Abstract
Broad-based cognitive deficits are an enduring and disabling symptom for many patients with severe mental illness, and these impairments are inadequately addressed by current medications. While novel drug targets for schizophrenia and depression have emerged from recent large-scale genome-wide association studies (GWAS) of these psychiatric disorders, GWAS of general cognitive ability can suggest potential targets for nootropic drug repurposing. Here, we (1) meta-analyze results from two recent cognitive GWAS to further enhance power for locus discovery; (2) employ several complementary transcriptomic methods to identify genes in these loci that are credibly associated with cognition; and (3) further annotate the resulting genes using multiple chemoinformatic databases to identify “druggable” targets. Using our meta-analytic data set (N = 373,617), we identified 241 independent cognition-associated loci (29 novel), and 76 genes were identified by 2 or more methods of gene identification. Actin and chromatin binding gene sets were identified as novel pathways that could be targeted via drug repurposing. Leveraging our transcriptomic and chemoinformatic databases, we identified 16 putative genes targeted by existing drugs potentially available for cognitive repurposing.
Similar content being viewed by others
Introduction
One central goal for genome-wide association studies (GWAS) is the identification of potential targets for clinically useful pharmacologic interventions; drugs whose targets have supporting genetic evidence of association to the indication are significantly more likely to successfully reach approval than those without such evidence [1]. While novel drug targets for major psychiatric illnesses have emerged from recent large-scale GWAS [2,3,4], broad-based cognitive deficits are an enduring and disabling feature for many patients with severe mental illness and are inadequately addressed by current medications [5]. Similarly, effective cognitive enhancing medications (“nootropics”) are limited for patients with dementias and other neurodegenerative disorders [6]. Thus, the genetic study of general cognitive ability (GCA) holds the potential for identifying novel targets for nootropic medications, which could have widespread applications [7].
The genetic architecture of GCA has been examined with increasingly large sample sizes over the last few years [8,9,10]. Physical health, illness, mortality [11], and psychiatric traits [12] have shown significant genetic correlations with individual differences in GCA. Dissecting the pleiotropic genetic architectures underlying GCA, educational attainment, and schizophrenia, we have recently shown that neurodevelopmental pathways and adulthood synaptic processes are dissociable etiologic mechanisms relating to genetic liability to psychosis [13].
Nevertheless, identifying specific genes functionally linked to GCA, with protein products that could be targeted by pharmacological agents, remains a core challenge. Using a pathway-based methodology [14], we previously reported that several genes encoding T- and L-type calcium channels, targeted by known pharmaceuticals, were associated with GCA [7]; however, that study was relatively underpowered. Now with much larger GWAS of cognition available [8, 9], increasingly large regions of the genome may demonstrate statistical association with GCA, requiring a principled approach to identify treatment-relevant genes within those regions. Fortunately, the recent release of large-scale brain eQTL/transcriptomic databases have substantially enhanced the assignment of regional GWAS signals to specific genes [15,16,17,18,19,20]. Simultaneously, recent advances in genetic epidemiology methods (e.g. Mendelian randomization) have enabled identification of potentially spurious eQTL associations that may be based on linkage rather than meaningful biology [21]. Thus, the convergence of adequately powered samples coupled with cutting-edge statistical and bioinformatics tools allows for novel genetic mechanisms underlying GCA to be discovered.
In order to turn the resulting GCA-associated gene sets into actionable nootropic drug targets, it is useful to limit the search to those genes which encode proteins that are known to be druggable [22]. However, novel technologies have nominated an increasing number of potentially druggable genes [23,24,25] beyond the 15% of the genome originally estimated based on fundamental pharmacologic principles [22]. Given this uncertainty, well-characterized existing compounds present the most robust evidence of demonstrable druggability for a given target, suggesting the efficiency of drug repurposing/repositioning as a primary research strategy [26]. Such drug repurposing efforts also serve to reduce the high rate of failure associated with novel drug discovery [27]. Applied to GWAS of psychiatric disorders, drug repurposing studies have pointed towards glutamatergic modulators and calcium channel blockers in schizophrenia and sex hormones in depression; such analyses consistently also point towards existing antipsychotic and antidepressant compounds, serving as a positive control for the drug repurposing approach [2, 4, 28, 29].
Here, we jointly analyzed the two largest GWAS of cognition to date [8, 9]. In doing so, we harmonized the genome-wide signals associated with GCA across these studies at the levels of both individual variants and broader genomic regions of loci and pathways. We also employed novel analytical methods not previously employed in cognitive GWAS studies to determine the direction of causality between GWAS hits for GCA and genetically correlated phenotypes. Large brain-based transcriptomic databases were then utilized to determine the biological underpinnings of the most credible and actionable cognitive GWAS signals. Finally, we sought converging evidence from large-scale chemoinformatic resources to identify the most promising nootropic drug targets for drug repurposing.
Materials and methods
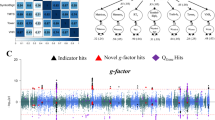
A broad study overview is provided in Fig. 1 and specific algorithm or software carried out in each analysis stage provided in Table S1, respectively. Further in-depth details are available in the ‘Materials and Methods’ section within the Supplementary Materials. The overall data analytic strategy follows a broad strategy of (i) Locus discovery (ii) Gene-based characterization (iii) Gene-to-drug annotations.
The overall data analytic strategy follows a broad strategy of (i) locus discovery, (ii) gene-based characterization, and (iii) gene-to-drug annotations. The green box at top summarizes locus discovery procedures and characterization of results. Yellow box summarizes downstream analysis of summary statistics, resulting in a set of genes available for druggability analysis displayed at figure bottom (red brackets). At each step, location of further details in Tables, Figures, and Supplementary Materials is specified. SMR summary statistics mendelian randomization, FUMA functional mapping and annotation of GWAS, eQTL expression quantitative trait locus, rQTL ribosomal occupancy qtl, sQTL splicing qtl, evQTL expression variation qtl.
Locus discovery
The core analysis combined summary statistics of the two largest GWAS of cognition to date [8, 9]. Savage et al (N = 269,867) analyzed 9,395,118 single-nucleotide polymorphisms (SNPs) for association to intelligence, and Davies et al. [9] (N = 283,531) analyzed 12,871,771 SNPs in relation to the somewhat broader general cognition phenotype. The latter set of summary statistics reported by Davies et al. [9] was reduced from the original N = 300,486 due to data access limitations. First, we carried out MTAG as meta-analysis. As in all genome-wide meta-analytic studies, we expected to gain and lose some loci. Hence, Winner’s Curse Adjustment via the FDR Inverse Quantile Transformation was carried out to evaluate if loci no longer significant were in fact due to winner’s curse. The two studies were noted to have a relatively large degree of sample overlap (89%); the relative increase in power available is more modest than would be expected for a meta-analysis of fully independent samples. While statistical inflation of the combined results is thought to be well-controlled by MTAG [30], the larger than usual sample overlap was not addressed by the original MTAG report. We therefore carried out data simulation in scenarios of 75 and 88.9% sample overlap to extend current understanding of how MTAG might appropriately handle the effects of sample overlap that is relevant to the results reported in the current study. Genome-wide GCA association results were passed through the FUMA v1.3.5 pipeline [31] (see Supplementary Materials: Materials and Methods) to identify GWAS-significant independent SNPs and genomic loci and perform other downstream analyses that we report in subsequent sections. The genome-wide significant threshold was set at P < 5e−8. MTAG results were characterized using genetic correlations with other psychiatric and physical traits via LD-hub [32], and gene property analysis [33] was performed to screen gene expression and localization in CNS tissue. Although many of the results from genetic correlation and gene expression screening are expected to be reported in prior cognitive GWAS [8, 9], they are conducted as “sanity check” to the genome-wide GCA results that are reported here.
