Abstract
Alzheimer’s disease (AD) is the most common cause of dementia. Mitochondrial dysfunction has been widely reported in AD due to its important role in cellular metabolism and energy production. Complex IV (cytochrome c oxidase, COX) of mitochondrial electron transport chain, is particularly vulnerable in AD. Defects of COX in AD have been well documented, but there is little evidence to support the genetic association of the COX-related genes with AD. In this study, we investigated the genetic association between 17 nuclear-encoded COX-related genes and AD in 1572 Han Chinese. The whole exons of these genes were also screened in 107 unrelated AD patients with a high probability of hereditarily transmitted AD. Variants in COX6B1, NDUFA4, SURF1, and COX10 were identified to be associated with AD. An integrative analysis with data of eQTL, expression and pathology revealed that most of the COX-related genes were significantly downregulated in AD patients and mouse models, and the AD-associated variants in COX6B1, SURF1, and COX10 were linked to altered mRNA levels in brain tissues. Furthermore, mRNA levels of Ndufa4, Cox5a, Cox10, Cox6b2, Cox7a2, and Lrpprc were significantly correlated with Aβ plaque burden in hippocampus of AD mice. Convergent functional genomics analysis revealed strong supportive evidence for the roles of COX6B1, COX10, NDUFA4, and SURF1 in AD. As the result of our comprehensive analysis of the COX-related genes at the genetic, expression, and pathology levels, we have been able to provide a systematic view for understanding the relationships of the COX-related genes in the pathology of AD.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is the most prevalent age-related neurodegenerative disease, and is characterized by progressive brain atrophy and neuronal cell loss leading to cognitive impairment [1,2,3]. The deposition of β-amyloid (Aβ) plaques and the accumulation of neurofibrillary tangles formed by hyperphosphorylated tau in the brain were recognized as the main neuropathological hallmarks of AD [3, 4]. It is widely accepted that genetic factors play an important role in the pathogenesis of AD [5, 6]. Over 100 mutations in three main genes, amyloid precursor protein (APP), presenilin 1 (PSEN1), and presenilin 2 (PSEN2) have been reported to cause familial AD [5, 6]. However, familial AD constitutes only 5% of total AD incidence. For the remaining 95%, genome-wide association studies (GWASs) in recent years have identified several AD susceptibility genes in populations of European ancestry [7], and the associations of part of these genes with AD were validated in Chinese populations in our previous study (including APOE, PICALM, BIN1, CLU, and MS4A4E [8]) and others [9]. However, the genes identified by GWAS can only explain about 16% of the total phenotypic variance, indicating there is missing heritability for AD [10]. Therefore, additional AD-related genes and variants are yet to be identified.
The mitochondrion is the center of cellular energy production, and its dysfunction contributes to the pathogenesis of neurodegenerative diseases [11,12,13]. An increasing amount of evidence has demonstrated that mitochondrial defects, including deficiency in oxidative phosphorylation (OXPHOS), increased reactive oxygen species (ROS) level, reduced ATP production, impaired balance of mitochondrial fission and fusion, are early events occurring in the brain of AD patients and animal models [11, 14,15,16,17,18,19,20,21,22,23,24]. On the other hand, the proteins of AD pathological pathways, including APP and Aβ, have been found to target mitochondria and disrupt mitochondrial function [11, 23, 25,26,27]. One recent study has found the enhancement of mitochondrial proteostasis could be helpful in reducing the toxic effect of Aβ [28].
Defects in the complexes of the mitochondrial respiratory chain, particularly those decreasing the efficiency of complex IV (also called cytochrome c oxidase (COX)), have been widely reported in AD patients and animal models [19, 20, 29,30,31,32]. In mammals, the COX is composed of 13 subunits [33, 34], and of these, 3 subunits are encoded by the mitochondrial genome (COX1, COX2, and COX3) and are the catalytic core of COX, while the other 10 subunits (COX4, COX5a, COX5b, COX6a, COX6b, COX6c, COX7a, COX7b, COX7c, and COX8) are nuclear-encoded and are structural subunits surrounding the catalytic core [33, 34]. Furthermore, there is another COX subunit NDUFA4, albeit with some controversy [35], newly identified in a recent study [36]. The expression levels of the nuclear-encoded COX subunits were found to be significantly reduced in brains and platelets of AD patients [20, 30, 37,38,39,40,41,42,43]; whereas, the mitochondrial-encoded 3 subunits were not significantly changed or even increased in AD patients [20, 30]. In addition to the subunits, there are many proteins participating in the assembling of COX, which are all nuclear-encoded proteins and are important regulators for the COX structure and function [33, 34, 44]. Among these assembly factors, mutations in COX10, COX15, SURF1, LRPPRC, SCO1, and SCO2 were identified as contributing to disorders of COX deficiency [44, 45]. One study in an Italian population, albeit with a limited sample size, has identified variants in the COX10 and COX15 genes to be associated with AD [46]. In general, the genetic association between the COX-related genes and AD has seldom been observed in human populations.
Considering the fact that decreased COX activity has been regarded as a characteristic of AD, we hypothesize that variants in the COX-related genes may confer genetic risk to AD. However, as yet there is no clear genetic evidence to support the association between the COX-related genes and AD. In this study, we analyzed variants in 17 COX-related genes (11 subunits and 6 assembly factors) in 713 AD patients and 859 controls from Han Chinese populations to look for their potential association with AD. We also performed an integrative analysis using publically available genetic data, dealing with gene expression and AD pathology, to clarify the potential role of the COX-related genes in AD. We were able to provide evidence supporting an active involvement of the COX-related genes in the development of AD.
Materials and methods
Subjects
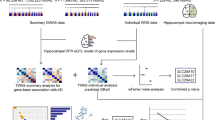
A two-stage case-control study was performed to analyze the association between common variants in the COX-related genes and AD in this study (Fig. 1a). All the samples were of Han Chinese origin and were analyzed for the APOE and other AD-related genes in our previous studies [8, 47,48,49,50]. In brief, the discovery stage (stage I) was based on samples collected in Southwest China from 333 AD cases (62.8% females, age 74.08 ± 8.89 years, 24.4% were APOE ε4 carriers) and 435 healthy individuals (50.6% females, 17.7% were APOE ε4 carriers). The validation stage (stage II) was based on samples collected in East China from 380 AD cases (54.2% females, age 76.18 ± 9.63 years, 38.8% were APOE ε4 carriers) and 424 healthy controls (71.0% females, 17.8% were APOE ε4 carriers).
