 through a yet to be identified mechanism. Taken together, we propose that miR-153 inhibits angiogenesis twofold. On the one hand, miR-153 inhibits HIF1α by directly binding to the HIF1A mRNA 3′-UTR. On the other hand, miR-153 disrupts the XBP1/HIF1α interaction by suppressing transcription of XBP1 via negative feedback.
In summary, we discovered a novel function of miR-153 in suppressing breast cancer angiogenesis and a specific regulatory mechanism of miR-153 by hypoxia. miR-153 inhibited the proliferation, migration, and tube formation of primary HUVECs and angiogenesis of the MDA-MB-231 formed xenograft tumor by inhibiting the HIF1α/VEGFA axis by binding to the 3′-UTR of HIF1A mRNA in breast cancer cells. Under hypoxia, ER stress was triggered and the IRE1α/XBP1 pathway was activated. Activated XBP1 transcription factor further induced miR-153 expression by binding to the promoter of a miR-153 host gene PTPRN. Our results imply that miR-153 has the potential to serve as an adjuvant of anti-angiogenesis therapy.
Materials and methods
Cell line, culture, and transfection
HEK293T cell line was cultured in the Dulbecco’s modified Eagle’s medium (DMEM) basic medium (Gibco, Grand Island, USA), which was added with fetal bovine serum (FBS, 10% as the final concentration, Gibco). MCF10A is a breast epithelial cell line, and was cultured in the DMEM/F-12 (1:1) basic medium (Gibco) containing horse serum (5% as the final concentration, Gibco), cholera enterotoxin (0.1 μg/ml), insulin (10 μg/ml), l-glutamine (2 mM), hydrocortisone (0.5 μg/ml), and epidermal growth factor (20 ng/ml). MDA-MB-231 and MDA-MB-231-Luc were maintained in the DMEM/F-12 (1:1) basic medium, which was added with 10 % FBS. HCC1937 was cultured in RPMI-1640 basic medium (Gibco) with 10% FBS. These above cell lines were authenticated by short tandem repeats (STRs) DNA profiling and were not contaminated by mycoplasma. Primary HUVECs were maintained in EGM-2 BulletKit (CC-3162, Lonza, USA). We use the Lipofectamine 2000 reagent (Invitrogen, USA) to transfect cells with plasmids, small interfering RNA (siRNA), miRNA mimics and miRNA inhibitor (Ribobio, Guangzhou, China). The sequences of siRNA targeting are shown in the Supplementary Table 1. The hypoxic condition was created by culturing cells in a hypoxia chamber flushed with 1% O2, 5% CO2, and 94% N2 mixture gas.
Western blot, antibodies, and reagents
The western blot assay was performed according the protocol described in our previous studies [49, 50]. The anti-HIF1α antibody was purchased from Epitomics (#2015-1, Burlingame, USA) and Cell Signaling (#3716, Beverly, USA). The antibodies of anti-GAPDH (sc-25778) and anti-XBP1 (sc-7160) were purchased from Santa Cruz Biotechnology. The antibodies of anti-β-actin (A5441) and anti-PTPRN (AV45381) were purchased from Sigma-Aldrich. The antibodies of anti-CD31 (ab28364) and anti-ATF6 (ab37149) were purchased from Abcam. The antibodies of anti-IRE1α (#3294), anti-BIP (#3177), and anti-PERK (#5683) were bought from Cell Signaling. TM (11089-65-9) and Cobalt (II) chloride (MB2442) were purchased from Melonepharma (Dalian, China). Actinomycin D (HY-17559) was purchased from MedChem Express (MCE, Shanghai, China).
