Abstract
Microbial metabolites are an emerging class of mediators influencing CD4+ T-cell function. To advance the understanding of direct causal microbial factors contributing to Crohn’s disease, we screened 139 predicted Crohn’s disease-associated microbial metabolites for their bioactivity on human CD4+ T-cell functions induced by disease-associated T helper 17 (Th17) polarizing conditions. We observed 15 metabolites with CD4+ T-cell bioactivity, 3 previously reported, and 12 unprecedented. A deeper investigation of the microbe-derived metabolite, ascorbate, revealed its selective inhibition on activated human CD4+ effector T cells, including IL-17A-, IL-4-, and IFNγ-producing cells. Mechanistic assessment suggested the apoptosis of activated human CD4+ T cells associated with selective inhibition of energy metabolism. These findings suggest a substantial rate of relevant T-cell bioactivity among Crohn’s disease-associated microbial metabolites, and evidence for novel modes of bioactivity, including targeting of T-cell energy metabolism.
Similar content being viewed by others
Introduction
Abnormalities of intestinal microbial composition are strongly associated with inflammatory bowel diseases.1,2,3 In the healthy state, much evidence suggests that small molecule products of the microbiota (microbial metabolites) are important modulators of normal host physiology.4,5,6 Among these, the literature includes only a handful of bioactive microbial metabolites, and few metabolites directly affect cellular targets of Crohn’s disease relevance.7,8 In Crohn’s disease, previous multi-omic studies suggest an association with alteration of microbial energy metabolism, enrichment of carbohydrate, lipid, co-factor and vitamin metabolism, and depletion of amino acid biosynthesis.9,10,11 Certain of these microbial metabolites are known to influence important aspects of host metabolism,12 activity of intestinal epithelial stem cells,7 and regulation of diverse immune cellular responses.8,13,14 For example, short-chain fatty acids (SCFAs) such as butyrate are known for their mechanisms of action on immune responses through the inhibition of histone deacetylase.14 For example, SCFAs such as butyrate are known for their mechanisms of action on immune responses through the inhibition of histone deacetylase.14 Tryptophan catabolites induce T-cell cytokine expression through a process involving ligand-activated aryl hydrocarbon receptor, AHR.15 However, most inferred microbial metabolites remain untested for bioactivity on disease-relevant host responses, and unresolved for their mechanisms of action.
In genetically susceptible individuals, Crohn’s disease results from inappropriate immune responses to the unique intestinal microbiota.16,17 Functional assessment of Crohn’s disease genetic loci revealed the involvement of CD4+ effector T helper signaling and the Th17-IL-23 pathway.18,19 Both clinical studies and pre-clinical colitis studies strongly link Th17 cell responses and the interleukin (IL)-17 pathway to the pathogenesis of Crohn’s disease.20,21,22,23,24 Given their pathogenic role, Th17 cells became a therapeutic target,25 but clinical trials revealed unexpected inefficacy and disease exacerbation.26,27,28,29,30 Subsequent work has uncovered dichotomous roles of the IL-17 pathway in Crohn’s disease progression, reflecting the differential roles of Th17 cells in disease induction and resolution.31 This affirms the importance of Th17 in Crohn’s disease, but suggests complexity in how the intestinal microbiome and its products may be modifiers of mucosal effector T-cell function.
To determine how Crohn’s disease-associated microbiome and its products modulate CD4+ T-cell responses, we applied a set of bioinformatics packages to tabulate a list of microbial metabolites predicted to be differentially abundant in Crohn’s disease. We functionally screened these metabolites by adding them to CD4+ T-cell cultures together with Th17-polarizing cytokines. This conceptually recapitulates the in vivo setting, in which lymphocytes simultaneously encounter bacterial products and inflammatory cytokines in the sub-mucosa or draining lymph nodes. Our screen revealed a substantial frequency (11%) of metabolites that modulated T-cell activity, with different patterns of inhibition and augmentation. Detailed study of one microbe-derived metabolite, ascorbate, demonstrated potent inhibition of activated effector CD4+ T cells associated with the disruption of energy metabolism. These findings support the idea that disease-associated microbial metabolites may have a significant role in the pathogenesis of Crohn’s disease.
Results
Strategy to identify microbial metabolites modulating T-cell responses
To predict Crohn’s disease-associated microbial metabolites, we performed a set of bioinformatics approach on the fecal microbiome dataset in PRISM cohort32 (supplementary information). This approach predicted the microbial metabolites that are differentially produced based on Crohn’s disease and healthy microbiome, and we selected the subset of 139 commercially available metabolites to test their functions on modulating CD4+ T-cell responses. Supplementary Table 1 show the list of tested metabolites. Figure 1a shown the summarized workflow of screen strategy. Briefly, purified human blood CD4+ T cells were cultured with different concentrations of each metabolite in the presence of standard 14 days of Th17 polarization culture conditions (anti-CD3, anti-CD28, IL-6, IL-1β, and IL-23).33 We assessed Th17 polarization by levels of secreted cytokines, and validated by the percentage of Th17 cells by flow cytometry (Supplementary Figure 2).
Microbial metabolites screen identifies bioactive metabolites modulating CD4+ T-cell cytokine profile. a Schematic of screening and analytic strategies. Toxic: cell death observed at all tested conditions; potential bioactive: achieved MANOVA statistical significance cutoff; putative bioactive: significant effects in first round but not second round of screening, potentially due to donor-specific variation; validated bioactive: reproducible effects in two individuals (achieved statistical significance cutoff by two-way ANOVA test). b Aggregate cytokine response (negative logarithm of the association p values calculated from multivariate analysis of variance) to candidate metabolites. p < 0.0004 represents the significant threshold after Holm–Sidak correction for multiple comparisons (blue dots). c Categories of response to metabolites (% of all tested metabolites). d Summary heatmap of multiplex cytokine expression data for the 12 novel validated bioactive metabolites. We show metabolite-treated samples (rows) at the optimal bioactive concentration. Asterisks (*) highlight metabolites with dose-dependent effects. Four metabolites with dose-dependent effects are marked with an asterisk (*). CoA coenzyme A, HMG-CoA 3-hydroxy-3-methyl-glutaryl-coenzyme A
Differential metabolite effects on T-cell cytokine profiles
The primary screening revealed 18 bioactive metabolites based on a relaxed cutoff in multivariate analysis of variance (MANOVA) (Fig. 1b). We performed a confirmatory screening in independent healthy individuals to validate the primary screen and refine dose-dependent responses (two-way ANOVA). Figure 1c summarizes results for all screened metabolites with respect to toxicity and secreted cytokine production. The toxicity was defined by the end points of cell death and reduced cell numbers. We tabulated metabolite conditions that significantly increased toxicity, and excluded them from downstream cytokine analyses. Notably, we did not observe any metabolites that significantly enhanced cell numbers (Supplementary Figure 3).
