Abstract
Attention-deficit/hyperactivity disorder (ADHD) and autism spectrum disorders (ASD) are strongly associated with educational attainment (EA), but little is known about their genetic relationship with school performance and whether these links are explained, in part, by the genetic liability of EA. Here, we aim to dissect the polygenic contribution of ADHD and ASD to school performance, early manifestation of psychopathology and other psychiatric disorders and related traits by their relationship with EA. To do so, we tested the association of polygenic scores for EA, ADHD and ASD with school performance, assessed whether the contribution of the genetic liability of ADHD and ASD to school performance is influenced by the genetic liability of EA, and evaluated the role of EA in the genetic overlap between ADHD and ASD with early manifestation of psychopathology and other psychiatric disorders and related traits in a sample of 4,278 school-age children. The genetic liability for ADHD and ASD dissected by their relationship with EA show differences in their association with school performance and early manifestation of psychopathology, partly mediated by ADHD and ASD symptoms. Genetic variation with concordant effects in ASD and EA contributes to better school performance, while the genetic variation with discordant effects in ADHD or ASD and EA is associated with poor school performance and higher rates of emotional and behavioral problems. Our results strongly support the usage of the genetic load for EA to dissect the genetic and phenotypic heterogeneity of ADHD and ASD, which could help to fill the gap of knowledge of mechanisms underlying educational outcomes.
Similar content being viewed by others
Introduction
Attention-deficit/hyperactivity disorder (ADHD) and autism spectrum disorder (ASD) are complex childhood-onset neurodevelopmental disorders that often co-occur, are highly polygenic and share common genetic architecture [1]. Both conditions are associated with educational outcomes including educational attainment (EA) [2, 3], which also has substantial polygenic contribution and is linked to a broad range of outcomes [2, 4,5,6,7,8,9,10].
Although the genetic relationship between ADHD and ASD with EA has been widely studied, its complex nature still remains largely unclear. A large proportion of ADHD and ASD risk variants are shared with EA, but while almost all ADHD risk loci are associated with worse educational outcomes, ASD genetic variants showed mixed direction of effects [3, 11]. Consistently, there is a strong negative genetic correlation between ADHD and EA and a moderate positive genetic correlation between ASD and EA [1]. Recent findings support the existence of a polygenic form of pleiotropy that contributes to the positive genetic correlation between ADHD and ASD, and is consistent with the discordant polygenic association between these neurodevelopmental disorders and EA [12]. Also, polygenic scores for ADHD are associated with lower levels of EA, while polygenic risk for ASD is associated with higher cognitive functionality and EA [3, 13]. These distinct patterns of association with EA align with mendelian randomization results showing causal effects with opposite direction between these neurodevelopmental disorders and EA [14].
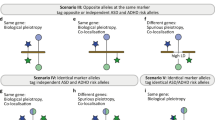
Phenotypic associations between school performance, considered an early life intermediate phenotype that may anticipate EA [6,7,8, 15,16,17], and ADHD or ASD align with their pattern of genetic correlation with EA [18, 19]. Children with ADHD are at increased risk of learning disabilities and communication disorders, obtain lower grades than their peers without ADHD and are at greater risk of failure to graduate from high school on time [20,21,22], while academic functioning and achievement in ASD is highly variable [23,24,25,26,27,28]. However, little is known about the genetic underpinnings for the relationship between these neurodevelopmental disorders and school performance and whether these links are explained, in part, by the genetic liability of EA. We aim to dissect the polygenic contribution of ADHD and ASD to school performance by its relationship with EA in a deeply phenotyped cohort of 4278 school-age children and adolescents (see the workflow in Fig. 1) by (i) testing the association of polygenic scores for EA, ADHD and ASD with school performance, (ii) assessing whether the contribution of the genetic liability of ADHD and ASD to school performance is influenced by the genetic liability of EA and (iii) evaluating the role of EA in the genetic overlap between ADHD and ASD with early manifestation of psychopathology and other psychiatric disorders and related traits.
GWAS summary statistics on educational attainment (EA) (orange rectangle) [4], attention-deficit/hyperactivity disorder (ADHD) (blue rectangle) [3] and autism spectrum disorder (ASD) (green rectangle) [13] were used. The genetic liability of ADHD and ASD were dissected by their relationship with EA (see material and methods). With these genomic partitions we conducted genome-wide polygenic scores (PGS) analyses and test their association with school performance and early manifestation of psychopathology in our deeply phenotyped cohort of 4278 school-age children and adolescents and performed analyses of partitioned heritability and annotation-based stratified genetic covariance. “+” and “−” in the figure refer to the direction of the effect of the alleles studied.
Materials and methods
Study sample
The study sample included 4278 school-age children and adolescents from the INSchool cohort (mean age = 10.03 years, SD = 2.95; 43.7% females) from 45 primary and secondary schools across Catalunya, Spain, with GWAS and data on school performance available. Inclusion criteria for the participants were European ancestry and written informed consent from parents/caregivers prior to participation. The project was approved by the Ethics Committee at the Hospital Universitari Vall d’Hebron and the Hospital maternoinfantil Sant Joan de Déu.
School performance
School performance data were provided by schools and assessed with school grades in three different subjects: mathematics, foreign language and primary language. School grades were on a four-point scale, being A: excellent performance, B: good performance, C: adequate performance and D: underperformance.
Clinical assessment
Childhood psychopathology was available for most of the school-aged sample (97%; n = 4135). Parents or surrogates completed the Child Behavior Checklist for ages 6–18 (CBCL-6/18) from Achenbach System of Empirically Based Assessment (ASEBA). ADHD symptoms were assessed with the Conners’ Rating Scale for parents (CPRS) [29]. Since no specific instrument assessing ASD symptoms was available in INSchool, we assessed ASD symptoms using a combination of 10 items from the Thought Problems, Attention Problems, Social Problems and the Withdrawn/Depressed scales of the CBCL-6/18, to generate an ASD-CBCL scale as previously described [30]. Details on the clinical evaluation can be found in Español-Martín et al. [31].
Genotyping, imputation and quality control
Genomic DNA was isolated from saliva samples or buccal swabs collected using Oragene DNA OG-500 or OC-175 kits, respectively (DNA Genotek) and genotyped in three different genotyping waves with the Illumina Infinium PsychChip_v1.0 array for wave 1 (n = 794) or the Infinium Global Screening Array-24 version_2 (GSA_v2) for waves 2 (n = 2735) and 3 (n = 749) (Illumina, CA, San Diego, USA). Pre-imputation quality control was done with the PLINK 2.0 [32] and included individual and variant filtering based on the following parameters: variant call rate >0.95 (before individual filtering), individual call rate >0.98, autosomal heterozygosity deviation (|Fhet| < 0.2), variant call rate >0.98 (after individual filtering), SNP Hardy-Weinberg equilibrium (HWE; P > 1e−10) and minor allele frequency (MAF) > 0.01. Genetic outliers were identified by principal component analysis (PCA) using PLINK_2.0 and the mixed ancestry 1000G reference panel [33]. Ancestry outliers were excluded if their principal component (PC) values for PC1 or PC2 were greater than 1 standard deviation from the mean-centering point for our sample, considering each GWAS wave separately. Related and duplicated samples were identified by the “KING-robust kinship estimator” analysis in PLINK_2.0 [34] and one individual was excluded from each pair of subjects with kinship coefficient >0.0442. Imputation was done with McCarthy tools, for data preparation, and the Michigan Imputation Server [35], using the Haplotype Reference Consortium (HRC_Version_r1.1_2016) reference panel (GRCh37/hg19). Post-imputation dosage files with imputation INFO score >0.8 and MAF > 0.01 were considered for subsequent analyses.
