Abstract
Disease activity in Interleukin-10-deficient (Il10−/−) mice, a model for IBD, depends on genetic background and microbiome composition. B6.129P2/JZtm-Il10tm1Cgn (B6-Il10−/−) mice are partially resistant to colitis, whereas mice carrying the Cdcs1C3Bir haplotype on chromosome 3, B6.Cg-Il10tm1CgnMMU3(D3Mit11-D3Mit348)/JZtm (BC-R3-Il10−/−), are susceptible. This study was performed to clarify Cdcs1 and candidate gene effects on the colitogenic potential of hematopoietic cells using bone marrow (BM) and T-cell transfer models. Acute and chronic graft versus host reaction was excluded by high-density genotyping, in vitro and in vivo approaches. BM-chimeras were created with animals housed in two barriers (I and II) with distinct microbiota composition as identified by sequencing. BM-chimeras of all groups developed comparable moderate-to-severe colitis in Barrier I, however, in Barrier II only recipients of BC-R3-Il10−/− BM. Subsequent adoptive T cell transfers pointed to a new subcongenic interval within Cdcs1 affecting their colitogenic potential. Transfers excluded Larp7 and Alpk1 but highlighted Ifi44 as potential candidate genes. In this model-system, colitis development after cell transfer heavily depends on microbiome, though Cdcs1 acts mainly independently in hematopoietic cells. A new subcongenic interval, provisionally named Cdcs1.4, modifies colitogenic T cell function. Within this locus, Ifi44 represents an important candidate gene for colitis expression.
Similar content being viewed by others
Introduction
Inflammatory bowel disease (IBD) summarizes a group of chronic intestinal disorders that predominantly includes ulcerative colitis and Crohn’s disease. The development of inflammation may be dependent on genetic background, microbes and environmental factors.1 Genetic and microbiota analyses were performed in IBD patients and to determine colitis susceptibility in mice.2,3,4,5 Several genetic loci were identified on different chromosomes,6 which suggested an influence of e.g. Proteobacteria, or Actinobacteria.7,8,9,10
Quantitative trait locus (QTL) analysis of mice susceptible to experimental IBD (C3H/HeJBir.129P2-Il10tm1Cgn) and mice mostly resistant (C57BL/6J.129P2-Il10tm1Cgn) highlighted 10 QTLs named cytokine deficiency-induced colitis susceptibility (Cdcs)1 to 10.3,11 The major locus Cdcs1 on chromosome 3 was further fine mapped. Therefore, Il10-deficient subcongenic strains were analyzed in detail and three independent regions (Cdcs1.1, Cdcs1.2 and Cdcs1.3) were identified.12 The impact of Cdcs1.3 (125.6–128.0 Mbp) was found independently in another model for colitis.13 The genetic and expression differences of both groups suggested that Alpha-kinase 1 (Alpk1) and La ribonucleoprotein domain family member 7 (Larp7) are potential candidate genes and modifiers in the immune response during experimental colitis. Furthermore, on the distal part of mouse chromosome 3 (MMU3) outside of Cdcs1.3, the Interferon-induced protein 44 (Ifi44) was identified as a potential candidate gene that may have an impact on immune cell proliferation and host-microbiota interactions.14,15 However, although Larp7, Ifi44 and some other genes were proposed as pivotal for the induction of inflammation, the only two Cdcs1 candidate genes with a proven effect on the colitis phenotype are Vcam1, an endothelial receptor,16 and Alpk1.17 Experiments using bone marrow-derived macrophages (BMDM), dendritic cells (BMDC) and T cells revealed a Cdcs1-dependent response to bacterial ligands as well as T cell hyperresponsiveness in vivo.18
The microbiota of IBD patients show reduced diversity, with decreases in Bacteroidetes and Firmicutes and the presence of E. coli (specifically AIEC).19 Consistent with this finding, Il10-deficient mice housed in germfree (GF) conditions did not develop colitis, whereas mice housed in specified pathogen free (SPF) conditions or in conventional conditions developed inflammation dependent on the microbial environment.20,21,22 Until now, a number of pathogens have been suggested to be associated with IBD, such as Yersinia enterocolitica and Mycobacterium avium subsp. paratuberculosis.23,24,25,26 Furthermore, the introduction of Enterococcus faecalis, Helicobacter spp. or Bilophila wadsworthia triggers inflammation.27,28,29 In contrast, the introduction of other bacterial strains, such as Lactobacillus or Bifidobacterium improve colitis development.30,31
This study demonstrates the impact of microbiome on the hematopoietic compartment in the Il10−/− mouse model with distinct effects on Cdcs1 mediated susceptibility. Furthermore, for T cell-dependent disease progression, the candidate genes Larp7 and Alpk1 were excluded but a new region, provisionally named Cdcs1.4, was highlighted as relevant. Within this genetic interval, Ifi44 was identified as a potential gene that modifies the outcome of T cell-induced experimental IBD.
Materials and methods
Mice
C57BL/6J.129P2-Il10tm1Cgn (B6-Il10−/−), C3H/HeJBir.129P2-Il10tm1Cgn (C3Bir-Il10−/−), B6.Cg-Il10tm1Cgn MMU3 (D3Mit49-D3Mit348)/JZtm (BC-R2-Il10−/−), B6.Cg-Il10tm1Cgn MMU3 (D3Mit11-D3Mit19)/JZtm (BC-R3-Il10−/−), B6.Cg-Il10tm1Cgn MMU3 (D3Mia129.3-D3Mit17)/JZtm (BC-R4-Il10−/−), B6.Cg-Il10tm1Cgn MMU3 (D3Mia126.1-D3Mit19)/JZtm (BC-R8-Il10−/−), C57BL/6N-Alpk1em2Wtsi (Alpk1−/−), C57BL/6NCrl (B6N) and C57BL/6J-Rag1tm1Mom/JZtm (B6-Rag1−/−) mice were obtained from the Central Animal Facility (Hannover Medical School, Hannover, Germany). This study was conducted in accordance with German law for animal protection and with the European Directive 2010/63/EU. All experiments were approved by the Local Institutional Animal Care and Research Advisory committee and permitted by the local government (No 10/0259; 15/2014; 17/2561). The animals were bred in two different hygienic areas (Barrier I and II). Barrier I is defined as an experimental SPF area that scientists can access, whereas Barrier II is an exclusive breeding area.
Both barriers are routinely monitored according to FELASA recommendations 51 and did not reveal any evidence of infectious agents except for occasional positive tests for Helicobacter species and Helicobacter hepaticus in Barrier I.
Genotyping
Genotyping of the genetic background of strains B6-Il10−/−, C3Bir-Il10−/− and BC-R2-Il10−/−, BC-R3-Il10−/−, BC-R4-Il10−/− and BC-R8-Il10−/− as well as characterization of the congenic fragment was performed using a MegaMUGA Array (Neogene Corporation, Lansing, Michigan) containing > 77,000 SNP markers. The genotyping results were analyzed using a database on MySQL Server Version 5.5.49-0ubuntu0.14.04.1, Webserver Apache/2.4.7 (Ubuntu), Datenbankclient libmysql Version 5.5.49, phpMyAdmin Version 4.0.10deb1. The positions of the SNP markers were estimated using GRC38/mm10.