Sixteen metabolites (11%) were toxic based on cell death at all tested conditions in both donors. A total of 15 metabolites (11%) displayed reproducible effects on T-cell cytokine production, and were classified as validated bioactive. Three metabolites (2%) showed inconsistent cytokine changes between two individual experiments, and were classified as putatively bioactive.
The 15 validated bioactive metabolites varied in their pattern of effects on cytokine production (Table 1). Three of these metabolites were previously reported for analogous effects on T cells13,34,35 (Supplementary Figure 4). Strikingly, the action of the other 12 metabolites are unprecedented, to our knowledge, for an effect on T-cell cytokine production. Figure 1d summarizes the relative cytokine secretion for the 12 novel bioactive metabolites. Unsupervised hierarchical clustering classified the metabolites into patterns of inhibitory and augmented responses. Among them, six metabolites are fatty acid intermediates or end products based on the Human Metabolome Database.36
Representative bioactive metabolites altering CD4+ T-cell cytokine secretion
We observed distinct patterns of altered cytokine production and we organized the data between main and supplementary figures based on the criteria of the dose-dependency. The first group of metabolites induced global changes in cytokine production. GDP mannose decreased the production of eight cytokines (pan-inhibition) at the dose range of 0.1–1 mM (Fig. 2a). Three other metabolites (HMG-CoA, CoA, and mevalonate) similarly decreased the production of all tested cytokines at high concentration (Supplementary Figure 5a). Instead of pan-inhibition, l-homocysteine (1 mM) augmented production of most tested cytokines (Supplementary Figure 5b).
Representative bioactive metabolites altering CD4+ T-cell cytokine secretion. a Pan-inhibition of cytokine secretion by GDP mannose. b Th17-specific cytokine inhibition by ascorbate. c, d Th1-specific cytokine augmentation by oleic acid (c) and linoleic acid (d). For each of the two donors (black and gray), we calculated mean ± SEM from three technical replicates. N.A. condition that causes cell death and excluded
Other metabolites showed selective effects on Th17 cytokines. Ascorbate decreased Th17-related cytokine production (Fig. 2b), whereas there was no statistically significant change for other cytokines in this Th17-polarizing condition. Three metabolites at indicated conditions (0.1 mM glutaryl-CoA, 1 mM hexanoyl-CoA, and 1 mM isopentenyl diphosphate) enhanced the production of IL-17A and IL-17F (Supplementary Figure 6a).
The third group of metabolites selectively altered production of certain non-Th17 cytokines. Oleic acid and linoleic acid increased interferon (IFN)γ in a dose-dependent fashion (Fig. 2c, d); in contrast, pyridoxine phosphate decreased the production of tumor necrosis factor (TNF)α, IFNγ, and IL-10 at 0.1 mM (Supplementary Figure 6b).
Ascorbate inhibits T-cell cytokines in both polarized and non-polarized conditions
Among all 12 metabolites, ascorbate is the only microbe-derived metabolite that humans cannot synthesize, which drew our attention to this metabolite for deeper validation and mechanistic assessment. To further validate ascorbate bioactivity under Th17-polarizing conditions, we performed four additional biological replicates, and found that ascorbate consistently inhibited the production of Th17 cytokines. Flow cytometry analysis demonstrated that ascorbate induced a dose-dependent reduction in IL-17A-producing CD4+ T cells, including both IL-17A single-positive T cells and IFNγ/IL-17A double-positive cells (Fig. 3a). Measurement of cytokine secretion by Luminex confirmed the ascorbate inhibitory effect. All the cardinal Th17 cytokines (IL-17A, IL-17F, and IL-22) were reduced by >50% after 100 μM ascorbate treatment (Fig. 3b). These data confirm that ascorbate inhibits the production of Th17 and Th17-related cytokines under Th17-polarizing conditions.
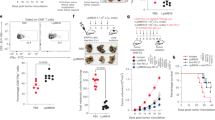
Ascorbate inhibits T-cell cytokines in both polarized and non-polarized conditions. a, b CD4+ T cells were cultured with 0–1 mM of ascorbate as described in Fig. 1. a Intracellular flow cytometry and (b) Luminex measurement of four biological replicates (four colors, mean ± SEM). c We cultured CD4+ T cells under T-cell activation condition with 0 or 0.1 mM of ascorbate. We used intracellular flow cytometry to determine the percentages of indicated CD4+ T cells. Mean ± SEM calculated from three biological replicates. N.A. condition that causes cell death and excluded
To investigate whether this inhibiting effect was specific to the Th17 subset, we cocultured CD4+ T cells with ascorbate under non-selective activating conditions (anti-CD3/CD28 without Th17-polaring cytokines (Fig. 3c). In the presence of ascorbate, IFNγ, IL-4, and IL-17A- producing CD4+ T cells were all reduced, whereas IL-10 producing cells were preserved. Both IL-17A-single-positive cells and IFNγ/IL-17A-double-positive cells were reduced in the presence of ascorbate. We concluded that ascorbate not only inhibited Th17 cells, but also reduced the activation of most other subsets of effector T cells (excluding IL-10-producing T cells).