The strategy for investigating the association between the COX-related genes and AD. a The workflow in this study. b Protein–protein interactions (PPI) of 17 proteins encoded by the COX-related genes. The PPI analysis was performed using the STRING database (https://string-db.org/cgi/input.pl), which presents known and predicted PPI [94]. Two clusters were obtained, one cluster (red nodes) included 11 subunits of COX; the other cluster (blue and green nodes) contained 6 assembly factors of COX
In addition, we took advantage of the exome sequencing data of 107 genetically unrelated AD patients (authors’ unpublished data) that were selected using an extreme phenotype sampling strategy [51], to identify potential disease-causal/rare/functional variants in the exonic regions of the COX-related genes by comparing to 368 control subjects (including 160 in-house non-dementia controls [52], Han Chinese in Beijing (CHB, N = 103) and Southern Han Chinese (CHS, N = 105) from the 1000 Genomes Project phase 3 [53, 54]). These genetically unrelated AD patients (46.7% females, age 64.6 ± 10.29 years, 37.4% were APOE ε4 carriers) either had an early onset age <= 55 years old and/or a positive family history (presence of disease in one or more first-degree family members). Several patients were identified to carry potentially pathogenic mutations in PSEN1 (one patient), PSEN2 (three patients), and GRN (one patient), but no patient had potentially pathogenic mutation in APP and MAPT (authors’ unpublished data). We focused on these patients with early onset AD and/or family history of the disease, instead of sporadic or late-onset AD, as these patients might be more susceptible to genetic influence and be easier to show the potential consequence of the COX-related gene mutations. This sampling strategy was different from that of the IGAP dataset where all cases with age-at-onset <65 years of age were excluded [7], but it could be used as an initial study for identifying potentially pathogenic variants to be validated in sporadic AD patients.
The AD patients were diagnosed by at least two clinical psychiatrists according to the revised National Institute of Neurological and Communicative Disorders and Stroke and the Alzheimer’s Disease and Related Disorders Association (NINCDS-ADRDA) criteria [55,56,57], and Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV). Sample collection complied with the declaration of Helsinki, with informed consent being obtained from all the subjects. This study was approved by the institutional review board of the Kunming Institute of Zoology, Chinese Academy of Sciences.
Genotyping and sequencing of genes encoding subunits and assembly factors of COX
Genomic DNA was extracted from peripheral blood by using AxyPrep Blood Genomic DNA Miniprep Kit (Axygen, USA). Seventeen genes encoding the subunits (11 genes) and assembly factors (6 genes) of the COX were selected to analyze the common variants and variations in the exonic regions (Fig. 1b and Supplementary Table S1). The linkage disequilibrium (LD) pattern of the −10 ~ +10 kb region of the targeted genes were analyzed by using Haploview software [58] to pick up SNPs that are capable of tagging more SNPs in the targeted region. A total of 41 SNPs each with a minor allele frequency (MAF) >10% in the CHB population in the International HapMap Project database (HapMap, http://hapmap.ncbi.nlm.nih.gov/ [59]) were selected (Supplementary Table S1). According to the LD pattern of Asian population in the 1000 Genomes Project phase 3 (http://www.1000genomes.org) [53, 54], these 41 SNPs are capable of tagging a total of 815 SNPs (r2 > 0.8) in the gene region (Supplementary Table S1). The genotyping analysis was performed by using the SNaPshot assay, as described in our previous studies [8, 47, 49, 50]. In brief, multiplex PCR was performed by using 41 pairs of primers (Supplementary Table S2). After cleaning up with SAP and ExoI (TaKaRa Biotechnology Co. Ltd., Dalian, China), the PCR products were subjected to single-base extension reaction by using respective extension primers (Supplementary Table S2) in a total volume of 10 μL containing 4 μL multiplex PCR products, 5 μL SNaPshot Multiplex Ready Reaction Mix and 0.4–0.8 μM pooled SNP-specific oligonucleotide primers (Supplementary Table S2). After purification, a mixture of 4 μL purified products and 6 μL ddH2O was analyzed by the 3730xl DNA analyzer (Applied Biosystems) at the Kunming Biodiversity Large-Apparatus Regional Center, Kunming Institute of Zoology. We used GeneMarker software (http://www.softgenetics.com/GeneMarker.html) to read the genotyping results.
For sequencing the exon and flanking regions of the COX-related genes, we used the next generation sequencing technologies as described in our previous studies [47, 52, 60]. In brief, the paired-end reads were generated on the HiSeq 4000 sequencer (Illumina). High-quality sequence reads were aligned with the reference genome hg19 by Burrows-Wheeler Aligner (BWA, http://bio-bwa.sourceforge.net/) [61] and Samtools (http://samtools.sourceforge.net/) [62, 63]. The Genome Analysis Toolkit (GATK, https://www.broadinstitute.org/gatk/) [64] was applied for SNP calling.
Association analyses of the variants
The frequency of variants was searched in publically available databases: the 1000 Genomes Project phase 3 (http://www.1000genomes.org) [53, 54] and the Asian samples in the Exome Aggregation Consortium (ExAC: http://exac.broadinstitute.org) [65]. Data from the International Genomics of Alzheimer’s Project (IGAP) [7] was retrieved to investigate whether the variants identified in this study are associated with AD in populations of European ancestry. The potential pathogenicity of each variant was estimated by using the Combined Annotation Dependent Depletion (CADD) database [66]. Quanto software [67] was used to evaluate the statistical power of the studied sample. The deviation from the Hardy–Weinberg equilibrium (HWE) for each SNP was calculated using the PLINK software [68]. The LD plot of the identified variants was checked in LocusZoom (http://locuszoom.org/) [69]. The allele and genotype frequencies of all the SNPs in AD patients and controls were analyzed by using the two-tailed Fisher’s exact test.
The association between the genotype and mRNA expression was analyzed by using the available brain expression quantitative trait loci (eQTL) database: BRAINEAC (http://www.braineac.org/) [70], BrainCloud (http://braincloud.jhmi.edu/) [71] and the Genotype-Tissue Expression project (GTEx: http://www.gtexportal.org/home/) [72].
Data-mining of mRNA expression in AD patients
The expression level of the COX-related genes in AD was retrieved from the AlzData database (www.alzdata.org) [73], in which we have integrated and normalized the original microarray data of 684 AD brain tissues and 562 control tissues from Gene Expression Omnibus (GEO: http://www.ncbi.nlm.nih.gov/sites/GDSbrowser) [19, 74,75,76,77,78,79,80,81,82,83,84,85,86]. We used the stage I data as defined in Xu et al. [73] that contains expression data of 269 AD brain tissues and 271 control brain tissues from four brain regions (entorhinal cortex, hippocampus, temporal cortex, and frontal cortex) to investigate the alteration of mRNA levels of the COX-related genes in AD, by using a two-tailed Student’s t-test with the GraphPad Prism software (GraphPad Software, La Jolla, CA, USA). The expression levels of the COX-related genes were further investigated in the stage II data as defined in Xu et al. [73], which contained the microarray data of 415 AD brain tissues and 291 control tissues from temporal cortex and frontal cortex.