Genome-wide polygenic scores
Genome-wide polygenic scores (PGS) were constructed in our in-house sample from INSchool considering each genotyping wave separately using PRS-CS [36], PLINK_2.0 and summary statistics from the largest available GWAS to date on EA (N = 766,345 individuals) [4], ADHD (N = 38,691 cases and 186,843 controls) [3] and ASD (N = 18,382 cases and 27,969 controls) [13]. All PGS were computed and standardized to a mean of 0 and a standard deviation of 1.
PGS for ADHD or ASD were constructed with five subsets of SNPs based on their contribution to EA: (i) The overall set of SNPs from Demontis et al. (PGSADHD; n = 6,774,224 SNPs) or Grove et al. (PGSASD; n = 9,112,386 SNPs) [3, 13]; (ii) variants not associated with EA (PGSADHD_noEA and PGSASD_noEA; PEA > 0.05); (iii) variants associated with EA (PGSADHD_EA and PGSASD_EA; PEA < = 0.05); (iv) variants associated with EA and showing consistent direction of the effect in EA and ADHD (PGSADHDconcordant; BetaADHD > 0 and BetaEA > 0 or BetaADHD < 0 and BetaEA < 0) or in EA and ASD (PGSASDconcordant; BetaASD > 0 and BetaEA > 0 or BetaASD < 0 and BetaEA < 0), and (v) variants associated with EA and showing opposite direction of the effect in EA and ADHD (PGSADHDdiscordant; BetaADHD > 0 and BetaEA < 0 or BetaADHD < 0 and BetaEA > 0) or in EA and ASD (PGSASDdiscordant; BetaASD > 0 and BetaEA < 0 or BetaASD < 0 and BetaEA > 0) (Fig. 1).
In addition, PGS constructed on the subset of SNPs shared between ADHDdiscordant and ASDconcordant partitions (n = 369,860 SNPs) were calculated using ADHD (PGSADHDshared) and ASD (PGSASDshared) weights separately.
Genome-wide polygenic score analyses stratified by sex
The same methods described for all individuals were used for the sex stratified analyses, except for the inclusion of school as a random effect in the models, which was not possible due to the reduction in sample size. An interaction test using the Wald-type test implemented in the metafor R Package was performed to test differences observed between sexes. In addition, results from the analyses stratified by sex where there was a significant effect in boys but not in girls were confirmed using a permutation-based approach to rule out the effect of a difference in sample sizes between boys and girls (n = 2409 boys and 1869 girls). A bootstrap resampling (1000 permutations) was used to select subsets of boys with the same sample size as the girls, and the association analyses were repeated for these subsets.
Statistical analysis
Genome-wide polygenic scores
The association between PGS and school performance was assessed with ordinal mixed-effect models considering D as the lowest and A as the highest category using the ordinal R package. In this context the effect estimate of the PGS produced by this model is an odds ratio, which provides the odds of an increase in school performance category per one PGS standard deviation. For instance, an odds ratio of 1.5 for a PGS in a specific subject performance would indicate that for an increase of one standard deviation in the PGS the odds of having a higher score in the specific subject is of 1.5. The percentage of variance attributable to each PGS was reported as the mean percentage of variance of each genotyping wave calculated as the increase in Nagelkerke’s pseudo-R2 between models with and without the PGS. For quantile plots, the target sample from each genotyping wave was divided into five quintiles of increasing PGS and PGS were compared across ranked quintiles considering the lower quintile as reference. To visualize how PGS affect school performance, predicted probabilities of being in each category (A, B, C or D) were calculated for each subject (mathematics, foreign language and primary language) and plotted using the effects R library [37].
The association between PGS and behavioral and emotional problems was tested using linear-mixed effects models. The CBCL and Conners’ parent rating scales were used as continuous variables and square-root transformed because of skewness.
All analyses were adjusted for age, sex, socioeconomic status (SES) and 20 genetic PC as fixed effects, as well as school as random effects to account for the multilevel nature of the data, considering each genotyping wave separately. SES was calculated using the Hollingshead Two-Factor Index based on parent’s education and occupation [38]. Results from each wave were then pooled using fixed-effects inverse variance weighted meta-analyses with the meta R library [39]. PGS P-values were corrected for multiple comparisons using Benjamini–Hochberg False Discovery Rate (FDR) method (adj-P < 0.05). Pairwise correlations between PGS from concordant and discordant genomic partitions were calculated using the non-parametric spearman rank correlation coefficient.
Mediation analyses
The effect of ADHD or ASD symptoms as mediators of the relationship between PGS of interest and school performance was assessed using a mediation analyses. We first estimated the effect of PGS on the mediator by using linear models and the effect of the mediator on school performance using ordinal mixed-effect models. If significant associations were detected in these two models (P < 0.05), the direct effect of PGS on school performance after taking into account the effect of the mediator was obtained using ordinal mixed-effect models. Mediators were square-root transformed to approximate normal distribution in all analyses where they were outcomes. All analyses were adjusted for age, sex, SES and 20 genetic PC as fixed effects, as well as school as random effects, considering each genotyping wave separately. Results from each wave were then pooled using fixed-effects inverse variance weighted meta-analyses with the meta R library [37]. After Bonferroni multiple testing correction (taking into account three subjects and two genomic partitions included in the mediation analysis for each disorder, P-value < 8.3e−03), we considered full mediation when the direct effect of the PGS on school performance becomes non-significant after taking into account the mediator and partial mediation when the direct effect remains significant, but to an attenuated degree.
Partitioned heritability
SNP-based heritability (h2SNP) estimates for ADHD and ASD were calculated using LDSC v1.0. [40, 41] for the three genome partitions described above: variants not associated with EA, variants with concordant and variants with discordant effects in EA. SNP-based heritability was computed for each annotation file using the GWAS summary statistics of ADHD and ASD [3, 13] and data from 1000G as reference panel [33]. Analyses were restricted to Hapmap3 SNPs and the major histocompatibility complex region was excluded. To improve the model performance, independent LD scores from the full baseline model v2.2 were used, which consist in a full annotation column and 97 independent functional annotations, which are available at LDSC repository (https://zenodo.org/record/7768714#.ZGaHZOzP3R1). Enrichment of genome-wide significant hits (P < 5e−08) of ADHD or ASD in the genome partitions described above were calculated using a Chi-square test and Manhattan plots were obtained using the qqmap R package.
Cell-type specific partitioned heritability
Cell-type specific analysis of partitioned heritability was run on LDSC v1.0. [40, 41] to explore the biological underpinnings of ADHD and ASD genomic partitions with significant h2SNP. These analyses were performed on new annotation files created from the intersection of each genomic partition and 13 brain-related tissues (Brain GTEx) [42] and three brain cell-type annotation files (neuron, astrocytes and oligodendrocytes) [43]. Reference annotation files were constructed as previously described [44] using two approaches: (i) “control” files - the intersection between each genomic partition and the provided control files in LDSC which include all genes in each study, and (ii) “anti-target” files – the intersection between the cell-type or brain specific annotation and the variants not included in the corresponding genomic partition. This “anti-target” analysis compares specific cell-type enrichment within a particular genomic partition and outside of this genomic partition, enabling us to assess the influence of EA on a particular cell-type. For each annotation file, we obtained an enrichment score (proportion of total SNPs in an annotation/percentage of the h2SNP) and then evaluated whether this cell-type enrichment was higher than the one associated to the “control” or “anti-target” enrichment. This was done by applying the LD score regression to specifically expressed genes (LDSC-SEG) approach using the --h2-cts argument [42]. P-values were corrected for multiple comparisons using FDR method (adj-P < 0.05).
