Abstract
Severe speech disorders lead to poor literacy, reduced academic attainment and negative psychosocial outcomes. As early as the 1950s, the familial nature of speech disorders was recognized, implying a genetic basis; but the molecular genetic basis remained unknown. In 2001, investigation of a large three generational family with severe speech disorder, known as childhood apraxia of speech (CAS), revealed the first causative gene; FOXP2. A long hiatus then followed for CAS candidate genes, but in the past three years, genetic analysis of cohorts ascertained for CAS have revealed over 30 causative genes. A total of 36 pathogenic variants have been identified from 122 cases across 3 cohorts in this nascent field. All genes identified have been in coding regions to date, with no apparent benefit at this stage for WGS over WES in identifying monogenic conditions associated with CAS. Hence current findings suggest a remarkable one in three children have a genetic variant that explains their CAS, with significant genetic heterogeneity emerging. Around half of the candidate genes identified are currently supported by medium (6 genes) to strong (9 genes) evidence supporting the association between the gene and CAS. Despite genetic heterogeneity; many implicated proteins functionally converge on pathways involved in chromatin modification or transcriptional regulation, opening the door to precision diagnosis and therapies. Most of the new candidate genes for CAS are associated with previously described neurodevelopmental conditions that include intellectual disability, autism and epilepsy; broadening the phenotypic spectrum to a distinctly milder presentation defined by primary speech disorder in the setting of normal intellect. Insights into the genetic bases of CAS, a severe, rare speech disorder, are yet to translate to understanding the heritability of more common, typically milder forms of speech or language impairment such as stuttering or phonological disorder. These disorders likely follow complex inheritance with polygenic contributions in many cases, rather than the monogenic patterns that underly one-third of patients with CAS. Clinical genetic testing for should now be implemented for individuals with CAS, given its high diagnostic rate, which parallels many other neurodevelopmental disorders where this testing is already standard of care. The shared mechanisms implicated by gene discovery for CAS highlight potential new targets for future precision therapies.
Similar content being viewed by others
Introduction
Speech acquisition is a biologically driven, inexorable developmental process in most infants. Yet up to 5% of children develop common speech disorders including stuttering, articulation and phonological impairments [Table 1]. These common conditions are highly tractable and tend to resolve, with or without intervention, by 7 years of age [1, 2]. By contrast, 1 in 1000 children follow a severely disrupted developmental path to an intractable speech disorder known as childhood apraxia of speech (CAS) [3]. In these individuals, early development is often marked by hypotonia, feeding difficulties, limited babbling, delayed onset of first words, and marked difficulty in acquiring speech which is unintelligible in the preschool years, when a diagnosis is usually made [4]. The condition was first described by pioneering British speech therapist Muriel Morley in 1957 who identified a childhood speech presentation akin to the speech praxis seen in adults following lesions to Broca’s area, with the crux of the diagnosis being difficulty accurately producing sound sequences [5].
Since the original description of CAS, there has been ongoing debate over the defining diagnostic features of the condition [6]. In 2007, the American Speech and Hearing Association supported an expert-based consensus which defined the three diagnostic features of CAS (Table 1) [7]. Whilst the condition is largely framed as a ‘motor’ speech disorder resulting from movement planning or programming deficits, language and literacy impairments also occur in over 90% of individuals [8,9,10]. Furthermore, neuroimaging points to perturbation of linguistic as well as motor pathways, in affected individuals [11].
Recently, the CAS phenotype has increasingly been associated with commonly occurring neurodevelopmental comorbidities, including motor and cognitive impairments, attention deficit hyperactivity disorder, seizures and autism spectrum disorders [8,9,10]. Similar to the presentation of these neurodevelopmental disorders (NDDs), speech and language disorders rarely occur in isolation, and rather are found in a broader context of perturbed neurodevelopment.
Until recently, understanding of the aetiology of CAS was limited. Parents of children with CAS would embark on a diagnostic odyssey to investigate the chronic and striking nature of the condition. Early studies have implicated copy number variants (CNVs), including chromosomal aneuplodies involving multiple genes, and single nucleotide variants (SNVs) in individual genes, to CAS.
A specific neurogenetic basis for CAS was first identified in 2001, with the seminal discovery that pathogenic missense SNVs in FOXP2 [12], a transcriptional repressor, were associated with CAS, initially inherited in a large multiplex family, but subsequently also found to arise de novo in sporadic cases. Functionally related transcription factors and downstream targets of FOXP2 were subsequently investigated, namely CNTNAP2 (MIM: 604569), FOXP1 (MIM: 605515) and TBR1 (MIM:606053). Although these genes have been associated with intellectual disability syndromes and ASD, they have not explained cases ascertained for primary or isolated speech or language disorder [13,14,15]. The next most promising candidate gene for CAS was GRIN2A, which is also associated with the epilepsy-aphasia syndromes, now termed developmental and/or epileptic encephalopathy with spike-wave activation in sleep [16], and including Llandau-Kleffner syndrome [17,18,19]. Yet again, as for CNTNAP2, FOXP1 and TBR1, pathogenic variants in GRIN2A have not been identified in cohorts ascertained for a primary diagnosis of speech or language disorder in the absence of epilepsy.