Ascorbate selectively inhibits activated but not resting CD4+ T cells
We next determined whether ascorbate regulated all CD4+ T cells, including resting T cells and activated T cells. We cultured CD4+ T cells in resting and activating conditions for 48 h in the absence or presence of 100 μM ascorbate. Ascorbate selectively inhibited the upregulation of the activation marker CD154 in activating conditions (Fig. 4a, b). Annexin V and 7-AAD staining further demonstrated that activated but not resting T cells underwent programmed cell death after 48 h of ascorbate treatment (Fig. 4c). These data illustrated that ascorbate selectively inhibited activated versus resting T cells.
Ascorbate selectively inhibits activated but not resting CD4+ T cells. We cultured CD4+ T cells in 10 ng/ml of IL-7 (resting) or activated with 1 μg/ml anti-CD3 and anti-CD28 antibody (activated) for 48 h. We assessed T-cell activation (CD154+) by flow cytometry. a Representative plot. b Summary of the three independent experiments. c We measured cell death by Annexin V flow cytometry from three independent experiments
Ascorbate selectively inhibits glycolytic-dependent activated T cells through GLUT1
Among the several metabolic roles of ascorbate is its inhibition of glycolytic energy metabolism concomitant cell death in glycolytic-dependent cells.37 Consistent with these findings, we observed an increase of oxygen consumption rate (OCR) and the significant increase of extracellular acidification rate (ECAR) in the activated T cells (Fig. 5a, b). The decreased ratio of OCR/ECAR between the resting and activated condition further validated the switch of glycolysis in the activated condition. In addition, in the ascorbate-treated condition, we observed a significant reduction of ECAR in the activated but not resting T cells, suggesting that ascorbate treatment may affect the overall increase of OCR/ECAR ratio via reduction in glycolysis (Fig. 5b).
Seahorse platform measurements of OCR and ECAR. a, b We cultured CD4+ T cells in 10 ng/ml of IL-7 (resting) or activated with 1 μg/ml anti-CD3 and anti-CD28 antibody (activated) for 48 h, with or without 100 μM ascorbate treatment. c, d Scramble and GLUT1-specific clone of siRNA was transfected via Nucleofector™ kit (Lonza). After transfection, cells were rested in 10 ng/ml of IL-7 (resting) or activated with 1 μg/ml anti-CD3 and anti-CD28 antibody (activated) for 48 h, under the conditions with or without 100 μM ascorbate treatment. e Ratio of oxidative to glycolytic energy production. Each seahorse experiment has been performed on three biological donors of cells and one of the representative plots was shown. Mean ± SEM calculated from three technical replicates
It is well known that TCR activation, specifically CD28 signaling, directly controls the metabolic switch to glycolysis by upregulating the expression of glucose transporter 1 (GLUT1), and remodeling activity of downstream biochemical pathways to favor glycolytic versus oxidative phosphorylation energy production in T cells.38,39,40 Since GLUT1 is also a transporter for the oxidized form of ascorbate,38 we hypothesized that ascorbate selectively inhibits glycolytic-dependent activated T cells through GLUT1. To evaluate the role of GLUT1 in ascorbate-induced inhibition, the GLUT1 and scramble small interfering RNA (siRNA) were transfected into activated CD4+ T cells and the knockdown efficiency was validated by GLUT1 western blot (Supplementary Figure 7). Like cells treated with ascorbate alone, we observed a significant decrease of ECAR (Fig. 5d, scramble) and significant increase of OCR/ECAR ratio in the ascorbate-treated scramble siRNA condition (Fig. 5e, scramble). In contrast, these changes were extinguished in the ascorbate-treated GLUT1 siRNA condition. This finding suggests that GLUT1 is required for the ascorbate-induced alteration of energy metabolism in activated T cells (Fig. 5d, e). Together, these data support that a selective inhibitory effect of ascorbate on activated T cells was related to a mechanism that regulated energy metabolism through a GLUT1-dependent action.
Ascorbate production by microbial taxa associated with Crohn’s disease
To investigate whether gut microbes can produce ascorbate, we performed a reverse metagenomic search for the bacterial genomes that contained ascorbate biosynthesis enzymes. The metabolic pathways utilized by microbes to produce ascorbate de novo are largely unknown, but biotechnology has suggested two microbial pathways to manufacture l-ascorbate from d-glucose.41 We therefore investigated the chemical reactions in these processes, and tabulated the metabolic enzymes and corresponding metagenes involved in ascorbate synthesis reactions. Using this tabulation, we performed a reverse metagenomics search of human fecal microbiota bearing some or all of the genes in the archetypal d-sorbitol and 2,5-diketo-d-gluconic acid pathways of ascorbate biosynthesis (Supplementary Tables 2 and 3). We identified five operational taxonomic units (OTUs) bearing the ascertainable metagenes of the d-sorbitol pathway, all of them in the Crohn’s disease-associated genus Burkholderia (Fig. 6a). We also identified 120 OTUs for the 2,5-diketo-d-gluconic acid ascorbate pathway; of these, the most prevalent taxa belonged to the Crohn’s disease-associated genera Pseudomonas and Erwinia (Fig. 6b). To directly assess ascorbate production by these microbes, we assembled a collection of three clinical isolates of Burkholderia and two strains of Pseudomonas aeruginosa. After aerobic culture to stationary phase, ascorbate production was detectable in both Pseudomonas strains (Fig. 6c), but not in the Burkholderia isolates (data not shown). These findings indicate that certain Crohn’s disease-associated microbiota are proficient at ascorbate biosynthesis, and point to the 2,5-diketo-gluconic acid pathway as a potential mechanism for this production.
