Abstract
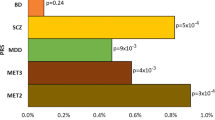
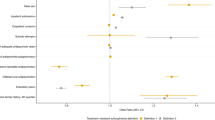
Genomic prediction of antipsychotic dose and polypharmacy has been difficult, mainly due to limited access to large cohorts with genetic and drug prescription data. In this proof of principle study, we investigated if genetic liability for schizophrenia is associated with high dose requirements of antipsychotics and antipsychotic polypharmacy, using real-world registry and biobank data from five independent Nordic cohorts of a total of N = 21,572 individuals with psychotic disorders (schizophrenia, bipolar disorder, and other psychosis). Within regression models, a polygenic risk score (PRS) for schizophrenia was studied in relation to standardized antipsychotic dose as well as antipsychotic polypharmacy, defined based on longitudinal prescription registry data as well as health records and self-reported data. Meta-analyses across the five cohorts showed that PRS for schizophrenia was significantly positively associated with prescribed (standardized) antipsychotic dose (beta(SE) = 0.0435(0.009), p = 0.0006) and antipsychotic polypharmacy defined as taking ≥2 antipsychotics (OR = 1.10, CI = 1.05–1.21, p = 0.0073). The direction of effect was similar in all five independent cohorts. These findings indicate that genotypes may aid clinically relevant decisions on individual patients´ antipsychotic treatment. Further, the findings illustrate how real-world data have the potential to generate results needed for future precision medicine approaches in psychiatry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 13 print issues and online access
$259.00 per year
only $19.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The GWAS summary statistics included in this study are publicly available. Data from the five cohorts can be made available upon request and with appropriate data transfer agreements. Due to national personal data protection regulations, raw data cannot be shared.
References
Yoshida K, Takeuchi H. Dose-dependent effects of antipsychotics on efficacy and adverse effects in schizophrenia. Behav Brain Res. 2021;402:113098.
Denny JC, Collins FS. Precision medicine in 2030-seven ways to transform healthcare. Cell. 2021;184:1415–9.
Njolstad PR, Andreassen OA, Brunak S, Borglum AD, Dillner J, Esko T, et al. Roadmap for a precision-medicine initiative in the Nordic region. Nat Genet. 2019;51:924–30.
Zheutlin AB, Dennis J, Karlsson Linner R, Moscati A, Restrepo N, Straub P, et al. Penetrance and pleiotropy of polygenic risk scores for schizophrenia in 106,160 patients across four health care systems. Am J Psychiatry. 2019;176:846–55.
Wettermark B, Zoega H, Furu K, Korhonen M, Hallas J, Norgaard M, et al. The Nordic prescription databases as a resource for pharmacoepidemiological research-a literature review. Pharmacoepidemiol Drug Saf. 2013;22:691–9.
Tschoner A, Engl J, Laimer M, Kaser S, Rettenbacher M, Fleischhacker WW, et al. Metabolic side effects of antipsychotic medication. Int J Clin Pract. 2007;61:1356–70.
Zadori D, Veres G, Szalardy L, Klivenyi P, Vecsei L. Drug-induced movement disorders. Expert Opin Drug Saf. 2015;14:877–90.
Barton BB, Segger F, Fischer K, Obermeier M, Musil R. Update on weight-gain caused by antipsychotics: a systematic review and meta-analysis. Expert Opin Drug Saf. 2020;19:295–314.
Cohen D, Bonnot O, Bodeau N, Consoli A, Laurent C. Adverse effects of second-generation antipsychotics in children and adolescents: a Bayesian meta-analysis. J Clin Psychopharmacol. 2012;32:309–16.
Ascher-Svanum H, Faries DE, Zhu B, Ernst FR, Swartz MS, Swanson JW. Medication adherence and long-term functional outcomes in the treatment of schizophrenia in usual care. J Clin Psychiatry. 2006;67:453–60.
Novick D, Haro JM, Suarez D, Perez V, Dittmann RW, Haddad PM. Predictors and clinical consequences of non-adherence with antipsychotic medication in the outpatient treatment of schizophrenia. Psychiatry Res. 2010;176:109–13.
Van Sant SP, Buckley PF. Pharmacotherapy for treatment-refractory schizophrenia. Expert Opin Pharmacother. 2011;12:411–34.
Kane JM, Correll CU. The role of clozapine in treatment-resistant schizophrenia. JAMA Psychiatry. 2016;73:187–8.
Wagner E, Siafis S, Fernando P, Falkai P, Honer WG, Roh A, et al. Efficacy and safety of clozapine in psychotic disorders-a systematic quantitative meta-review. Transl Psychiatry. 2021;11:487.
Gallego JA, Bonetti J, Zhang J, Kane JM, Correll CU. Prevalence and correlates of antipsychotic polypharmacy: a systematic review and meta-regression of global and regional trends from the 1970s to 2009. Schizophr Res. 2012;138:18–28.