Correlation between mRNA levels of the COX-related genes and pathology level in AD mouse models
The spatial-temporal expression data and pathological features of AD transgenic and wide-type (WT) mice were retrieved from Mouseac (www.mouseac.org) [87], including 219 brain tissues from five kinds of AD mouse models with familial AD mutations and 114 brain tissues of age-matched WT mice. The expression levels of the COX-related genes in the hippocampus and cortex tissues from mice at four life stages (2, 4, 8, and 18 months) were compared between AD mouse models and WT mice. The pathology level of Aβ plaques was quantified. The correlation between mRNA levels of the COX-related genes and pathology level was measured using the Pearson correlation test with the GraphPad Prism software (GraphPad Software, La Jolla, CA, USA).
Convergent functional genomics (CFG) for the COX-related genes in AD
A CFG approach was applied to integrate multiple lines of evidence for the association between the COX-related genes and AD [73, 88, 89]. The CFG score for each gene was estimated based on the following evidence: (1) variant(s) in the gene was associated with AD in the combined Han Chinese population (P < 0.05); (2) variant(s) in the gene was associated with AD in the IGAP population (P < 0.05) [7]; (3) the AD-associated SNP(s) was eQTLs that related to altered gene expression in brain (P < 0.05) [70,71,72]; (4) the gene showed an differential expression in AD patients (P < 0.05) [73]; (5) the gene showed an differential expression in AD mice (P < 0.05) [87]; (6) the mRNA level was correlated with AD pathology in AD mice (P < 0.05) [87]. When any of the above mentioned features were present, one point was assigned, leading to a CFG score ranging from 0 to 6 points. Note that the actual inclusion criteria for the above evidence for the CFG analysis were somewhat arbitrary, as the cutoff of significant P-value was set based on the manner of P-value distribution for each line of evidence [73]. For instance, evidence from the current genetic association analysis and from IGAP was given equal weight despite a much higher likelihood of false-positive association in the IGAP data without correction for multiple hypothesis testing [7]. Therefore, some targets with equally weighted significance at single dimension could be false positive. Nonetheless, the truly disease-related gene would be expected to be cross-validated by multi-dimension of evidence, regardless of the level of significance at single-dimension [73].
Results
Genetic variants in the COX-related genes were associated with AD in Han Chinese
In order to perform a comprehensive analysis of the genetic association between the COX-related genes and AD in Chinese, we selected 17 COX-related genes (11 subunit genes and 6 assembly factors) and screened for common variants in a two-stage case-control study of 1572 Han Chinese subjects (Supplementary Table S1). All of the 41 SNPs were in Hardy–Weinberg equilibrium (Supplementary Table S1). The MAF values of these SNPs ranged from 9.2 to 49.3%. The power to detect the odds ratio (OR) value as 1.25 for risk allele was expected to range from 46.22 to 84.33% under a log additive model, with a false positive rate being under 5%.
In the Stage I study of the cohort from Southwest China, SNPs in NDUFA4, SURF1, COX10, COX6B1, COX5B, COX7C, and COX7A2 were associated with genetic risk of AD (Table 1 and Supplementary Table S3). In the Stage II study of an independent Han Chinese population from East China, the trends of the SNPs in NDUFA4, SURF1, COX10, and COX6B1 were validated, although they did not reach a statistically significant level (Table 1 and Supplementary Table S4). As the sample size of both study cohorts were relatively small, we pooled these samples together to increase the statistical power. In the combined Chinese AD cohorts, SNPs in NDUFA4 (rs12671052, OR = 0.746, P = 0.006; rs12667441, OR = 0.738, P = 0.005), SURF1 (rs571549, OR = 1.230, P = 0.007), COX10 (rs9891988, OR = 0.851, P = 0.026; rs2530377, OR = 0.836, P = 0.018; rs11078233, OR = 1.181, P = 0.023) and COX6B1 (rs10420252, OR = 0.802, P = 0.042) remained to be associated with AD (Table 2 and Supplementary Table S5). Considering the fact that the potential population stratification of the Han regional samples [90] might affect this result, the data of the Han Chinese in Beijing (CHB, N = 103) and Southern Han Chinese (CHS, N = 105) were retrieved from 1000 Genomes Project phase 3 [53, 54] and were combined with the Han Chinese controls in this study as a control of the general populations. We found that the association between SNPs in NDUFA4, SURF1, COX10, COX6B1, and AD remained to be significant in the combined population (Table 2). Note that the significance did not survive the Bonferroni test, in part due to the relatively small sample size.
Exons sequencing of the COX-related genes identified variants associated with AD risk
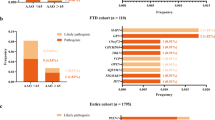
In order to further explore whether there are potential mutations or undetected rare variants in the COX-related genes to be associated with AD, the coding and flanking regions of the COX-related genes were sequenced in 107 AD cases with extreme phenotypes (onset age <= 55 years old and/or with family history of AD), and were compared with a combined control population containing 368 individuals [52,53,54]. The genotyping data from 4327 East Asians from the ExAC (http://exac.broadinstitute.org) [65] was used as another control. A total of 72 variants were identified in the exons and flanking regions of these genes (Table 3 and Supplementary Table S6). Among them, 9 variants in COX6B1, COX6B2, COX10, COX4I1, COX15, and SURF1 showed significantly different frequencies between the cases and the controls (Table 3). Analysis of the LD pattern revealed that these variants were not linked with those AD-associated SNPs identified in the SNaPshot assay (Supplementary Figure S1). Among these variants, rs7991 in the COX6B1 gene was associated with AD risk (Table 3). Interestingly, the association between rs7991 and AD was also observed in the IGAP dataset [7], though the association was not significant after multiple testing correction, suggesting a potential risk effect of this variant in different populations (Table 3). Pathogenicity assessments of these variants by using the CADD database [66] indicated that rs7991 (COX6B1), rs11190255 (COX15), NC_000017.10:g.14110383C>T (COX10) and rs62637580 (SURF1) belong to the top 10% most deleterious variants in the human genome (Table 3). To sum up, through sequencing the coding and flanking regions of the COX-related genes in AD patients with a high probability of hereditarily transmitted AD, we identified several potentially pathogenic variants in the COX-related genes associated with AD risk.
mRNA expression levels of the COX-related genes were significantly decreased in brains of AD patients
Considering the fact that the significantly decreased activity of COX is a characteristic of AD [20, 30, 37,38,39,40,41,42,43], we compared mRNA expression profiles of 269 AD brain tissues and 271 control brain tissues from four brain regions (entorhinal cortex, hippocampus, temporal cortex, and frontal cortex), which were integrated and normalized by the AlzData database (www.alzdata.org) ([73] and references therein), to investigate whether the mRNA expression levels of the COX-related genes were altered in AD brains. Among the 17 analyzed genes, the mRNA levels of 13 genes were significantly decreased in AD patients (Table 4). In particular, we observed a remarkable downregulation of mRNA levels of the COX5B, COX7C, COX7A2, NDUFA4, COX6C, COX8A, COX5A, COX4I1, COX6A1, and COX6B1 genes in the entorhinal cortex and hippocampus tissues of AD patients (Table 4 and Fig. 2), which were the most affected brain regions in AD [2]. Furthermore, the mRNA levels of COX5B, NDUFA4, COX6C, COX8A, COX5A, COX4I1, and COX6B1 were also significantly downregulated in the temporal cortex of AD patients, and COX5B and COX8A were significantly decreased in all four brain tissues of AD patients (Table 4). It is of note that all of the above significantly regulated genes in entorhinal cortex and hippocampus tissues of AD patients were subunits of COX [33], while the mRNA expression level of assembly factors were not consistently altered in AD brains. Further analysis of the Stage II data of the Alzdata database ([73] and references therein) (containing 415 AD brain tissues and 291 control tissues from temporal cortex and frontal cortex) gave similar results and revealed one more gene, COX10, to be significantly decreased in the frontal cortex of AD patients (Supplementary Table S7).