Mixed lymphocyte culture (MLC)
Spleens from B6-Il10−/−, BC-R2-Il10−/− and BC-R3-Il10−/− mice were removed, and a single-cell suspension was prepared. Red blood cells were lysed, and the cells were suspended in RPMI supplemented with 10% FCS, 50 U/mL penicillin, 50 µg/mL streptomycin, 4 mM L-glutamine, 1 mM sodium pyruvate, and 0.05 mM β-mercaptoethanol. For the induction of proliferative responses, 105 irradiated (30 Gy) spleen cell suspensions/100 µl (stimulator cells) were mixed with 105 lymph node cells per 100 µl (responder cells) in triplicate wells of Costar 96-well, round-bottom plates and cultured at 37 °C with 5% CO2 for 72 h. In parallel, triplicates of 105 responder cells were cultured for 72 h without stimulator cells. Triplicates of 105 responder cells were also stimulated with Concanavalin A (2 µg/mL) for 48 h. Subsequently, 37 KBq H-3 thymidine/50 µl (Perkin Elmer, Waltham, Massachusetts, USA) was added per well, and the plates were incubated for an additional 16 h. To determine the extent of induced cell divisions, we harvested cell nuclei after the respective incubation periods with a FilterMate (Perkin Elmer LAS, Germany) and assessed incorporation of H-3 thymidine by liquid scintillation counting (Microbeta).
Bone marrow chimeras
Donor mice were euthanized by CO2 inhalation followed by cervical dislocation. BM was isolated from the tibia and femur by flushing with RPMI (Biochrom AG, Berlin, Germany). The cell suspension was passed through a 70-µm cell strainer (BD Biosciences, San Jose, USA) and washed twice with RPMI. The cell number was determined and adjusted to 1 × 108/mL in PBS. Six- to seven-week-old recipient mice were lethally irradiated (9 Gy) using a Synergy linear accelerator (Elekta, Stockholm, Sweden) and were intravenously injected with 1 x 107 donor BM cells. Recipient mice received 5 mL Cotrim K-ratiopharm®/l via drinking water for 1 week (240 mg/5 mL syrup, Ratiopharm GmbH, Ulm, Germany). The animals had a recovery period of 3 weeks and an additional period of 4 weeks for the induction of colitis via hematopoietic cells.
Cell Sorting
Single cell suspensions of splenocytes were achieved using a 40 μm cell strainer (Greiner Bio-One), and then the red blood cell lysis spleen cells were washed twice in MACS buffer and counted with a Scil Vet abcTM hematology analyzer (Scil Animal Care Company, Viernheim, Germany). For cell sorting, 10 × 107 cells were incubated with 100 μl of antibody mix (anti CD4-APC, anti CD8-PE-Cy7 both from Biolegend and anti CD62L-PE, anti B220-VioBlue (VB) from Miltenyi Biotec GmbH). Cells were stained for 20 min with the respective antibodies, washed and then resuspended in MACS buffer with an end concentration between 30 × 106 and 50 × 106 cells per mL. Cell sorting was performed at the Cell Sorting Core Facility of the MHH using FACSAria Fusion or a FACSAria IIu, (Becton Dickinson). Cells were selected using the positive markers CD4-APC and CD62L-PE and negative markers B220-VB and CD8-PE-Cy7. The purity of the sorted cells was determined for each sample, and it was over 95%. Isolated CD4+ CD62L+ T cells were used for RNA and protein isolation or for T cell transfer experiments.
Transfer colitis
A total of 250,000 isolated naive CD4+ CD62L+ T cells from B6-Il10−/− and BC-R2-Il10−/−, BC-R3-Il10−/−, BC-R4-Il10−/− and BC-R8-Il10−/− mice as well as B6N and Alpk1−/− mice were transferred into B6-Rag1−/− mice. Four weeks (BC-R2-Il10−/−, BC-R3-Il10−/− mice), 6-7 weeks (BC-R4-Il10−/− and BC-R8-Il10−/− mice) or 8 weeks (B6N and Alpk1−/− mice) after the transfer, colon samples were obtained for histological, flow cytometry and qPCR analyses.
Magnetic resonance imaging
Intestinal MR images were acquired using a high-field, small-animal scanner (PharmaScan 70/16, Bruker BioSpin, Rheinstetten, Germany). A dedicated rat brain coil was used as a whole-body mouse coil for image acquisitions. Multislice multiecho (MSME) MR sequences were utilized. A MSME sequence (TR 2658 ms and TE 11, 22, 33 ms, flip angle [FA] 180°) with a slice thickness of 0.8 mm was used to generate high-resolution coronal images (2D matrix: 256 × 256, pixel size: 0.117 × 0.195 mm). A T2W fat-suppression setting was chosen since both wall thickness and edema can be assessed in the same image (for MRI parameters.52)
Image analysis
Image analysis was performed using ImageJ (Wayne Rasband). To evaluate the severity of colitis, we determined colon wall thickness as described earlier.52
Histology
During the necropsy of the colon, the head, liver, bone marrow and abdominal wall were obtained. Samples were fixed in neutral buffered 4% formalin, embedded in paraffin, sectioned at 5–6 µm, and stained with hematoxylin and eosin. The hind legs and head were additionally decalcified using Userapid® (Medite GmbH, Burgdorf, Germany) solution for three days. The hind leg was cut longitudinally into halves to reveal the bone marrow cavity of the femur and tibia. The head was cut transversally to subsequently obtain the skin tissue and mucosal surfaces on one slide. Histology slides were blindly scored based on previously described colitis scores with histopathologic lesions graded separately for the proximal, middle and distal colon.53,54 Skin and mucosal surfaces in the abdominal wall and head were investigated, especially for vacuolar degeneration on cells in the stratum basale and stratum spinosum, hyperkeratosis and thickened dermis.36
Microbiota analysis
The cecum content of healthy individuals (B6-Il10−/− and BC-R3-Il10−/− mice) was obtained from different cages under sterile conditions, immediately snap frozen and stored at −80 °C until further processing.
DNA extraction, amplification of the V1-V2 region of the 16S rRNA gene and sequencing on a MiSeq Illumina platform was performed as previously described.55 In brief, after sequencing, paired-end raw reads were then merged and quality filtered yielding 63.065 ± 20.069 sequences per sample that were subsequently directly assigned a taxonomic affiliation down to the genus level using RDP’s naive Bayesian classifier.57 All subsequent analyses were performed on the genus level. Diversity indices were calculated from data that were rarified to equal depth (38,396 sequences) using the function rarefy_even_depth (rngseed = TRUE) from the R phyloseq package.58 Sequences that were not reliably assigned a genus by the classifier were binned into higher-order taxonomy. Non-metric multidimensional scaling analysis was completed in R (package: vegan) on percentage normalized and square-root transformed data.