Gut microbes producing ascorbate. a Enzymes involved in the sorbitol pathway or (b) in the 2,5-diketo-d-gluconic acid pathway of ascorbate biosynthesis. c We cultured bacteria for 24 h and then collected supernatants to measure ascorbate levels. Since all cultures had reached equivalent McFarland readings at 24 h, the ascorbate levels represent production from comparable cell numbers. OTUs operational taxonomic units, Left numbers of OTUs bearing metagenes for the indicated number of pathway enzymes, Right bar plots of the numbers of OTUs expressing indicated enzymes in bacterial genera
Discussion
By screening microbial metabolites using primary human CD4+ T cells in Th17-polarizing culture conditions, we identified three known and 12 novel microbial metabolites with the potential to module T-cell cytokine production. A detailed study of one novel microbial-derived metabolite, ascorbate, revealed inhibition of Th17 and activated T-cell formation by a mechanism involving blockade of glycolytic energy metabolism. We metagenomically inferred and functionally validated examples of Crohn’s disease-associated microbiota proficient for ascorbate production. These findings suggest a substantial frequency of relevant T-cell bioactivity among Crohn’s disease-associated microbial metabolites, and evidence for novel modes of bioactivity, including targeting of T-cell energy metabolism.
This study emphasizes the importance of environmental metabolic cues to modulate CD4+ T-cell functions, notably via differentiation and activation of specific subsets of CD4+ T cells with distinct cytokine expression patterns.42,43 Three bioactive metabolites identified in our study confirmed previous observations of these metabolites as selective mediators of such differentiation and activation.13,34,35 The bioactivity of 12 additional metabolites for human CD4+ T cells did not have literature precedents to our knowledge. The diversity of their effects on T-cell cytokine function suggests that their mechanistic assessment may reveal new insights on the action of microbial metabolites in CD4+ T-cell function, and the role of these actions in disease biology.
We studied the mechanism of ascorbate, since it was unknown for inhibitory action on activated CD4+ T cells until a recent independent study.44 Among its several biochemical roles, ascorbic acid in its oxidized form (dehydroascorbate), transported by the GLUT hexose transporter family, inhibits glycolytic energy metabolism by oxidative inactivation of glyceraldehyde-3-phosphate dehydrogenase, an apex enzyme in glycolytic energy metabolism.37 The present study demonstrated that in active CD4+ T cells, the predominant action of ascorbate was as an inhibitor of glycolytic energy production. Concordantly, we also observed that this effect was dependent on GLUT1, the major T-cell hexose transporter isoform,38,45 and the transporter class responsible for uptake of oxidized ascorbate.46,47
Intracellular dehydryoascorbate is an efficient substrate for enzymatic glutathione oxidation,48 and at high levels, it rapidly increases reduced glutathione (GSSH) levels leading to downstream endogenous reactive oxygen species (ROS).49 GAPDH is a sensitive target for oxidative inhibition via two pathways. First, ROS induces S-glutathionylation or S-sulfonylation of the cysteine residue on GAPDH, diminishing GAPDH activity.50,51,52 In addition, accumulation of endogenous ROS induces DNA damage leading to activation of PARP, depleting levels of NAD+, the substrate for GAPDH.37,53 Both mechanisms cause the reduction of GAPDH activity, which leads to the impaired glycolysis, and cell death in selectively glycolysis-dependent cells.
It is well known that the early phase of T-cell activation induces a metabolic switch to and dependence upon aerobic glycolysis.38,39,40 This provides a plausible mechanism by which ascorbate, via interruption of glycolytic energy production, may account for the selective functional inhibition and apoptosis of activated CD4+ T cells. However, a recent report observed that after completion of Th17 induction, these cells shift again to oxidative phosphorylation as their preferential energy source.54 Incorporating this point, we speculate that the potential in vivo action of ascorbate on Th17 cells may be limited to the stages of Th17 induction and activation and to their relevant anatomic sites (such as mesenteric nodes in the case of intestinal Th17 cells).
A distinct action of ascorbate on energy metabolism is its augmentation of oxidative phosphorylation via alpha-ketoglutarate-dependent mitochondrial ascorbic acid recycling. This is mediated by reduced ascorbate, which is transported at the surface or mitochondrial membrane by the sodium transporter family members SVCT1 and 2.55,56,57,58 However, in our study, the ascorbate effect on OCR was modest and not altered by GLUT1 knockdown (Fig. 5), suggesting that augmented TCA cycling was a minor mode of action for ascorbate on T-cell energy metabolism. A recent study reported SVCT2 expression and sodium-dependent ascorbate uptake in T cells, and ascorbate-mediated inhibition of T-cell activation.44 This study was distinguished from the present one by the requirement for supraphysiologic levels of ascorbate (0.5 mM), and the role of energy metabolism was not delineated. It is possible that the inhibitory mechanism of high ascorbate is distinct, and suggests a role for mitochondrial SVCT2, since it is a low-affinity ascorbate receptor at high divalent cation levels characteristic of the cytoplasm.57
We also observed that ascorbate inhibits the production of IFNγ-, IL-4, and IL-17A, but not IL-10. Unlike other CD4+ T-cell subsets, induced Tregs (which inhibit effector T-cell responses in part through IL-10 release) do not substantially shift to glycolytic energy metabolism,39,40 and this may account for their resistance to ascorbate inhibition. Moreover, in mice, ascorbate facilitates the demethylation of Foxp3 enhancer and promotes Treg cell function.59,60 Taken together, these observations suggest that ascorbate as an environmental cue may selectively inhibit effector CD4+ T-cell responses.
An issue raised by these observations is the physiological role of dietary ascorbate versus local microbial ascorbate production. Unlike most eutherians, humans and primates require ascorbate intake, because they genetically lack the enzyme responsible for the last step in ascorbate synthesis, l-gulonolactone oxidase.61 Previous work indicates that microbial ascorbate is not the major nutritional source for vitamin C in humans,62 suggesting the importance of dietary intake of ascorbate in human. However, dietary ascorbate is largely absorbed before transit to the colon, so local microbial production may be a significant contributor to luminal ascorbate in the distal intestine. The predicted ascorbate-producing genera are Proteobacteria, an uncommon intestinal phylum most prevalent in the colon. P. aeruginosa and related species are notable for their production of redox-active small molecules that modify the cellular redox state, acting as inter-microbial signals to control biofilm formation.63 Ascorbate may represent an additional class of redox-active molecules involved in such inter-microbial interaction. Microbial genetics and ecologic studies are necessary to assess the control and bioavailability of ascorbate in the gut, and test its potential role in microbial ecology.