The mRNA levels of the COX-related genes were significantly decreased in entorhinal cortex and hippocampus tissues of AD patients. a–j The mRNA expression data of the COX-related genes in entorhinal cortex and hippocampus tissues were retrieved from the NCBI Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo) and were normalized and analyzed in the AlzData database (www.alzdata.org) [73]. The expression data for the entorhinal cortex came from 39 AD patients and 39 controls in the 4 GEO datasets: GSE26927 [75], GSE48350 [80], GSE26972 [84], and GSE5281 [19]. The expression data for the hippocampus came from 74 AD patients and 65 controls in the 5 GEO datasets: GSE28146 [74], GSE48350 [80], GSE5281 [19], GSE29378 [78], and GSE36980 [76]. The difference between AD patients and controls was quantified by using a two-tailed Student’s t-test. Error bars represent the standard error of the mean. *P-value <0.05; **P-value <0.01; ***P-value <0.001; ****P-value <0.0001
Taken together, though a systematic analysis of mRNA expression profiles of AD brain tissues, we found that most of the subunits of the COX were downregulated at the mRNA levels in AD brain tissues and this was consistent with the reduced COX activity in AD brains as reported in previous studies [20, 37, 39,40,41].
The AD risk SNPs in the COX-related genes were associated with altered gene expression level
Based on the finding that the mRNA levels of the COX-related genes were significantly downregulated in AD brains, we investigated whether the AD risk SNPs in these genes would account for the altered gene expression. Through data-mining of the brain eQTL data from the BRAINEAC database (http://www.braineac.org/) [70], which provides the brain eQTL data across 10 brain tissues of 134 neurological normal individuals, we found that the carriers of allele A of SNP rs10420252 have significantly decreased COX6B1 mRNA expression level in the brain (Table 2 and Fig. 3a). Investigation of the BrainCloud database (http://braincloud.jhmi.edu/; [71]) identified that the AD risk genotype CC of rs10778233 was associated with a reduced COX10 mRNA expression level in prefrontal cortex (Fig. 3b). An analysis of the eQTL data in the CTEx project [72] (GTEx, http://www.gtexportal.org/home/) revealed that the AD risk genotype CC of rs571549 was prominently associated with an increased SURF1 mRNA expression level in hippocampus and cortex tissues (Fig. 3c), which was consistent with the upregulated SURF1 mRNA level in the frontal cortex tissues of AD patients (Table 4 and Fig. 3d). Collectively, these results suggested that variants in the COX-related genes may be associated with AD by affecting gene expression.
The AD-associated SNPs in the COX-related genes were eQTLs and affected mRNA expression in brain tissues. a The carriers of the A allele of SNP rs10420252 showed a significantly decreased COX6B1 mRNA expression level. Data were retrieved from the brain eQTL database BRAINEAC [70], and was analyzed by using a two-tailed Student’s t-test. b Genotype CC of SNP rs10778233 was associated with a decreased COX10 mRNA expression level in cortex. Data were retrieved from the BrainCloud database (http://braincloud.jhmi.edu/) [71] and was analyzed by using a linear regression test. c Genotype CC of SNP rs571549 was prominently associated with an increased SURF1 mRNA expression level in frontal cortex, cortex, and hippocampus. Data were retrieved from the CTEx project (GTEx, http://www.gtexportal.org/home/) [72] and was analyzed by using a linear regression test. d The mRNA expression level of SURF1 was increased in frontal cortex tissues from AD patients relative to controls. The data were retrieved from the 7 datasets of GEO: GSE48350 [80], GSE66333 [81], GSE53890 [77], GSE5281 [19], GSE36980 [76], GSE12685 [86], and GSE15222 [85], and were normalized in the AlzData database (www.alzdata.org) [73]. Error bars represent the standard error of the mean. *P-value <0.05; **P-value <0.01; two-tailed Student’s t-test
The mRNA expression levels of the COX-related genes were correlated with the level of Aβ plaques in AD mouse models
In order to explore whether the mRNA expression levels of the COX-related genes were associated with AD pathology in species other than in man, we retrieved and analyzed the gene expression data and pathology data of AD mouse models from Mouseac (www.mouseac.org) [87]. By analyzing the gene expression levels in hippocampus and cortex tissues of wild type and HO _TASTPM mice (with homogenous APP K670N/M671L and PSEN1M146V mutations) at different life stages, we found that most of the COX-related genes were downregulated in hippocampus at the late life stage of the AD mouse models, though only some of them (Cox6a1, Cox7a2, Cox7c, Surf1, and Lrpprc) reached a significant level (Fig. 4). An analysis of the mRNA expression data in the cortex tissues gave similar results (Supplementary Figure S2). Strong negative correlations between the mRNA expression levels of Lrpprc, Cox7a2, Ndufa4, Cox5a, and Cox10 and the level of Aβ plaques were observed in hippocampus and cortex tissues of mouse models with the APP K670N/M671L mutation, while the Cox6b2 gene exhibited a significantly positive correlation (Fig. 4 and Supplementary Figure S2). Overall, these observations further supported the association between COX-related genes and AD, especially at the pathological level.