Gene-based characterization
In the second phase of the analysis, we utilized a series of approaches to identify potential genes emanating from the MTAG genome-wide results. These ranged from targeted transcriptome wide approaches, to co-located gene expression eQTL mapping, to more general gene-set and pathway-based methods. The objective is to elucidate a broadly inclusive list of genes associated with GCA that could be subjected to downstream gene-drug annotations. First, S-PrediXcan and S-TissueXcan [34] Transcriptome-Wide Analysis (TWAS) were used to characterize gene expression mediated associations in GTEx(v7) tissues. The latter was used to identify potentially functional genes in general, and the former was used to identify genes that are expressed specifically in the brain. We also carried out Summary Statistics Based Mendelian Randomization (SMR) and Heterogeneity in Dependent Instruments (HEIDI) analyses [21] where SNPs are used as instruments to identify gene expression effects on a GCA with estimated SNP-gene expression and SNP-phenotype effects entered into a formal mediation model. At the same time, the HEIDI test identifies SNP-gene expression effects and SNP-phenotype effects that are correlated with each other through LD rather than biologically related via horizontal or vertical pleiotropy. SMR/HEIDI allows us to prioritize genes that might have an immediate relationship between its function and phenotype variance and de-prioritize genes that tend to be in regions with long range LD, which might require further experimental follow up to clarify their function. Next, we carried out molecular quantitative trait locus mapping to identify genomic regions where association signals and gene expression “hot spots” in brain tissue tend to be co-located. Annotation databases that are part of the FUMA pipeline were utilized for this step (See Supplementary Materials: Materials and Methods). Additional molecular annotations from the PsychENCODE database such as splicing variants, expression variation QTL, and ribosomal occupancy QTL were also examined. Gene association tests and gene set analysis were carried out via MAGMA [33]. Results of gene-based tests from MTAG genome-wide results (via MAGMA gene mapping), SMR, and S-TissueXcan were included in gene-set analysis. Genes surviving multiple testing correction in the gene-mapping analysis and nominally significant genes that were part of significant gene sets were selected for downstream drug-gene annotations.
Gene-drug annotations
“Druggable” genes from the Drug-Gene Interaction database (DGIdb v.2), Psychoactive Drug Screening Database KiDB, and a recent review on “druggability” [35] were consolidated. Genes that were identified by transcriptome wide analysis, molecular mapping QTL, and gene-set analysis were filtered on the consolidated druggable gene list from drug databases. At the final stage of the analysis we annotated high confidence genes using the Broad Institute Connectivity Map, Drug Re-purposing Database [36] that provides more in-depth details such as drug names, mechanism of action, and drug indicatons. We matched drug mechanisms of action (MOA) with direction of effects obtained from earlier transcriptome analyses. Genes that are filtered in this manner are deemed as high likelihood for being potential therapeutic targets for nootropic repurposing.
Results
We have prepared a study flowchart (see Fig. 1) as well as a methodological overview and results roadmap (see Table S1) to aid in navigating through the reported results.
Locus discovery: MTAG of general cognitive ability
A total of 8,990,900 SNPs present in both sets of summary statistics were extracted for use in the MTAG meta-analysis of the two largest GWAS of cognition [8, 9]. Since both sets of GWAS summary statistics indexed GCA, we constrained MTAG analysis to give a single output; MTAG was further constrained such that the heritability of both sets of summary statistics were set to equivalent and genetic covariance set to 1 – an approach not unlike fixed effect meta-analysis. Potential inflationary effects of sample overlap, as tested by simulations in scenarios of 75 and 88.9% sample overlap, showed no inflation in either scenario (β75% = 1.011; β88.9% = 1.015) and good agreement between GWAS of the full sample, and MTAG of the GWAS of subsamples with substantial sample overlap (Adj. R275% = 0.94; Adj. R288.9% = 0.96; see Fig. S1). MTAG’s maxFDR analysis revealed low probability of false positives (maxfdr = 4.5e–7). The resulting mean chi-square values after carrying out MTAG were as follows: mean χ2Savage = 1.624, mean χ2Davies = 1.544, and mean χ2MTAG = 1.783. The average projected GWAS equivalent sample size after MTAG analysis was 373,617, which shows substantial power improvement over input GWAS.
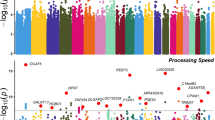
Clumping procedures were carried out on 8,990,900 SNPs (see Methods). Genome-wide significant loci were defined based on R2 > 0.6, and loci within 250 kb of each other were merged. A total of 241 loci were GWAS significant for the MTAG analysis (Fig. 2A–D), while 214 loci and 124 loci were GWAS significant for Savage et al. [8] and Davies et al. [9] respectively (aee Table S2). It should be noted that 38 loci reported as significant in Savage et al. [8] and 8 loci in Davies et al. [9] were no longer significant in the MTAG analysis (Table S3). Winner’s curse analysis suggested that these loci were likely false positives in the original studies (Table S4). A total of 39 MTAG-significant loci were not reported as significant in the input GWAS (Table S3). We then looked up reports that have used multi-trait strategies to enhance power for GCA [10, 13] and found that of the 39 loci, 8 loci were also reported by Hill et al. [10], 1 locus was reported by Lam et al. [13], and 1 locus was reported by both of these studies (Fig S2). Region plots of fully novel loci are reported in Figs. S3–S30.
Genome-wide characterization
A preliminary screen of the genome-wide MTAG via the MAGMA gene-property analysis confirms that the genetic architecture of GCA is closely related to gene expression within various brain tissues after Bonferroni correction (Fig. 2E, F). At the same time, we observe that GCA is also related to genes expressed in prenatal brain tissue implicating a role for early neurodevelopment.
Genetic correlations were estimated between GCA and 855 phenotypes from LD-hub [32] and UK Biobank. MTAG summary statistics were merged and aligned with HapMap3 SNPs excluding the MHC region for genetic correlation analysis (1,190,946 SNPs remained). In total, 297 phenotypes showed significant genetic correlation with cognition at Bonferroni corrected P < 0.05. Consistent with prior reports [7, 10, 37, 38], traits genetically correlated with GCA included education, reproduction, longevity, personality, smoking behavior, anthropometric, brain volume, psychiatric, dementias, lung function, sleep, glycemic, autoimmune, cardio-metabolic, cancer and several ICD-10 medical phenotypes (Table S5). Several novel traits that have not been previously reported to be genetically correlated with GCA are displayed in Fig S3; these include several gastric, vascular, and bronchial diagnoses and medications.
Gene-based characterization
In the next several sections, we report results of a series of analyses that aimed to nominate genes associated with GCA via a series of transcriptomic and pathway-based approaches applied to SNP-based summary statistics obtained in the earlier MTAG analysis. As described (see Fig.1; Table S1, “Gene-Based Characterization”), a variety of complementary transcriptomic analysis were conducted to convert SNP/locus associations into directional, biologically interpretable gene effects on GCA. In the sections below, we take an inclusive approach to generate a broad list of candidate genes which will be subjected to further downstream gene-drug annotations. Moreover, by utilizing a range of complementary functional characterization approaches, the resulting gene list can be prioritized in terms of total strength of supporting evidence (which will be summarized in Table S18).