Flow cytometry
The large intestine was removed and flushed well with PBS. The colon was opened longitudinally, transferred to buffer 1 (HBSS, 3.5% FCS and 100 mM DTT) and incubated at 37 °C for 20 min Next, buffer 1 was discarded, and the intestine was incubated in buffer 2 (HBSS, 3.5% FCS and 0.5 M EDTA) at 37 °C for 15 min, the incubation was repeated twice. The remaining tissue was treated with buffer 3 (RPMI, 10% FCS, 5 mg/mL DNase and 125 U/mg Collagenase D) for 60 min The cell suspensions were pooled, centrifuged, washed and resuspended in 40% Percoll solution. A 70% Percoll solution was carefully overlaid with the cell suspensions and centrifuged for 20 min at 850 rcf. The cells in interphase were collected, washed twice and resuspended in T cell medium (DMEM, 10% FCS, 1% penicillin/streptomycin and 50 μM β-mercaptoethanol) to a density of 4 × 106 cells/mL and seeded in a flat-bottom, 96-well plate (Greiner bio one). Seeded cells were incubated for 90 min at 37 °C with 5% CO2 together with a cell stimulation cocktail (eBioscience, 1:500) according to the manufacturer's instructions. After the addition of Brefeldin A (eBioscience, 1:1000) to each well, another incubation period of 90 min followed. Subsequently, the cells were thoroughly resuspended, and surface and intracellular staining was performed. Up to 1 × 106 cells per well were incubated with an antibody mix for 20 min (anti-CD90-APC-Cy7 and anti-CD4-Viogreen). Next, the cells were washed twice and treated with a True-Nuclear™ Transcription Factor Buffer set (BioLegend, San Diego, USA) according to the manufacturer’s instructions, including incubation with intracellular dyes (anti-IL17A-AF700; anti-RORγt-APC; anti-IFNγ-PE and anti-TNFα-Pacific blue; all BioLegend). The stained cells were analyzed using a Gallios (Beckmann-Coulter) 10-color flow cytometer with Kaluza detection software, followed by analysis with Kaluza analysis software Version 1.3.
RNA isolation and quantitative real-time PCR
Total RNA was purified from colon samples, isolated CD4+CD62L+ T cells or lymphocytes using a RNeasy Kit (Qiagen, Hilden, Germany), including an additional step of on-column DNase digestion (RNase-Free DNase Set, Qiagen, Hilden, Germany). Both kits were used according to the manufacturer's instructions. Organ samples were lysed and homogenized in the RLT lysis buffer by using an ultrasound processor (Ultrasound processor Up50H). The obtained RNA concentration was determined with a NanoDrop® (Peqlab Biotechnologie GmbH). Up to 1 μg total RNA was used for cDNA synthesis with a QuantiTect Reverse Transcription Kit (Qiagen) following the manufacturer’s protocol. For quantitative real-time PCR analysis, a StepOnePlusTM Real-Time PCR cycler (Applied Biosystems) was used in combination with TaqMan®-based assays (Actb: Mm00607939_s1; Tbx21: Mm00450960_m1; Tnfα: Mm00443258_m1; Ifnγ: Mm01168134_m1; Rort: Mm01261022_m1; Alpk1: Mm01320380_m1; Applied Biosystems) or TaqMan® Assay Primer and Probe for Ifi44 (forward: 5′-GGGAATGAAGAAGGCACAGA-3′, reverse: 5′-AGAATTGCGATTGGTCCTTG-3′ and probe: 5′-FAM-GACTGTGCCAGGGGAAAAGCACA-BHQ1-3′ (Eurofins, Ebersberg, Germany)) and Larp7 (forward: 5′-GTGAAGATTGTGAGCGGAGAG-3′, reverse: 5′-TTCATTACTGCCTGAGCATCC-3′ (Eurofins, Ebersberg, Germany) and probe: 5′-FAM-AGAAGGAGATACTGAATGCCATGCCCGATT-BHQ-1-3′ (biomers.net, Ulm, Germany)). Each sample was measured in triplicate. The plate preparation and pipetting were performed under a sterile working flow (UV Sterilizing PCR Workstation, Peqlab). Beta actin served as an endogenous control, and relative gene expression was calculated using the 2-ΔΔCt method.
Microarray analysis
The RNA of naive T cells was extracted using an RNeasy kit (Qiagen) according to the manufactures’ instruction, and the RNA quality and amount were determined (Agilent 2100 Bioanalyzer). Labeled cDNA was hybridized on the Agilent SurePrint G3 Whole Mouse Genome (4x180k) Quadruplicate Array. The referenced microarray data used in this study were generated by the Research Core Unit Transcriptomics (RCUTAS) of the MHH. Raw data were processed by the RCUTAS and imported into Excel 2010 (Microsoft). For further analysis, the CoreModule v1.5.2 implemented in Excel 2010 was used. The differences in gene expression were determined by the fold change and the intensity cutoff parameter. The former parameter was set to be greater than two, and the latter was set to be greater than 50. For candidate gene acquisition, the focus was set to the distal part of chromosome 3 ( > 130 Mbp).
Western blot of CD4+CD62L+ T cells
Isolated naive T cells were resuspended in the lysis buffer and lysed with two short ultrasound impulses. Undiluted proteins were denaturated in a thermomixer (Mixing Block MB-102, BIOER Technology Co). For sodium dodecyl-sulfate-polyacrylamide (SDS) gel electrophoresis, a Mini PROTEAN® Tetra-Electrophoresis cell (Bio-Rad Laboratories) was used. SDS-gelelectrophoresis was performed on 10% gels, and the Precision plus Protein™ Standard (Bio-Rad Laboratories) was used as a protein ladder. The separated proteins were transferred to a nitrocellulose membrane (GE Healthcare) using a semidry system (“Pegasus”, Gesellschaft für Phorese, Analytik und Separation mbH). After blocking, primary antibodies (GAPDH (GenScript USA Inc., NJ USA) and IFI44 (MyBiosource, San Diego, USA)) were incubated overnight. After incubation with the secondary antibody, the membrane was incubated with a chemiluminescence solution. IFI44 was stained and detected, and then a stripping step was performed, which was required before GAPDH detection. GAPDH was used as an internal control. Signals were detected with the ChemiDoc™ Touch Imaging system (Bio-Rad Laboratories).
Statistical analysis
All analyses were performed in a blinded fashion. If not otherwise stated, values are the means ± SEM. All statistical analyses were performed using GraphPad Prism6® software (GraphPad Software, La Jolla, USA). T-tests were performed to compare data between BM chimeras. P < 0.05 was considered significant. *P < 0.05, **P < 0.01, and ***P < 0.001.
Results
Confirmation of the genetic authenticity of congenic substrains
Genotyping of BC-R2-Il10−/−, BC-R3-Il10−/− BC-R4-Il10−/− and BC-R8-Il10−/− using the MegaMUGA array verified that B6-Il10−/− and subcongenic mice shared the same genetic background, with the exception of different Cdcs1 fragments on MMU3 derived from C3Bir-Il10−/−. The congenic Cdcs1 fragment on MMU3 in subcongenic mice is defined by 735 out of 1365 single nucleotide polymorphism (SNP) markers informative for MMU3 of C3Bir-Il10−/− and B6-Il10−/−. The BC-R2-Il10−/− fragment was defined by rs30408500 at position 88.367708 Mbp to rs37014574 at 127.809720 Mbp. The BC-R3-Il10−/− fragment is ~60 Mb and extends from rs33209355 at position 98.778412 Mbp to rs36403089 at 159.518497 Mbp at the telomeric end of the chromosome. BC-R4-Il10−/− animals carried the smallest fragment from rs30507350 at position 129.212872 Mbp to rs31152536 at 144.340395 Mbp. The BC-R8-Il10−/− fragment spanned from rs46255391 at 125749892 bp to rs51899299 at position 155.749021 Mbp (Fig. 1). Thus, the genetic authenticity of the strains was confirmed and consistent with previous studies.12,32 Approximately 1265 annotated genes are located within the transferred Cdcs1 fragment according to the NCBI Genome Data Viewer. Three of those genes (Larp7, Alpk1 and Ifi44) were identified as candidate genes involved in inflammatory development.12,13,15 Larp7 and Alpk1 are located in Cdcs1.3 (MMU3: 127.536714 to 127.553349 Mbp and 127.670308 to 127.780553 Mbp, mm10) and identical in BC-R2-Il10−/−, BC-R3-Il10−/−, BC-R8-Il10−/− and C3Bir-Il10−/− mice but differed in BC-R4-Il10−/− and B6-Il10−/− mice. Alleles of Ifi44 (MMU3:151.730922 to 151.749959 Mbp, mm10) are derived from C3Bir-Il10−/− in BC-R3-Il10−/− and BC-R8-Il10−/− and from B6-Il10−/− in BC-R2-Il10−/− and BC-R4-Il10−/−.