Clinically, ascorbate levels and the ratio of oxidized versus reduced form of ascorbate are distinct in inflammatory bowel disease (IBD) patients versus healthy subjects.64 Genetic variations in the sodium-dependent ascorbate transporter gene, SLC23A1, have been significantly associated with susceptibility to Crohn’s disease.65 The disease variant of human SLC23A1 results in lower transporter activity and decreased intracellular ascorbate. Decreased ascorbate transport limits the capacity to prevent oxidative tissue damage and impair recovery from mucosal injury.66 However, the mechanistic basis of these disease-associated phenotypes has not yet been probed in experimental animal models, because except for primates, eutherians are proficient for ascorbate synthesis. Such studies, and testing the potential in vivo roles of microbial ascorbate on CD4+ T cells and Crohn’s disease, await genetic engineering of mice bearing the human-homologous l-gulonolactone oxidase deficiency.
Collectively, this study demonstrates the utility of a screening strategy to probe the T-cell bioactivity of Crohn’s disease -associated microbial metabolites. These results suggest that the balance of luminal metabolites could be important to maintain physiological CD4+ T-cell functions of the intestinal mucosa. The implications for future studies include testing in vivo whether levels of bioactive metabolites correlate with disease state, the abundance of cognate producing microbial taxa, and cognate changes in local T-cell function attributable to these metabolites.
Methods
Human blood sample collection and T-cell isolation
We recruited healthy donors at UCLA immune assessment core, and collected blood samples according to an IRB protocol approved by the institutional review committee of University of California, Los Angeles (protocol #10-001689). Human CD4+ T cells were isolated using Rosettesep Human CD4+ T-cell enrichment cocktails (StemCell™ technology) following manufacturer’s instructions. Purified CD4+ T cells were confirmed to be >95% pure by flow cytometry and the T-cell subset composition was monitored (Supplementary Figure 1). We cultured T cells with soluble anti-CD28 (0.2 μg/ml) antibodies on 96-well plates pre-coated with anti-CD3 antibodies (1 μg/ml). IL-2 was added at 4.8 U/ml (2 ng/ml). We induced Th17 differentiation with 50 ng/ml IL-6, 50 ng/ml IL-1β, and 50 ng/ml IL-23.33 We refed T cells at day 7 with medium containing the same Th17-polarizing cytokines and further cultured for another 7 days. We collected supernatants for multiplex Luminex cytokine analysis, and analyzed T cells by intracellular flow cytometry.
Cell survival assay
The survival of metabolites-conditioned cells was examined by SYTOX™ Red dead cell stain (Molecular Probes, cat. no. S34859), which is a nucleic acid stain that penetrates cells with compromised plasma membranes. We used the SYTOX™ Red dead cell stain after cell surface marker staining, and detected SYTOX Red levels by flow cytometry.
Intracellular flow cytometry
At the end of the 14-day culture, T cells were stimulated with Leukocyte activation cocktail with BD GolgiPlug (BD Biosciences, cat. no. 550583) for 4 h and washed in FACS buffer. BV650-conjugated CD3 (clone OKT3) and PE-CF594-conjugated CD4 (clone OKT4) antibody were used to detect these cell surface markers. We performed intracellular fixation and permeabilization with a commercial buffer set (eBiosciences, cat. no. 88-8824). The following antibodies were used to detect intracellular cytokines: APC-conjugated IL-17A (clone eBio64DEC17), FITC-conjugated IFNγ (clone 4S.B3), PE-conjugated IL-10 (clone JES3-9D7), and PE-Cy7-conjugated IL-4 (clone 8D4-8) were purchased from eBioscience or Biolegend.
Cell respiration measurements
We activated isolated CD4+ T cells on plates coated with 10 μg/ml anti-CD3 and 10 μg/ml anti-CD28, and stimulated in the presence of 10 ng/ml IL-2 as the activated condition, and 10 ng/ml IL-7 as the resting condition. After 48 h, activated or resting cells were collected, washed with unbuffered DMEM (2 g/l glucose), and seeded into CELL-TAK™ (Corning cat. no. 354240)-covered 96-well plates (Agilent Seahorse XF96) (0.5 million cells per well) for OCR and ECAR measurement. We measured cellular OCRs and ECARs as described previously.38 Oxygen consumption was blocked by oligomycin (1.5 μM), an ATP synthase inhibitor; the ionophore FCCP (carbonyl cyanide-p-trifluoromethoxyphenylhydrazone; 2 and 3 uM) assayed maximal respiratory capacity of mitochondria, whereas rotenone plus antimycin A (3 μM), a mitochondrial inhibitor, was used to block mitochondrial respiration. Three independent experiments were performed, and three technical replicates per experiment were measured to calculate the mean ± SEM for each time point.
Bioinformatic prediction of Crohn’s disease-associated microbial metabolites
Using a previously reported PRISM dataset32 of microbial composition at the level of OTUs from 16S ribosomal RNA gene amplicon sequencing data, we tabulated microbial count data for healthy and Crohn’s disease subjects at the subject level, and converted them to imputed metagene data using PICRUSt.67 We then determined the relative abundance of enzymatic pathways using HUMAnN2,68 with pathway end products identified using UniRef and MetaCyc databases. Making the assumption that metabolite abundance is the sum of abundances of the generating enzymes, we calculated the predicted abundance of each metabolite using the average of most abundant route through each pathway.
Disease-associated metabolites were determined using a multivariate model comparing predicted relative compound levels in health versus Crohn’s disease subjects. Specifically, we performed a generalized linear mixed model (glmmPQL, implemented in R), correcting for age and medications (antibiotics, immunosuppressants, and anti-inflammatory agents). Disease-associated metabolites from this analysis were ranked by multi-testing-corrected q value. From this raw list, the top molecules for which commercial sourcing could be identified were tabulated (Supplementary Table 1), and procured for testing.