S-PrediXcan/S-TissueXcan
Genome-wide joint transcriptomic modeling carried out via S-TissueXcan [34] analysis in all 48 GTExv7 tissues yielded 444 significant genes after Bonferroni correction (Table S6). We also conducted brain tissue-specific transcriptome wide modeling via S-PrediXcan and found that 194 genes were significant in one or more brain tissue annotations (Table S7a–k).
SMR and HEIDI
Using Summary-stats-based Mendelian Randomization (SMR [21]), we were able to identify 166 genes that were genome-wide significant (Bonferroni corrected), whose gene expression levels were contributing to variation of GCA (Table S8). As discussed in the Methods and Materials section (Supplementary Materials), SMR analysis tends to be more conservative than other gene identification methodologies [34]. Hence, we also nominated 1212 genes nominally significant genes (PSMR < 0.05) for follow up in the later gene annotation (Table S9). Importantly, SMR analysis indicated that the colocalization of GWAS and eQTL signals in 412 genes was likely artifactual due to linkage disequilibrium, rather than causal mediation, as indicated by PHEIDI < 0.01. These were excluded from subsequent druggability analysis.
Brain-based molecular QTL mapping
Next, we utilized eQTL mapping approaches to identify expressed genes in and around genome-wide significant regions within the MTAG results. We leveraged on databases packaged with FUMA [31] such as BRAINEAC, GTEx, COMMONMIND and XQTLServer for initial eQTL mapping. eQTL mapping from the FUMA [31] pipeline revealed 421 significantly expressed genes within GWAS significant regions (FDR corrected p values; Table S10). Additional molecular QTL mapping of PsychENCODE prefrontal cortex with GCA SNPs identified 592 genes with eQTL (Table S11), 42 genes implicated in ribosomal occupancy QTLs (Table S12), 119 genes with expression variation QTLs (Table S13) and 638 genes with splicing activity QTLs (Tables S7a–S14c).
MAGMA association tests and gene set analysis
MAGMA gene-based analysis revealed that 652 of 18,730 genes were significantly associated with GCA after Bonferroni correction (Table S15). MAGMA gene-set analyses were carried out using gene lists derived from the MAGMA gene-based results, as well as from lists of genes significant in the SMR analysis (only using PsychENCODE results), and S-TissueXcan (Using GTExv7) output. Full gene-set analysis results are presented in Table S16.
Gene set results derived from the MAGMA gene-based results were highly consistent with findings of previous cognitive GWAS: gene sets that have been associated with neuropsychiatric disorders such as schizophrenia and ASD were highly significant, congruent with significant genetic correlations between GCA and these disorders. Relatedly, gene sets reflecting neurodevelopmental processes implicated in schizophrenia and ASD, including the CHD8, FMRP, and RBFOX pathways, were also implicated in GCA [39]. Also consistent with prior reports, a series of neuronal and dendritic development, differentiation, and regulation gene sets were also associated with GCA [10].
There were also several classes of gene sets emerging from our data that are novel with respect to GCA; notably, these results emerged in the context of the SMR (Table 1a) and S-TissueXcan (Table 1b) results, demonstrating the value of leveraging multiple approaches to post-GWAS gene identification. First, genes responsible for cellular response to small molecules such as sugars and cytokines appear to be implicated. Cell signal transductions mediated by small monomeric GTPases also appear to be relevant for GCA. In addition, gene sets underpinning cell structure and binding mechanisms, including adhesion, protein complexes, actin and chromatin binding were identified.
Given the strong signals derived from gene-set analysis, we extracted all nominally significant genes (P < 0.05) within each significant MAGMA gene set for further downstream annotations. A total of 449 genes were identified as part of significant MAGMA pathways (Table S16), and were subjected to further chemoinformatic analysis for druggability as described below.
Identifying drug-gene targets for nootropic re-purposing
A total of 2017 genes (see Fig. 1) were identified via the gene characterization approaches described above. To consolidate evidence for these 2017 nominated genes, we performed the following steps: First, we combined genes identified by MAGMA gene-set and gene-mapping analysis, genes identified via eQTL methods, and genes identified via transcriptome wide analysis (Bonferroni corrected). 695 SMR identified genes, 166 genes identified by S-PrediXcan brain tissue eQTL analysis were added to the annotation. 1876 “druggable” genes were identified by merging across several drug databases (see Supplementary Materials: Materials and Methods, Table S17). After applying a preliminary filter based on three requirements (druggable genes, genes with PHEIDI > 0.01, and genes that were identified by two or more gene-identification approaches), 91 genes remained. It should be noted that gene sets representing methylation processes, DNA complex, and nucleosomes, while significantly associated with GCA, do not contain any genes that are targeted by known drugs, based upon our druggability criteria described in the Materials and Methods section.
We further annotated these genes with information from the Broad Institute CMAP Drug Repurposing Database [36]. We filtered out drug indications for “Oncology” mainly for drug delivery concerns, yielding a final list of 76 “high-confidence” genes that were deemed druggable (Fig. 3). According to the Cell-type Specific Expression Analysis (CSEA) [40], these genes are expressed in a broad range of CNS cell types, including cortical, subcortical, and cerebellar neurons, as well as astrocytes and oligodendrocytes (Table S20); moreover, expression of these genes is broadly demonstrated across developmental epochs (Table S21). These genes were annotated with eQTL directions (i.e., up- or down-regulation associated with higher GCA) for each gene; eQTL directions were obtained from earlier analysis, including brain-eQTLs from S-PrediXcan, SMR, PsychENCODE eQTL, RNA-seq Ribosomal and Splicing eQTL mapping, and overall S-TissueXcan GTEX eQTL analysis (Table S18). Effect sizes that indicated up-regulation of the gene associated with higher GCA were denoted as “↑,” while those where down-regulation was associated with higher GCA were denoted as “↓.” We predicted the “mechanism of action” from the overall eQTL direction to determine if a given gene might require either an “Agonist” or “Antagonist” to enhance GCA. This was achieved by taking the sum of eQTL directions across tissues (see Table S18). If overall eQTL indicates up-regulation, it would more likely require an agonist and vice-versa. We eliminated “Ambiguous” gene targets that have an equal number of tissues that show up- and down-regulated gene expression.
Annotations from the CMAP Drug Re-purposing Database, which include drug names, mechanism of action (MOA), and drug indications, were merged with these “high-confidence” genes (Table S19). Of the 76 “high-confidence” genes, manual curation of the CMAP annotations compared to the predicted mechanism of action based on eQTL directions yielded a final set of 16 most likely targets for drug repurposing; these potential drugs and their current indications with various types of physical or psychiatric conditions are listed in Table 2. Notably, the relationships between some of the drug MOA and gene targets do not always appear to be direct. For example, adrenergic receptor agonists can indirectly activate calcium channels of which CACNA2D2 is a constituent.
Discussion
We report the largest joint GWAS analysis of GCA at an estimated sample size of 373,617. A total of 241 significant genetic loci were identified via MTAG procedures, of which 39 were not reported in the input cognitive GWASs, and 29 were fully novel to this report, despite substantial sample overlap between the two input GWAS. Notably, within these 29 loci are several genes that have been previously associated with psychiatric and brain related phenotypes, as well as underlying biological mechanisms that may underlie severe cognitive deficits (see Figs. S3–S30). Though not exhaustive, the list of such genes includes CTNNA2, LMF1, ZNF536, BAIAP2, and GOSR1, discussed below.