Advances in microarray technology have also led to numerous chromosomal deletions being associated with CAS, but typically in the presence of cognitive impairment or ASD, such as 16p11.2 deletion [20, 21]. Some of these CNVs have drawn attention to possible genes in the pathogenesis of CAS such as 18q12.3 microdeletions encompassing SETBP1 [22], 12p13.33 microdeletions including ELKS/ERC1 [23], 2p15-p16.1 microdeletions encompassing and proximal to BCL11A [24], 7q11.23 duplication syndrome [25] implicating a number of genes and 17q21.31 deletion or Koolen-de Vries syndrome encompassing KANSL1 [26].
Most recently, advances in massively parallel sequencing technologies and bioinformatic algorithms have allowed rapid identification of genes not previously implicated in speech dysfunction. Here we review the rapidly unfolding Mendelian genetic bases for CAS. Specifically, we have reviewed data on gene discovery cohorts applying exome or genome sequencing to cohorts ascertained for primary speech disorder CAS [Search strategy box below].
Search strategy and selection criteria
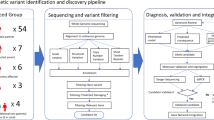
We searched PubMed for articles published between Jan 1, 2001, and March 15, 2023, using the search terms “childhood apraxia of speech”, “dyspraxia”, “speech”, “exome sequencing” and “genome sequencing”. There were no language restrictions. We selected articles that had ascertained cohorts with CAS and applied next generation sequencing approaches and analysis to report novel genes associated with CAS. We also searched for articles describing the function and implications of pathogenic variants in the genes identified, for literature on other neurodevelopmental disorders associated with these genes. The final reference list was generated based on the relevance to the topics covered in this review.
Discussion/analysis of recent literature
Three CAS gene discovery cohort studies were identified, each relatively small given the rarity of the disorder, but growing in cohort size over time: n = 19 probands, Eising et al., 2019; n = 33 probands, Hildebrand et al., 2020; n = 70 probands, and Kaspi et al., 2023. In the first study, 8/19 (~42%) probands were found to have a pathogenic or likely pathogenic gene variant via genome sequencing [8]. In the second study, 11/34 (~32%) probands had highly plausible pathogenic variants identified by a combination of exome and genome sequencing, and chromosomal microarray analysis [9]. In the third study, 18/70 (~26%) probands had a high confidence pathogenic variant detected via genome sequencing or chromosomal microarray analysis [10]. There was no apparent benefit at this stage for WGS over WES in identifying monogenic conditions associated with CAS. The overall clinical genetic diagnostic yield across the three cohorts was 30% (36/122 probands) (see Fig. 1a).
a Genetic causes identified for CAS over time. WGS, whole genome sequencing; WES, whole exome sequencing. More patients have undergone WGS to date. The yield to date has been similar for WGS and WES. No variants have been reported in non-coding regions to date. b CAS candidate genes and co-occurring neurodevelopmental phenotypes. An association between a gene and a phenotype was denoted if the association was higher than the prevalence in the general population. c Overlap in phenotype in children with CAS, comparing those with a pathogenic variant and those with no known cause. Phenotypic features of CAS cohorts with (n = 29) and without (n = 74) pathogenic genetic variants, based on data from Hildebrand et al. (2020) and Kaspi et al. (2023). Authors were approached for any updated diagnoses for study participants. Confirmation of diagnoses for cognitive impairment, ASD, receptive and expressive language impairment, and gross and fine motor impairment was based on psychometric testing/clinical report. Seizure diagnosis is based on parent report.
These studies provided the first neurobiological insights into the mechanisms of speech disorders, including the key finding that pathogenic variants are enriched in genes involved in transcriptional regulation and chromatin remodelling in the developing brain (Table 2). Importantly, these genes also showed significant clustering within a module of genes highly co-expressed in the human embryonic brain, in regions known to subserve speech function [8]. Hence the speech disorders field now has the first evidence that CAS is a neurodevelopmental disorder due to dysregulation of genes expressed in white-matter tracts critical for development of speech [8, 9, 27].
Unlike FOXP2, which had no disease association prior to being linked to CAS, many newer genes associated with CAS were already implicated in other NDDs such as intellectual disability (ID), ASD and epilepsy (see Table 2 for outline of known associated conditions). These findings [8,9,10] align with the well-established genetic overlap between other neurodevelopmental phenotypes [28] and indicate that CAS can be added to these overlapping profiles [see Fig. 1b].