For prediction of ascorbate-producing microbiota, we used the reverse of this bioinformatics pipeline, with the following modifications. Microbial metabolic pathways and enzymes involved in ascorbate biosynthesis were defined using KEGG and literature reports. For those enzymes annotated in KEGG or MetaCyc databases, we performed a prediction of microbiota genomically bearing them using PICRUSt-precalculated gene content. We filtered these results for human fecal-associated OTUs, generating lists of OTUs bearing various numbers of ascorbate pathway genes. Finally, we filtered for OTUs validated for significant differential association in Crohn’s disease versus healthy subjects.
The computational resources for all analytic steps are available at https://bitbucket.org/biobakery/biobakery/wiki/Home.
Metabolite preparation
All chemical were of cell culture or higher grade, and purchased from Sigma-Aldrich. The majority of chemicals were dissolved in aqueous solution; water-insoluble metabolites were dissolved in DMSO.
Luminex multiplex cytokine production
We measured cytokines from CD4+ T-cell culture supernatants using Luminex. The following 11 cytokines were included in the Luminex panels and acquired on a Luminex® FLEXMAP 3D: Th17 cytokines, IL-17A, IL-17F, and IL-22; Th1 cytokines, TNFα, and IFNγ; and Th2 cytokines, IL-5, and IL-13; Treg cytokine, IL-10; other related cytokines, IL-9, IL-21, and MIP-3a. We normalized the Luminex readouts for the cell count values.
Detecting ascorbate in bacterial culture
P. aeruginosa included two strains, PA-01 and PA-14, obtained from the ATCC. Burkholderia cepacia included three clinical isolates (2-1, 3-1, 3-2; source strains ZK2853-ZK2862) from two cystic fibrosis patients,69 a gift from the collection of Robert Kolter (Harvard Medical School). Bacteria were cultured in 1 ml Luria-Bretani broth at 35 ± 2 °C under aerobic condition, and the bacterial cell numbers were monitored by McFarland standard.70 After pelleting bacterial cells (5000 r.p.m., 10 min), we filtered the supernatants using centrifugal filters per manufacturer instructions (Corning® Costar® Spin-X® centrifuge tube filters, 0.22 μm). We measured ascorbate levels in supernatant filtrates by an enzyme-based assay (Abcam, ab65656).
Statistical analysis
To estimate contributors of experimental variability and control for potential technical effects, we performed principal variance component analysis (PVCA). The technical factors assessed included metabolite, experimental run, donor, and technical replicates in different plates. We used principal components analysis (PCA) to reduce data dimensionality data from the 11 cytokines, and to maintain majority of the variability in the data. We used VCA to fit a mixed effect model using subject-specific factors as random effects to estimate and partition the total variability. Supplementary Figure 3 shows one example. We ran PVCA for every experiment to ensure the stability of technical variation.
We used MANOVA to identify putative bioactive metabolites. In each condition treated with a specific metabolite concentration, we compared the levels of the 11 cytokines to those found in the vehicle control condition. We defined a metabolite as bioactive if p values <0.05; we confirmed bioactivity in a different healthy donor to confirm metabolite bioactivity.
To further validate biological effects and define specific cytokine responses to bioactive metabolites, we employed two-way ANOVA with metabolite and donor factors using the incorporated data from at least two independent experiments. We used the Holm–Sidak’s post-test to correct for multiple comparisons. We defined statistical significance as adjusted p values <0.05.
We generated cytokine heatmaps from Luminex cytokine data by calculating the mean of log2 fold change between metabolite-untreated versus metabolite-treated samples (treated/untreated). We used the heatmap.2 function in gplot package to generate dendrogram and colorplot in R version 3.2.4.
We performed statistical analyses with Prism software (GraphPad) or using the R program (R version 3.1.1). We used the nonparametric Mann–Whitney test to compare two experimental conditions. We defined statistically significant results are indicated (*p < 0.05, ***p < 0.001). We displayed error bars using mean ± SEM calculated from three or more experimental replicates.
References
Integrative HMPRNC. The Integrative Human Microbiome Project: dynamic analysis of microbiome-host omics profiles during periods of human health and disease. Cell Host Microbe 16, 276–289 (2014).
Huttenhower, C., Kostic, A. D. & Xavier, R. J. Inflammatory bowel disease as a model for translating the microbiome. Immunity 40, 843–854 (2014).
Sartor, R. B. & Wu, G. D. Roles for intestinal bacteria, viruses, and fungi in pathogenesis of inflammatory bowel diseases and therapeutic approaches. Gastroenterology 152, 327–339.e324 (2017).
Brestoff, J. R. & Artis, D. Commensal bacteria at the interface of host metabolism and the immune system. Nat. Immunol. 14, 676–684 (2013).
Dorrestein, P. C., Mazmanian, S. K. & Knight, R. Finding the missing links among metabolites, microbes, and the host. Immunity 40, 824–832 (2014).
Thorburn, A. N., Macia, L. & Mackay, C. R. Diet, metabolites, and “western-lifestyle” inflammatory diseases. Immunity 40, 833–842 (2014).
Kaiko, G. E. et al. The colonic crypt protects stem cells from microbiota-derived metabolites. Cell 167, 1137 (2016).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 15, 382–392 (2014).
Tong, M. et al. Reprograming of gut microbiome energy metabolism by the FUT2 Crohn’s disease risk polymorphism. ISME J. 8, 2193–2206 (2014).
Lewis, J. D. et al. Inflammation, antibiotics, and diet as environmental stressors of the gut microbiome in pediatric Crohn’s disease. Cell Host Microbe 18, 489–500 (2015).
Gao, Z. et al. Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes 58, 1509–1517 (2009).
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Chang, P. V., Hao, L., Offermanns, S. & Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl Acad. Sci. USA 111, 2247–2252 (2014).
Mascanfroni, I. D. et al. Metabolic control of type 1 regulatory T cell differentiation by AHR and HIF1-alpha. Nat. Med. 21, 638–646 (2015).
Hooper, L. V., Littman, D. R. & Macpherson, A. J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Knights, D. et al. Complex host genetics influence the microbiome in inflammatory bowel disease. Genome Med. 6, 107 (2014).