Catenin (Cadherin-Associated Protein), Alpha 2 (CTNNA2), encodes for brain expressed alpha-catenin crucial for synaptic contact, previously implicated with excitement seeking, risk-taking, hyperactivity, substance use, bipolar, and antisocial disorders [41]. CTNNA2 was found to be neuronal-specific, and abundant in the dorsolateral prefrontal cortex and hippocampus in primate brains; CTNNA2 and other catenin genes are involved in folding and lamination of the cerebral cortex [42]. Loss of function mutations within CTNNA2 are thought to be one of the factors underlying pachygyria, characterized by loss of neurite stability and migration [43]. Lipase Maturation Factor 1 (LMF1) lipoproteins are mostly made in the brain; neurons and astrocytes coordinate lipoprotein metabolism within the brain [44] and evidence for lipoprotein lipase activity mediated by LMF genes implicate regulation of brain energy balance underlying cognition [45]. Zinc-Finger 536 (ZNF536) was identified as a candidate schizophrenia gene and a double-knockout zebrafish line shows behavioral and neuroanatomical (decreased forebrain volume) changes [46]. ZNF536 has also been implicated in the maintenance of neural progenitor cells and neuronal differentiation within the prefrontal cortex [47, 48]. Brain-specific angiogenesis inhibitor 1-associated protein 2 (BAIAP2) is a multi-domain scaffolding and adapter protein implicated in the regulation of membrane and actin dynamics at subcellular structures, and is an abundant component of the postsynaptic density at excitatory synapses and an important regulator of actin-rich dendritic spines [49]. BAIAP2 has been shown to be potentially involved in the etiology of attention deficit/hyperactivity disorder [50]. Golgi SNAP receptor complex member 1 (GOSR1) is involved in transport from the ER to the Golgi apparatus as well as in intra-Golgi transport [51]. The gene has been identified in large-scale genetics of brain phenotypes and is also reportedly associated with cognitive traits in follow-on analysis [52]. Similarly, evidence has also been reported that GOSR1 is implicated in architecture of epigenetic and neuronal aging rates in human brain regions [53]. Further follow up is necessary to validate these genes in larger samples and biological experiments.
Gene property analysis revealed significant tissue expression overrepresented in CNS tissue compared with expression in other types of tissues, consistent with earlier reports [7,8,9, 39]. Notably, some of these genes appear significantly expressed during the prenatal state, indicating potential neurodevelopmental impact of genes associated with GCA. We focused on identifying genes associated with GCA that could be “actionable” in terms of identifying pharmacological agents that could be re-purposed for nootropic utilization based on GWAS results. In an earlier study, MAGMA pathway analysis was carried out on drug-based pathway annotations using a smaller GWAS of GCA [7], where we reported several T and L-type calcium channels as potential targets for nootropic agents. Here, we were able to leverage recent novel developments, including newly available brain eQTL data and complementary transcriptomic methodologies, enabling estimation of directionality (i.e., up- vs. down-regulation of expression) of gene effects on cognition. Notably, our study is the first cognitive GWAS to employ HEIDI, an approach that allows pleiotropy (either vertical or horizontal) to be differentiated from spurious associations due to linkage. HEIDI tests against the null hypothesis that a single causal variant affects both gene expression and trait variation, and so HEIDI-significant genes are less likely to be causal and require closer inspection and further biological experiments to unravel any true functional effects of the genes. Therefore, we have filtered gene results based on a nominal threshold of PHEDI < 0.01. Additionally, several novel classes of gene sets, such as cell binding, cell metabolism, and cell structure not previously reported as associated with GCA, created an additional pool of genes available for further investigation. While many of the genes in these novel sets were not easily druggable, identification of methylation and chromatin binding gene sets further highlights the link between GCA, neurodevelopment, and psychopathology; loss of function variants in multiple genes in these sets result in increased risk for schizophrenia, autism, and other neurodevelopmental disorders [54,55,56,57,58].
The most crucial stage of the current report involved the identification of genes that are potential drug targets. Using filtering methods that were detailed earlier, the 76 “high-confidence” druggable genes were selected for further annotation. Of these, 16 genes were identified as “most likely druggable” based on their predicted function from eQTL results and the CMAP Drug Re-purposing database [36]. These selected genes could be further classified into broad gene classes (i) Serotonergic genes, (ii) Carbonic Anhydrase, (iii) Phosphodiesterase, (iv) Ion channel, (v) Glutamatergic/GABA-ergic, and (vi) Others (See Table 2); details of these gene classes are provided in the Extended Discussion in the supplement. Though we have focused the discussion of results explicitly on identifying potential targets for nootropic purposes, the converse could also be relevant—where there might be commonly administered drugs that appear to result in cognitive deficits. Indeed, we note that at least two drugs with known cognitive side effects, topiramate [59] and gabapentin [60], are predicted by our model to cause cognitive deficits.
The results here have generated leads for further investigation into potential drug targets and how they might subserve nootropic repurposing. However, there are limitations to the evidence that we report. First, although this report comprises the largest and most well-powered genome-wide analysis of GCA, there continues to be potential to expand sample size to increase power. The modest increase in novel loci reported in the current study could be accounted for by substantial sample overlap in the earlier GWAS reports. Nevertheless, it is notable that results of MTAG simulation of 75 and 88.9% sample overlap were largely similar to those reported in Turley et al., [30], which used height to show that MTAG was robust to 50% sample overlap. Here, under simulation conditions, we were able to show that MTAG can effectively control for inflation due to largely overlapping samples and is a useful tool for merging and combining summary statistics from large-scale GWAS into a single high-powered set of summary statistics. Relatedly, the present study focused on general cognitive ability, as this broad phenotype has the largest available sample size for GWAS; studies of individual cognitive subdomains (such as memory, processing speed, etc.) tend to be far less powered. Nevertheless, it is worth noting that there is some overlap of our results with prior, smaller studies of individual cognitive domains [61, 62]. As just one example, one of our short list of “most likely druggable” genes, GRIN2A, was previously reported to link schizophrenia with performance on an antisaccade task [63].
Additionally, the transcriptomic reference databases (e.g., PsychENCODE) that we employed are an order of magnitude larger than those previously available, but brain annotations remain somewhat smaller than other QTL annotations (e.g., blood). Second, identifying eQTL for a particular phenotype is challenging– as with most summary statistics approaches, it is not always possible to directly confirm that the proposed eQTL is necessarily leading to variation in the phenotype [64]. Direct experimentation is required to rule out potential extraneous factors that might be pleiotropic to both phenotypic variation and eQTL effects. Third, the issue of LD complexity within a GWAS region makes identification of a gene that is deemed associated with the phenotype challenging [65], further computational and functional work is needed to enhance the precision to which genes are prioritized or identified as truly functional within these genomic regions. Here, we attempted to identify functionally relevant genes by examining the convergence across a range of complementary methodologies to overcome some of the limitations noted above. In addition, we used the HEIDI test to explicitly exclude genes marked by linkage that might be inaccurately labeled as “causal”. Nevertheless, the challenge of regions of extensive LD encompassing many genes should be addressed in future studies, perhaps incorporating recently developed methods for examining three-dimensional properties of the genome [66]. Moreover, we attempted to combine results across multiple QTL tissues and annotations (Table S18), in order to match gene functions with drug MOA; however, we acknowledge that as yet there are no formal methodologies developed to statistically harmonize these signals.