Genotyping of B6-Il10−/−, C3Bir-Il10−/− and congenic mice on MMU3. SNP analysis of B6-Il10−/−, C3Bir-Il10−/− and BC-R2-Il10−/− BC-R3-Il10−/− BC-R4-Il10−/− and BC-R8-Il10−/− mice was performed (n = 1–6). The SNP and position (Mbp) of MMU3 are shown. The Cdcs1 region is highlighted in light gray. Of the 1365 informative SNP markers between C3Bir and B6, 10 selected SNP positions are shown for B6-Il10−/−, C3Bir-Il10−/− and congenic mice. Annotations are based on the GRCm38/mm10 mouse genome assembly. The candidate genes Larp7, Alpk1 (red line) and Ifi44 (green line) are highlighted
No detectable graft versus host reaction
To exclude the possibility of an acute graft versus host (aGvH) reaction between B6-Il10−/− and the subcongenic strains during bone marrow (BM) transplantation, mixed lymphocyte culture (MLC) experiments were performed with BC-R2-Il10−/− (Supplement Figure 1A) and BC-R3-Il10−/− mice carrying the longest fragment of Cdcs1. Isolated lymphocytes from B6-Il10−/− mice were cultured with irradiated splenic cells isolated from BC-R3-Il10−/− mice (Fig. 2a). As expected, no proliferation was found in the co-culture of B6-Il10−/− and BC-R3-Il10−/− cells. When the cells in the co-cultures were swapped, similar results were observed (Fig. 2b). These data indicate that lymphocytes from B6-Il10−/− mice showed no aGvH reaction against BC-R3-Il10−/− animals and vice versa.
No graft versus host reaction was detectable. An acute as well as a chronic GvH reaction was excluded using mixed lymphocyte cultures (MLC) and histological analysis, respectively. a B6-Il10−/− responder cells were incubated with medium only, with medium containing ConA, or BC-R3-Il10−/− stimulator cells. No increased proliferation was detected when stimulator cells were incubated with irradiated splenic cells from BC-R3-Il10−/− animals, while strong proliferation was observed when B6-Il10−/− responder cells were incubated with ConA. Means and standard deviations from 6 independent experiments are presented. b MLC of BC-R3-Il10−/− mice. Means and standard deviations from 6 independent experiments are presented. c Representative histological pictures of the skin, mucosa, liver and bone marrow from BM chimeras. No signs of cGvH, such as hyperkeratosis, subepidermal clefts or thickened dermis, in the epidermal and mucosal tissue and no ductopenia or portal fibrosis in the liver tissue were detectable. The BM is engrafted, and several hematopoietic progenitor cells are visible
A chronic GvH (cGvH) reaction was excluded histologically. BM chimeras of B6-Il10−/−, BC-R2-Il10−/− and BC-R3-Il10−/− animals were created, and 7 weeks later, the abdominal skin, liver, bone marrow and mucosal surfaces of the head were evaluated for lymphocyte infiltration, vacuolar degeneration on cells in the stratum basale and stratum spinosum or thickened dermis in the skin and hyperkeratosis in the head (Supplement Figure 1C and Figure 2C). In all analyzed slices, no indicators of a cGvH reaction were detected. Therefore, the intestinal inflammation of BM chimeras is not due to a GvH reaction.
Distinct microbiota-Cdcs1-haplotype interaction modifies the colitogenic potential of hematopoietic cells in BM chimeras
In the first experiment, BM chimeras of B6-Il10−/− and BC-R3-Il10−/− animals (Table 1) were generated with mice bred in Barrier I (experimental barrier). The onset of colitis was determined in vivo by a MRI scan and finally confirmed by histological scoring of the colon (Fig. 3a, b). The controls as well as BM chimeras of all groups revealed a thickened colon wall, hyperplasia of the crypts, strong infiltration of lymphocytes up to the L. muscularis, peritonitis, a reduced number of goblet cells and edema in the T. submucosa (Fig. 3c), indicating that animals from all groups developed colitis. Due to high variation within the groups, no difference between the groups and no impact of C3Bir-derived Cdcs1 could be detected. However, the experiment was repeated with animals bred in Barrier II (breeding barrier), where the spontaneous colitis onset in B6-Il10−/− mice was reduced. In Barrier II, no inflammatory changes were observed in the colons of B6-Il10−/− and BC-R3-Il10−/− control mice. In addition, only minor signs of colitis were detected in the MRI scan (Fig. 4a) or in histology (Fig. 4b, c) following transplantation of B6-Il10−/− BM cells into B6-Il10−/− or BC-R3-Il10−/− animals, while transplantation of BC-R3-Il10−/− BM cells into either B6-Il10−/− or BC-R3-Il10−/− mice led to the development of intestinal inflammation. These mice showed increased colon thickness in the MRI scan (Fig. 4a) and were histologically characterized by cell infiltration in the lamina propria and submucosa, abnormal crypt architecture and hyperplasia (Fig. 4c). Thus, the hematopoietic compartment of BC-R3-Il10−/− mice is a potential inducer of inflammation in the intestine but seems to need an additional trigger to induce colitis. However, the microbiota seems to have a strong impact on colitis induction.
Cdcs1-dependent inflammation in SPF-housed BM chimera mice. MRI (a) and histological scoring (b) of BM chimeras bred in Barrier II were performed 7 weeks after generation (n = 3–4). Significant differences in unpaired t-test are indicated by *P < 0.05; ***P < 0.001. c H&E staining of colon tissue obtained from BM chimeras
Microbiota composition is associated with inflammation in BM chimeras
Next, a microbiome analysis of cecal contents obtained from macroscopically healthy B6-Il10−/− and BC-R3-Il10−/− mice housed in Barrier I and Barrier II was performed. The microbiota analysis revealed distinct diversity between the two barriers (Fig. 5a, b) and between the mice bred in Barrier I (Fig. 5b). However, the overlap of the microbial composition between B6-Il10−/− and BC-R3-Il10−/− mice housed in Barrier I was much lower than that observed in Barrier II (Fig. 5c). Firmicutes were the most prevalent phylum, and the majority belonged to the family Lachnospiraceae. The second prominent phylum detected was Bacteroidetes, while bacteria of the phyla Proteobacteria, Actinobacteria or Verrucomicrobia were found only rarely (Fig. 5d). Furthermore, almost no bacteria of the genera Ruminococcus (2.25 ± 2.49 sequences), Roseburia (2.63 ± 2.26 sequences) or Akkermansia (0.63 ± 0.92 sequences) were identified, and no Helicobacter spp. were observed in Barrier II.