Lahteenvuo M, Tiihonen J. Antipsychotic polypharmacy for the management of schizophrenia: evidence and recommendations. Drugs. 2021;81:1273–84.
Kinon BJ. The group of treatment resistant schizophrenias. Heterogeneity in treatment resistant schizophrenia (TRS). Front Psychiatry. 2018;9:757.
van Schaik RHN, Muller DJ, Serretti A, Ingelman-Sundberg M. Pharmacogenetics in psychiatry: an update on clinical usability. Front Pharmacol. 2020;11:575540.
Pardinas AF, Smart SE, Willcocks IR, Holmans PA, Dennison CA, Lynham AJ, et al. Interaction testing and polygenic risk scoring to estimate the association of common genetic variants with treatment resistance in schizophrenia. JAMA Psychiatry. 2022;79:260–9.
Paternoster L, Tilling K, Davey Smith G. Genetic epidemiology and Mendelian randomization for informing disease therapeutics: Conceptual and methodological challenges. PLoS Genet. 2017;13:e1006944.
Kappel DB, Legge SE, Hubbard L, Willcocks IR, O'Connell KS, Smith RL, et al. Genomic Stratification of Clozapine Prescription Patterns Using Schizophrenia Polygenic Scores. Biol Psychiatry. 2023;93:149–56.
Campos AI, Mulcahy A, Thorp JG, Wray NR, Byrne EM, Lind PA, et al. Understanding genetic risk factors for common side effects of antidepressant medications. Commun Med. 2021;1:45.
Kiiskinen T, Helkkula P, Krebs K, Karjalainen J, Saarentaus E, Mars N, et al. Genetic predictors of lifelong medication-use patterns in cardiometabolic diseases. Nat Med. 2023;29:209–18.
Word Health Organization. The ICD-10 classification of mental and behavioural disorders: diagnostic criteria for research. Geneva: World Health Organization; 1993.
Nesvag R, Jonsson EG, Bakken IJ, Knudsen GP, Bjella TD, Reichborn-Kjennerud T, et al. The quality of severe mental disorder diagnoses in a national health registry as compared to research diagnoses based on structured interview. BMC Psychiatry. 2017;17:93.
McIntyre RS, Berk M, Brietzke E, Goldstein BI, López-Jaramillo C, Kessing LV, et al. Bipolar disorders. Lancet. 2020;396:1841–56.
Kurki MI, Karjalainen J, Palta P, Sipila TP, Kristiansson K, Donner KM, et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature. 2023;613:508–18.
Tanskanen A, Taipale H, Koponen M, Tolppanen AM, Hartikainen S, Ahonen R, et al. From prescription drug purchases to drug use periods - a second generation method (PRE2DUP). BMC Med Inform Decis Mak. 2015;15:21.
Leitsalu L, Haller T, Esko T, Tammesoo ML, Alavere H, Snieder H, et al. Cohort profile: Estonian Biobank of the Estonian Genome Center, University of Tartu. Int J Epidemiol. 2015;44:1137–47.
Trubetskoy V, Pardinas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Vilhjalmsson BJ, Yang J, Finucane HK, Gusev A, Lindstrom S, Ripke S, et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am J Hum Genet. 2015;97:576–92.
Ge T, Chen CY, Ni Y, Feng YA, Smoller JW. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat Commun. 2019;10:1776.
Genomes Project C, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Leucht S, Samara M, Heres S, Davis JM. Dose equivalents for antipsychotic drugs: the DDD method. Schizophr Bull. 2016;42:S90–4.
WHO. Guidelines for ATC Classification and DDD Assignment 2022. Oslo, Norway: WHO Collaborating Centre for Drug Statistics Methodology; 2021.
Beath K. metaplus: an R package for the analysis of robust meta-analysis and meta-regression. R J. 2016;8:5–16.
Hettige NC, Cole CB, Khalid S, De Luca V. Polygenic risk score prediction of antipsychotic dosage in schizophrenia. Schizophr Res. 2016;170:265–70.
Murray GK, Lin T, Austin J, McGrath JJ, Hickie IB, Wray NR. Could polygenic risk scores be useful in psychiatry?: A review. JAMA Psychiatry. 2021;78:210–9.
Smeland OB, Andreassen OA. Polygenic risk scores in psychiatry - Large potential but still limited clinical utility. Eur Neuropsychopharmacol. 2021;51:68–70.
Lewis ACF, Green RC, Vassy JL. Polygenic risk scores in the clinic: translating risk into action. HGG Adv. 2021;2:100047.
Cai N, Revez JA, Adams MJ, Andlauer TFM, Breen G, Byrne EM, et al. Minimal phenotyping yields genome-wide association signals of low specificity for major depression. Nat Genet. 2020;52:437–47.