The mRNA expression levels of the COX-related genes were correlated with the level of Aβ plaque in hippocampus tissues of AD mouse models. The gene expression data and pathology data were retrieved from Mouseac (www.mouseac.org) [87]. Data shown were the age-related mRNA expression (left) and the correlation between mRNA level and Aβ plaque (right) of each of 15 genes (a–o). Sco1 and Sco2 had no data available in Mouseac [87]. The age-related mRNA expression level was measured in hippocampus tissues of wild type (n = 7) and HO _TASTPM mice (with homogenous APP K670N/M671L and PSEN1M146V mutations, n = 3) at different life stages (2, 4, 8, and 18 months). Error bars represent the standard error of the mean. *P-value <0.05; **P-value <0.01; ****P-value <0.0001; ns, not significant; two-tailed Student’s t-test. The correlation between mRNA level and Aβ plaques in mice with APPK670N/M671L mutation (n = 44) was measured using the Pearson correlation test. A P-value <0.05 was considered as statistically significant, ns: not significant
CFG analysis prioritized COX6B1, COX10, NDUFA4, and SURF1 genes as being associated with AD
In order to identify the highly AD-relevant genes, the COX-related genes were subjected to a CFG analysis [73, 88, 89]. By using multiple lines of evidence (including the genetic association in Han Chinese and in IGAP populations [7], eQTLs [70,71,72], gene expression in AD patients [73] and AD mice, and pathology correlation in AD mice [87]), each gene was scored and ranked (Table 5). COX6B1 and COX10 showed the highest CFG score of 5, followed by NDUFA4 and SURF1 with a CFG score of 4, indicating that these genes were possibly the most significant of the 17 COX-related genes considered to be associated with AD (Table 5).
Discussion
Despite the fact that the downregulation of the COX-related genes and defects of the COX activity have been recognized as characteristics of AD [20, 30, 37,38,39,40,41,42,43], there has been little evidence produced to show the genetic association between the COX-related genes and AD [46], especially in populations of Asian origin. We present here an association study exploring the involvement of the COX-related genes in contributing genetic risk to AD. By screening for the common variants (Tag SNPs) and variations in the exon regions of 17 COX-related genes in Han Chinese populations, followed by an integrative analysis with data of eQTL, expression and pathology, we have been able to show an active role of the COX-related genes in AD. To our knowledge, this is the first extensive study evaluating the association of the COX-related genes with AD.
Among the 11 subunit genes of COX, the COX6A1, COX6B1, and NDUFA4 genes showed associations with AD genetic risk (Table 2). Further analysis of the exon regions in patients with a high probability of hereditarily transmitted AD also identified variants in COX6B1, COX6B2, and COX4I1 to be associated with increased AD risk (Table 3). Most of these genes, except for COX6B2, were prominently downregulated in the entorhinal cortex and hippocampus tissues of AD patients (Table 4 and Fig. 2). NDUFA4, a newly recognized subunit of COX [36], exhibited a similar mRNA expression pattern with the other COX subunits (Fig. 2 and Table 4), and was negatively correlated with Aβ plaque burden in AD mouse models (Fig. 4). The data further supported the previous reports for decreased mRNA levels of nuclear-encoded subunits of COX and impaired COX activity in AD brains [20, 37, 39,40,41]. It is of note that three of the subunit genes, COX6A1, COX6B1, and COX6B2, identified as being risk genes in the genetic analysis were members of COX subunit 6. The mRNA expression levels of COX6A1 and COX6B1 were decreased in AD patients and AD mouse models (Table 4, Figs. 2 and 4). However, eQTL analysis showed that the risk genotype of rs10420252 was associated with an increased COX6B1 mRNA level in AD brain (Fig. 3). Interestingly, COX6B2 exhibited a remarkably increased mRNA expression level in the hippocampus tissues of AD patients and mouse models, and there was a positive correlation between its mRNA expression level and the pathological level of Aβ plaque burden in hippocampus and cortex tissues of AD mouse models (Figs. 2 and 4, and Supplementary Figure S2). The increased mRNA expression level of COX6B2 in AD was in contrast to those of COX6A1 and other COX subunits, suggesting that there might be underlying mechanisms to regulate the balance of expression of COX subunits in AD.
Assembly factors are key regulators for the COX structure and function [33, 34, 44]. Many diseases, including Leigh syndrome, myopathy, encephalopathy, to name a few, are caused by mutations of the COX assembly factors [44]. In this study, we screened variants in LRPPRC, SURF1, SCO1, SCO2, COX15, and COX10, which were reported to be mutational hotspots for the COX-related diseases [44]. Variants in SURF1, COX10, and COX15 were associated with AD in Han Chinese population (Table 2 and Table 3). However, the mRNA levels of the three genes were not consistently altered in AD patients. An increased mRNA expression level of the SURF1 gene was observed in the frontal cortex tissues of AD patients (Table 4 and Fig. 3). Consistently, the risk genotype of rs571549 in the SURF1 gene was found to be associated with a significantly increased mRNA level of SURF1 in frontal cortex, cortex, and hippocampus tissues (Fig. 3). The mRNA expression of COX10 was decreased in the frontal cortex of AD patients (Supplementary Table S7). The mRNA level of this gene was also decreased in the hippocampus and cortex tissues from the AD mouse models, and was negatively correlated with Aβ plaque burden in both hippocampus and cortex tissues (Fig. 4 and Supplementary Figure S2). Accordingly, carriers with the risk genotype of rs11078233 had a decreased mRNA level of COX10 in brain tissues (Fig. 3). All of the data indicated that variants in the COX-related genes may be associated with AD through affecting the mRNA expression.
By using an integrative analysis of mRNA expression data of AD patients ([73] and references therein) and mouse models [87], we obtained a more comprehensive dataset (684 AD brain tissues and 562 control tissues; and the brain expression and pathology data of 44 AD mice) to explore the mRNA expression alterations of the COX-related genes in AD. Almost all of the COX-related genes, except for COX15, SURF1, and COXB2, which have been discussed above, were significantly downregulated in AD patients (Table 4 and Supplementary Table S7). In particular, Cox7a2 and Lrpprc were significantly downregulated in mouse models, and we observed a negative correlation of the mRNA expression levels and Aβ plaque burden levels in hippocampus and cortex tissues (Fig. 4 and Supplementary Figure S2). Although, we found no association of genetic variants in these two genes with AD risk in this study, we could not exclude a possibility of rare functional variants or SNPs in the regulatory regions of these genes that would confer genetic risk to AD. Further investigation is needed to clarify this important issue.
Among all the identified variants in this study, only rs7991 in the COX6B1 gene showed an association with AD in the IGAP dataset [7], but the association could not survive after multiple testing correction. The inconsistence of the associations of other SNPs and genes with AD between our sample and the IGAP dataset [7] could be explained by different genetic structures between populations of Asian and European origins that may have different AD susceptible genes/variants [8]. Note that none of the identified common variants showed an association with AD after Bonferroni correction if we followed a stringent P-value <0.001 (0.05/41), partially because of the limited sample analyzed here. However, the association of these variants with AD could not be simply neglected, as we did identify some variants that could alter gene expression and showed different frequencies between cases and controls. These results provided genetic evidence to further support the critical role of COX in AD as reported by previous studies [20, 30, 37,38,39,40,41,42,43].