In some ways, the association of CAS with genes known to cause ID, ASD and epilepsy is not surprising given that these neurodevelopmental phenotypes have long been associated with speech and language pathology; however, a primary speech phenotype of CAS had been considered separate from the larger group of NDDs. Now genetic findings show this distinction may not be valid. Furthermore, recent studies of individuals with FOXP2 variants show that they also experience broader, subtle, neurodevelopmental phenotypes beyond speech dysfunction [29]. Thus, whilst FOXP2 remains the most ‘speech specific’ gene to be identified [29], there may not exist a “pure” speech apraxia gene. As such, speech and neurodevelopmental phenotypes should be considered as existing across a phenotypic spectrum rather than as categorical diagnoses, mirroring findings in genetic understanding of other diseases, such as epilepsy.
Finally, there is currently a strong bias in comparing next generation sequencing findings for ID, ASD and epilepsy, with tens of thousands of probands reported in the literature, compared to just over 120 probands with CAS. Thus, surprisingly, the published CAS cohort studies have shown a comparably high genetic diagnostic yield for individuals with these speech phenotypes, despite them arguably being milder relative to ID, ASD and epilepsy. This suggests clinical genetic testing is also warranted for children with CAS given that genome-wide testing is increasingly routine and often funded for children with other NDDs. Routine clinical genetic testing will be important as although many of the gene variants reported to date are de novo and predicted pathogenic according to ACMG guidelines, most have been found only in individual probands and identification of the same gene in unrelated patients with the same phenotype will confirm that gene’s contribution. As outlined below, unrelated patients have been identified for three of the candidate genes for CAS across the small CAS-ascertained cohort studies alone.
Although genetic heterogeneity is a feature of gene discovery findings in CAS cohorts, pathogenic variants in a handful of genes, namely SETBP1, SETD1A and DDX3X, each account for multiple cases across the cohorts studied to date [8,9,10]. SETBP1 stands out as being particularly intriguing, with pathogenic loss-of-function (LoF) variants detected in all three cohorts [8,9,10] With emerging evidence for CAS in SETBP1 haploinsufficiency disorder, a speech and language study of 28 individuals with SETBP1 LoF variants then confirmed the diagnosis of CAS, seen in 80% of individuals studied, as a core part of the phenotype [30]. When comparing children’s performance across developmental domains, it was also clear that communication was most impaired relative to social skills, daily living skills, motor abilities and adaptive functioning, supporting SETBP1 having a central role in speech and language development [30]. Further, studies of common genetic variants suggest SETBP1 may also be important for communication abilities in the general population. Associations between single nucleotide polymorphisms (SNPs) in SETBP1 and scores on a test examining syntactic complexity were reported in a genome wide association study of language disorder in a geographically isolated Russian cohort aged 3–18 years [31]. SNPs in SETBP1 have also been associated with phonological working memory in a reading-impaired cohort [32].
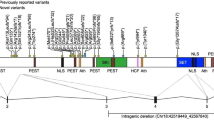
In addition to evidence for the strength of association between CAS and the candidate genes across the three CAS-ascertained gene discovery cohorts discussed here, Table 2 further outlines the strength of independent evidence currently found to support the candidate genes. At this time, nine of the candidate genes have a high level of independent supporting evidence (FOXP2, KAT6A, MKL2/MRTFB, SETBP1, CDK13, EBF3, MEIS2, RBFOX3, SHANK3), six have medium (CHD3, SETD1A, WDR5, DDX3X, ZNF142, BRPF1) and the remainder have low levels of independent evidence, but we expect expanded clinical genetic testing will reveal additional cases for many of the other candidate genes implicated [8,9,10].
Alternative genetic mechanisms for CAS
If high impact de novo sequence variants and CNVs account for about one third of individuals with CAS, the question that follows is what genes or mechanisms account for the remaining unsolved cases. Whole genome sequencing has not been completed for all cases studied, meaning non-coding variants have not been routinely interrogated and may account for some undiagnosed cases. Mosaicism, increasingly implicated in neurodevelopmental diseases such as intellectual disability, epilepsy and autism [33], may be low level and brain-limited and may underpin CAS in some individuals where it may be limited to key networks; however, detection would require sequencing of brain tissue, which is generally inaccessible.
From existing data, there is evidence that the cohort of CAS individuals with an identified pathogenic de novo gene variant is enriched for individuals with cognitive impairment and co-morbid language and motor diagnoses, compared to those without a genetic diagnosis (Fig. 1c). These data suggest that different genetic mechanisms may apply to those cases with CAS currently without a specific single gene diagnosis.