Andreassen OA, Hindley G, Frei O, Smeland OB. New insights from the last decade of research in psychiatric genetics- discoveries, challenges and clinical implications. World Psychiatry. 2023;22:4–24.
Zhang JP, Robinson D, Yu J, Gallego J, Fleischhacker WW, Kahn RS, et al. Schizophrenia polygenic risk score as a predictor of antipsychotic efficacy in first-episode psychosis. Am J Psychiatry. 2019;176:21–8.
Frank J, Lang M, Witt SH, Strohmaier J, Rujescu D, Cichon S, et al. Identification of increased genetic risk scores for schizophrenia in treatment-resistant patients. Mol Psychiatry. 2015;20:150–1.
Acknowledgements
This work was partly performed on the TSD (Tjeneste for Sensitive Data) facilities, owned by the University of Oslo, operated and developed by the TSD service group at the University of Oslo, IT-Department (USIT). Computations were also performed on resources provided by UNINETT Sigma2—the National Infrastructure for High Performance Computing and Data Storage in Norway (NS9666S) and the High-Performance Computing Center of University of Tartu. This project has received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No 964874. We also acknowledge support from the Research Council of Norway (296030, 223273), from the US NIMH (grant R01 MH123724), the Estonian Research Council (PRG184), and the Swedish Research Council (grant number 2021-02732). We acknowledge the Estonian Biobank Research Team (Andres Metspalu, Tõnu Esko, Reedik Mägi, Mari Nelis and Georgi Hudjashov). We want to acknowledge the participants and investigators of FinnGen study. The FinnGen project is funded by two grants from Business Finland (HUS 4685/31/2016 and UH 4386/31/2016) and the following industry partners: AbbVie Inc., AstraZeneca UK Ltd, Biogen MA Inc., Bristol Myers Squibb (and Celgene Corporation & Celgene International II Sàrl), Genentech Inc., Merck Sharp & Dohme LCC, Pfizer Inc., GlaxoSmithKline Intellectual Property Development Ltd., Sanofi US Services Inc., Maze Therapeutics Inc., Janssen Biotech Inc, Novartis AG, and Boehringer Ingelheim International GmbH. Following biobanks are acknowledged for delivering biobank samples to FinnGen: Auria Biobank (www.auria.fi/biopankki), THL Biobank (www.thl.fi/biobank), Helsinki Biobank (www.helsinginbiopankki.fi), Biobank Borealis of Northern Finland (https://www.ppshp.fi/Tutkimus-ja-opetus/Biopankki/Pages/Biobank-Borealis-briefly-in-English.aspx), Finnish Clinical Biobank Tampere (www.tays.fi/en-US/Research_and_development/Finnish_Clinical_Biobank_Tampere), Biobank of Eastern Finland (www.ita-suomenbiopankki.fi/en), Central Finland Biobank (www.ksshp.fi/fi-FI/Potilaalle/Biopankki), Finnish Red Cross Blood Service Biobank (www.veripalvelu.fi/verenluovutus/biopankkitoiminta), Terveystalo Biobank (www.terveystalo.com/fi/Yritystietoa/Terveystalo-Biopankki/Biopankki/) and Arctic Biobank (https://www.oulu.fi/en/university/faculties-and-units/faculty-medicine/northern-finland-birth-cohorts-and-arctic-biobank). All Finnish Biobanks are members of BBMRI.fi infrastructure (www.bbmri.fi). Finnish Biobank Cooperative -FINBB (https://finbb.fi/) is the coordinator of BBMRI-ERIC operations in Finland. The Finnish biobank data can be accessed through the Fingenious® services (https://site.fingenious.fi/en/) managed by FINBB.
Author information
Authors and Affiliations
Consortia
Contributions
OA, KO, and EK conceived the study and were involved in study design. EK, SS, AK, MA, GE, and JP conducted analyses. EK drafted the initial manuscript. All authors (including banner authors of the Estonian Biobank Research team and FinnGen) contributed to data interpretation and editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
OAA reported grants from Stiftelsen Kristian Gerhard Jebsen, South-East Regional Health Authority, Research Council of Norway, and European Union’s Horizon 2020 during the conduct of the study; personal fees from cortechs.ai (stock options), Lundbeck (speaker’s honorarium), and Sunovion (speaker’s honorarium) and Janssen (speaker’s honorarium) outside the submitted work. HT reports personal fees from Gedeon Richter, Janssen, Lundbeck and Otsuka, and grants from Eli Lilly and Janssen, outside of the submitted work. No other disclosures were reported.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Koch, E., Kämpe, A., Alver, M. et al. Polygenic liability for antipsychotic dosage and polypharmacy - a real-world registry and biobank study. Neuropsychopharmacol. (2024). https://doi.org/10.1038/s41386-023-01792-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41386-023-01792-0