Microbiota analysis of Barrier I and Barrier II. The microbiota of the cecal contents from healthy B6-Il10−/− and BC-R3-Il10−/− mice housed in Barrier I and II was analyzed. a Differences in alpha-diversity (observed genera and Shannon diversity) between Barrier I and Barrier II were evaluated. b Differences in alpha-diversity (observed genera and Shannon diversity) between strains were evaluated. c NMDS diagram of B6-Il10−/− and BC-R3-Il10−/− animals from Barrier I and Barrier II. Each dot represents the microbiome composition of an animal: red dots = B6-Il10−/−, blue dots = BC-R3-Il10−/−, dark color = Barrier I, and light color = Barrier II (n = 4). d Bacterial composition (at the phylum and genus level) of mice housed in Barrier I and Barrier II was determined. Here, “unclassified” represents sequences that were not reliably assigned a genus by the classifier and binned into higher-order taxonomy
Although the major phylum detected was still Firmicutes, the ratio between Firmicutes and Bacteroidetes changed (Barrier I: B6-Il10−/− 2:1 and BC-R3-Il10−/− 2:1; Barrier II: B6-Il10−/− 7:1 and BC-R3-Il10−/− 8:1), and a greater amount of Proteobacteria was identified in Barrier I, especially in BC-R3-Il10−/− mice. Moreover, elevated levels of Ruminococcus (1.55% of total bacteria), Roseburia (0.06% of total bacteria) and Akkermansia (0.06% of total bacteria) were observed in Barrier I compared with those in Barrier II. In addition, Clostridia spp. of the cluster XIVa was decreased. These differences in the microbial composition of Barrier I and II animals might be responsible for the increased inflammatory phenotype of mice bred in Barrier I. Thus, our data suggest that the composition of the microflora has an impact on the induction of spontaneous colitis in the Il10−/− mouse model.
T cell transfer induced colitis is not modified by Larp7 and Alpk1
Because the hematopoietic compartment of congenic mice seems to be a potential inducer of inflammation in the intestine, a T cell transfer was performed using naive T cells isolated from BC-R4-Il10−/− and BC-R8-Il10−/− mice transferred into B6-Rag1−/− mice. These congenic mouse strains were selected to discriminate the effects of Alpk1 and Larp7 alleles derived from B6-Il10−/− in BC-R4-Il10−/− and from C3Bir-Il10−/− in BC-R8-Il10−/− mice (Fig. 1). T cell transfer was performed, and 6-7 weeks later, a histopathological analysis of the colon revealed higher histological scores in recipients of BC-R8-Il10−/− cells compared to recipients of BC-R4-Il10−/− cells (Fig. 6a). Increased lymphocytic infiltration, crypt hyperplasia and goblet cell loss, crypt abscesses and crypt architecture destruction were present in the recipients of BC-R8-Il10−/− cells (Fig. 6a). Thus, genes within Cdcs1 expressed in naive T cells from BC-R8-Il10−/− mice seem to induce increased colonic inflammation. mRNA expression of Larp7 in lymphocytes revealed similar expression levels in all substrains (Fig. 6b), although different expression was observed for Alpk1. B6-Il10−/− and BC-R4-Il10−/−, which share the same genetic fragment, showed slight Alpk1 expression levels while BC-R8-Il10−/− and C3Bir-Il10−/− mice showed increased expression levels (Fig. 6b). To investigate the impact of Alpk1 in T cell transfer experiments, B6-Alpk1−/− mice were used. Eight weeks after transfer, mice were sacrificed and the colons were histologically analyzed (Fig. 6c). Both groups developed intestinal inflammations, and no histopathological differences were detected in the transfer of naive T cells isolated from B6 or B6-Alpk1−/− mice. Thus, neither Larp7 nor Alpk1 seem to be involved in colitis development in a T cell-dependent manner.
T cell-mediated colitis is independent of Larp7 or Alpk1. a Histological scoring of B6-Rag1−/− animals that received cells isolated from BC-R4-Il10−/− and BC-R8-Il10−/− mice (n = 9–11, **P ≤ 0.01, samples from two independent experiments). Representative H&E stained histological slides of the distal colon. b Gene expression levels of Larp7 and Alpk1 in isolated lymphocytes. c Histological scoring and representative H&E stained histological slides of the distal colon of B6-Rag1−/− mice that received cells isolated from B6 and Alpk1−/− mice (n = 11–12, samples from two experiments)
Ifi44 as a candidate gene in T cell transfer-induced colitis
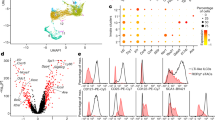
The severe colitis phenotype caused by BC-R8-Il10−/− but not by BC-R4-Il10−/− T cells might be caused by the C3Bir-fragment between 125.7 to 129.0 Mbp or the distal part starting at 144.3 Mbp. To resolve this question T cell transfer was performed using naive T cells isolated from B6-Il10−/−, BC-R2-Il10−/− or BC-R3-Il10−/− mice, and recipient animals were necropsied 4 weeks after transfer. The histopathological analysis of the colon revealed colitis in all animals (Fig. 7a). Lesions observed in the colon were characterized by massive lymphocytic infiltration, crypt hyperplasia and goblet cell loss. However, recipients that received adoptive transfer of BC-R3-Il10−/− T cells displayed a significantly higher histopathological colitis phenotype, which included epithelial ulceration, crypt abscesses and destruction of the crypt architecture. To more precisely characterize the T cell response in the colon, we determined the cytokine and transcription factor expression levels by qPCR and flow cytometry. The gene expression levels of Tbx21, a master transcription factor for the Th1 response, Ifn and Tnfα were higher in the colon tissue of animals that received cells from B6-Il10−/−, BC-R2-Il10−/− and BC-R3-Il10−/− mice than those in untreated B6-Rag1−/− mice (Fig. 7b). No differences were observed in Rort expression. Il17a was not detected by qPCR in whole colon tissue. However, stimulated BC-R3-Il10−/− T cells showed enhanced IL17A production and a slightly increased amount of RORγt-positive cells (Fig. 7c). Altogether, T cells from all strains mainly cause a Th1 response after adoptive transfer, although BC-R3-Il10−/− T cells exhibited a higher proportion of Th17 response (IL17A+)-positive cells. Because intestinal inflammation in mice that had received T cells isolated from BC-R2-Il10−/− mice was reduced compared with that of BC-R3-Il10−/− mice, the focus was put on the distal region of MMU3 starting at rs47978238 (144.877822 Mbp). Because this fragment was contained in the original Cdcs1 interval,3 we provisionally denominate this interval as Cdcs1.4.