It is likely that inherited variants will account for a sizeable portion of CAS, but elucidation of these variants will require large cohorts coupled with deep phenotyping of family members. Interestingly, many families report a family history of speech difficulties, which might be explained by inherited variants that exhibit variable expressivity and phenotypic heterogeneity due to variability in the genetic background, similar to multi-hit models in other neurodevelopmental disorders [34].
The fact that many children with CAS exhibit comorbidities with ASD and ADHD suggests an additional genetic overlap with these conditions, which are mostly attributed to polygenic aetiology, as is the case for ASD [35]. A limitation in this nascent field of speech genetics is the lack of available population-based cohorts with both high quality genetic and phenotyping data. There is a concerted effort by the address this issue via the GenLang consortia (https://www.genlang.org); yet, to date, the cohorts in GenLang typically include language and literacy data, with recent fruitful GWAS publications identifying loci associated with language and literacy traits [36,37,38], but do not include speech-specific data.
One remaining challenge for the field is the lack of clinical variables which robustly predict who will have a monogenic cause or polygenic contributions (accepting that for monogenic diseases there may be modifier genetic contributions). From existing data, there is some evidence that individuals with any degree of cognitive impairment, or those more likely to have co-morbid language and motor diagnoses, have a greater likelihood of monogenic disease (Fig. 1c). However, it is clear that larger cohorts of individuals with CAS are required to provide adequate power to generate accurate genetic diagnostic prediction models to confirm these findings.
Conclusions
After almost two decades with only one established gene for CAS, over 30 new genes of relevance have been identified in the past three years. Critically, about one third of children sequenced have received a molecular genetic diagnosis for their CAS, supporting implementation in clinical testing alongside other neurodevelopmental disorders. Around half of the candidate genes identified are currently supported by medium to strong evidence supporting the association between the gene and CAS. Almost all genes identified were previously described to cause neurodevelopmental conditions including ID, ASD and epilepsy. Hence the phenotypic spectrum of these conditions has been expanded to include individuals with a distinctly milder presentation of primary speech disorder. Whilst there is genetic heterogeneity, the genes coalesce on a small number of biological pathways, largely involved in chromatin remodelling or transcriptional regulation, providing new targets for precision medicines. Although genetic diagnoses in CAS to date have largely been de novo high impact variants in neurodevelopmental genes, the genetic architecture of CAS is likely to also encompass polygenic inheritance of common variants and rare inherited variants with incomplete penetrance and variable expressivity.
Data availability
Data for this review was collated from the three manuscripts meeting inclusion criteria (Kaspi et al., 2023; Hildebrand et al., 2020; Eising et al., 2020), or from additional manuscripts providing further supporting evidence of the association between apraxia of speech (phenotype) and specific genotypes which are all cited in the reference list. Hence all data is publicly available and replication possible using methods from the review.
References
Eadie P, Morgan A, Ukoumunne OC, Ttofari Eecen K, Wake M, Reilly S. Speech sound disorder at 4 years: prevalence, comorbidities, and predictors in a community cohort of children. Dev Med Child Neurol. 2015;57:578–84.
Reilly S, McKean C, Morgan A, Wake M. Identifying and managing common childhood language and speech impairments. BMJ. 2015;350:h2318.
Shriberg LD, Aram DM, Kwiatkowski J. Developmental apraxia of speech: I. Descriptive and theoretical perspectives. J Speech Lang Hear Res. 1997;40:273–85.
Morgan AT, Fisher SE, Scheffer IE, Hildebrand MJ FOXP2-related speech and language disorder; In: Adam MP, Mirzaa GM, Pagon RA, Wallace SE, Bean LJH, Gripp KW, et al. (Eds.), GeneReviews® [Internet]. University of Washington, Seattle; 2023. https://www.ncbi.nlm.nih.gov/books/NBK368474/.
Morley M. The development and disorders of speech in childhood. London: Churchill Livingston; 1957.
Morgan AT, Webster R. Aetiology of childhood apraxia of speech: a clinical practice update for paediatricians. J Paediatr Child Health. 2018;54:1090–5.
American Speech-Language-Hearing Association. Childhood apraxia of speech, 2007. [Technical Report].
Eising E, Carrion-Castillo A, Vino A, Strand EA, Jakielski KJ, Scerri TS, et al. A set of regulatory genes co-expressed in embryonic human brain is implicated in disrupted speech development. Mol Psychiatry. 2019;24:1065–78.
Hildebrand MS, Jackson VE, Scerri TS, Van Reyk O, Coleman M, Braden RO, et al. Severe childhood speech disorder: gene discovery highlights transcriptional dysregulation. Neurology. 2020;94:e2148–e67.