Genetic covariance
Annotation-stratified genetic covariance between ADHD or ASD with multiple disorders and traits was assessed with GNOVA (https://github.com/xtonyjiang/GNOVA) [45], which calculates the genetic covariance between two phenotypes considering the LD structure and sample overlap. Covariance analyses considering the overall set of SNPs for ADHD or ASD were also performed [3, 13]. Available summary statistics from GWAS on 14 neuropsychiatric disorders, cognition and personality traits previously correlated with ADHD and/or ASD were considered, including ADHD [3], ASD [13], anorexia nervosa [46], antisocial behavior [47], anxiety disorder, bipolar disorder [48], extraversion [49], intelligence quotient (IQ) [50], loneliness and isolation, major depression [51], risk tolerance [52], schizophrenia [53], subjective well-being [54] and substance use disorder (SUD) [55]. GWAS summary statistics for anxiety disorder as well as loneliness and isolation were downloaded from Ben Neale UK Biobank GWAS analysis webpage (http://www.nealelab.is/uk-biobank). P-values were corrected for multiple comparisons using FDR (adj-P < 0.05).
Results
Contribution of the genetic liability for ADHD, ASD and EA on school performance
PGS for educational attainment (PGSEA) were associated with higher odds of better performance in mathematics (OR = 1.43, 95% CI = 1.35–1.52; adj-P = 1.0e−28), foreign language (OR = 1.32, 95% CI = 1.24–1.41; adj-P = 1.0e−17) and primary language (OR = 1.40, 95% CI = 1.32–1.50; adj-P = 2.1e−24), while PGS for ADHD showed the opposite pattern of associations, with PGSADHD associated with worse performance in all three subjects: mathematics (OR = 0.81, 95% CI = 0.76–0.85; adj-P = 2.3e−12), foreign language (OR = 0.81, 95% CI = 0.76–0.85; adj-P = 2.8e−12) and primary language (OR = 0.80, 95% CI = 0.76–0.85; adj-P = 4.5e−12) (Table 1 and Supplementary Table 1). No association was found between PGS for ASD (PGSASD) and school performance.
PGS comparisons across ranked quintiles showed that the odds for better performance in children at the highest quintile for PGSEA was, on average, over twice than in children in the first quintile for the three subjects (Fig. 2a and Supplementary Table 2a). Conversely, children at higher quintiles for PGSADHD showed lower odds for better performance than children in the first quintile (Fig. 2a and Supplementary Table 2b). Consistently, the probability of reaching better scores (A or B) increased with higher PGSEA or lower PGSADHD. Likewise, the probability of reaching worse scores (C or D) increased with higher PGSADHD or lower PGSEA (Fig. 2b, c).
a Quantile plots of meta-analysis odds ratios (OR) with 95% confidence intervals for PGSEA (in orange) and PGSADHD (in blue). The target sample was divided into quintiles, and school performance of each quintile was compared to the first quintile using ordinal mixed-effect models with age, sex, SES and 20 principal genetic components as fixed effects and school as random effects. Significant comparisons (PFDR < 0.05) are indicated with an asterisk; b Density plots of the contribution of PGSEA and PGSADHD to school performance grades for the three subjects under study. c Probability plots showing the probability of reaching each school performance grade (A, B, C or D) in each of the subjects. Per each subject we show four sections of probability corresponding to each school performance grade according to the PGSEA (in orange) and PGSADHD (in blue).
Polygenic dissection of the contribution of ADHD and ASD to school performance by their relationship with EA
We constructed PGS for these neurodevelopmental disorders with four subsets of SNPs based on to their contribution to EA, including variants not associated with EA (PGSADHD_noEA/PGSASD_noEA) and variants associated with EA (PGSADHD_EA/PGSASD_EA), which were divided in concordant (PGSADHDconcordant/PGSASDconcordant), showing consistent direction of effect in EA and ADHD/ASD, and discordant variants (PGSADHDconcordant/PGSASDconcordant), showing opposite direction of effect in EA and ADHD/ASD (Fig. 1).
Both PGSADHD_noEA and PGSADHD_EA were associated with worse school performance in the three subjects (adj-P < = 3.0e−06) (Table 1 and Supplementary Table 3). When focused on variants associated with EA, we found that PGSADHDdiscordant were associated with poor school performance in mathematics (OR = 0.77, 95% CI = 0.72–0.81; adj-P = 1.3e−17), foreign language (OR = 0.81, 95% CI = 0.76–0.86; adj-P = 2.5e−11) and primary language (OR = 0.77, 95% CI = 0.72–0.82; adj-P = 5.2e−16) (Table 1 and Supplementary Table 3). Conversely, PGSADHDconcordant showed the opposite direction of effect and were associated, to a lesser extent than PGSADHDdiscordant, with better performance in primary language (OR = 1.07, 95% CI = 1.01–1.13; adj-P = 0.042) (Table 1 and Supplementary Table 3). PGSADHDdiscordant accounted for the largest proportion of the variance explained for school performance compared to the other PGSADHD, ranging from 1.8% in foreign language to 2.6% in mathematics (Fig. 3a and Table 1). It explained on average over 28 times more variance for school performance than PGSADHDconcordant across the three subjects (Fig. 3a and Table 1).
PGSADHD were constructed considering four subsets of SNPs: variants not associated with EA (PEA > 0.05; PGSADHD_noEA), variants associated with EA (PEA < = 0.05; PGSADHD_EA), variants showing concordant (PGSADHDconcordant) and discordant (PGSADHDdiscordant) direction of the effect in EA and ADHD. a Percentage of variance, as mean of Nagelkerke’s pseudo-R2 calculated in each genotyping wave, explained by PGSADHD according to the four partitions described above; b Quantile plots of meta-analysis odds ratios (OR) with 95% confidence intervals for PGSADHD according to the four partitions described above. The target sample was divided into quintiles and school performance of each quintile was compared to the first quintile using ordinal mixed-effect models with age, sex, SES and 20 principal genetic components as fixed effects and school as random effects. Significant comparisons (PFDR < 0.05) are indicated with an asterisk.
PGS comparisons across ranked quintiles showed the expected trend of lower odds for better school performance in individuals with higher PGSADHD across all partitions but PGSADHDconcordant (Fig. 3b and Supplementary Table 3). The strongest effect was identified for PGSADHDdiscordant, with children at the highest quintile doubling the odds for underperformance at school of children in the first quintile (Fig. 3b and Supplementary Table 3). Consistently, higher PGSADHDdiscordant decreased the probability of reaching an A or B score while the probability of reaching a C or D score increased in all three subjects (Supplementary Fig. 1). Besides, PGSADHDconcordant showed a trend towards the opposite direction of the association, with higher PGSADHDconcordant associated with higher probability of school success, especially in primary language, where children at the highest quintile showed 1.24 times better performance than children in the first quintile (Fig. 3b and Supplementary Table 3).
Then we assessed whether ADHD symptoms explained the associations between polygenic risk for ADHD and school performance. Both PGSADHDnoEA and PGSADHDdiscordant were associated with increased ADHD symptoms (P = 2.9e−12 and P = 7.1e−13, respectively) (Supplementary Table 4), and ADHD symptoms lower odds for better school performance in all subjects (P < 1.5e−121) (Supplementary Table 5). The effect of both PGS on school performance decreased after considering the effect of ADHD symptoms, suggesting that ADHD symptoms partially mediate the effect of PGSADHDdiscordant (from 16% in primary language to 37% in foreign language) and fully mediate the effect of PGSADHDnoEA (from 52% in primary language to 55% in mathematics) on school performance (Supplementary Tables 6 and 7).