We also observed several counter-intuitive findings with respect to directionality of effects: for example, with respect to carbonic anhydrase inhibition. It is plausible that many molecular functions in the brain observe either a U-shape or inverted U-shape curve, such that effects of up- or down-regulation are not strictly linear. Moreover, the results reported here are with reference to GWAS conducted in the general population and may be more complicated when applied to a disease population. For instance, calcium channel blockers might rescue cognitive impairments in schizophrenia, but blocking calcium channel function in the general population could be detrimental to synaptic function. At the same time, our GWAS cohorts included older adults, and some findings may be a function of cryptic pathologic processes occurring in these apparently normal subjects. Further work is necessary to replicate evidence reported here into disease populations, along with more precise data on biological mechanisms underlying cognitive function to ensure that compounds identified as nootropic are indeed applicable in disease contexts.
Conclusions
We performed the largest genetic analysis for GCA. Aside from identifying 29 fully novel loci in the current study, the effort has included the most well powered analysis for identifying GCA-related genes that are “druggable” and for nominating potential drugs that could be repurposed for nootropic indications. Gene set analysis identified known neurodevelopmental and synaptic related pathways, as well as novel cell structure and binding pathways underlying GCA, which we exploited for drug identification purposes. Utilizing multiple chemoinformatic and drug repurposing databases, along with eQTL and GWAS data, we identified several gene classes contributing to GCA, including serotonergic and glutamatergic/GABA-ergic genes, voltage-gated ion channels, phosphodiesterase components, and carbonic anhydrases. Our efforts show that within these classes, specific drug candidates for nootropic repurposing appear most promising. Further work is necessary to confirm the role of these genes and receptors, to specify their biological mechanisms influencing cognition, and to consider potential CNS effects (including blood-brain barrier permeability) of the putative nootropic compounds nominated by this approach.
Funding and Disclosures
This work has been supported by grants from the National Institutes of Health (R01 MH117646 to TL; R01 MH079800 and P50 MH080173 to AKM; R01 MH080912 to DCG; K23 MH077807 to KEB; K01 MH085812 to MCK). Data collection for the TOP cohort was supported by the Research Council of Norway, South-East Norway Health Authority, and KG Jebsen Foundation. The NCNG study was supported by Research Council of Norway Grants 154313/V50 and 177458/V50. The NCNG GWAS was financed by grants from the Bergen Research Foundation, the University of Bergen, the Research Council of Norway (FUGE, Psykisk Helse), Helse Vest RHF and Dr Einar Martens Fund. The Helsinki Birth Cohort Study has been supported by grants from the Academy of Finland, the Finnish Diabetes Research Society, Folkhälsan Research Foundation, Novo Nordisk Foundation, Finska Läkaresällskapet, Signe and Ane Gyllenberg Foundation, University of Helsinki, Ministry of Education, Ahokas Foundation, Emil Aaltonen Foundation. We thank the Lothian Birth Cohort participants and research team members. We thank the staff from the Wellcome Trust Clinical Research Facility at the Western General Hospital Edinburgh. Phenotype collection in the Lothian Birth Cohort 1921 was supported by the UK Biotechnology and Biological Sciences Research Council (BBSRC; 15/SAG09977), a Royal Society-Wolfson Research Merit Award (to Ian Deary), and The Chief Scientist Office of the Scottish Government (CZH/4/213, CZG/3/2/79, CZB/4/505, ETM/55). Phenotype collection in the Lothian Birth Cohort 1936 was supported by Age UK (The Disconnected Mind project). Genotyping of the cohorts was funded by the BBSRC (BB/F019394/1, 15/S18386). The work was undertaken by The University of Edinburgh Center for Cognitive Ageing and Cognitive Epidemiology (CCACE), part of the cross council Lifelong Health and Wellbeing Initiative (G0700704/84698, MR/K026992/1), for which funding from the BBSRC and Medical
Research Council (MRC) is gratefully acknowledged. WDH is supported by a grant from Age UK (Disconnected Mind Project). The CAMH work was supported by the CAMH Foundation and the Canadian Institutes of Health Research. The Duke Cognition Cohort (DCC) acknowledges K. Linney, J.M. McEvoy, P. Hunt, V. Dixon, T. Pennuto, K. Cornett, D. Swilling, L. Phillips, M. Silver, J. Covington, N. Walley, J. Dawson, H. Onabanjo, P. Nicoletti, A. Wagoner, J. Elmore, L. Bevan, J. Hunkin and R. Wilson for recruitment and testing of subjects. DCC also acknowledges the Ellison Medical Foundation New Scholar award AG-NS-0441-08 for partial funding of this study as well as the National Institute of Mental Health of the National Institutes of Health under award number K01MH098126. The UCLA Consortium for Neuropsychiatric Phenomics (CNP) study acknowledges the following sources of funding from the NIH: Grants UL1DE019580 and PL1MH083271 (RMB), RL1MH083269 (TDC), RL1DA024853 (EL) and PL1NS062410. The ASPIS study was supported by National Institute of Mental Health research grants R01MH085018 and R01MH092515 to Dr. Dimitrios Avramopoulos. Support for the Duke Neurogenetics Study was provided the National Institutes of Health (R01 DA033369 and R01 AG049789 to ARH) and by a National Science Foundation Graduate Research Fellowship to MAS. Recruitment, genotyping and analysis of the TCD healthy control samples were supported by Science Foundation Ireland (grants 12/IP/1670, 12/IP/1359 and 08/IN.1/B1916).
Data access for several cohorts used in this study was provided by the National Center for Biotechnology Information (NCBI) database of Genotypes and Phenotypes (dbGaP). dbGaP accession numbers for these cohorts were:
Cardiovascular Health Study (CHS): phs000287.v4.p1, phs000377.v5.p1, and phs000226.v3.p1
Framingham Heart Study (FHS): phs000007.v23.p8 and phs000342.v11.p8
Multi-Site Collaborative Study for Genotype-Phenotype Associations in Alzheimer’s Disease (GENADA): phs000219.v1.p1
Long Life Family Study (LLFS): phs000397.v1.p1
Genetics of Late Onset Alzheimer’s Disease Study (LOAD): phs000168.v1.p1
Minnesota Center for Twin and Family Research (MCTFR): phs000620.v1.p1
Philadelphia Neurodevelopmental Cohort (PNC): phs000607.v1.p1
The acknowledgment statements for these cohorts are found below:
Framingham Heart Study: The Framingham Heart Study is conducted and supported by the National Heart, Lung, and Blood Institute (NHLBI) in collaboration with Boston University (Contract No. N01-HC-25195 and HHSN268201500001I). This manuscript was not prepared in collaboration with investigators of the Framingham Heart Study and does not necessarily reflect the opinions or views of the Framingham Heart Study, Boston University, or NHLBI. Funding for SHARe Affymetrix genotyping was provided by NHLBI Contract N02-HL-64278. SHARe Illumina genotyping was provided under an agreement between Illumina and Boston University.