Kaspi A, Hildebrand MS, Jackson VE, Braden R, van Reyk O, Howell T, et al. Genetic aetiologies for childhood speech disorder: novel pathways co-expressed during brain development. Mol Psychiatry. 2023;28:1647–63.
Liegeois FJ, Turner SJ, Mayes A, Bonthrone AF, Boys A, Smith L, et al. Dorsal language stream anomalies in an inherited speech disorder. Brain. 2019;142:966–77.
Lai CS, Fisher SE, Hurst JA, Vargha-Khadem F, Monaco AP. A forkhead-domain gene is mutated in a severe speech and language disorder. Nature. 2001;413:519–23.
Braden RO, Amor DJ, Fisher SE, Mei C, Myers CT, Mefford H, et al. Severe speech impairment is a distinguishing feature of FOXP1-related disorder. Dev Med Child Neurol. 2021;63:1417–26.
Rodenas-Cuadrado P, Ho J, Vernes SC. Shining a light on CNTNAP2: complex functions to complex disorders. Eur J Hum Genet. 2014;22:171–8.
Deriziotis P, O’Roak BJ, Graham SA, Estruch SB, Dimitropoulou D, Bernier RA, et al. De novo TBR1 mutations in sporadic autism disrupt protein functions. Nat Commun. 2014;5:4954.
Specchio N, Wirrell EC, Scheffer IE, Nabbout R, Riney K, Samia P, et al. International League Against Epilepsy classification and definition of epilepsy syndromes with onset in childhood: Position paper by the ILAE Task Force on Nosology and Definitions. Epilepsia. 2022;63:1398–442.
Turner SJ, Mayes AK, Verhoeven A, Mandelstam SA, Morgan AT, Scheffer IE. GRIN2A: an aptly named gene for speech dysfunction. Neurology. 2015;84:586–93.
Carvill GL, Regan BM, Yendle SC, O’Roak BJ, Lozovaya N, Bruneau N, et al. GRIN2A mutations cause epilepsy-aphasia spectrum disorders. Nat Genet. 2013;45:1073–6.
Lesca G, Rudolf G, Bruneau N, Lozovaya N, Labalme A, Boutry-Kryza N, et al. GRIN2A mutations in acquired epileptic aphasia and related childhood focal epilepsies and encephalopathies with speech and language dysfunction. Nat Genet. 2013;45:1061–6.
Laffin JJ, Raca G, Jackson CA, Strand EA, Jakielski KJ, Shriberg LD. Novel candidate genes and regions for childhood apraxia of speech identified by array comparative genomic hybridization. Genet Med. 2012;14:928–36.
Mei C, Fedorenko E, Amor DJ, Boys A, Hoeflin C, Carew P, et al. Deep phenotyping of speech and language skills in individuals with 16p11.2 deletion. Eur J Hum Genet. 2018;26:676–86.
Coe BP, Witherspoon K, Rosenfeld JA, van Bon BW, Vulto-van Silfhout AT, Bosco P, et al. Refining analyses of copy number variation identifies specific genes associated with developmental delay. Nat Genet. 2014;46:1063–71.
Thevenon J, Callier P, Andrieux J, Delobel B, David A, Sukno S, et al. 12p13.33 microdeletion including ELKS/ERC1, a new locus associated with childhood apraxia of speech. Eur J Hum Genet. 2013;21:82–8.
Peter B, Matsushita M, Oda K, Raskind W. De novo microdeletion of BCL11A is associated with severe speech sound disorder. Am J Med Genet A. 2014;164A:2091–6.
Mervis CB, Klein-Tasman BP, Huffman MJ, Velleman SL, Pitts CH, Henderson DR, et al. Children with 7q11.23 duplication syndrome: psychological characteristics. Am J Med Genet A. 2015;167:1436–50.
Morgan AT, Haaften LV, van Hulst K, Edley C, Mei C, Tan TY, et al. Early speech development in Koolen de Vries syndrome limited by oral praxis and hypotonia. Eur J Hum Genet. 2018;26:75–84.
St John M, Tripathi T, Morgan AT, Amor DJ. To speak may draw on epigenetic writing and reading: Unravelling the complexity of speech and language outcomes across chromatin-related neurodevelopmental disorders. Neurosci Biobehav Rev. 2023;152:105293.
Jensen M, Girirajan S. Mapping a shared genetic basis for neurodevelopmental disorders. Genome Med. 2017;9:109.
Morison LD, Meffert E, Stampfer M, Steiner-Wilke I, Vollmer B, Schulze K, et al. In-depth characterisation of a cohort of individuals with missense and loss-of-function variants disrupting FOXP2. J Med Genet. 2023;60:597–607.
Morgan A, Braden R, Wong MMK, Colin E, Amor D, Liégeois F, et al. Speech and language deficits are central to SETBP1 haploinsufficiency disorder. Eur J Hum Genet. 2021;29:1216–25.