Dual-luciferase report assays
To analyze the function of the miR-153 (MI0000463) binding site at the 3′-UTR of HIF1A mRNA, we synthesized complementary DNA oligonucleotides that consisted of the test or the mutant sequence (the sequence of HIF1A-3′-UTR-WT is 5′-CCCAAGCTTATAATATAGAAAGATATGCATATATCTAGAAGGTACTAGTCCG-3′; and the sequence of HIF1A-3′-UTR-Mut is 5′-CCCAAGCTTATAATATAGAAAGACGCTTCTATATCTAGAAGGTACTAGTCCG-3′). Then, we cloned them into the pMIR-REPORTTM system. The HEK293T cells were transfected with the negative control or the miR-153 mimics (50 nM as the final concentration) and the pMIR-HIF1A-3′-UTR or pMIR-HIF1A-3′-UTR-Mutant luciferase reporter construct (800 ng per well) together with pCMV-Renilla control (8 ng per well). Every kind of treatment was performed in triplicate. Forty-eight hours later, the cell lysate was collected and the luciferase activities were detected by using the system of dual-luciferase reporter assay (Promega, USA).
To detect the function of the XBP1 binding motif in the PTPRN (NM_001199763.1) promoter, we amplified the PTPRN promoters with or without the XBP1 binding motif from the genomic DNA of HEK293T cell line. For the WT PTPRN promoter (with the XBP1 binding motif), the primers of PCR are as follow: the forward is 5′-CGGGGTACCAATACTACCAAACCTGAAACAACCCAA-3′; the reverse is 5′-CCCAAGCTTCCAGAACGATAAACCCGAAGACA -3′. For the mutant PTPRN promoter (without the XBP1 binding motif), the primers of PCR are as follow: the forward is 5′-CGGGGTACCATACACTGGAGAAATACAGGGAAAT-3′; the reverse is 5′-CCCAAGCTTAGAACGATAAACCCGAAGACAAG-3′. These two promoters were further cloned into the pGL3-Basic system (Promega). The pGL3-PTPRN-promoter-WT or the pGL3-PTPRN-promoter-Mut was transfected into the HEK293T cells together with the pCMV-Renilla control. Following the transfection for 48 h, the cells were put into the hypoxia chamber for 12 h, and then the luciferase activity was detected.
The real-time polymerase chain reaction
The total mRNAs were isolated by using the reagent of TRIzol® (15596-026, Invitrogen). In order to get the complementary DNA, we performed the reverse transcription assay with the kit of Iscript complementary DNA synthesis (170-8891; Bio-Rad, USA) for HIF1A, GLUT1, VEGFA, GAPDH, PTPRN, PTPRN2, and 18S rRNA, and with the TaqMan® MicroRNA Reverse Transcription Kit (4366596, Thermo Fisher Scientific, Fremont, CA, USA) for miRNA. For the quantitative PCR (qPCR), the SYBR Green Select Master Mix system (4472908, Applied Biosystems, USA) was used on the ABI-7900HT system (Applied Biosystems). The reverse transcription primers of miR-153 and U6, and the primers of miR-153, U6, HIF1A, GLUT1, VEGFA, PTPRN, PTPRN2, and 18S rRNA for qPCR are shown in the Supplementary Table 2.
In vitro VEGFA quantification
MDA-MB-231 (0.8 × 105 cells per well) and HCC1937 (1 × 105 cells per well) cells were transfected with the negative control or the miR-153 mimics (50 nM). Following the transfection for 48 h, the cells were exposed to 1% O2 for 24 h. Then, the supernatants were collected and VEGFA was quantified by using the Quantikine VEGFA the enzyme-linked immunosorbent assay (ELISA) kit (DVE00, R&D Systems, UK).
TCGA database and clinical samples
We downloaded the data of human breast cancer from TCGA database, and analyzed the correlation between miR-153 and VEGFA mRNA in 86 normal breast tissue samples and in 732 breast cancer tissue samples. The sample size is chosen to ensure the adequate power to detect the meaningful correlation trend. The computer code used in the study is available from the corresponding authors on request. Eighteen clinical breast cancer tissue samples were obtained from the First Affiliated Hospital of Kunming Medical University, which was approved by the human ethics committee of Kunming Institute of Zoology, Chinese Academy of Sciences. The informed consent was obtained from the patients. To detect the expression correlation between miR-153 and VEGFA mRNA, we purified the total RNA of these tissue samples and used the qPCR to detect the levels miR-153 and VEGFA mRNA. Their expression correlation was analyzed by SPSS 18.0 and the correlation regression curve were fitted by GraghPad Prism 6.0 software.