When we subset the genetic liability for ASD based on its role in EA, PGSASD showed a different pattern of association with school performance than PGSADHD. In line with the lack of association between PGSASD and school performance, no association was found either for PGSASD_noEA or PGSASD_EA (Table 1 and Supplementary Table 3). However, the genetic liability with concordant and discordant effects between ASD and EA showed significant association with school performance with opposite direction of effect. PGSASDdiscordant were associated with poor school performance in mathematics (OR = 0.83, 95% CI = 0.78–0.88; adj-P = 7.5e−10), foreign language (OR = 0.84, 95% CI = 0.79–0.89; adj-P = 3.3e−08) and primary language (OR = 0.82, 95% CI = 0.77–0.87; adj-P = 1.6e−10) (Fig. 4a, Table 1 and Supplementary Table 3). Whereas, PGSASDconcordant were associated with better performance in mathematics (OR = 1.17, 95% CI = 1.11–1.24; adj-P = 1.3e−07), foreign language (OR = 1.11, 95% CI = 1.04–1.17; adj-P = 1.1e−03) and primary language (OR = 1.15, 95% CI = 1.09–1.23; adj-P = 5.2e−06) (Table 1 and Supplementary Table 3). PGSASDdiscordant accounted for the largest proportion of the variance explained for school performance (from 1.12% in foreign language to 1.95% in mathematics) followed by PGSASDconcordant (from 1.18% in foreign language to 1.28% in primary language) (Fig. 4a and Table 1).
PGSASD were constructed considering four subsets of SNPs: variants not associated with EA (PEA > 0.05; PGSASD_noEA), variants associated with EA (PEA < = 0.05; PGSASD_EA), variants showing concordant (PGSASDconcordant) and discordant (PGSASDdiscordant) direction of the effect in EA and ASD. a Percentage of variance, as mean of Nagelkerke’s pseudo-R2 calculated in each genotyping wave, explained by PGSASD according to the four partitions described above; b Quantile plots of meta-analysis odds ratios (OR) with 95% confidence intervals for PGSASD according to the four partitions described above. The target sample was divided into quintiles and school performance of each quintile was compared to the first quintile using ordinal mixed-effect models with age, sex, SES and 20 principal genetic components as fixed effects and school as random effects. Significant comparisons (PFDR < 0.05) are indicated with an asterisk.
PGS comparisons across ranked quintiles showed that children at the highest quintile for PGSASDdiscordant had lower odds for better performance than children in the first quintile (P < = 5.3e−05), while children at higher quintiles for PGSASDconcordant showed better performance than children in the first quintile in all three subjects (P < = 0.018) (Fig. 4b and Supplementary Table 2c). Consistently, the probability of reaching an A or B score increased with higher PGSASDconcordant or lower PGSASDdiscordant, while the probability of reaching a C or D score increased with higher PGSASDdiscordant or lower PGSASDconcordant (Supplementary Fig. 2).
Also, ASD symptoms emerged as partial mediators of the effect of the ASD genetic liability on school performance. PGSASDconcordant and PGSASDdiscordant were significantly associated with increased ASD symptoms (P = 6.3e−04 and P = 1.1e−4, respectively) (Supplementary Table 4), and ASD symptoms with lower school performance in all three subjects (P < = 4.2e−33) (Supplementary Table 5). The effect of PGSASDdiscordant on poor school performance decreased when taking into account the effect of ASD symptoms, suggesting that they mediate from 6% to 9% of the effect of PGSASDdiscordant on school performance (Supplementary Tables 6 and 7). By contrast, for PGSASDconcordant we found evidence of inconsistent mediation or suppressor effects [56, 57] that occur when two predictors (PGSASDconcordant and ASD symptoms) with different direction of effect on the outcome (school performance) are simultaneously considered. PGSASDconcordant and ASD symptoms operate consistently as mutual suppressor for school performance, meaning that taking into account the effect of ASD symptoms into the regression equation increases the positive effect of PGSASDconcordant on school performance (from 13% in mathematics to 24% in foreign language) (Supplementary Tables 6 and 7).
When stratifying by sex, we found no differences on the effect of the genetic liability for ADHD or ASD dissected by its relationship with EA on school performance (Pinteraction > 0.05) (Supplementary Table 8).
Given the significant negative correlation found between PGSADHDdiscordant and PGSASDconcordant (ρ = −0.29, P = 8.8e−82; Supplementary Fig. 3), a sensitivity analysis was performed to further assess the effect of the overlapping variation between ADHD and ASD on school performance. PGS constructed on the subset of shared variants (PGSADHDshared and PGSASDshared) were associated with school performance with opposite direction of the effect in the three subjects under study. While PGSADHDshared contributed to poor school performance, PGSASDshared were associated with better performance (Supplementary Table 9), results supporting that the same set of SNPs may drive the discordant association of both disorders with school performance.
Polygenic dissection of the contribution of ADHD and ASD to childhood psychopathology by their relationship with EA
The majority of behavioral and emotional problems showed a distinct pattern of association with the genetic liability for ADHD and ASD by their role in EA, being most of them positively associated with the subset of variants with discordant, but not concordant, direction of effect (i.e., inattention, hyperactivity/impulsivity or rule-breaking behavior) (Fig. 5). Many of those displaying association with discordant variation were also associated with PGSnoEA. Also, some traits were associated with the genetic liability for ADHD (i.e., ADHD symptoms, oppositional problems, social problems and aggressive behavior) or ASD (i.e., ASD symptoms) regardless of EA, with concordant effect sizes being generally smaller than those for the discordant set of variants or variants not associated with EA (Fig. 5 and Supplementary Table 10).
Z-scores from regression analyses for school performance in the three subjects, CBCL scales or Conners’ parent rating scales and (a) PGSADHDdiscordant (x axis) and PGSADHDconcordant (y axis) or (b) PGSASDdiscordant (x axis) and PGSASDconcordant (y axis). Point size corresponds to -log10(p-value) with concordant PGS in red and discordant PGS in blue. Red dotted lines indicate significant results after 5% FDR correction.
Partitioned heritability
Partitioned heritability estimates for ADHD showed significant h2SNP enrichment in ADHDdiscordant and ADHDnoEA variants, but negative h2SNP in the ADHDconcordant SNP set, indicating that most of the SNPs in this group have null effects on ADHD or suggesting a model misspecification (Supplementary Table 11). Consistently, we found enrichment of genome-wide significant hits for ADHD [3] within the ADHDdiscordant and ADHDnoEA, (P < 2.2e−16), but not the ADHDconcordant, SNP subsets (Supplementary Fig. 4a–c). For ASD, only the ASDconcordant subset of variants showed significant h2SNP enrichment (Supplementary Table 11) and none of the SNP partitions showed enrichment of ASD hits [13] (Supplementary Fig. 4d–f).
To further explore the biological underpinnings of ADHD and ASD genomic partitions with significant h2SNP we used partitioned heritability and LD score regression applied to specifically expressed genes (LDSC-SEG) [42]. When we compared specific cell-type enrichment inside and outside a particular genomic partition considering the “anti-target” reference annotations, we found that the ADHDdiscordant partition, but not the ADHDnoEA, was enriched by ADHD h2SNP in all studied cell-types and brain areas (Supplementary Fig. 5 and Table 12). We also found enrichment in ASD h2SNP within the ASDconcordant partition from neurons and four brain areas (i.e., anterior cingulate cortex, cerebellum, frontal cortex, hypothalamus; Supplementary Fig. 5 and Table 12). No significant results, were observed when comparing the enrichment of h2SNP with the “control” annotation for any partition (Supplementary Table 12).