Khor, B., Gardet, A. & Xavier, R. J. Genetics and pathogenesis of inflammatory bowel disease. Nature 474, 307–317 (2011).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Yen, D. et al. IL-23 is essential for T cell-mediated colitis and promotes inflammation via IL-17 and IL-6. J. Clin. Invest. 116, 1310–1316 (2006).
Fujino, S. et al. Increased expression of interleukin 17 in inflammatory bowel disease. Gut 52, 65–70 (2003).
Kleinschek, M. A. et al. Circulating and gut-resident human Th17 cells express CD161 and promote intestinal inflammation. J. Exp. Med. 206, 525–534 (2009).
Bogaert, S. et al. Differential mucosal expression of Th17-related genes between the inflamed colon and ileum of patients with inflammatory bowel disease. BMC Immunol. 11, 61 (2010).
Kobayashi, T. et al. IL23 differentially regulates the Th1/Th17 balance in ulcerative colitis and Crohn’s disease. Gut 57, 1682–1689 (2008).
Neurath, M. F. New targets for mucosal healing and therapy in inflammatory bowel diseases. Mucosal Immunol. 7, 6–19 (2014).
Hueber, W. et al. Secukinumab, a human anti-IL-17A monoclonal antibody, for moderate to severe Crohn’s disease: unexpected results of a randomised, double-blind placebo-controlled trial. Gut 61, 1693–1700 (2012).
Sandborn, W. J. et al. Ustekinumab induction and maintenance therapy in refractory Crohn’s disease. N. Engl. J. Med. 367, 1519–1528 (2012).
Sands, B. E. et al. OP025: a randomized, double-blind placebo-controlled phase 2a induction study of MEDI2070 (anti-p19 antibody) in patients with active Crohn’s disease who have failed anti-TNF antibody therapy. J. Crohn’s Colitis 9, S15–S16 (2015).
Targan, S. R. et al. Mo2083: a randomized, double-blind, placebo-controlled study to evaluate the safety, tolerability, and efficacy of AMG 827 in subjects with moderate to severe Crohn’s disease. Gastroenterology 143, e26 (2012).
Colombel, J. F., Sendid, B., Jouault, T. & Poulain, D. Secukinumab failure in Crohn’s disease: the yeast connection? Gut 62, 800–801 (2013).
Maxwell, J. R. et al. Differential roles for interleukin-23 and interleukin-17 in intestinal immunoregulation. Immunity 43, 739–750 (2015).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13, R79 (2012).
Wilson, N. J. et al. Development, cytokine profile and function of human interleukin 17-producing helper T cells. Nat. Immunol. 8, 950–957 (2007).
Wang, C. et al. CD5L/AIM regulates lipid biosynthesis and restrains Th17 cell pathogenicity. Cell 163, 1413–1427 (2015).
Pacheco, R. et al. Glutamate released by dendritic cells as a novel modulator of T cell activation. J. Immunol. 177, 6695–6704 (2006).
Wishart, D. S. et al. HMDB 3.0--The Human Metabolome Database in 2013. Nucleic Acids Res. 41, D801–D807 (2013).
Yun, J. et al. Vitamin C selectively kills KRAS and BRAF mutant colorectal cancer cells by targeting GAPDH. Science 350, 1391–1396 (2015).
Macintyre, A. N. et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 20, 61–72 (2014).
Buck, M. D., O’Sullivan, D. & Pearce, E. L. T cell metabolism drives immunity. J. Exp. Med. 212, 1345–1360 (2015).
Slack, M., Wang, T. & Wang, R. T cell metabolic reprogramming and plasticity. Mol. Immunol. 68, 507–512 (2015).
Hancock, R. D. & Viola, R. Biotechnological approaches for L-ascorbic acid production. Trends Biotechnol. 20, 299–305 (2002).
DuPage, M. & Bluestone, J. A. Harnessing the plasticity of CD4(+) T cells to treat immune-mediated disease. Nat. Rev. Immunol. 16, 149–163 (2016).
Sallusto, F. Heterogeneity of human CD4(+) T cells against microbes. Annu. Rev. Immunol. 34, 317–334 (2016).
Hong, J. M., Kim, J. H., Kang, J. S., Lee, W. J. & Hwang, Y. I. Vitamin C is taken up by human T cells via sodium-dependent vitamin C transporter 2 (SVCT2) and exerts inhibitory effects on the activation of these cells in vitro. Anat. Cell Biol. 49, 88–98 (2016).
Maratou, E. et al. Glucose transporter expression on the plasma membrane of resting and activated white blood cells. Eur. J. Clin. Invest. 37, 282–290 (2007).
Rumsey, S. C. et al. Glucose transporter isoforms GLUT1 and GLUT3 transport dehydroascorbic acid. J. Biol. Chem. 272, 18982–18989 (1997).
Vera, J. C., Rivas, C. I., Fischbarg, J. & Golde, D. W. Mammalian facilitative hexose transporters mediate the transport of dehydroascorbic acid. Nature 364, 79–82 (1993).
May, J. M., Mendiratta, S., Hill, K. E. & Burk, R. F. Reduction of dehydroascorbate to ascorbate by the selenoenzyme thioredoxin reductase. J. Biol. Chem. 272, 22607–22610 (1997).
Winkler, B. S., Orselli, S. M. & Rex, T. S. The redox couple between glutathione and ascorbic acid: a chemical and physiological perspective. Free Radic. Biol. Med. 17, 333–349 (1994).
Hwang, N. R. et al. Oxidative modifications of glyceraldehyde-3-phosphate dehydrogenase play a key role in its multiple cellular functions. Biochem. J. 423, 253–264 (2009).
Shenton, D. & Grant, C. M. Protein S-thiolation targets glycolysis and protein synthesis in response to oxidative stress in the yeast Saccharomyces cerevisiae. Biochem. J. 374, 513–519 (2003).
Ravichandran, V., Seres, T., Moriguchi, T., Thomas, J. A. & Johnston, R. B. Jr. S-thiolation of glyceraldehyde-3-phosphate dehydrogenase induced by the phagocytosis-associated respiratory burst in blood monocytes. J. Biol. Chem. 269, 25010–25015 (1994).