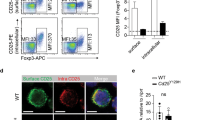
T cell-mediated colitis development identified Ifi44 as a candidate gene. a Histological scoring of animals that received cells isolated from B6-Il10−/−, BC-R3-Il10−/− and BC-R2-Il10−/− mice (n = 6–12, **P ≤ 0.01, samples from two independent experiments). Representative H&E stained histological slides of the distal colon that received naive T cells isolated from B6-Il10−/−, BC-R3-Il10−/− and BC-R2-Il10−/− animals. b The gene expression levels of Tbx21, Rort, Ifn and Tnf measured in the proximal colon of B6-Rag1−/− animals compared with those in B6-Rag1−/− animals, which received cells from B6-Il10−/−, BC-R3-Il10−/− and BC-R2-Il10−/− mice. The relative quantification (RQ) of gene expression was measured by qPCR and referred to a reference sample set to 1 (n = 4-12; mean ± SEM; samples from two independent experiments). c The percentage of IL17A+, RORYt+, IFNγ+ and TNFα+ CD3+CD4+ T cells of animals, which received T cells isolated from B6-Il10−/−, BC-R3-Il10−/− or BC-R2-Il10−/− mice, was determined by flow cytometry (n = 3–9; mean ± SEM; * P ≤ 0.05, samples from two independent experiments). d Heat map of genes differently expressed due to the distal MMU3. Naive T cells from BC-R3-Il10−/− or BC-R2-Il10−/− mice were compared with B6-Il10−/− samples (n = 2, fold change > 2, intensity cutoff > 50). e Relative quantification (RQ) in gene expression of Ifi44 in colitogenic T cells is based on a reference sample set to 1 (n = 4–8, mean ± SEM; * = P < 0. 05, *** = < 0.001, samples from two independent experiments). f Densitometric quantification of IFI44 by western blot of lysates from naive T cells isolated from B6-Il10−/−, BC-R3-Il10−/− or BC-R2-Il10−/− normalized to the internal control GAPDH (n = 5). Representative western blot of IFI44 and GAPDH protein expression
To identify candidate genes in the Cdcs1.4 interval, microarray data were obtained from naive T cells, which revealed 5 genes were upregulated and 4 genes were downregulated in BC-R3-Il10−/− but not in BC-R2-Il10−/− T cells compared with B6-Il10−/− T cells (Fig. 7d). Only Ifi44 and Ifi44l were located in the Cdcs1.4 region. Ifi44 was previously found to be expressed at a higher level in C3H mice in mucosal response to bacterial colonization.14 A qPCR analysis of the gene expression of Ifi44 in T cells re-isolated after adoptive transfer confirmed the low expression in B6-Il10−/− and BC-R2-Il10−/− mice but increased expression in BC-R3-Il10−/− T cells (Fig. 7e). In addition, a western blot analysis of IFI44 in naive T cells was consistent with the microarray results (Fig. 7f). Thus, the distal part of MMU3 seems to contribute to intestinal inflammation in a T-cell-dependent manner. This new Cdcs1.4 region (144 to 155 Mbp) includes Ifi44 as a candidate gene involved in a detrimental Th17 response when IL10 control is absent.
Discussion
IBD is a multifactorial human disease suggested to be dependent on genetic background, microbes and environmental factors. Il10-deficient mice are a suitable animal model for IBD.1 Several Il10-deficient inbred mouse strains have shown that colitis severity depends on the genetic background of the respective strain. Linkage analysis using F2 and N2 populations derived from B6-Il10−/− and C3Bir-Il10−/− mice revealed 10 QTLs (Cdcs1 to Cdcs10).3,11 Thus, Cdcs1 on chromosome 3 was identified as the major modifier for colitis susceptibility.3,6,11 BC-R2-Il10−/−, BC-R3-Il10−/−, BC-R4-Il10−/− and BC-R8-Il10−/− are subcongenic inbred strains on a B6 background, and they received their Cdcs1 fragment from the IBD-prone inbred strain C3Bir-Il10−/−. These subcongenic lines have been developed to enable fine mapping of Cdcs1 using microsatellite markers and estimations of fragment length 12 and they are valuable for demonstrating that Cdcs1 is a very complex locus that is split into at least three subfragments (Cdcs1.1, 1.2 and 1.3). In the present study, all subfragments were detected at the expected length, illustrating the stable inheritance of these fragments.
To investigate the impact of the hematopoietic compartment regarding the impact of Cdcs1, bone marrow transplantation experiments between congenic BC-R2-Il10−/− or BC-R3-Il10−/− and B6-Il10−/− animals were performed. However, congenic mice harbor the risk of a GvH reaction during BM chimera experiments because of potentially incompatible donor genetic elements. A GvH is primarily caused by the remaining T cells in the graft reacting against surface proteins such as MHC molecules of the host,33 although it can also evolve due to minor histocompatibility complexes potentially located in the congenic element. In humans, approximately 40% of stem cell transplantations with a matched MHC between the donor and recipient lead to a GvH caused by minor histocompatibility complexes.34
An MHC-dependent acute GvH reaction was excluded in this study by the MegaMUGA SNP array, which confirmed the B6-derived MHC-haplotype on MMU17 in all congenic mice. Furthermore, MLC did not reveal increased proliferation of B6-Il10−/− T cells against those isolated from BC-R2-Il10−/− or BC-R3-Il10−/− mice and vice versa. A chronic GvH reaction due to the congenic element would almost be histologically indistinguishable from Cdcs1-mediated intestinal inflammation.35 However, cGvH reaction is characterized by apoptotic bodies, cytoid bodies, hyperkeratosis, subepidermal clefts and thickened dermis in the skin and the head.36 Histological determination of abdominal skin and epidermal and mucosal surfaces on the head revealed no signs of cGvH in the present study. Therefore, a GvH reaction did not represent a likely cause of the intestinal inflammation in BM chimeras.
Although the Cdcs1 locus is associated with a disturbed immune response,12,32 a final confirmation of the relevant cell population is still missing. In this study, the hematopoietic compartment of BC-R3-Il10−/− mice harboring the longest C3Bir-derived Cdcs1 locus was a potent inducer of inflammation in the intestine. This finding is in line with previously data showing that after stimulation by CBir1 flagellin, BMDM of C3Bir mice produced reduced levels of proinflammatory cytokines, while T cells showed increased proliferation compared with those from B6 mice.18 In addition, genetic modifiers in other IBD-related models mapped to the same genomic region (Hiccs, Cdcs1TRUC, Gpdc1),13,37,38,39,40 and one of these studies identified the hematopoietic cell population as the pivotal compartment for colitis induction.13 Therefore, we produced BM chimeras using recipient mice bred in two different hygienic barriers, including a strict breeding barrier (Barrier II) and an experimental SPF barrier (Barrier I). The strict breeding barrier displayed lower microbial complexity, a higher Firmicutes/Bacteriodetes ratio, and an increase in Proteobacteria. The two major bacteria phyla detected in humans and mice are Firmicutes and Bacteriodetes,7,41 followed by Proteobacteria and Actinobacteria.7 However, an altered Firmicutes/Bacteroidetes ratio as well as an increase in Proteobacteria and Actinobacteria was described in IBD patients.7,9,10 In the literature Akkermansia (with the only species Akkermansia muciniphila) is found to both worsen intestinal inflammation and improve metabolic disorders.8,42,43 A recently published work found no impact of A. muciniphila on the induction of chronic intestinal inflammation.44 Furthermore, the number of Clostridia cluster XIVa was decreased in mice housed in Barrier I. The Clostridia cluster XIVa favors the development of Tregs 45 and may produce short chain fatty acids, which seem to have anti-inflammatory properties.46,47 In addition, the bacterial diversity was reduced in patients with UC.5,48,49
In the present study, compositional differences of the gut microbiota were associated with intestinal inflammation, suggesting that gut bacteria play a role in colitis development in the chimera model. While hematopoietic cells carrying the susceptibility Cdcs1 haplotype exerted similar detrimental effects under both barrier conditions, the cells carrying the resistant haplotype induced colitis under high microbiome complexity conditions only. Therefore, the detrimental Cdcs1 effect mediated by hematopoietic cells is independent of the microbiome composition of the host. However, this finding also confirms our previous hypothesis 1 that microbiome effects can supersede genetic susceptibility in the Il10-deficiency model.