Kornilov SA, Rakhlin N, Koposov R, Lee M, Yrigollen C, Caglayan AO, et al. Genome-wide association and exome sequencing study of language disorder in an isolated population. Pediatrics. 2016;137:e20152469.
Perdue MV, Mascheretti S, Kornilov SA, Jasińska KK, Ryherd K, Einar Mencl W, et al. Common variation within the SETBP1 gene is associated with reading-related skills and patterns of functional neural activation. Neuropsychologia. 2019;130:44–51.
D’Gama AM, Walsh CA. Somatic mosaicism and neurodevelopmental disease. Nat Neurosci. 2018;21:1504–14.
Pizzo L, Jensen M, Polyak A, Rosenfeld JA, Mannik K, Krishnan A, et al. Rare variants in the genetic background modulate cognitive and developmental phenotypes in individuals carrying disease-associated variants. Genet Med. 2019;21:816–25.
Warrier V, Zhang X, Reed P, Havdahl A, Moore TM, Cliquet F, et al. Genetic correlates of phenotypic heterogeneity in autism. Nat Genet. 2022;54:1293–304.
Price KM, Wigg KG, Eising E, Feng Y, Blokland K, Wilkinson M, et al. Hypothesis-driven genome-wide association studies provide novel insights into genetics of reading disabilities. Transl Psychiatry. 2022;12:495.
Doust C, Fontanillas P, Eising E, Gordon SD, Wang Z, Alagöz G, et al. Discovery of 42 genome-wide significant loci associated with dyslexia. Nat Genet. 2022;54:1621–9.
Eising E, Mirza-Schreiber N, de Zeeuw EL, Wang CA, Truong DT, Allegrini AG, et al. Genome-wide analyses of individual differences in quantitatively assessed reading- and language-related skills in up to 34,000 people. Proc Natl Acad Sci USA. 2022;119:e2202764119.
Dodd B, Reilly S, Ttofari Eecen K, Morgan AT. Articulation or phonology? Evidence from longitudinal error data. Clin Linguist Phon. 2018;32:1027–41.
Kefalianos E, Onslow M, Packman A, Vogel A, Pezic A, Mensah F, et al. The History of stuttering by 7 years of age: follow-up of a prospective community cohort. J Speech Lang Hear Res. 2017;60:2828–39.
Spiteri E, Konopka G, Coppola G, Bomar J, Oldham M, Ou J, et al. Identification of the transcriptional targets of FOXP2, a gene linked to speech and language, in developing human brain. Am J Hum Genet. 2007;81:1144–57.
Nitarska J, Smith JG, Sherlock WT, Hillege MMG, Nott A, Barshop WD, et al. A functional switch of NuRD chromatin remodeling complex subunits regulates mouse cortical development. Cell Rep. 2016;17:1683–98.
Snijders Blok L, Rousseau J, Twist J, Ehresmann S, Takaku M, Venselaar H, et al. CHD3 helicase domain mutations cause a neurodevelopmental syndrome with macrocephaly and impaired speech and language. Nat Commun. 2018;9:4619.
Voss AK, Collin C, Dixon MP, Thomas T. Moz and retinoic acid coordinately regulate H3K9 acetylation, Hox gene expression, and segment identity. Dev Cell. 2009;17:674–86.
St John M, Amor DJ, Morgan AT. Speech and language development and genotype-phenotype correlation in 49 individuals with KAT6A syndrome. Am J Med Genet A. 2022;188:3389–400.
Andrews JC, Mok JW, Kanca O, Jangam S, Tifft C, Macnamara EF, et al. De novo variants in MRTFB have gain-of-function activity in Drosophila and are associated with a novel neurodevelopmental phenotype with dysmorphic features. Genet Med. 2023;25:100833.
Piazza R, Magistroni V, Redaelli S, Mauri M, Massimino L, Sessa A, et al. SETBP1 induces transcription of a network of development genes by acting as an epigenetic hub. Nat Commun. 2018;9:2192.
Kummeling J, Stremmelaar DE, Raun N, Reijnders MRF, Willemsen MH, Ruiterkamp-Versteeg M, et al. Characterization of SETD1A haploinsufficiency in humans and Drosophila defines a novel neurodevelopmental syndrome. Mol Psychiatry. 2021;26:2013–24.
Baillat D, Shiekhattar R. Functional dissection of the human TNRC6 (GW182-related) family of proteins. Mol Cell Biol. 2009;29:4144–55.
Bryan AF, Wang J, Howard GC, Guarnaccia AD, Woodley CM, Aho ER, et al. WDR5 is a conserved regulator of protein synthesis gene expression. Nucleic Acids Res. 2020;48:2924–41.