Whether defects of mitochondria contribute to the pathology of AD is still under debate [20, 91]. Converging lines of evidence have demonstrated that mitochondrial dysfunction occurs before the development of AD pathology [16, 21, 22, 24, 30, 31, 92]. In the early stages of AD, downregulation of many of the OXPHOS related genes has been observed [19, 30,31,32]. Furthermore, oxidative stress as a result of OXPHOS defects has contributed to neuronal death in AD brains [12, 93]. On the other hand, there are multiple lines of direct evidence showing that mitochondria are the direct target of Aβ, thus the mitochondrial defects observed in AD should be the consequence of Aβ toxicity [23, 25,26,27]. A focused functional characterization of the risk COX-related genes in the context of the above studies is essential to define the roles of mitochondrial genes and mitochondrial dysfunction in AD pathology.
Finally, by using a CFG approach which integrating multiple lines of evidence, we have highlighted COX6B1, COX10, NDUFA4, and SURF1 as highly AD-relevant genes. Taken together, through an extensive association analysis of the COX-related genes at the genetic, expressional, and pathological levels, we have provided comprehensive evidence to support the important role of the COX-related genes in the development of AD. Further studies with independent populations for validation and functional assays are necessary to confirm the present conclusion and to characterize the mechanisms of the COX-related genes in AD pathogenesis.
References
Alzheimer’s Association. 2016 Alzheimer’s disease facts and figures. Alzheimers Dement. 2016;12:459–509.
Huang Y, Mucke L. Alzheimer mechanisms and therapeutic strategies. Cell. 2012;148:1204–22.
Querfurth HW, LaFerla FM. Alzheimer’s disease. N Engl J Med. 2010;362:329–44.
De Strooper B, Karran E. The cellular phase of Alzheimer’s disease. Cell. 2016;164:603–15.
Karch CM, Goate AM. Alzheimer’s disease risk genes and mechanisms of disease pathogenesis. Biol Psychiatry. 2015;77:43–51.
Bertram L, Lill CM, Tanzi RE. The genetics of Alzheimer disease: back to the future. Neuron. 2010;68:270–81.
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013;45:1452–8.
Wang HZ, Bi R, Hu QX, Xiang Q, Zhang C, Zhang DF, et al. Validating GWAS-identified risk Loci for Alzheimer’s disease in Han Chinese populations. Mol Neurobiol. 2016a;53:379–90.
Tan L, Yu JT, Zhang W, Wu ZC, Zhang Q, Liu QY, et al. Association of GWAS-linked loci with late-onset Alzheimer’s disease in a northern Han Chinese population. Alzheimers Dement. 2013;9:546–53.
Ridge PG, Mukherjee S, Crane PK, Kauwe JS. Alzheimer’s disease: analyzing the missing heritability. PLoS ONE. 2013;8:e79771.
Devine MJ, Kittler JT. Mitochondria at the neuronal presynapse in health and disease. Nat Rev Neurosci. 2018;19:63–80.
Lin MT, Beal MF. Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature. 2006;443:787–95.
Mattson MP, Gleichmann M, Cheng A. Mitochondria in neuroplasticity and neurological disorders. Neuron. 2008;60:748–66.
Calkins MJ, Manczak M, Mao P, Shirendeb U, Reddy PH. Impaired mitochondrial biogenesis, defective axonal transport of mitochondria, abnormal mitochondrial dynamics and synaptic degeneration in a mouse model of Alzheimer’s disease. Hum Mol Genet. 2011;20:4515–29.
Cho DH, Nakamura T, Fang J, Cieplak P, Godzik A, Gu Z, et al. S-nitrosylation of Drp1 mediates beta-amyloid-related mitochondrial fission and neuronal injury. Science. 2009;324:102–5.
Du H, Guo L, Yan S, Sosunov AA, McKhann GM, Yan SS. Early deficits in synaptic mitochondria in an Alzheimer’s disease mouse model. Proc Natl Acad Sci USA. 2010;107:18670–5.
DuBoff B, Feany M, Gotz J. Why size matters - balancing mitochondrial dynamics in Alzheimer’s disease. Trends Neurosci. 2013;36:325–35.
Knott AB, Perkins G, Schwarzenbacher R, Bossy-Wetzel E. Mitochondrial fragmentation in neurodegeneration. Nat Rev Neurosci. 2008;9:505–18.
Liang WS, Reiman EM, Valla J, Dunckley T, Beach TG, Grover A, et al. Alzheimer’s disease is associated with reduced expression of energy metabolism genes in posterior cingulate neurons. Proc Natl Acad Sci USA. 2008;105:4441–6.
Mastroeni D, Khdour OM, Delvaux E, Nolz J, Olsen G, Berchtold N, et al. Nuclear but not mitochondrial-encoded oxidative phosphorylation genes are altered in aging, mild cognitive impairment, and Alzheimer’s disease. Alzheimers Dement. 2017;13:510–9.
Pedros I, Petrov D, Allgaier M, Sureda F, Barroso E, Beas-Zarate C, et al. Early alterations in energy metabolism in the hippocampus of APPswe/PS1dE9 mouse model of Alzheimer’s disease. Biochim Biophys Acta. 2014;1842:1556–66.
Reddy PH, Tripathi R, Troung Q, Tirumala K, Reddy TP, Anekonda V, et al. Abnormal mitochondrial dynamics and synaptic degeneration as early events in Alzheimer’s disease: implications to mitochondria-targeted antioxidant therapeutics. Biochim Biophys Acta. 2012;1822:639–49.
Rhein V, Song X, Wiesner A, Ittner LM, Baysang G, Meier F, et al. Amyloid-beta and tau synergistically impair the oxidative phosphorylation system in triple transgenic Alzheimer’s disease mice. Proc Natl Acad Sci USA. 2009;106:20057–62.
Yao J, Irwin RW, Zhao L, Nilsen J, Hamilton RT, Brinton RD. Mitochondrial bioenergetic deficit precedes Alzheimer’s pathology in female mouse model of Alzheimer’s disease. Proc Natl Acad Sci USA. 2009;106:14670–5.
Anandatheerthavarada HK, Biswas G, Robin MA, Avadhani NG. Mitochondrial targeting and a novel transmembrane arrest of Alzheimer’s amyloid precursor protein impairs mitochondrial function in neuronal cells. J Cell Biol. 2003;161:41–54.
Muirhead KE, Borger E, Aitken L, Conway SJ, Gunn-Moore FJ. The consequences of mitochondrial amyloid beta-peptide in Alzheimer’s disease. Biochem J. 2010;426:255–70.
Spuch C, Ortolano S, Navarro C. New insights in the amyloid-Beta interaction with mitochondria. J Aging Res. 2012;2012:324968.
Sorrentino V, Romani M, Mouchiroud L, Beck JS, Zhang H, D’Amico D, et al. Enhancing mitochondrial proteostasis reduces amyloid-beta proteotoxicity. Nature. 2017;552:187–93.
Feldhaus P, Fraga DB, Ghedim FV, De Luca RD, Bruna TD, Heluany M, et al. Evaluation of respiratory chain activity in lymphocytes of patients with Alzheimer disease. Metab Brain Dis. 2011;26:229–36.