Primary HUVEC identification
We isolated the primary human umbilical vein endothelial cells from neonatal umbilical cord vein, which was approved by the human ethics committee of Kunming Institute of Zoology, Chinese Academy of Sciences. The informed consent was obtained from the maternal. Expression of CD31 in primary HUVECs was detected by both the flow cytometry and the immunofluorescence staining assays. For the flow cytometry assay, primary HUVECs were trypsinized into single cells, and then were washed once by using the phosphate-buffered saline (PBS) with 3% FBS (named as PF solution). Cells were stained with the FITC IgG isotype control (400905) or FITC anti-hCD31 (303103) antibody (Biolegend, USA) on ice for 30 min. After centrifugation at 1500 rpm for 5 min, the cells were washed twice by using the PF solution. Then they were analyzed by using flow cytometry. For the immunofluorescence staining, primary HUVECs were seeded on coverslips (BD Biosciences, USA). On the next day, we used methanol to fix the cells at –20 °C for 5 min, and used 2% bovine serine albumin to block the slides for 20 min. After washing with PBS, cells were separately incubated with the normal mouse IgG (sc-2025, Santa Cruz) or the primary monoclonal mouse anti-CD31 antibody (MAB-0031, Maixin, Fujian, China) overnight at 4 °C. Subsequently, cells were incubated with the FITC-conjugated secondary antibody (115-095-003, Jackson ImmunoResearch, USA) for 1 h. Nuclei were stained by using DAPI for 2 min. The coverslips were mounted with the 90% glycerol and examined with fluorescence microscopy.
Cell proliferation and migration assays
To detect the proliferation of primary HUVECs, we used the Click-iT®EdU Alexa Fluor 647 Imaging Kits (Invitrogen) according to the manufacturer’s protocol. Briefly, HUVECs were seeded on coverslips (BD Biosciences) at 0.5 × 105 cells per well. On the next day, the supernatants were discarded and the cells were cultured with the CM derived from hypoxia-treated MDA-MB-231 or HCC1937 cells that had been transfected with miR-153 mimics with or without exogenous 20 ng/ml VEGFA (CC-4114B, Lonza). Six hours later, the cells were incubated with EdU in CM for 5 h, followed by fixing and staining. For each sample, three random fields were observed by using fluorescence microscopy, and the total numbers of cells and EdU-positive cells were counted. To detect the migration of primary HUVECs, we performed the wound-healing assay. Twenty-four hours after seeding, the supernatants of HUVECs were discarded and the cells were scratched and cultured with the above described CM for 24 h. Wound closure was imaged under microscopy. For each image, the gap width was analyzed by Image Pro-Plus 6.0 (Media Cybernetics, USA).
Tube formation assays
HUVECs (1 × 104) in CM with or without exogenous VEGFA (20 ng/ml, Lonza) were seeded onto Matrigel (BD Biosciences)-coated μ-Slide angiogenesis (ibidi GmbH, Munich, Germany). At 6 h after seeding, images were taken under microscopy, and then were analyzed with Image Pro-Plus 6.0 software. The total tube length was measured.
HIF1α overexpression
The pcDNA3.1-HIF1α plasmid were kindly provided by Professor Lingqiang Zhang at the Academy of Military Medical Sciences (Beijing, China), and the pCDH-CMV-MCS-EF1-puro-3 × FLAG-3 × HA lentiviral vector was obtained from Prof. Wen Liu at Xiamen University (Xiamen, Fujian, China). We amplified the full-length HIF1A gene (NM_001243084.1) without the miR-153 binding sites, which coding sequence from the pcDNA3.1-HIF1α by using the PCR primers of 5′-CGGGATCCGCCACCATGGAGGGCGCCGGCGGCGCG-3′ (forward) and 5′-TCCCCCGGGTCAGTTAACTTGATCCAAAGC-3′ (reverse), and then subcloned it into the pCDH lentiviral vector. The packaging plasmids including pMDLg/pRRE, pRSV-Rev, pCMV-VSV-G, and pCDH-HIF1α expression plasmid were co-transfected into the HEK293T cells (2 × 106 in 10 cm plate) to produce lentivirus. Following the transfection for 72 h, the lentivirus was collected and used to infect the MDA-MB-231 cells. Forty-eight hours later, puromycin (2 μg/ml) was used to screen the cell populations that are resistant to drug.