Cardiovascular Health Study: This research was supported by contracts HHSN268201200036C, HHSN268200800007C, N01-HC-85079, N01-HC-85080, N01-HC-85081, N01-HC-85082, N01-HC-85083, N01-HC-85084, N01-HC-85085, N01-HC-85086, N01-HC-35129, N01 HC-15103, N01 HC-55222, N01-HC-75150, N01-HC-45133, and N01-HC-85239; grant numbers U01 HL080295 and U01 HL130014 from the National Heart, Lung, and Blood Institute, and R01 AG-023629 from the National Institute on Aging, with additional contribution from the National Institute of Neurological Disorders and Stroke. A full list of principal CHS investigators and institutions can be found at https://chs-nhlbi.org/pi. This manuscript was not prepared in collaboration with CHS investigators and does not necessarily reflect the opinions or views of CHS, or the NHLBI. Support for the genotyping through the CARe Study was provided by NHLBI Contract N01-HC-65226. Support for the Cardiovascular Health Study Whole Genome Study was provided by NHLBI grant HL087652. Additional support for infrastructure was provided by HL105756 and additional genotyping among the African-American cohort was supported in part by HL085251, DNA handling and genotyping at Cedars-Sinai Medical Center was supported in part by National Center for Research Resources grant UL1RR033176, now at the National Center for Advancing Translational Technologies CTSI grant UL1TR000124; in addition to the National Institute of Diabetes and Digestive and Kidney Diseases grant DK063491 to the Southern California Diabetes Endocrinology Research Center.
Multi-Site Collaborative Study for Genotype-Phenotype Associations in Alzheimer’s Disease: The genotypic and associated phenotypic data used in the study were provided by the GlaxoSmithKline, R&D Limited. Details on data acquisition have been published previously in: Li H, Wetten S, Li L, St Jean PL, Upmanyu R, Surh L, Hosford D, Barnes MR, Briley JD, Borrie M, Coletta N, Delisle R, Dhalla D, Ehm MG, Feldman HH, Fornazzari L, Gauthier S, Goodgame N, Guzman D, Hammond S, Hollingworth P, Hsiung GY, Johnson J, Kelly DD, Keren R, Kertesz A, King KS, Lovestone S, Loy-English I, Matthews PM, Owen MJ, Plumpton M, Pryse-Phillips W, Prinjha RK, Richardson JC, Saunders A, Slater AJ, St George-Hyslop PH, Stinnett SW, Swartz JE, Taylor RL, Wherrett J, Williams J, Yarnall DP, Gibson RA, Irizarry MC, Middleton LT, Roses AD. Candidate single-nucleotide polymorphisms from a genome-wide association study of Alzheimer disease. Arch Neurol., Jan;65(1):45-53, 2008 (PMID: 17998437). Filippini N, Rao A, Wetten S, Gibson RA, Borrie M, Guzman D, Kertesz A, Loy-English I,
Williams J, Nichols T, Whitcher B, Matthews PM. Anatomically-distinct genetic associations of
APOE epsilon4 allele load with regional cortical atrophy in Alzheimer’s disease. Neuroimage,
Feb 1;44(3):724-8, 2009. (PMID: 19013250).
Genetics of Late Onset Alzheimer’s Disease Study: Funding support for the “Genetic Consortium for Late Onset Alzheimer’s Disease” was provided through the Division of Neuroscience, NIA. The Genetic Consortium for Late Onset Alzheimer’s Disease includes a genome-wide association study funded as part of the Division of Neuroscience, NIA. Assistance with phenotype harmonization and genotype cleaning, as well as with general study coordination, was provided by Genetic Consortium for Late Onset Alzheimer’s Disease. A list of contributing investigators is available at https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000168.v1.p1
Long Life Family Study: Funding support for the Long Life Family Study was provided by the Division of Geriatrics and Clinical Gerontology, National Institute on Aging. The Long Life Family Study includes GWAS analyses for factors that contribute to long and healthy life. Assistance with phenotype harmonization and genotype cleaning as well as with general study coordination, was provided by the Division of Geriatrics and Clinical Gerontology, National Institute on Aging. Support for the collection of datasets and samples were provided by Multicenter Cooperative Agreement support by the Division of Geriatrics and Clinical Gerontology, National Institute on Aging (UO1AG023746; UO1023755; UO1023749; UO1023744; UO1023712). Funding support for the genotyping which was performed at the Johns Hopkins University Center for Inherited Disease Research was provided by the National Institute on Aging, National Institutes of Health.
Minnesota Center for Twin and Family Research: This project was led by William G. Iacono, PhD. And Matthew K. McGue, PhD (Co-Principal Investigators) at the University of Minnesota, Minneapolis, MN, USA. Co-investigators from the same institution included: Irene J. Elkins, Margaret A. Keyes, Lisa N. Legrand, Stephen M. Malone, William S. Oetting, Michael B. Miller, and Saonli Basu. Funding support for this project was provided through NIDA (U01 DA 024417). Other support for sample ascertainment and data collection came from several grants: R37 DA 05147, R01 AA 09367, R01 AA 11886, R01 DA 13240, R01 MH 66140.
Philadelphia Neurodevelopmental Cohort: Support for the collection of the datasets was provided by grant RC2MH089983 awarded to Raquel Gur, MD, and RC2MH089924 awarded to Hakon Hakonarson, MD, PhD. All subjects were recruited through the Center for Applied Genomics at The Children’s Hospital in Philadelphia.
Funding: Primary support for this study came from the National Institute of Mental Health (R01 MH117646, PI: Lencz).
Software and Online Resources: Multi-Trait Analysis of GWAS (MTAG v1.08) https://github.com/omeed-maghzian/mtag, FDR Inverse Quantile Transformation (FIQT v1.0) https://github.com/bacanusa/FIQT, Functional mapping and annotation of genetic associations (FUMA v1.3.5) https://github.com/Kyoko-wtnb/FUMA-webapp, LD-hub v1.9.3 http://ldsc.broadinstitute.org/, Multi-marker Analysis of GenoMic Annotation (MAGMA v1.07b) https://ctg.cncr.nl/software/magma, S-PrediXcan/S-TissueXcan (v0.6) https://github.com/hakyimlab/MetaXcan; https://github.com/hakyimlab/MetaXcan/wiki, Summary-based Mendelian Randomization) and HEIDI (Heterogeneity in Dependent Instruments) tests (v1.02) https://cnsgenomics.com/software/smr/, LeafCutter (v1.0) https://github.com/davidaknowles/leafcutter, Drug-Gene Interaction database (DGIdb v.2) http://www.dgidb.org/, Psychoactive Drug Screening Database KiDB https://pdsp.unc.edu/databases/kidb.php, Broad Institute Connectivity Map, Drug Re-purposing Database https://www.broadinstitute.org/drug-repurposing-hub
Authors have no competing or conflicts of interests pertaining to the study.
Data availability
Cognitive GWASs utilized in the current manuscript are publicly available at https://ctg.cncr.nl/software/summary_statistics and https://www.ccace.ed.ac.uk/node/335. GWAS summary statistics that are reported in the current manuscript would be made available publicly upon publication of the manuscript. eQTL annotation data and drug annotation data are available in the respective weblinks included in the “Software and Online Resources” section.
References
King EA, Davis JW, Degner JF. Are drug targets with genetic support twice as likely to be approved? Revised estimates of the impact of genetic support for drug mechanisms on the probability of drug approval. PLoS Genet. 2019;15:e1008489.