Lunnon K, Keohane A, Pidsley R, Newhouse S, Riddoch-Contreras J, Thubron EB, et al. Mitochondrial genes are altered in blood early in Alzheimer’s disease. Neurobiol Aging. 2017;53:36–47.
Manczak M, Park BS, Jung Y, Reddy PH. Differential expression of oxidative phosphorylation genes in patients with Alzheimer’s disease: implications for early mitochondrial dysfunction and oxidative damage. Neuromolecular Med. 2004;5:147–62.
Valla J, Schneider L, Niedzielko T, Coon KD, Caselli R, Sabbagh MN, et al. Impaired platelet mitochondrial activity in Alzheimer’s disease and mild cognitive impairment. Mitochondrion. 2006;6:323–30.
Kadenbach B, Huttemann M. The subunit composition and function of mammalian cytochrome c oxidase. Mitochondrion. 2015;24:64–76.
Soto IC, Fontanesi F, Liu J, Barrientos A. Biogenesis and assembly of eukaryotic cytochrome c oxidase catalytic core. Biochim Biophys Acta. 2012;1817:883–97.
Kadenbach B. Regulation of mammalian 13-subunit cytochrome c oxidase and binding of other proteins: role of NDUFA4. Trends Endocrinol Metab. 2017;28:761–70.
Balsa E, Marco R, Perales-Clemente E, Szklarczyk R, Calvo E, Landazuri MO, et al. NDUFA4 is a subunit of complex IV of the mammalian electron transport chain. Cell Metab. 2012;16:378–86.
Bosetti F, Brizzi F, Barogi S, Mancuso M, Siciliano G, Tendi EA, et al. Cytochrome c oxidase and mitochondrial F1F0-ATPase (ATP synthase) activities in platelets and brain from patients with Alzheimer’s disease. Neurobiol Aging. 2002;23:371–6.
Cardoso SM, Proenca MT, Santos S, Santana I, Oliveira CR. Cytochrome c oxidase is decreased in Alzheimer’s disease platelets. Neurobiol Aging. 2004;25:105–10.
Kish SJ, Bergeron C, Rajput A, Dozic S, Mastrogiacomo F, Chang LJ, et al. Brain cytochrome oxidase in Alzheimer’s disease. J Neurochem. 1992;59:776–9.
Maurer I, Zierz S, Moller HJ. A selective defect of cytochrome c oxidase is present in brain of Alzheimer disease patients. Neurobiol Aging. 2000;21:455–62.
Mutisya EM, Bowling AC, Beal MF. Cortical cytochrome oxidase activity is reduced in Alzheimer’s disease. J Neurochem. 1994;63:2179–84.
Parker WD Jr., Mahr NJ, Filley CM, Parks JK, Hughes D, Young DA, et al. Reduced platelet cytochrome c oxidase activity in Alzheimer’s disease. Neurology. 1994;44:1086–90.
Valla J, Berndt JD, Gonzalez-Lima F. Energy hypometabolism in posterior cingulate cortex of Alzheimer’s patients: superficial laminar cytochrome oxidase associated with disease duration. J Neurosci. 2001;21:4923–30.
Diaz F. Cytochrome c oxidase deficiency: patients and animal models. Biochim Biophys Acta. 2010;1802:100–10.
Koopman WJ, Distelmaier F, Smeitink JA, Willems PH. OXPHOS mutations and neurodegeneration. EMBO J. 2013;32:9–29.
Vitali M, Venturelli E, Galimberti D, Benerini Gatta L, Scarpini E, Finazzi D. Analysis of the genes coding for subunit 10 and 15 of cytochrome c oxidase in Alzheimer’s disease. J Neural Transm. 2009;116:1635–41.
Bi R, Kong LL, Xu M, Li GD, Zhang DF, Li T, et al. The Arc gene confers genetic susceptibility to Alzheimer’s disease in Han Chinese. Mol Neurobiol. 2018;55:1217–26.
Bi R, Zhang W, Yu D, Li X, Wang HZ, Hu QX, et al. Mitochondrial DNA haplogroup B5 confers genetic susceptibility to Alzheimer’s disease in Han Chinese. Neurobiol Aging. 2015;36:1604 e1607–16.
Wang HZ, Bi R, Zhang DF, Li GD, Ma XH, Fang Y, et al. Neprilysin confers genetic susceptibility to Alzheimer’s disease in Han Chinese. Mol Neurobiol. 2016b;53:4883–92.
Zhang DF, Li J, Wu H, Cui Y, Bi R, Zhou HJ, et al. CFH variants affect structural and functional brain changes and genetic risk of Alzheimer’s disease. Neuropsychopharmacology. 2016;41:1034–45.
MacArthur DG, Manolio TA, Dimmock DP, Rehm HL, Shendure J, Abecasis GR, et al. Guidelines for investigating causality of sequence variants in human disease. Nature. 2014;508:469–76.
Wang D, Fan Y, Malhi M, Bi R, Wu Y, Xu M, et al. Missense variants in HIF1A and LACC1 contribute to leprosy risk in Han Chinese. Am J Hum Genet. 2018;102:794–805.
Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, et al. An integrated map of genetic variation from 1,092 human genomes. Nature. 2012;491:56–65.
Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Jack CR Jr., Albert MS, Knopman DS, McKhann GM, Sperling RA, Carrillo MC, et al. Introduction to the recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011;7:257–62.
Khachaturian ZS. Revised criteria for diagnosis of Alzheimer’s disease: National Institute on Aging-Alzheimer’s Association diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011;7:253–6.
McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM. Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology. 1984;34:939–44.
Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21:263–5.
International HapMap Consortium. The International HapMap Project. Nature. 2003;426:789–96.
Zhang DF, Fan Y, Wang D, Bi R, Zhang C, Fang Y, et al. PLD3 in Alzheimer’s disease: a modest effect as revealed by updated association and expression analyses. Mol Neurobiol. 2015;53:4034–45.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics. 2011;27:2987–93.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310–5.
Gauderman WJ. Sample size requirements for matched case-control studies of gene-environment interaction. Stat Med. 2002;21:35–50.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP, et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics. 2010;26:2336–7.
Ramasamy A, Trabzuni D, Guelfi S, Varghese V, Smith C, Walker R, et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat Neurosci. 2014;17:1418–28.
Colantuoni C, Lipska BK, Ye T, Hyde TM, Tao R, Leek JT, et al. Temporal dynamics and genetic control of transcription in the human prefrontal cortex. Nature. 2011;478:519–23.
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45:580–5.
Xu M, Zhang DF, Luo R, Wu Y, Zhou H, Kong LL, et al. A systematic integrated analysis of brain expression profiles reveals YAP1 and other prioritized hub genes as important upstream regulators in Alzheimer’s disease. Alzheimers Dement. 2018;14:215–29.