Snijders Blok L, Verseput J, Rots D, Venselaar H, Innes AM, Stumpel C, et al. A clustering of heterozygous missense variants in the crucial chromatin modifier WDR5 defines a new neurodevelopmental disorder. HGG Adv. 2023;4:100157.
Fontana P, Ginevrino M, Bejo K, Cantalupo G, Ciavarella M, Lombardi C, et al. A ZFHX4 mutation associated with a recognizable neuropsychological and facial phenotype. Eur J Med Genet. 2021;64:104321.
Liang K, Gao X, Gilmore JM, Florens L, Washburn MP, Smith E, et al. Characterization of human cyclin-dependent kinase 12 (CDK12) and CDK13 complexes in C-terminal domain phosphorylation, gene transcription, and RNA processing. Mol Cell Biol. 2015;35:928–38.
Morison LD, van Reyk O, Forbes E, Rouxel F, Faivre L, Bruinsma F, et al. CDK13-related disorder: a deep characterization of speech and language abilities and addition of 33 novel cases. Eur J Hum Genet. 2023;31:793–804.
Snijders Blok L, Madsen E, Juusola J, Gilissen C, Baralle D, Reijnders MR, et al. Mutations in DDX3X are a common cause of unexplained intellectual disability with gender-specific effects on wnt signaling. Am J Hum Genet. 2015;97:343–52.
Treiber T, Mandel EM, Pott S, Györy I, Firner S, Liu ET, et al. Early B cell factor 1 regulates B cell gene networks by activation, repression, and transcription- independent poising of chromatin. Immunity. 2010;32:714–25.
Chao HT, Davids M, Burke E, Pappas JG, Rosenfeld JA, McCarty AJ, et al. A syndromic neurodevelopmental disorder caused by de novo variants in EBF3. Am J Hum Genet. 2017;100:128–37.
Feng H, Khalil S, Neubig RR, Sidiropoulos C. A mechanistic review on GNAO1-associated movement disorder. Neurobiol Dis. 2018;116:131–41.
Wirth T, Garone G, Kurian MA, Piton A, Millan F, Telegrafi A, et al. Highlighting the dystonic phenotype related to GNAO1. Mov Disord. 2022;37:1547–54.
Lohmann K, Masuho I, Patil DN, Baumann H, Hebert E, Steinrücke S, et al. Novel GNB1 mutations disrupt assembly and function of G protein heterotrimers and cause global developmental delay in humans. Hum Mol Genet. 2017;26:1078–86.
Yao M, Gu Y, Yang Z, Zhong K, Chen Z. MEIS1 and its potential as a cancer therapeutic target (Review). Int J Mol Med. 2021;48:181.
Douglas G, Cho MT, Telegrafi A, Winter S, Carmichael J, Zackai EH, et al. De novo missense variants in MEIS2 recapitulate the microdeletion phenotype of cardiac and palate abnormalities, developmental delay, intellectual disability and dysmorphic features. Am J Med Genet A. 2018;176:1845–51.
Markenscoff-Papadimitriou E, Binyameen F, Whalen S, Price J, Lim K, Ypsilanti AR, et al. Autism risk gene POGZ promotes chromatin accessibility and expression of clustered synaptic genes. Cell Rep. 2021;37:110089.
Nagy D, Verheyen S, Wigby KM, Borovikov A, Sharkov A, Slegesky V, et al. Genotype-phenotype comparison in POGZ-related neurodevelopmental disorders by using clinical scoring. Genes (Basel). 2022;13:154.
Johnson JL, Stoica L, Liu Y, Zhu PJ, Bhattacharya A, Buffington SA, et al. Inhibition of Upf2-dependent nonsense-mediated decay leads to behavioral and neurophysiological abnormalities by activating the immune response. Neuron. 2019;104:665–79.e8.
Kamal N, Khamirani HJ, Mohammadi S, Dastgheib SA, Dianatpour M, Tabei SMB. ZNF142 mutation causes neurodevelopmental disorder with speech impairment and seizures: novel variants and literature review. Eur J Med Genet. 2022;65:104522.
Khan K, Zech M, Morgan AT, Amor DJ, Skorvanek M, Khan TN, et al. Recessive variants in ZNF142 cause a complex neurodevelopmental disorder with intellectual disability, speech impairment, seizures, and dystonia. Genet Med. 2019;21:2532–42.
Christensen MB, Levy AM, Mohammadi NA, Niceta M, Kaiyrzhanov R, Dentici ML, et al. Biallelic variants in ZNF142 lead to a syndromic neurodevelopmental disorder. Clin Genet. 2022;102:98–109.
Ravindran E, Hu H, Yuzwa SA, Kraemer N, Ninnemann O, Musante L, et al. Homozygous ARHGEF2 mutation causes intellectual disability and midbrain-hindbrain malformation. PLoS Genet. 2017;13:e1006746.