So H-C, Chau CK-L, Chiu W-T, Ho K-S, Lo C-P, Yim SH-Y, et al. Analysis of genome-wide association data highlights candidates for drug repositioning in psychiatry. Nat Neurosci. 2017;20:1342–9.
Breen G, Li Q, Roth BL, O’Donnell P, Didriksen M, Dolmetsch R, et al. Translating genome-wide association findings into new therapeutics for psychiatry. Nat Neurosci. 2016;19:1392–6.
Lencz T, Malhotra AK. Targeting the schizophrenia genome: a fast track strategy from GWAS to clinic. Mol Psychiatry. 2015;20:820–6.
Green MJ, Girshkin L, Kremerskothen K, Watkeys O, Quidé Y. A systematic review of studies reporting data-driven cognitive subtypes across the psychosis spectrum. Neuropsychol Rev. 2019. https://doi.org/10.1007/s11065-019-09422-7.
Sun M-K. Potential therapeutics for vascular cognitive impairment and dementia. Curr Neuropharmacol. 2018;16:1036–44.
Lam M, Trampush JW, Yu J, Knowles E, Davies G, Liewald DC, et al. Large-scale cognitive GWAS meta-analysis reveals tissue-specific neural expression and potential nootropic drug targets. Cell Rep. 2017;21:2597–613.
Savage JE, Jansen PR, Stringer S, Watanabe K, Bryois J, Leeuw CAde, et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat Genet. 2018;50:912–9.
Davies G, Lam M, Harris SE, Trampush JW, Luciano M, Hill WD, et al. Study of 300,486 individuals identifies 148 independent genetic loci influencing general cognitive function. Nat Commun. 2018;9:2098.
Hill WD, Marioni RE, Maghzian O, Ritchie SJ, Hagenaars SP, McIntosh AM, et al. A combined analysis of genetically correlated traits identifies 187 loci and a role for neurogenesis and myelination in intelligence. Mol Psychiatry. 2019;24:169–81.
Deary IJ, Harris SE, Hill WD. What genome-wide association studies reveal about the association between intelligence and physical health, illness, and mortality. Curr Opin Psychol. 2018;27:6–12.
Hill WD, Harris SE, Deary IJ. What genome-wide association studies reveal about the association between intelligence and mental health. Curr Opin Psychol. 2018;27:25–30.
Lam M, Hill WD, Trampush JW, Yu J, Knowles E, Davies G, et al. Pleiotropic meta-analysis of cognition, education, and schizophrenia differentiates roles of early neurodevelopmental and adult synaptic pathways. Am J Hum Genet. 2019;105:334–50.
Gaspar HA, Breen G. Drug enrichment and discovery from schizophrenia genome-wide association results: an analysis and visualisation approach. Sci Rep. 2017;7:12460.
PsychENCODE Consortium. Revealing the brain’s molecular architecture. Science. 2018;362:1262–3.
Qi T, Wu Y, Zeng J, Zhang F, Xue A, Jiang L, et al. Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood. Nat Commun. 2018;9:2282.
GTEx Consortium, Laboratory, Data Analysis &Coordinating Center (LDACC)—Analysis Working Group, Statistical Methods groups—Analysis Working Group, Enhancing GTEx (eGTEx) groups, NIH Common Fund, NIH/NCI. et al. Genetic effects on gene expression across human tissues. Nature. 2017;550:204–13.
Fromer M, Roussos P, Sieberts SK, Johnson JS, Kavanagh DH, Perumal TM, et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat Neurosci. 2016;19:1442–53.
Ng B, White CC, Klein H-U, Sieberts SK, McCabe C, Patrick E, et al. An xQTL map integrates the genetic architecture of the human brain’s transcriptome and epigenome. Nat Neurosci. 2017;20:1418–26.
Wang D, Liu S, Warrell J, Won H, Shi X, Navarro FCP, et al. Comprehensive functional genomic resource and integrative model for the human brain. Science. 2018;362:eaat8464.
Zhu Z, Zhang F, Hu H, Bakshi A, Robinson MR, Powell JE, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48:481–7.
Hopkins AL, Groom CR. The druggable genome. Nat Rev Drug Discov. 2002;1:727–30.
Sheils TK, Mathias SL, Kelleher KJ, Siramshetty VB, Nguyen D-T, Bologa CG, et al. TCRD and Pharos 2021: mining the human proteome for disease biology. Nucleic Acids Res. 2021;49:D1334–46.
Kozlovskii I, Popov P. Spatiotemporal identification of druggable binding sites using deep learning. Commun Biol. 2020;3:618.
Oprea TI, Bologa CG, Brunak S, Campbell A, Gan GN, Gaulton A, et al. Unexplored therapeutic opportunities in the human genome. Nat Rev Drug Discov. 2018;17:317–32.
Pushpakom S, Iorio F, Eyers PA, Escott KJ, Hopper S, Wells A, et al. Drug repurposing: progress, challenges and recommendations. Nat Rev Drug Discov. 2019;18:41–58.
Scannell JW, Blanckley A, Boldon H, Warrington B. Diagnosing the decline in pharmaceutical R&D efficiency. Nat Rev Drug Discov. 2012;11:191–200.
Ruderfer DM, Charney AW, Readhead B, Kidd BA, Kähler AK, Kenny PJ, et al. Polygenic overlap between schizophrenia risk and antipsychotic response: a genomic medicine approach. Lancet Psychiatry. 2016;3:350–7.
Gaspar HA, Gerring Z, Hübel C, Major Depressive Disorder Working Group of the Psychiatric Genomics Consortium, Middeldorp CM, Derks EM, et al. Using genetic drug-target networks to develop new drug hypotheses for major depressive disorder. Transl Psychiatry. 2019;9:117.
Turley P, Walters RK, Maghzian O, Okbay A, Lee JJ, Fontana MA, et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat Genet. 2018;50:229–37.
Watanabe K, Taskesen E, Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Zheng J, Erzurumluoglu AM, Elsworth BL, Kemp JP, Howe L, Haycock PC, et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics. 2017;33:272–9.
de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11:e1004219.
Barbeira AN, Dickinson SP, Bonazzola R, Zheng J, Wheeler HE, Torres JM, et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat Commun. 2018;9:1825.
Finan C, Gaulton A, Kruger FA, Lumbers RT, Shah T, Engmann J, et al. The druggable genome and support for target identification and validation in drug development. Sci Transl Med. 2017;9:eaag1166.
Corsello SM, Bittker JA, Liu Z, Gould J, McCarren P, Hirschman JE, et al. The Drug Repurposing Hub: a next-generation drug library and information resource. Nat Med. 2017;23:405–8.
Trampush JW, Yang MLZ, Yu J, Knowles E, Davies G, Liewald DC, et al. GWAS meta-analysis reveals novel loci and genetic correlates for general cognitive function: a report from the COGENT consortium. Mol Psychiatry. 2017;22:336–45.
Hagenaars SP, Harris SE, Davies G, Hill WD, Liewald DCM, Ritchie SJ, et al. Shared genetic aetiology between cognitive functions and physical and mental health in UK Biobank (N=112 151) and 24 GWAS consortia. Mol Psychiatry. 2016;21:1624–32.
Hill WD, Davies G, Harris SE, Hagenaars SP, Liewald DC, Penke L, et al. Molecular genetic aetiology of general cognitive function is enriched in evolutionarily conserved regions. Transl Psychiatry. 2016;6:e980–980.