Genetic covariance between ADHD and ASD and related disorders and traits dissected by their relationship with EA
ASD displayed positive covariance with ADHDnoEA and ADHDconcordant, while ADHD had positive covariance with ASDdiscordant and negative covariance with ASDconcordant, confirming the opposite genetic overlap between both disorders and EA (Supplementary Table 13 and Fig. 6).
When we assessed the genetic covariance between ADHD or ASD and a set of neuropsychiatric disorders, cognition and personality traits, we found similar patterns for most of the traits and SNP partitions. Estimated covariances for variants not associated with EA were similar to genetic correlations previously described for ADHD or ASD across the whole genome [3, 13], except for IQ. The majority of tested phenotypes showed opposite patterns of genetic covariance within the sets of variants with concordant or discordant effects for both disorders. Anorexia nervosa, IQ and subjective well-being displayed positive covariance with concordant but negative covariance with discordant genetic load. Conversely, SUD, loneliness and isolation and antisocial behavior displayed positive covariance with discordant and negative covariance with concordant genetic variation for both disorders. In contrast, for several traits the genetic covariance with ADHD (i.e., risk tolerance or schizophrenia) or ASD (i.e., schizophrenia) did not depend on EA, and others had a significant covariance only with either the concordant (i.e., bipolar disorder) or the discordant (i.e., anxiety or major depression) set of variants according to their effect on EA (Supplementary Table 13 and Fig. 6).
Discussion
Our results reveal that, despite sharing a common genetic architecture, the genetic liability for ADHD and ASD show differences in their association with school performance and early manifestation of psychopathology by their relationship with EA. This highlights the potential of leveraging overlapping polygenic signals with EA to dissect clinical and genetic heterogeneity in mental health, as described for schizophrenia [58].
The strong association between PGS for adult EA and better school grades in childhood and adolescence confirms the lifetime nature of the EA genetic background, supporting that school performance is an early life intermediate phenotype for future outcomes in adulthood. These results align with previous evidence of higher polygenic scores for EA associated with speech and reading skills even before school, as well as with loftier academic aspiration and attainment extending into adulthood [8].
We confirm the strong and inverse association between polygenic ADHD risk and school performance [59], which involves genetic variation shared with EA, especially when variants with discordant effects in both traits were considered. We found that the genetic liability for ADHD not associated with EA also impacts on school performance, suggesting that observed genetic associations between ADHD and poor academic outcomes may not only reflect shared genetic variation with EA, but also specific ADHD genetic liability, independently of EA. These results are in line with findings where ADHD associations with literacy-related impairments involve genetic variation shared with, but also independent of, EA [59]. Interestingly, the ADHDnoEA genomic partition was found associated with higher IQ, as well as with several psychiatric disorders positively correlated with EA and/or IQ in previous studies, such as ASD, anorexia nervosa, schizophrenia or bipolar disorder [7, 50, 60, 61]. These apparent contradictory findings could be explained by the fact that school performance is not only explained by cognitive traits but also by other factors such as self-perceived abilities, psychopathology or well-being, even after accounting for IQ [6, 62, 63]. In addition, apart from the genetic liability associated with high IQ, ADHDnoEA may also account for noncognitive factors, such as ADHD symptoms, which can change across the lifespan [64], and might affect early-life school outcomes without having an effect on later-life educational attainment. We found that this genomic partition also correlates negatively with subjective well-being and positively with SUD, loneliness and isolation, antisocial behavior, risk tolerance, anxiety or depression, that may contribute negatively on school performance independently of IQ. Interestingly, similar results were described for late-life EA, where beyond heritable cognitive skills, there are noncognitive influences that have a polygenic architecture, impact on EA and correlate with personality traits and increased risk for certain psychiatric disorders [7].
ADHD shows negative genetic correlation with EA, with the vast majority of risk variants being associated with low levels of EA [3]. Consistently, we found that the overlapping genetic variation with discordant effects in both phenotypes has the strongest negative effect on school performance, which is partially mediated by ADHD symptoms. It also accounts for most of the ADHD genetic background described to date [3], and is associated with higher rates of children psychopathology. Conversely, concordant variation between ADHD and EA has no impact on school performance and contributes, to less extent than discordant variation, to ADHD symptoms and externalizing behaviors. These results confirm that a large proportion of ADHD risk loci is shared with EA and has opposite direction of the effect, and provide further evidence on the adverse effects of ADHD on long-term academic outcomes [65].
The polygenic association of ASD and school performance was to a large extent attributable to genetic effects that are shared with EA. Both the concordant and discordant variation that make up the genetic load for EA have polygenic contributions to school performance, which are partially explained by ASD symptoms but shows divergent patterns of genetic effects. While genetic variation with discordant effects in ASD and EA was associated with impaired performance at school and early manifestation of psychopathology, PGS for ASD constructed on variants with concordant effects were associated with improved school performance and ASD symptoms, but not with other emotional or behavioral problems. This complex relationship may underlie the variability in academic achievement observed across the autism spectrum [23], strongly supports that ASD is genetically heterogeneous and demonstrates that ASD and EA share many genetic loci but without a clear pattern of sign concordance. Consistently, recent findings showed that most genetic variants associated with ASD are shared with EA, with both positive and negative direction of the effect [11], which may explain the lack of association found between school performance and genome-wide PGS for ASD. The combination of ASD discordant and concordant variation with opposite effects in school performance may result in the cancellation of the association signal, which aligns with observational studies showing that academic achievement in ASD is more variable than in ADHD [23], and strongly supports our study design that leverages EA genetic data to detect opposite polygenic effects that would otherwise be missed.
We also found that a great proportion of the ASD genetic liability is not associated either with EA nor with school performance. Genetic variants included in this ASDnoEA genomic partition may contribute to ASD symptoms different from cognition with no impact on school performance, such as difficulties in social interaction. This idea was supported by the analysis of genomic covariance, where we found that this genomic partition was negatively correlated with extraversion but not associated with IQ. For most of the psychiatric disorders and related traits, the pattern of genetic covariance was opposite for concordant and discordant overlapping variation, the latter being more similar to that previously described for ADHD or ASD [2, 3, 13]. These results align with recent findings reporting association between poor school achievement and greater risk of a subsequent mental disorder, except for eating disorders, where the association was of similar magnitude but in the opposite direction [66]. Our findings are consistent with the epidemiological and genetic correlations described between ADHD or ASD and other psychiatric conditions [2] and provide further evidence that genetic data on EA, in aggregate, aid to identifying individuals at higher risk not only for poor school performance but also for worse disease course and clinical outcomes. The genetic covariance results also suggest that concordant genomic partitions are associated with positive outcomes, as reflected by positive genetic covariance with subjective well-being and negative genetic covariance with different psychiatric disorders and traits (i.e., SUD, antisocial behavior or loneliness and isolation). However, this pattern of association was not observed in schizophrenia, bipolar disorder and ASD, which are positively correlated with concordant partitions. This is in line with a recent study supporting that bipolar disorder, schizophrenia and ASD may belong to the same psychopathology factor (i.e., thought problems factor) at the genomic level [67].