Tumorigenesis in BALB/c nude mice
We purchased thirteen 6- to 7-week-old female BALB/c nude mice and twenty-four 6- to 7-week-old female BALB/c nude mice from Slaccas (Changsha, Hunan, China) and Vital River Laboratory Animal Technology Co. Ltd (Beijing, China), respectively. Animal feeding and experiments were approved by the animal ethics committee of Kunming Institute of Zoology, Chinese Academy of Sciences. MDA-MB-231-Luc, MDA-MB-231-pCDH-vector, or MDA-MB-231-pCDH-HIF1α cells (3 × 105 in Matrigel (BD Biosciences)) were implanted into the mammary fat pads of the mice. The size of tumor was measured and was calculated by using the equation: tumor volume (cm3) = π (length × width2)/6. The mice bearing different cells were distributed into two groups through generating random number based on animal’s number, when the tumor volume is close to 50 mm3. The experimenter who was responsible for animal grouping and tumor size detecting did not know the mice belong to which group. One group was administered with agomir control, the other group was administered with miR-153 agomir. The agomir control and miR-153 agomir (1 nmol per tumor, Ribobio) were subcutaneously administered around the tumor every 3 days. For bioluminescent detection, each mouse was first given d-luciferin (D12505, Bridgen, Beijing, China) through intraperitoneal injection. After 5 min, it was anesthetized with ketamine and xylazine. The bioluminescent images were collected by using the system of IVIS Lumina XR (Caliper life Sciences, USA). Each tumor tissue was divided into two parts, one is used to purify the total RNA, the other is used for immunohistochemistry. Some tumors were only used in immunohistochemistry for their too small sizes. For the results of qPCR, the sample will be excluded if its value is higher or lower than the mean ± 3 × s.d.
Immunohistochemistry for CD31
The xenograft tumor tissues were fixed in 3.7% formalin solution. The immunohistochemistry assay was performed on 4-μm-thick paraffin sections after the pressure-cooking for antigen retrieval. The anti-CD31 primary antibody (1:400, Abcam, ab28364) was used. Subsequently, we incubated the slides with the anti-mouse/rabbit ultra sensitive polymer system (PV-8000, ZSGB-BIO, Beijing, China). Signals for all slides were visualized by DAB staining. The slides were mounted after hematoxylin staining. Ten image fields for each slide, except for those are too small, were randomly collected by the persons who did not participate in this study by using microscopy, and the number of microvessel with positive CD31 expression was counted.
Chromatin immunoprecipitation assays
After hypoxia treatment, the diluted DNA–protein complex derived from MCF10A cells was incubated with the rabbit IgG or the anti-XBP1 antibody (Santa Cruz) in 1.5 ml centrifuge tube for 4 h at 4 °C. Then, each tube of sample was added with herring sperm DNA and protein A/G PLUS-Agarose (sc-2003, Santa Cruz), and continued to incubate overnight at 4 °C. Chromosomal DNA was amplified by PCR after purification. The PCR primers for amplifying the interest region on the PTPRN gene promoter (–2238 to –2225 from ATG) were as follows: 5′-GGACTACAGGTACGTGTTAC-3′ and 5′-GAGCCCAGGAGATTGAGA-3′.
Statistical analysis
All data in this study were analyzed by using the software of SPSS 18.0 (SPSS Inc., USA). Each experiment was performed at least three times, and the results are represented as the means ± s.d. The differences between two groups were analyzed by using the two-sided t-test, and the variance is similar between the groups, which are being statistically compared. The value of P smaller than 0.05 were considered statistically significant.
References
Folkman J. How is blood vessel growth regulated in normal and neoplastic tissue? G.H.A. Clowes memorial Award lecture. Cancer Res. 1986;46:467–73.
Urra H, Hetz C. A novel ER stress-independent function of the UPR in angiogenesis. Mol Cell. 2014;54:542–4.