Blalock EM, Buechel HM, Popovic J, Geddes JW, Landfield PW. Microarray analyses of laser-captured hippocampus reveal distinct gray and white matter signatures associated with incipient Alzheimer’s disease. J Chem Neuroanat. 2011;42:118–26.
Durrenberger PF, Fernando FS, Kashefi SN, Bonnert TP, Seilhean D, Nait-Oumesmar B, et al. Common mechanisms in neurodegeneration and neuroinflammation: a BrainNet Europe gene expression microarray study. J Neural Transm. 2015;122:1055–68.
Hokama M, Oka S, Leon J, Ninomiya T, Honda H, Sasaki K, et al. Altered expression of diabetes-related genes in Alzheimer’s disease brains: the Hisayama study. Cereb Cortex. 2014;24:2476–88.
Lu T, Aron L, Zullo J, Pan Y, Kim H, Chen Y, et al. REST and stress resistance in ageing and Alzheimer’s disease. Nature. 2014;507:448–54.
Miller JA, Woltjer RL, Goodenbour JM, Horvath S, Geschwind DH. Genes and pathways underlying regional and cell type changes in Alzheimer’s disease. Genome Med. 2013;5:48.
Narayanan M, Huynh JL, Wang K, Yang X, Yoo S, McElwee J, et al. Common dysregulation network in the human prefrontal cortex underlies two neurodegenerative diseases. Mol Syst Biol. 2014;10:743.
Berchtold NC, Cribbs DH, Coleman PD, Rogers J, Head E, Kim R, et al. Gene expression changes in the course of normal brain aging are sexually dimorphic. Proc Natl Acad Sci USA. 2008;105:15605–10.
Simpson JE, Ince PG, Minett T, Matthews FE, Heath PR, Shaw PJ, et al. Neuronal DNA damage response-associated dysregulation of signalling pathways and cholesterol metabolism at the earliest stages of Alzheimer-type pathology. Neuropathol Appl Neurobiol. 2016;42:167–79.
Simpson JE, Ince PG, Shaw PJ, Heath PR, Raman R, Garwood CJ, et al. Microarray analysis of the astrocyte transcriptome in the aging brain: relationship to Alzheimer’s pathology and APOE genotype. Neurobiol Aging. 2011;32:1795–807.
Tan MG, Chua WT, Esiri MM, Smith AD, Vinters HV, Lai MK. Genome wide profiling of altered gene expression in the neocortex of Alzheimer’s disease. J Neurosci Res. 2010;88:1157–69.
Berson A, Barbash S, Shaltiel G, Goll Y, Hanin G, Greenberg DS, et al. Cholinergic-associated loss of hnRNP-A/B in Alzheimer’s disease impairs cortical splicing and cognitive function in mice. EMBO Mol Med. 2012;4:730–42.
Webster JA, Gibbs JR, Clarke J, Ray M, Zhang W, Holmans P, et al. Genetic control of human brain transcript expression in Alzheimer disease. Am J Hum Genet. 2009;84:445–58.
Williams C, Mehrian Shai R, Wu Y, Hsu YH, Sitzer T, Spann B, et al. Transcriptome analysis of synaptoneurosomes identifies neuroplasticity genes overexpressed in incipient Alzheimer’s disease. PLoS ONE. 2009;4:e4936.
Matarin M, Salih DA, Yasvoina M, Cummings DM, Guelfi S, Liu W, et al. A genome-wide gene-expression analysis and database in transgenic mice during development of amyloid or tau pathology. Cell Rep. 2015;10:633–44.
Niculescu AB 3rd, Segal DS, Kuczenski R, Barrett T, Hauger RL, Kelsoe JR. Identifying a series of candidate genes for mania and psychosis: a convergent functional genomics approach. Physiol Genom. 2000;4:83–91.
Niculescu AB, Le-Niculescu H. Convergent Functional Genomics: what we have learned and can learn about genes, pathways, and mechanisms. Neuropsychopharmacology. 2010;35:355–6.
Yao YG, Kong QP, Bandelt HJ, Kivisild T, Zhang YP. Phylogeographic differentiation of mitochondrial DNA in Han Chinese. Am J Hum Genet. 2002;70:635–51.
Fukui H, Diaz F, Garcia S, Moraes CT. Cytochrome c oxidase deficiency in neurons decreases both oxidative stress and amyloid formation in a mouse model of Alzheimer’s disease. Proc Natl Acad Sci USA. 2007;104:14163–8.
Leuner K, Hauptmann S, Abdel-Kader R, Scherping I, Keil U, Strosznajder JB, et al. Mitochondrial dysfunction: the first domino in brain aging and Alzheimer’s disease? Antioxid Redox Signal. 2007;9:1659–75.
Xie H, Hou S, Jiang J, Sekutowicz M, Kelly J, Bacskai BJ. Rapid cell death is preceded by amyloid plaque-mediated oxidative stress. Proc Natl Acad Sci USA. 2012;110:7904–9.
von Mering C, Huynen M, Jaeggi D, Schmidt S, Bork P, Snel B. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res. 2003;31:258–61.
Acknowledgements
We thank Ian Logan for helpful comments and language editing of the manuscript. We thank all participants in this study. We thank Miss Hui-Zhen Wang and Li-li Kong, and Mr. Guo-Dong Li and Dong Wang for technical assistance.
Funding:
This study was supported by the National Natural Science Foundation of China (31730037 to Y.-G.Y., 81601124 to R.B., and 81560230 to H.-Y.J.), and Yunnan Province (2015FB180 to W.Z.), the Key Research Program of Frontier Sciences, Chinese Academy of Sciences (QYZDJ-SSW-SMC005 to Y.-G.Y.), and the Project for International Collaboration of the Bureau of International Collaboration, CAS (GJHZ1846 to Y.-G.Y.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Bi, R., Zhang, W., Zhang, DF. et al. Genetic association of the cytochrome c oxidase-related genes with Alzheimer’s disease in Han Chinese. Neuropsychopharmacol 43, 2264–2276 (2018). https://doi.org/10.1038/s41386-018-0144-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41386-018-0144-3
This article is cited by
-
Mitochondrial respiratory chain component NDUFA4: a promising therapeutic target for gastrointestinal cancer
Cancer Cell International (2024)
-
Convergent transcriptomic and genomic evidence supporting a dysregulation of CXCL16 and CCL5 in Alzheimer’s disease
Alzheimer's Research & Therapy (2023)
-
Functional genomics identify causal variant underlying the protective CTSH locus for Alzheimer’s disease
Neuropsychopharmacology (2023)
-
Ndufa4 Regulates the Proliferation and Apoptosis of Neurons via miR-145a-5p/Homer1/Ccnd2
Molecular Neurobiology (2023)
-
Guidelines for bioinformatics of single-cell sequencing data analysis in Alzheimer’s disease: review, recommendation, implementation and application
Molecular Neurodegeneration (2022)