The main strength of our study is to disentangle the polygenic effects of ADHD and ASD on school performance using a third trait, EA, allowing us to better understand their polygenic contribution and early manifestation of psychopathology in a deeply phenotyped population-based cohort. However, several considerations need to be contemplated. First, we found that discordant polygenic signatures for both disorders correlate with early manifestation of psychopathology and poor school performance, which confirms genetic influences transcending diagnostic boundaries and suggests that prodromal symptoms may negatively affect school performance. However, we did not infer causality and cannot exclude reverse causation, where poor school achievement increases the risk for psychopathology later in life [68,69,70,71,72]; Second, although our results are robust when accounting for potential confounders, including school and SES, we cannot discard residual confounding by other factors, such as treatment [73], meaning that we may underestimate the negative polygenic effect of ADHD and ASD on school performance; Third, the main findings held across the three subjects under study, which aligns with previous results on population-wide registers [73, 74]. Strikingly, there was also a trend towards better performance in primary language for ADHD concordant variation. Further analyses in larger samples will be needed to confirm whether this association is subject specific or can be extended to performance in foreign language or mathematics; Fourth, results from ADHD partitioned SNP heritability should be considered with caution since negative h2SNP for the ADHDconcordant genomic partition could indicate a model misspecification; Finally, there are consistent evidences indicating that EA and school performance are influenced by demographic and indirect genetic effects [75]. Educational outcomes are particularly susceptible to bias arising from assortative mating or dynastic effects when studied in unrelated individuals, which may lead to inaccurate estimations of direct genetic effects [76,77,78,79]. Although the aim of this study was to dissect the polygenic contribution of ADHD and ASD to school performance by its relationship with EA, rather than estimating overall genetic effects, our results should be interpreted with caution in light of these population mechanisms. Further family-based study designs accounting for these population phenomena are required to provide more accurate genetic associations as well as to assess indirect genetic effects on school performance in the offspring.
In conclusion, we provide evidence showing that school performance is an early life intermediate phenotype associated with adult EA. By dissecting the genetic liability of ADHD and ASD by their effect on EA, we disentangled their polygenic effects on school performance and early manifestation of psychopathology, and provided new insights into overlapping genetic signatures with other comorbid disorders and traits. Overall, our findings expand on previous studies and strongly support the usage of the genetic load for EA to deepen insights into the genetic relationship between these neurodevelopmental disorders and school performance, to fill the gap of knowledge of the biological mechanisms underlying educational outcomes and to target more vulnerable individuals for early interventions.
Data availability
Raw data from this article is not publicly available because of limitations in ethical approvals and the summary data will be available from the corresponding autor upon reasonable request.
References
Mattheisen M, Grove J, Als TD, Martin J, Voloudakis G, Meier S, et al. Identification of shared and differentiating genetic architecture for autism spectrum disorder, attention-deficit hyperactivity disorder and case subgroups. Nat Genet. 2022;54:1470–8.
Anttila V, Bulik-Sullivan B, Finucane HK, Walters RK, Bras J, Duncan L, et al. Analysis of shared heritability in common disorders of the brain. Science. 2018;360:eaap8757.
Demontis D, Walters GB, Athanasiadis G, Walters R, Therrien K, Nielsen TT, et al. Genome-wide analyses of ADHD identify 27 risk loci, refine the genetic architecture and implicate several cognitive domains. Nat Genet. 2023;55:198–208.
Lee JJ, Wedow R, Okbay A, Kong E, Maghzian O, Zacher M, et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat Genet. 2018;50:1112–21.
McGue M, Rustichini A, Iacono WG. Cognitive, Noncognitive, and Family Background Contributions to College Attainment: A Behavioral Genetic Perspective. J Pers. 2017;85:65–78.
Krapohl E, Rimfeld K, Shakeshaft NG, Trzaskowski M, McMillan A, Pingault JB, et al. The high heritability of educational achievement reflects many genetically influenced traits, not just intelligence. Proc Natl Acad Sci USA. 2014;111:15273–8.
Demange PA, Malanchini M, Mallard TT, Biroli P, Cox SR, Grotzinger AD, et al. Investigating the genetic architecture of noncognitive skills using GWAS-by-subtraction. Nat Genet. 2021;53:1. 2021;53:35–44.
Ward ME, McMahon G, St Pourcain B, Evans DM, Rietveld CA, Benjamin DJ, et al. Genetic Variation Associated with Differential Educational Attainment in Adults Has Anticipated Associations with School Performance in Children. PLoS One. 2014;9:e100248.
Okbay A, Wu Y, Wang N, Jayashankar H, Bennett M, Nehzati SM, et al. Polygenic prediction of educational attainment within and between families from genome-wide association analyses in 3 million individuals. Nat Genet. 2022;54:4. 2022;54:437–449.
Okbay A, Beauchamp JP, Fontana MA, Lee JJ, Pers TH, Rietveld CA, et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature. 2016;533:539–42.
Hope S, Jebsen KG, Lin A, Bahrami S, Rødevand L, Frei O, et al. Bidirectional genetic overlap between autism spectrum disorder and cognitive traits. Transl Psychiatry. 2022. https://doi.org/10.21203/RS.3.RS-1427702/V1.
Verhoef E, Grove J, Shapland CY, Demontis D, Burgess S, Rai D. et al. Discordant associations of educational attainment with ASD and ADHD implicate a polygenic form of pleiotropy. Nat Commun. 2021;12:1–14.
Grove J, Ripke S, Als TD, Mattheisen M, Walters RK, Won H, et al. Identification of common genetic risk variants for autism spectrum disorder. Nat Genet. 2019;51:431.
Dardani C, Riglin L, Leppert B, Sanderson E, Rai D, Howe LD, et al. Is genetic liability to ADHD and ASD causally linked to educational attainment? Int J Epidemiol. 2022;50:2011–23.
Hicks BM, Angus Clark D, Deak JD, Schaefer JD, Liu M, Jang S, et al. Polygenic scores for smoking and educational attainment have independent influences on academic success and adjustment in adolescence and educational attainment in adulthood. PLoS One. 2021;16:e0255348.
Zajacova A, Lawrence EM. The relationship between education and health: reducing disparities through a contextual approach. Annu Rev Public Health. 2018;39:273.
Raghupathi V, Raghupathi W. The influence of education on health: An empirical assessment of OECD countries for the period 1995-2015. Archives of Public Health. 2020;78:1–18.
Darney D, Reinke WM, Herman KC, Stormont M, Ialongo NS. Children with co-occurring academic and behavior problems in first grade: distal outcomes in twelfth grade. J Sch Psychol. 2013;51:117–28.
Taanila A, Ebeling H, Tiihala M, Kaakinen M, Moilanen I, Hurtig T, et al. Association between childhood specific learning difficulties and school performance in adolescents with and without ADHD symptoms: a 16-year follow-up. J Atten Disord. 2014;18:61–72.
Barkley RA, Fischer M, Edelbrock CS, Smallish L. The adolescent outcome of hyperactive children diagnosed by research criteria: I. An 8-year prospective follow-up study. J Am Acad Child Adolesc Psychiatry. 1990;29:546–57.
Frazier TW, Youngstrom EA, Glutting JJ, Watkins MW. ADHD and achievement: meta-analysis of the child, adolescent, and adult literatures and a concomitant study with college students. J Learn Disabil. 2007;40:49–65.
Español-Martín G, Pagerols M, Prat R, Rivas C, Ramos-Quiroga JA, Casas M, et al. The impact of attention-deficit/hyperactivity disorder and specific learning disorders on academic performance in Spanish children from a low-middle- and a high-income population. Front Psychiatry. 2023;14:1136994.
Keen D, Webster A, Ridley G. How well are children with autism spectrum disorder doing academically at school? An overview of the literature. Autism. 2016;20:276–94.
Estes A, Rivera V, Bryan M, Cali P, Dawson G. Discrepancies between academic achievement and intellectual ability in higher-functioning school-aged children with autism spectrum disorder. J Autism Dev Disord. 2011;41:1044–52.
McDougal E, Riby DM, Hanley M. Profiles of academic achievement and attention in children with and without Autism Spectrum Disorder. Res Dev Disabil. 2020;106:103749.
Chiang HM, Cheung YK, Hickson L, Xiang R, Tsai LY. Predictive factors of participation in postsecondary education for high school leavers with autism. J Autism Dev Disord. 2012;42:685–96.