Albini A, Indraccolo S, Noonan DM, Pfeffer U. Functional genomics of endothelial cells treated with anti-angiogenic or angiopreventive drugs. Clin Exp Metastas-. 2010;27:419–39.
Senger DR, Galli SJ, Dvorak AM, Perruzzi CA, Harvey VS, Dvorak HF. Tumor cells secrete a vascular permeability factor that promotes accumulation of ascites fluid. Sci (New Y, NY). 1983;219:983–5.
Senger DR, Van de Water L, Brown LF, Nagy JA, Yeo KT, Yeo TK, et al. Vascular permeability factor (VPF, VEGF) in tumor biology. Cancer Metastas- Rev. 1993;12:303–24.
Hirota K, Semenza GL. Regulation of angiogenesis by hypoxia-inducible factor 1. Crit Rev Oncol Hematol. 2006;59:15–26.
Pugh CW, Ratcliffe PJ. Regulation of angiogenesis by hypoxia: role of the HIF system. Nat Med. 2003;9:677–84.
Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10:126–39.
Kota J, Chivukula RR, O’Donnell KA, Wentzel EA, Montgomery CL, Hwang HW, et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009;137:1005–17.
Lee YS, Nakahara K, Pham JW, Kim K, He Z, Sontheimer EJ, et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell. 2004;117:69–81.
Xu Q, Sun Q, Zhang J, Yu J, Chen W, Zhang Z. Downregulation of miR-153 contributes to epithelial-mesenchymal transition and tumor metastasis in human epithelial cancer. Carcinogenesis. 2013;34:539–49.
Li W, Zhai L, Zhao C, Lv S. MiR-153 inhibits epithelial-mesenchymal transition by targeting metadherin in human breast cancer. Breast Cancer Res Treat. 2015;150:501–9.
Shan N, Shen L, Wang J, He D, Duan C. MiR-153 inhibits migration and invasion of human non-small-cell lung cancer by targeting ADAM19. Biochem Biophys Res Commun. 2015;456:385–91.
Yuan Y, Du W, Wang Y, Xu C, Wang J, Zhang Y, et al. Suppression of AKT expression by miR-153 produced anti-tumor activity in lung cancer. Int J Cancer. 2015;136:1333–40.
Wu X, Li L, Li Y, Liu Z. MiR-153 promotes breast cancer cell apoptosis by targeting HECTD3. Am J Cancer Res. 2016;6:1563–71.
Zou Y, Liu W, Zhang J, Xiang D. miR-153 regulates apoptosis and autophagy of cardiomyocytes by targeting Mcl-1. Mol Med Rep. 2016;14:1033–9.
Liu R, Shi P, Nie Z, Liang H, Zhou Z, Chen W, et al. Mifepristone suppresses basal triple-negative breast cancer stem cells by down-regulating KLF5 expression. Growth. 2016;17:18.
Wu Z, He B, He J, Mao X. Upregulation of miR-153 promotes cell proliferation via downregulation of the phosphatase and tensin homolog (PTEN) tumor suppressor gene in human prostate cancer. Prostate. 2013;73:596–604.
Zhang L, Pickard K, Jenei V, Bullock MD, Bruce A, Mitter R, et al. miR-153 supports colorectal cancer progression via pleiotropic effects that enhance invasion and chemotherapeutic resistance. Cancer Res. 2013;73:6435–47.
Chen WJ, Zhang EN, Zhong ZK, Jiang MZ, Yang XF, Zhou DM, et al. MicroRNA-153 expression and prognosis in non-small cell lung cancer. Int J Clin Exp Pathol. 2015;8:8671–5.
Ghasemi A, Fallah S, Ansari M. MiR-153 as a tumor suppressor in glioblastoma multiforme is downregulated by DNA methylation. Clin Lab. 2016;62:573–80.
Xu J, Liao X, Wong C. Downregulations of B-cell lymphoma 2 and myeloid cell leukemia sequence 1 by microRNA 153 induce apoptosis in a glioblastoma cell line DBTRG-05MG. Int J Cancer. 2010;126:1029–35.
Bao B, Rodriguez-Melendez R, Zempleni J. Cytosine methylation in miR-153 gene promoters increases the expression of holocarboxylase synthetase, thereby increasing the abundance of histone H4 biotinylation marks in HEK-293 human kidney cells. J Nutr Biochem. 2012;23:635–9.