A candidate gene analysis revealed that various genes differed in the sequence and expression patterns between susceptible and resistant mice and showed that some of these genes may be pivotal in the induction of colonic inflammation. Ifi44, Larp7 and Alpk1 are genes located on MMU3, and Larp7 and Alpk1 are specifically located within Cdcs1.3. In this study the pivotal role of Larp7 as well as Alpk1 in the induction of T cell-dependent colitis induction was excluded. Larp7 was found to be similarly expressed in lymphocytes in all substrains, thus indicating a minor role in T cell proliferation or activation. Although Alpk1 was expressed at a higher level in lymphocytes harboring the susceptible Cdcs1 fragment, the T cell transfer experiments using naive T cells from knockout mice showed no differences in colitis development. This result is consistent with the findings of Ryzhakov et al., who very recently published similar results.17 However, they were able to show that Alpk1 impacted IL-12 production in phagocytes in a Helicobacter hepaticus-induced colitis model. Thus, Alpk1 seems to be a good candidate gene for colitis development.
In this study, the naive T cells of BC-R8-Il10−/− mice exerted a pronounced colitogenic effect due to the C3Bir haplotype of Cdcs1 compared with the T cells from BC-R4-Il10−/− mice with the B6 haplotype. These findings suggest a colitogenic region of Cdcs1 that is proximal or distal to the congenic fragment of BC-R4-Il10−/− animals. T cell transfer experiments using BC-R2-Il10−/− and BC-R3-Il10−/− mice excluded the relevance of the proximal region in this experimental setup, and a distal colitogenic region called Cdcs1.4 was newly identified. This colitogenic effect is likely mediated by a Th17-mediated response. Therefore, within the hematopoietic compartment, CD4+ T cells play a pivotal role in Cdcs1-mediated hyperresponsiveness. A gene-array analysis revealed Ifi44 as a candidate gene located at 151.730922 to 151.749959 Mbp, and its expression is higher in naive T cells of BC-R3-Il10−/− mice. Until now, IFI44 was identified to have antiproliferative activity,50 and it seems to be involved in an appendicitis and appendectomy colitis model 15 and genotype-specific host responses to commensal gut bacteria.14
In summary, this study clearly indicates the pivotal role of Cdcs1 in the induction of colitis by hematopoietic cells. Microbiome effects determine colitis severity in this model, while Cdcs1 likely exerts its effect in the hematopoietic compartment independent of microbiome composition. These effects certainly underlie the complexity of IBD development. Furthermore, Larp7 and Alpk1 could be excluded as candidate genes but Ifi44 seems to have an impact in T cell-mediated colitis development.
References
Keubler, L. M., Buettner, M., Hager, C. & Bleich, A. A multihit model: colitis lessons from the interleukin-10-deficient mouse. Inflamm. Bowel Dis. 21, 1967–1975 (2015).
Ek, W. E., D'Amato, M. & Halfvarson, J. The history of genetics in inflammatory bowel disease. Ann. Gastroenterol. 27, 294–303 (2014).
Mahler, M. et al. Genetics of colitis susceptibility in IL-10-deficient mice: backcross versus F2 results contrasted by principal component analysis. Genomics 80, 274–282 (2002).
Nishida, A. et al. Gut microbiota in the pathogenesis of inflammatory bowel disease. Clin. J. Gastroenterol. 11, 1–10 (2017).
Schaubeck, M. et al. Dysbiotic gut microbiota causes transmissible Crohn's disease-like ileitis independent of failure in antimicrobial defence. Gut 65, 225–237 (2016).
Buettner, M. & Bleich, A. Mapping colitis susceptibility in mouse models: distal chromosome 3 contains major loci related to Cdcs1. Physiol. Genom. 45, 925–930 (2013).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl Acad. Sci. USA 104, 13780–13785 (2007).
Ganesh, B. P., Richter, J. F., Blaut, M. & Loh, G. Enterococcus faecium NCIMB 10415 does not protect interleukin-10 knock-out mice from chronic gut inflammation. Benef. Microbes 3, 43–50 (2012).
Matsuoka, K. & Kanai, T. The gut microbiota and inflammatory bowel disease. Semin. Immunopathol. 37, 47–55 (2015).
Walters, W. A., Xu, Z. & Knight, R. Meta-analyses of human gut microbes associated with obesity and IBD. FEBS Lett. 588, 4223–4233 (2014).
Farmer, M. A. et al. A major quantitative trait locus on chromosome 3 controls colitis severity in IL-10-deficient mice. Proc. . Natl Acad. Sci. USA 98, 13820–13825 (2001).
Bleich, A. et al. Cdcs1 a major colitis susceptibility locus in mice; subcongenic analysis reveals genetic complexity. Inflamm. Bowel Dis. 16, 765–775 (2010).
Boulard, O., Kirchberger, S., Royston, D. J., Maloy, K. J. & Powrie, F. M. Identification of a genetic locus controlling bacteria-driven colitis and associated cancer through effects on innate inflammation. J. Exp. Med. 209, 1309–1324 (2012).
Brodziak, F., Meharg, C., Blaut, M. & Loh, G. Differences in mucosal gene expression in the colon of two inbred mouse strains after colonization with commensal gut bacteria. PLoS ONE 8, e72317 (2013).
Cheluvappa, R., Eri, R., Luo, A. S. & Grimm, M. C. Modulation of interferon activity-associated soluble molecules by appendicitis and appendectomy limits colitis-identification of novel anti-colitic targets. J. Interferon Cytokine Res 35, 108–115 (2015).
Jurisic, G. et al. Quantitative lymphatic vessel trait analysis suggests Vcam1 as candidate modifier gene of inflammatory bowel disease. Genes Immun. 11, 219–231 (2010).
Ryzhakov, G. et al. Alpha kinase 1 controls intestinal inflammation by suppressing the IL-12/Th1 axis. Nat. Commun. 9, 3797 (2018).
Beckwith, J., Cong, Y., Sundberg, J. P., Elson, C. O. & Leiter, E. H. Cdcs1, a major colitogenic locus in mice, regulates innate and adaptive immune response to enteric bacterial antigens. Gastroenterology 129, 1473–1484 (2005).
Kostic, A. D., Xavier, R. J. & Gevers, D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146, 1489–1499 (2014).
Bleich, A. & Mahler, M. Environment as a critical factor for the pathogenesis and outcome of gastrointestinal disease: experimental and human inflammatory bowel disease and helicobacter-induced gastritis. Pathobiology 72, 293–307 (2005).
Kuhn, R., Lohler, J., Rennick, D., Rajewsky, K. & Muller, W. Interleukin-10-deficient mice develop chronic enterocolitis. Cell 75, 263–274 (1993).
Mahler, M. & Leiter, E. H. Genetic and environmental context determines the course of colitis developing in IL-10-deficient mice. Inflamm. Bowel Dis. 8, 347–355 (2002).
Feller, M. et al. Mycobacterium avium subspecies paratuberculosis and Crohn's disease: a systematic review and meta-analysis. Lancet Infect. Dis. 7, 607–613 (2007).
Grant, I. R. Zoonotic potential of Mycobacterium avium ssp. paratuberculosis: the current position. J. Appl. Microbiol. 98, 1282–1293 (2005).