Dijkhuis R, de Sonneville L, Ziermans T, Staal W, Swaab H. Autism Symptoms, Executive Functioning and Academic Progress in Higher Education Students. J Autism Dev Disord. 2020;50:1353–63.
Shattuck PT, Narendorf SC, Cooper B, Sterzing PR, Wagner M, Taylor JL. Postsecondary education and employment among youth with an autism spectrum disorder. Pediatrics. 2012;129:1042–9.
Conners CK. The Conners’ rating scales–revised: Technical manual. North Tonawanda, NY: Multi-Health Systems; 1997.
So P, Greaves-Lord K, Van Der Ende J, Verhulst FC, Rescorla L, De Nijs PF. Using the Child Behavior Checklist and the Teacher’s Report Form for identification of children with autism spectrum disorders. Autism. 2013;17:595–607.
Español-Martín G, Pagerols M, Prat R, Rivas C, Sixto L, Valero S, et al. Strengths and Difficulties Questionnaire: Psychometric Properties and Normative Data for Spanish 5- to 17-Year-Olds. Assessment. 2021;28:1445–58.
Chang CC, Chow CC, Tellier LCAM, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Auton A, Abecasis GR, Altshuler DM, Durbin RM, Bentley DR, Chakravarti A, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Manichaikul A, Mychaleckyj JC, Rich SS, Daly K, Sale M, Chen WM. Robust relationship inference in genome-wide association studies. Bioinformatics. 2010;26:2867–73.
Das S, Forer L, Schönherr S, Sidore C, Locke AE, Kwong A, et al. Next-generation genotype imputation service and methods. Nat Genet. 2016;48:1284–7.
Ge T, Chen CY, Ni Y, Feng YCA, Smoller JW. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat Commun. 2019;10:1. 2019;10:1–10.
Fox J. Effect Displays in R for Generalised Linear Models. J Stat Softw. 2003;8:1–27.
Hollingshead AB. Four Factor Index of Social Status. Yale. J Sociol. 2011;8:21–51.
Balduzzi S, Rücker G, Schwarzer G. How to perform a meta-analysis with R: a practical tutorial. Evid Based Ment Health. 2019;22:153–60.
Bulik-Sullivan B, Loh PR, Finucane HK, Ripke S, Yang J, Patterson N, et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:3. 2015;47:291-5.
Finucane HK, Bulik-Sullivan B, Gusev A, Trynka G, Reshef Y, Loh PR, et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet. 2015;47:11. 2015;47:1228-35.
Finucane HK, Reshef YA, Anttila V, Slowikowski K, Gusev A, Byrnes A, et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat Genet. 2018;50:4. 2018;50:621-9.
Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, et al. A Transcriptome Database for Astrocytes, Neurons, and Oligodendrocytes: A New Resource for Understanding Brain Development and Function. J Neurosci. 2008;28:264.
Andreu-Bernabeu Á, Díaz-Caneja CM, Costas J, De Hoyos L, Stella C, Gurriarán X, et al. Polygenic contribution to the relationship of loneliness and social isolation with schizophrenia. Nat Commun. 2022;13:1. 2022;13:1–11.
Lu Q, Li B, Ou D, Erlendsdottir M, Powles RL, Jiang T, et al. A Powerful Approach to Estimating Annotation-Stratified Genetic Covariance via GWAS Summary Statistics. Am J Hum Genet. 2017;101:939–64.
Watson HJ, Yilmaz Z, Thornton LM, Hübel C, Coleman JRI, Gaspar HA, et al. Genome-wide association study identifies eight risk loci and implicates metabo-psychiatric origins for anorexia nervosa. Nat Genet. 2019;51:8. 2019;51:1207–1214.
Tielbeek JJ, Uffelmann E, Williams BS, Colodro-Conde L, Gagnon É, Mallard TT, et al. Uncovering the genetic architecture of broad antisocial behavior through a genome-wide association study meta-analysis. Mol Psychiatry. 2022;27:11. 2022;27:4453–4463.
Mullins N, Forstner AJ, O’Connell KS, Coombes B, Coleman JRI, Qiao Z, et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat Genet. 2021;53:6. 2021;53:817–829.
van den Berg SM, de Moor MHM, Verweij KJH, Krueger RF, Luciano M, Arias Vasquez A, et al. Meta-analysis of Genome-Wide Association Studies for Extraversion: Findings from the Genetics of Personality Consortium. Behav Genet. 2016;46:170–82.
Savage JE, Jansen PR, Stringer S, Watanabe K, Bryois J, De Leeuw CA, et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat Genet. 2018;50:912–9.
Howard DM, Adams MJ, Clarke TK, Hafferty JD, Gibson J, Shirali M, et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat Neurosci. 2019;22:3. 2019;22:343–352.
Karlsson Linnér R, Biroli P, Kong E, Meddens SFW, Wedow R, Fontana MA, et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat Genet. 2019;51:2. 2019;51:245–257.
Trubetskoy V, Pardiñas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:7906. 2022;604:502–508.
Baselmans BML, Jansen R, Ip HF, van Dongen J, Abdellaoui A, van de Weijer MP, et al. Multivariate genome-wide analyses of the well-being spectrum. Nat Genet. 2019;51:3. 2019;51:445–451.
Hatoum AS, Colbert SMC, Johnson EC, Huggett SB, Deak JD, Pathak GA, et al. Multivariate genome-wide association meta-analysis of over 1 million subjects identifies loci underlying multiple substance use disorders. Nat Mental Health. 2023;1:210–23.
MacKinnon DP, Fairchild AJ, Fritz MS. Mediation analysis. Annu Rev Psychol. 2007;58:593–614.
Paulhus DL, Robins RW, Trzesniewski KH, Tracy JL. Two Replicable Suppressor Situations in Personality Research. Multivariate Behav Res. 2004;39:303–28.
Bansal V, Mitjans M, Burik CAP, Linnér RK, Okbay A, Rietveld CA, et al. Genome-wide association study results for educational attainment aid in identifying genetic heterogeneity of schizophrenia. Nat Commun. 2018;9:1. 2018;9:1–12.
Verhoef E, Demontis D, Burgess S, Shapland CY, Dale PS, Okbay A, et al. Disentangling polygenic associations between attention-deficit/hyperactivity disorder, educational attainment, literacy and language. Transl Psychiatry. 2019;9:1. 2019;9:1–12.
Hill WD, Marioni RE, Maghzian O, Ritchie SJ, Hagenaars SP, McIntosh AM, et al. A combined analysis of genetically correlated traits identifies 187 loci and a role for neurogenesis and myelination in intelligence. Mol Psychiatry. 2018;24:2. 2018;24:169–181.
Bang L, Bahrami S, Hindley G, Smeland OB, Rødevand L, Jaholkowski PP, et al. Genome-wide analysis of anorexia nervosa and major psychiatric disorders and related traits reveals genetic overlap and identifies novel risk loci for anorexia nervosa. Transl Psychiatry. 2023;13:1. 2023;13:1–9.
Greven CU, Harlaar N, Kovas Y, Chamorro-Premuzic T, Plomin R. More than just IQ: school achievement is predicted by self-perceived abilities--but for genetic rather than environmental reasons. Psychol Sci. 2009;20:753–62.
Deary IJ, Strand S, Smith P, Fernandes C. Intelligence and educational achievement. Intelligence. 2007;35:13–21.
Franke B, Michelini G, Asherson P, Banaschewski T, Bilbow A, Buitelaar JK, et al. Live fast, die young? A review on the developmental trajectories of ADHD across the lifespan. Eur Neuropsychopharmacol. 2018;28:1059–88.