Dougherty JD, Schmidt EF, Nakajima M, Heintz N. Analytical approaches to RNA profiling data for the identification of genes enriched in specific cells. Nucleic Acids Res. 2010;38:4218–30.
Terracciano A, Esko T, Sutin AR, de Moor MHM, Meirelles O, Zhu G, et al. Meta-analysis of genome-wide association studies identifies common variants in CTNNA2 associated with excitement-seeking. Transl Psychiatry. 2011;1:e49.
Smith A, Bourdeau I, Wang J, Bondy CA. Expression of Catenin family members CTNNA1, CTNNA2, CTNNB1 and JUP in the primate prefrontal cortex and hippocampus. Brain Res Mol Brain Res. 2005;135:225–31.
Schaffer AE, Breuss MW, Caglayan AO, Al-Sanaa N, Al-Abdulwahed HY, Kaymakçalan H, et al. Biallelic loss of human CTNNA2, encoding αN-catenin, leads to ARP2/3 complex overactivity and disordered cortical neuronal migration. Nat Genet. 2018;50:1093–101.
Wang H, Eckel RH. What are lipoproteins doing in the brain? Trends Endocrinol Metab. 2014;25:8–14.
Wang H, Eckel RH. Lipoprotein lipase in the brain and nervous system. Annu Rev Nutr. 2012;32:147–60.
Thyme SB, Pieper LM, Li EH, Pandey S, Wang Y, Morris NS, et al. Phenotypic landscape of schizophrenia-associated genes defines candidates and their shared functions. Cell. 2019;177:478–91.e20.
Berto S, Nowick K. Species-specific changes in a primate transcription factor network provide insights into the molecular evolution of the primate prefrontal cortex. Genome Biol Evol. 2018;10:2023–36.
Qin Z, Ren F, Xu X, Ren Y, Li H, Wang Y, et al. ZNF536, a novel zinc finger protein specifically expressed in the brain, negatively regulates neuron differentiation by repressing retinoic acid-induced gene transcription. Mol Cell Biol. 2009;29:3633–43.
Kang J, Park H, Kim E. IRSp53/BAIAP2 in dendritic spine development, NMDA receptor regulation, and psychiatric disorders. Neuropharmacology. 2016;100:27–39.
Bonvicini C, Faraone SV, Scassellati C. Attention-deficit hyperactivity disorder in adults: a systematic review and meta-analysis of genetic, pharmacogenetic and biochemical studies. Mol Psychiatry. 2016;21:872–84.
Tai G, Lu L, Wang TL, Tang BL, Goud B, Johannes L, et al. Participation of the syntaxin 5/Ykt6/GS28/GS15 SNARE complex in transport from the early/recycling endosome to the trans-Golgi network. Mol Biol Cell. 2004;15:4011–22.
Zhao B, Luo T, Li T, Li Y, Zhang J, Shan Y, et al. Genome-wide association analysis of 19,629 individuals identifies variants influencing regional brain volumes and refines their genetic co-architecture with cognitive and mental health traits. Nat Genet. 2019;51:1637–44.
Lu AT, Hannon E, Levine ME, Crimmins EM, Lunnon K, Mill J, et al. Genetic architecture of epigenetic and neuronal ageing rates in human brain regions. Nat Commun. 2017;8:1–14.
Lencz T, Yu J, Khan RR, Carmi S, Lam M, Ben-Avraham D, et al. Novel Ultra-Rare Exonic Variants Identified in a Founder Population Implicate Cadherins in Schizophrenia. MedRxiv. 2020:2020.05.29.20115352.
Singh T, Kurki MI, Curtis D, Purcell SM, Crooks L, McRae J, et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat Neurosci. 2016;19:571–7.
Satterstrom FK, Kosmicki JA, Wang J, Breen MS, De Rubeis S, An J-Y, et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell. 2020;180:568–84.e23.
De Rubeis S, He X, Goldberg AP, Poultney CS, Samocha K, Cicek AE, et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature. 2014;515:209–15.
Vitriolo A, Gabriele M, Testa G. From enhanceropathies to the epigenetic manifold underlying human cognition. Hum Mol Genet. 2019;28:R226–34.
Mula M. Topiramate and cognitive impairment: evidence and clinical implications. Ther Adv Drug Saf. 2012;3:279–89.
Salimzade A, Hosseini-Sharifabad A, Rabbani M. Comparative effects of chronic administrations of gabapentin, pregabalin and baclofen on rat memory using object recognition test. Res Pharm Sci. 2017;12:204–10.
Ibrahim-Verbaas CA, Bressler J, Debette S, Schuur M, Smith AV, Bis JC, et al. GWAS for executive function and processing speed suggests involvement of the CADM2 gene. Mol Psychiatry. 2016;21:189–97.
Debette S, Ibrahim Verbaas CA, Bressler J, Schuur M, Smith A, Bis JC, et al. Genome-wide studies of verbal declarative memory in nondemented older people: the Cohorts for Heart and Aging Research in Genomic Epidemiology consortium. Biol Psychiatry. 2015;77:749–63.
Greenwood TA, Lazzeroni LC, Maihofer AX, Swerdlow NR, Calkins ME, Freedman R, et al. Genome-wide Association of Endophenotypes for Schizophrenia From the Consortium on the Genetics of Schizophrenia (COGS) Study. JAMA Psychiatry. 2019;76:1274–84.
Wainberg M, Sinnott-Armstrong N, Mancuso N, Barbeira AN, Knowles DA, Golan D, et al. Opportunities and challenges for transcriptome-wide association studies. Nat Genet. 2019;51:592–9.
Mancuso N, Freund MK, Johnson R, Shi H, Kichaev G, Gusev A, et al. Probabilistic fine-mapping of transcriptome-wide association studies. Nat Genet. 2019;51:675–82.
Rajarajan P, Borrman T, Liao W, Schrode N, Flaherty E, Casiño C, et al. Neuron-specific signatures in the chromosomal connectome associated with schizophrenia risk. Science. 2018;362:eaat4311.
Acknowledgements
The authors would also like to thank Heiko Runz (Biogen Inc.) for his comments and suggestions to this manuscript.
Author information
Authors and Affiliations
Contributions
ML ran the analyses and drafted the manuscript. ML and TL conceived the idea and designed the study. TL supervised the drafting and analyses reported in the manuscript. All authors read and provided scientific feedback, and participated in finalizing the draft of the manuscript.
Corresponding author
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lam, M., Chen, CY., Ge, T. et al. Identifying nootropic drug targets via large-scale cognitive GWAS and transcriptomics. Neuropsychopharmacol. 46, 1788–1801 (2021). https://doi.org/10.1038/s41386-021-01023-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41386-021-01023-4
This article is cited by
-
Opportunities for use of neuroimaging in de-risking drug development and improving clinical outcomes in psychiatry: an industry perspective
Neuropsychopharmacology (2024)
-
The effects of methylphenidate and atomoxetine on Drosophila brain at single-cell resolution and potential drug repurposing for ADHD treatment
Molecular Psychiatry (2024)
-
Multivariate genetic analysis of personality and cognitive traits reveals abundant pleiotropy
Nature Human Behaviour (2023)
-
The impact of rare protein coding genetic variation on adult cognitive function
Nature Genetics (2023)
-
Collective genomic segments with differential pleiotropic patterns between cognitive dimensions and psychopathology
Nature Communications (2022)