Poplawski A, Hu K, Lee W, Natesan S, Peng D, Carlson S, et al. Molecular insights into the recognition of N-terminal histone modifications by the BRPF1 bromodomain. J Mol Biol. 2014;426:1661–76.
Yan K, Rousseau J, Littlejohn RO, Kiss C, Lehman A, Rosenfeld JA, et al. Mutations in the chromatin regulator gene BRPF1 cause syndromic intellectual disability and deficient histone acetylation. Am J Hum Genet. 2017;100:91–104.
Larsson C, Ali MA, Pandzic T, Lindroth AM, He L, Sjoblom T. Loss of DIP2C in RKO cells stimulates changes in DNA methylation and epithelial-mesenchymal transition. BMC Cancer. 2017;17:487.
Fukuda T, Naiki T, Saito M, Irie K. hnRNP K interacts with RNA binding motif protein 42 and functions in the maintenance of cellular ATP level during stress conditions. Genes Cells. 2009;14:113–28.
Tahiliani M, Mei P, Fang R, Leonor T, Rutenberg M, Shimizu F, et al. The histone H3K4 demethylase SMCX links REST target genes to X-linked mental retardation. Nature. 2007;447:601–5.
Jensen LR, Amende M, Gurok U, Moser B, Gimmel V, Tzschach A, et al. Mutations in the JARID1C gene, which is involved in transcriptional regulation and chromatin remodeling, cause X-linked mental retardation. Am J Hum Genet. 2005;76:227–36.
Tzschach A, Lenzner S, Moser B, Reinhardt R, Chelly J, Fryns JP, et al. Novel JARID1C/SMCX mutations in patients with X-linked mental retardation. Hum Mutat. 2006;27:389.
Leonardi E, Aspromonte MC, Drongitis D, Bettella E, Verrillo L, Polli R, et al. Expanding the genetics and phenotypic spectrum of Lysine-specific demethylase 5C (KDM5C): a report of 13 novel variants. Eur J Hum Genet. 2023;31:202–15.
Lan F, Collins RE, De Cegli R, Alpatov R, Horton JR, Shi X, et al. Recognition of unmethylated histone H3 lysine 4 links BHC80 to LSD1-mediated gene repression. Nature. 2007;448:718–22.
Hunt D, Leventer RJ, Simons C, Taft R, Swoboda KJ, Gawne-Cain M, et al. Whole exome sequencing in family trios reveals de novo mutations in PURA as a cause of severe neurodevelopmental delay and learning disability. J Med Genet. 2014;51:806–13.
Lin YS, Wang HY, Huang DF, Hsieh PF, Lin MY, Chou CH, et al. Neuronal splicing regulator RBFOX3 (NeuN) regulates adult hippocampal neurogenesis and synaptogenesis. PLoS ONE. 2016;11:e0164164.
Lal D, Reinthaler EM, Altmuller J, Toliat MR, Thiele H, Nürnberg P, et al. RBFOX1 and RBFOX3 mutations in rolandic epilepsy. PLoS ONE. 2013;8:e73323.
Lee JH, Tate CM, You JS, Skalnik DG. Identification and characterization of the human Set1B histone H3-Lys4 methyltransferase complex. J Biol Chem. 2007;282:13419–28.
Monteiro P, Feng G. SHANK proteins: roles at the synapse and in autism spectrum disorder. Nat Rev Neurosci. 2017;18:147–57.
Brignell A, Gu C, Holm A, Carrigg B, Sheppard DA, Amor DJ, et al. Speech and language phenotype in Phelan-McDermid (22q13.3) syndrome. Eur J Hum Genet. 2021;29:564–74.
Roll-Mecak A, Vale RD. Structural basis of microtubule severing by the hereditary spastic paraplegia protein spastin. Nature. 2008;451:363–7.
Larrieu D, Brunet M, Vargas C, Hanoun N, Ligat L, Dagnon L, et al. The E3 ubiquitin ligase TRIP12 participates in cell cycle progression and chromosome stability. Sci Rep. 2020;10:789.
Wang R, Bhatt AB, Minden-Birkenmaier BA, Travis OK, Tiwari S, Jia H, et al. ZBTB18 restricts chromatin accessibility and prevents transcriptional adaptations that drive metastasis. Sci Adv. 2023;9:eabq3951.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
AM, DA and MH drafted the manuscript. MSJ developed figures. All authors contributed further to the tables and manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Morgan, A.T., Amor, D.J., St John, M.D. et al. Genetic architecture of childhood speech disorder: a review. Mol Psychiatry (2024). https://doi.org/10.1038/s41380-024-02409-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41380-024-02409-8