Ullah, M. F., Khan, H. Y., Zubair, H., Shamim, U. & Hadi, S. M. The antioxidant ascorbic acid mobilizes nuclear copper leading to a prooxidant breakage of cellular DNA: implications for chemotherapeutic action against cancer. Cancer Chemother. Pharmacol. 67, 103–110 (2011).
Franchi, L. et al. Inhibiting oxidative phosphorylation in vivo restrains Th17 effector responses and ameliorates murine colitis. J. Immunol. 198, 2735–2746 (2017).
Fiorani, M. et al. The mitochondrial transporter of ascorbic acid functions with high affinity in the presence of low millimolar concentrations of sodium and in the absence of calcium and magnesium. Biochim. Biophys. Acta 1848, 1393–1401 (2015).
Tsukaguchi, H. et al. A family of mammalian Na+-dependent L-ascorbic acid transporters. Nature 399, 70–75 (1999).
Savini, I., Rossi, A., Pierro, C., Avigliano, L. & Catani, M. V. SVCT1 and SVCT2: key proteins for vitamin C uptake. Amino Acids 34, 347–355 (2008).
Li, X., Cobb, C. E. & May, J. M. Mitochondrial recycling of ascorbic acid from dehydroascorbic acid: dependence on the electron transport chain. Arch. Biochem. Biophys. 403, 103–110 (2002).
Sasidharan Nair, V., Song, M. H. & Oh, K. I. Vitamin C facilitates demethylation of the Foxp3 enhancer in a Tet-dependent manner. J. Immunol. 196, 2119–2131 (2016).
Yue, X. et al. Control of Foxp3 stability through modulation of TET activity. J. Exp. Med. 213, 377–397 (2016).
Nishikimi, M., Fukuyama, R., Minoshima, S., Shimizu, N. & Yagi, K. Cloning and chromosomal mapping of the human nonfunctional gene for L-gulono-gamma-lactone oxidase, the enzyme for L-ascorbic acid biosynthesis missing in man. J. Biol. Chem. 269, 13685–13688 (1994).
Naidu, K. A. Vitamin C in human health and disease is still a mystery? An overview. Nutr. J. 2, 7 (2003).
Costa, K. C., Glasser, N. R., Conway, S. J. & Newman, D. K. Pyocyanin degradation by a tautomerizing demethylase inhibits Pseudomonas aeruginosa biofilms. Science 355, 170–173 (2017).
Vera, J. C. et al. Resolution of the facilitated transport of dehydroascorbic acid from its intracellular accumulation as ascorbic acid. J. Biol. Chem. 270, 23706–23712 (1995).
Amir Shaghaghi, M., Bernstein, C. N., Serrano Leon, A., El-Gabalawy, H. & Eck, P. Polymorphisms in the sodium-dependent ascorbate transporter gene SLC23A1 are associated with susceptibility to Crohn disease. Am. J. Clin. Nutr. 99, 378–383 (2014).
Hengstermann, S. et al. Altered status of antioxidant vitamins and fatty acids in patients with inactive inflammatory bowel disease. Clin. Nutr. 27, 571–578 (2008).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 8, e1002358 (2012).
Sokol, P. A., Darling, P., Woods, D. E., Mahenthiralingam, E. & Kooi, C. Role of ornibactin biosynthesis in the virulence of Burkholderia cepacia: characterization of pvdA, the gene encoding L-ornithine N(5)-oxygenase. Infect. Immun. 67, 4443–4455 (1999).
Mc, F. J. The nephelometer:aN instrument for estimating the number of bacteria in suspensions used for calculating the opsonic index and for vaccines. J. Am. Med. Assoc. XLIX, 1176–1178 (1907).
Acknowledgements
This study was supported by the Crohn’s and Colitis Foundation 323814, and National Institutes of Health grants PO1-DK46763 (J.B., D.P.B.M.), P30-CA016042 (UCLA Jonsson Comprehensive Cancer Center), and NCATS UCLA CTSI UL1TR001881. We thank all the volunteers for their participation in this study. We particularly thank the Immune Assessment Core at University of California, Los Angeles, for their support in scaling up the screens and running Luminex experiments. We also thank members from Dr. Michael Teitell Laboratory, Dr. Laurent Vergnes, and Dr. Linsey Stiles from Cellular Bioenergetics Core at University of California, Los Angeles, for the help with the Seahorse experiments.
Author information
Authors and Affiliations
Contributions
Y.-L.C., M.R. and J.B. were responsible for experimental design, data interpretation, and manuscript preparation. Y.-L.C., G.S., N.H., and S.M. performed the experiments. Y.-L.C. and D.C. contributed to data analysis, and C.H. and H.V. predicted metabolites from metagenomics analysis. R.H., K.W.S., R.B.S., G.W., J.L., F.B., D.P.B.M., N.S., and J.B. in the CCFA consortium contributed to study design. J.B., T.S., R.X., and C.H. contributed to information integration and analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Chang, YL., Rossetti, M., Vlamakis, H. et al. A screen of Crohn’s disease-associated microbial metabolites identifies ascorbate as a novel metabolic inhibitor of activated human T cells. Mucosal Immunol 12, 457–467 (2019). https://doi.org/10.1038/s41385-018-0022-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-018-0022-7
This article is cited by
-
Vitamins as regulators of calcium-containing kidney stones — new perspectives on the role of the gut microbiome
Nature Reviews Urology (2023)
-
The evolution of vitamin C biosynthesis and transport in animals
BMC Ecology and Evolution (2022)
-
Food for thought – ILC metabolism in the context of helminth infections
Mucosal Immunology (2022)
-
Highly selective and efficient electrochemical sensing of ascorbic acid via CuO/rGO nanocomposites deposited on conductive fabric
Applied Physics A (2022)
-
Gut instincts in neuroimmunity from the eighteenth to twenty-first centuries
Seminars in Immunopathology (2022)