Mandemakers W, Abuhatzira L, Xu H, Caromile LA, Hebert SS, Snellinx A, et al. Co-regulation of intragenic microRNA miR-153 and its host gene Ia-2 beta: identification of miR-153 target genes with functions related to IA-2beta in pancreas and brain. Diabetologia. 2013;56:1547–56.
Dioufa N, Kassi E, Papavassiliou AG, Kiaris H. Atypical induction of the unfolded protein response by mifepristone. Endocrine. 2010;38:167–73.
Wang Y, Gao L, Li Y, Chen H, Sun Z. Nifedipine protects INS-1 beta-cell from high glucose-induced ER stress and apoptosis. Int J Mol Sci. 2011;12:7569–80.
He X, Bi XY, Lu XZ, Zhao M, Yu XJ, Sun L, et al. Reduction of mitochondria-endoplasmic reticulum interactions by acetylcholine protects human umbilical vein endothelial cells from hypoxia/reoxygenation injury. Arter Throm Vas. 2015;35:1623–34.
Yang B, Xu Y, Hu Y, Luo Y, Lu X, Tsui CK, et al. Madecassic acid protects against hypoxia-induced oxidative stress in retinal microvascular endothelial cells via ROS-mediated endoplasmic reticulum stress. Biomed Pharmacother. 2016;84:845–52.
Boelens J, Lust S, Offner F, Bracke ME, Vanhoecke BW. Review. The endoplasmic reticulum: a target for new anticancer drugs. Vivo. 2007;21:215–26.
Boyce M, Yuan J. Cellular response to endoplasmic reticulum stress: a matter of life or death. Cell Death Differ. 2006;13:363–73.
Healy SJM, Gorman AM, Mousavi-Shafaei P, Gupta S, Samali A. Targeting the endoplasmic reticulum-stress response as an anticancer strategy. Eur J Pharmacol. 2009;625:234–46.
Chitnis N, Pytel D, Diehl JA. UPR-inducible miRNAs contribute to stressful situations. Trends Biochem Sci. 2013;38:447–52.
Chitnis NS, Pytel D, Bobrovnikova-Marjon E, Pant D, Zheng H, Maas NL, et al. miR-211 is a prosurvival microRNA that regulates chop expression in a PERK-dependent manner. Mol Cell. 2012;48:353–64.
Liu F, Liu B, Qian J, Wu G, Li J, Ma Z. miR-153 enhances the therapeutic effect of gemcitabine by targeting Snail in pancreatic cancer. Acta Biochim Biophys Sin (Shanghai). 2017;49:520–9.
Liu R, Shi P, Nie Z, Liang H, Zhou Z, Chen W, et al. Mifepristone suppresses basal triple-negative breast cancer stem cells by down-regulating KLF5 expression. Theranostics. 2016;6:533–44.
Wang Z, Liu C, MiR-153 regulates metastases of gastric cancer through Snail. Tumour Biol. 2015;37:15509–15.
Zeng HF, Yan S, Wu SF. MicroRNA-153-3p suppress cell proliferation and invasion by targeting SNAI1 in melanoma. Biochem Biophys Res Commun. 2017;487:140–5.
Li Q, Song W, Wang W, Yao S, Tian C, Cai X, et al. Suppression of epithelial-mesenchymal transition in hepatocellular carcinoma cells by Kruppel-like factor 4. Oncotarget. 2016;7:29749–60.
Zuo J, Wang D, Shen H, Liu F, Han J, Zhang X, MicroRNA-153 inhibits tumor progression in esophageal squamous cell carcinoma by targeting SNAI1. Tumour Biol. 2016;37:16135–40.
Yang W, Shen Y, Wei J, Liu F. MicroRNA-153/Nrf-2/GPx1 pathway regulates radiosensitivity and stemness of glioma stem cells via reactive oxygen species. Oncotarget. 2015;6:22006–27.