Arnold LE, Hodgkins P, Kahle J, Madhoo M, Kewley G. Long-Term Outcomes of ADHD: Academic Achievement and Performance. J Atten Disord. 2020;24:73–85.
Weckström T, Elovainio M, Pulkki-Råback L, Suokas K, Komulainen K, Mullola S, et al. School achievement in adolescence and the risk of mental disorders in early adulthood: a Finnish nationwide register study. Mol Psychiatry. 2023;2023:1–7.
Waldman ID, Poore HE, Luningham JM, Yang J. Testing structural models of psychopathology at the genomic level. World Psychiatry. 2020;19:350.
Pedersen SD, Østergaard SD, Petersen L. The association between school exam grades and subsequent development of bipolar disorder. Acta Neuropsychiatr. 2018;30:209–17.
Kendler KS, Ohlsson H, Mezuk B, Sundquist K, Sundquist J. A Swedish National Prospective and Co-relative Study of School Achievement at Age 16, and Risk for Schizophrenia, Other Nonaffective Psychosis, and Bipolar Illness. Schizophr Bull. 2016;42:77–86.
Gyllenberg D, Ristikari T, Kelleher I, Kääriälä A, Gissler M. School performance and later diagnoses of nonaffective psychoses, bipolar disorder, and depression. Acta Psychiatr Scand. 2022;146:420–9.
Rahman MA, Todd C, John A, Tan J, Kerr M, Potter R, et al. School achievement as a predictor of depression and self-harm in adolescence: linked education and health record study. Br J Psychiatry. 2018;212:215–21.
Sörberg Wallin A, Koupil I, Gustafsson JE, Zammit S, Allebeck P, Falkstedt D. Academic performance, externalizing disorders and depression: 26,000 adolescents followed into adulthood. Soc Psychiatry Psychiatr Epidemiol. 2019;54:977–86.
Jangmo A, Stålhandske A, Chang Z, Chen Q, Almqvist C, Feldman I, et al. Attention-Deficit/Hyperactivity Disorder, School Performance, and Effect of Medication. J Am Acad Child Adolesc Psychiatry. 2019;58:423–32.
Sunde HF, Kleppestø TH, Gustavson K, Nordmo M, Reme B-A, Fartein AT. et al. The ADHD deficit in school performance across sex and parental education: A prospective sibling-comparison register study of 344,152 Norwegian adolescents. JCPP Adv.2022;2:e12064.
Howe LJ, Nivard MG, Morris TT, Hansen AF, Rasheed H, Cho Y, et al. Within-sibship genome-wide association analyses decrease bias in estimates of direct genetic effects. Nat Genet. 2022;54:5. 2022;54:581–592.
Nivard MG, Belsky D, Harden KP, Baier T, Andreassen OA, Ystrom E, et al. Neither nature nor nurture: Using extended pedigree data to understand indirect genetic effects on offspring educational outcomes. 2023. https://doi.org/10.31234/OSF.IO/BHPM5.
Wang B, Baldwin JR, Schoeler T, Cheesman R, Barkhuizen W, Dudbridge F, et al. Robust genetic nurture effects on education: A systematic review and meta-analysis based on 38,654 families across 8 cohorts. Am J Hum Genet. 2021;108:1780–91.
Hugh-Jones D, Verweij KJH, St. Pourcain B, Abdellaoui A. Assortative mating on educational attainment leads to genetic spousal resemblance for polygenic scores. Intelligence. 2016;59:103–8.
Gonggrijp BMA, Silventoinen K, Dolan CV, Boomsma DI, Kaprio J, Willemsen G. The mechanism of assortative mating for educational attainment: a study of Finnish and Dutch twins and their spouses. Front Genet. 2023;14:1150697.
Acknowledgements
The authors are grateful to families, students, and staff of the public primary schools (i.e., Joan Maragall, María Bores, Marqués de la Pobla, Martinet, Pins del Vallès, Puiggracios, Sant Jordi, Ramon Llull, Rivo Rubeo, Tagamanent and Teresa Berguedà), public secondary schools (i.e., Angeleta Ferrer i Sensat, Antoni Pous i Argila, Cal Gravat, Duc de Montblanc, Institut del Ter, Jaume Callís, Lacetània, Lluís de Peguera, Molí de la Vila, Montsuar, Pius Font i Quer, Vallbona d’Anoia, and Vil la Romana), and private schools (i.e., Airina, L’Ave Maria, Casals – Gràcia, Episcopal Lleida, La Farga, FEDAC Manresa, FEDAC Vic, Garbí Pere Vergés Esplugues, Institucio Igualada, Joviat, Oms i de Prat, Pies Mataró, Pureza de Maria, Regina Carmeli, Sagrats Cors Centelles, La Salle Manlleu, La Salle Manresa, Sant Miquel dels Sants, Thau Barcelona and Vedruna Escorial Vic) who kindly contribute in this research. The genotyping service was carried out at the Genotyping Unit-CEGEN in the Spanish National Cancer Research Centre (CNIO), supported by Instituto de Salud Carlos III (ISCIII), Ministerio de Ciencia e Innovación. CEGEN is part of the initiative IMPaCTGENóMICA (IMP/00009) cofunded by ISCIII and the European Regional Development Fund (ERDF).
Funding
This work was supported by the Agència de Gestió d’Ajuts Universitaris i de Recerca (AGAUR, 2017SGR-1461, 2021SGR-00840), the Instituto de Salud Carlos III (P19/01224, PI20/00041, PI22/00464, PI23/00404, PI23/00026 and FI18/00285 to L.V.R, CP22/00128 to M.S.A and CP22/00026 to S.A), the Network Center for Biomedical Research (CIBER) to J.C.D.; the European Regional Development Fund (ERDF); the ECNP Network ‘ADHD across the Lifespan’; “la Marató de TV3” (202228-30 and 202228-31), “Fundació ‘la Caixa’, Diputació de Barcelona, Pla Estratègic de Recerca i Innovació en Salut” (PERISSLT006/17/285); “Fundació Privada d’Investigació Sant Pau” (FISP); and Ministry of Health of Generalitat de Catalunya.
Author information
Authors and Affiliations
Contributions
JCD, RB, MSA, SA, JARQ, MC and MR conceived the project. RB, GEM, CdC, LM, MP, EP, RP, JP, MC, JARQ participated in the clinical assessment and in the recruitment of participants. JCD, NL, LVR, PC, LA and VMC participated in the RNA isolation and preparation of DNA samples. JCD, RB, MSA, SA and MR undertook the statistical analyses, participated in the study design and the discussion of results. JCD, MSA, SA and MR participated in the manuscript preparation. All authors contributed to the interpretation of the findings and revised and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
JARQ was on the speakers’ bureau and/or acted as consultant for Biogen, Idorsia,Janssen-Cilag, Novartis, Takeda, Bial, Sincrolab, Neuraxpharm, Novartis, BMS, Medice, Rubió, Uriach, Technofarma and Raffo in the last 3 years. He also received travel awards (air tickets + hotel) for taking part in psychiatric meetings from Idorsia, Janssen-Cilag, Rubió, Takeda, Bial and Medice. The Department of Psychiatry chaired by him received unrestricted educational and research support from the following companies in the last 3 years: Exeltis, Idorsia, Janssen- Cilag, Neuraxpharm, Oryzon, Roche, Probitas, Psious, and Rubió. All other authors declare no biomedical financial interests or conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cabana-Domínguez, J., Bosch, R., Soler Artigas, M. et al. Dissecting the polygenic contribution of attention-deficit/hyperactivity disorder and autism spectrum disorder on school performance by their relationship with educational attainment. Mol Psychiatry (2024). https://doi.org/10.1038/s41380-024-02582-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41380-024-02582-w