Lamps, L. W. et al. Pathogenic Yersinia DNA is detected in bowel and mesenteric lymph nodes from patients with Crohn's disease. Am. J. Surg. Pathol. 27, 220–227 (2003).
Saebo, A., Vik, E., Lange, O. J. & Matuszkiewicz, L. Inflammatory bowel disease associated with Yersinia enterocolitica O:3 infection. Eur. J. Intern. Med. 16, 176–182 (2005).
Balish, E. & Warner, T. Enterococcus faecalis induces inflammatory bowel disease in interleukin-10 knockout mice. Am. J. Pathol. 160, 2253–2257 (2002).
Devkota, S. et al. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10-/- mice. Nature 487, 104–108 (2012).
Fox, J. G. et al. A novel urease-negative Helicobacter species associated with colitis and typhlitis in IL-10-deficient mice. Infect. Immun. 67, 1757–1762 (1999).
Madsen, K. L., Doyle, J. S., Jewell, L. D., Tavernini, M. M. & Fedorak, R. N. Lactobacillus species prevents colitis in interleukin 10 gene-deficient mice. Gastroenterology 116, 1107–1114 (1999).
McCarthy, J. et al. Double blind, placebo controlled trial of two probiotic strains in interleukin 10 knockout mice and mechanistic link with cytokine balance. Gut 52, 975–980 (2003).
Buchler, G. et al. Strain-specific colitis susceptibility in IL10-deficient mice depends on complex gut microbiota-host interactions. Inflamm. Bowel Dis. 18: 943–954 (2012).
Kuroiwa, T. et al. Hepatocyte growth factor ameliorates acute graft-versus-host disease and promotes hematopoietic function. J. Clin. Invest. 107, 1365–1373 (2001).
Ferrara, J. L., Levine, J. E., Reddy, P. & Holler, E. Graft-versus-host disease. Lancet 373, 1550–1561 (2009).
Shulman, H. M. et al. NIH Consensus development project on criteria for clinical trials in chronic graft-versus-host disease: II. The 2014 Pathology Working Group Report. Biol. Blood. Marrow Transplant. 21, 589–603 (2015).
Pintar, T. et al. Skin and kidney histological changes in graft-versus-host disease (GVHD) after kidney transplantation. Bosn. J. Basic. Med. Sci. 11, 119–123 (2011).
Borm, M. E. et al. A major quantitative trait locus on mouse chromosome 3 is involved in disease susceptibility in different colitis models. Gastroenterology 128, 74–85 (2005).
Ermann, J. et al. Severity of innate immune-mediated colitis is controlled by the cytokine deficiency-induced colitis susceptibility-1 (Cdcs1) locus. Proc. Natl Acad. Sci. USA 108, 7137–7141 (2011).
Hillhouse, A. E., Myles, M. H., Taylor, J. F., Bryda, E. C. & Franklin, C. L. Quantitative trait loci in a bacterially induced model of inflammatory bowel disease. Mamm. Genome 22, 544–555 (2011).
Hornquist, C. E. et al. G(alpha)i2-deficient mice with colitis exhibit a local increase in memory CD4+ T cells and proinflammatory Th1-type cytokines. J. Immunol. 158, 1068–1077 (1997).
Nguyen, T. L., Vieira-Silva, S., Liston, A. & Raes, J. How informative is the mouse for human gut microbiota research? Dis. Model Mech. 8, 1–16 (2015).
Derrien, M. et al. Modulation of mucosal immune response, tolerance, and proliferation in mice colonized by the mucin-degrader Akkermansia muciniphila. Front. Microbiol. 2, 166 (2011).
Shin, N. R. et al. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 63, 727–735 (2014).
Ring, C. et al. Akkermansia muciniphila strain ATCC BAA-835 does not promote short-term intestinal inflammation in gnotobiotic interleukin-10-deficient mice. Gut Microbes. 1–16 (2018) https://doi.org/10.1080/19490976.2018.1511663.
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011).
Barcenilla, A. et al. Phylogenetic relationships of butyrate-producing bacteria from the human gut. Appl. Environ. Microbiol. 66, 1654–1661 (2000).
Van den Abbeele, P. et al. Butyrate-producing Clostridium cluster XIVa species specifically colonize mucins in an in vitro gut model. Isme. J. 7, 949–961 (2013).
Alipour, M. et al. Mucosal Barrier Depletion and Loss of Bacterial Diversity are Primary Abnormalities in Paediatric Ulcerative Colitis. J Crohns Colitis 10, 462–17 (2015).
Larmonier, C. B. et al. Reduced colonic microbial diversity is associated with colitis in NHE3-deficient mice. Am. J. Physiol. Gastrointest. Liver Physiol. 305, G667–G677 (2013).
Hallen, L. C. et al. Antiproliferative activity of the human IFN-alpha-inducible protein IFI44. J. Interferon Cytokine Res. 27, 675–680 (2007).
rodents, Fwgorogfhmo et al. FELASA recommendations for the health monitoring of mouse, rat, hamster, guinea pig and rabbit colonies in breeding and experimental units. Lab. Anim. 48, 178–192 (2014).
Michael, S. et al. Quantitative phenotyping of inflammatory bowel disease in the IL-10-deficient mouse by use of noninvasive magnetic resonance imaging. Inflamm. Bowel Dis. 19, 185–193 (2013).
de Buhr, M. F. et al. Cd14, Gbp1, and Pla2g2a: three major candidate genes for experimental IBD identified by combining QTL and microarray analyses. Physiol. Genom. 25, 426–434 (2006).
Bleich, A. et al. Refined histopathologic scoring system improves power to detect colitis QTL in mice. Mamm. Genome 15, 865–871 (2004).
Rath, S., Heidrich, B., Pieper, D. H. & Vital, M. Uncovering the trimethylamine-producing bacteria of the human gut microbiota. Microbiome 5, 54 (2017).
Cole, J. R. et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42(Database issue), D633–D642 (2014).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Acknowledgements
This work was supported by the Hannover Biomedical Research School (HBRS), the Center for Infection Biology (ZIB), and a stipend to Inga Brüsch. Marius Vital and Dietmar H. Pieper were supported by iMed, the Helmholtz Association’s Initiative on Personalized Medicine. We thank Britta Trautewig, Anna Smoczek, Andrea Liese, Anja Siebert, Silke Kahl, Iris Plumeier and Elena Wiebe for providing excellent technical assistance.
Author information
Authors and Affiliations
Contributions
I.B., M.Bue., P.M., M.Ba. and A.B. conceived and designed the experiments and wrote the manuscript. I.B., P.M., K.S., S.B. and M.Bue. performed the experiments and analyzed the data. I.B. and P.M. contributed equally to the work. M.V. and D.P. performed and analyzed microbiome experiments. M.M. performed MRI evaluation. S.G. provided support for the evaluation of the histology. J.H. performed and analyzed MLC experiments. D.W. performed genetic data confirmation. M.Bue. and A.B. supervised the work and contributed equally.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Bruesch, I., Meier, P., Vital, M. et al. Analysis of Cdcs1 colitogenic effects in the hematopoietic compartment reveals distinct microbiome interaction and a new subcongenic interval active in T cells. Mucosal Immunol 12, 691–702 (2019). https://doi.org/10.1038/s41385-019-0133-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-019-0133-9