Bartoszewska S, Kochan K, Piotrowski A, Kamysz W, Ochocka RJ, Collawn JF, et al. The hypoxia-inducible miR-429 regulates hypoxia-inducible factor-1alpha expression in human endothelial cells through a negative feedback loop. FASEB J. 2015;29:1467–79.
Cha ST, Chen PS, Johansson G, Chu CY, Wang MY, Jeng YM, et al. MicroRNA-519c suppresses hypoxia-inducible factor-1alpha expression and tumor angiogenesis. Cancer Res. 2010;70:2675–85.
Dai L, Lou W, Zhu J, Zhou X, Di W. MiR-199a inhibits the angiogenic potential of endometrial stromal cells under hypoxia by targeting HIF-1alpha/VEGF pathway. Int J Clin Exp Pathol. 2015;8:4735–44.
Janaszak-Jasiecka A, Bartoszewska S, Kochan K, Piotrowski A, Kalinowski L, Kamysz W, et al. miR-429 regulates the transition between Hypoxia-Inducible Factor (HIF)1A and HIF3A expression in human endothelial cells. Sci Rep. 2016;6:22775.
Kang SG, Lee WH, Lee YH, Lee YS, Kim SG. Hypoxia-inducible factor-1alpha inhibition by a pyrrolopyrazine metabolite of oltipraz as a consequence of microRNAs 199a-5p and 20a induction. Carcinogenesis. 2012;33:661–9.
Wu Z, Cai X, Huang C, Xu J, Liu A. miR-497 suppresses angiogenesis in breast carcinoma by targeting HIF-1alpha. Oncol Rep. 2016;35:1696–702.
Chen X, Iliopoulos D, Zhang Q, Tang Q, Greenblatt MB, Hatziapostolou M, et al. XBP1 promotes triple-negative breast cancer by controlling the HIF1alpha pathway. Nature. 2014;508:103–7.
Paridaens A, Laukens D, Vandewynckel YP, Coulon S, Van Vlierberghe H, Geerts A, et al. Endoplasmic reticulum stress and angiogenesis: is there an interaction between them? Liver Int. 2014;34:e10–18.
Chen C, Sun X, Ran Q, Wilkinson KD, Murphy TJ, Simons JW, et al. Ubiquitin-proteasome degradation of KLF5 transcription factor in cancer and untransformed epithelial cells. Oncogene. 2005;24:3319–27.
Wu J, Ding Y, Chen CH, Zhou Z, Ding C, Chen H, et al. A new oridonin analog suppresses triple-negative breast cancer cells and tumor growth via the induction of death receptor 5. Cancer Lett. 2016;380:393–402.
Acknowledgements
We sincerely thank Professor Lingqiang Zhang from the Academy of Chinese Military Medical Sciences and Professor Wen Liu from Xiamen University for providing reagents. This study was supported by a Strategic Priority Research Program of the Chinese Academy of Sciences (XDA12010303), National Natural Science Foundation of China (31771516, U1502222, U1602221, 81772847, and 81660438), and the Yunnan Applied Basic Research Key Projects (2015FA027).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Liang, H., Xiao, J., Zhou, Z. et al. Hypoxia induces miR-153 through the IRE1α-XBP1 pathway to fine tune the HIF1α/VEGFA axis in breast cancer angiogenesis. Oncogene 37, 1961–1975 (2018). https://doi.org/10.1038/s41388-017-0089-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-017-0089-8
This article is cited by
-
Decoding contextual crosstalk: revealing distinct interactions between non-coding RNAs and unfolded protein response in breast cancer
Cancer Cell International (2024)
-
Endoplasmic reticulum stress: a novel targeted approach to repair bone defects by regulating osteogenesis and angiogenesis
Journal of Translational Medicine (2023)
-
Protein disulfide-isomerase A4 confers glioblastoma angiogenesis promotion capacity and resistance to anti-angiogenic therapy
Journal of Experimental & Clinical Cancer Research (2023)
-
Sevoflurane Confers Protection Against the Malignant Phenotypes of Lung Cancer Cells via the microRNA-153-3p/HIF1α/KDM2B Axis
Biochemical Genetics (2023)
-
The exosome: a review of current therapeutic roles and capabilities in human reproduction
Drug Delivery and Translational Research (2023)