Abstract
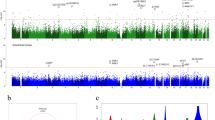
Human hippocampal volume has been separately associated with single nucleotide polymorphisms (SNPs), DNA methylation and gene expression, but their causal relationships remain largely unknown. Here, we aimed at identifying the causal relationships of SNPs, DNA methylation, and gene expression that are associated with hippocampal volume by integrating cross-omics analyses with genome editing, overexpression and causality inference. Based on structural neuroimaging data and blood-derived genome, transcriptome and methylome data, we prioritized a possibly causal association across multiple molecular phenotypes: rs1053218 mutation leads to cg26741686 hypermethylation, thus leads to overactivation of the associated ANKRD37 gene expression in blood, a gene involving hypoxia, which may result in the reduction of human hippocampal volume. The possibly causal relationships from rs1053218 to cg26741686 methylation to ANKRD37 expression obtained from peripheral blood were replicated in human hippocampal tissue. To confirm causality, we performed CRISPR-based genome and epigenome-editing of rs1053218 homologous alleles and cg26741686 methylation in mouse neural stem cell differentiation models, and overexpressed ANKRD37 in mouse hippocampus. These in-vitro and in-vivo experiments confirmed that rs1053218 mutation caused cg26741686 hypermethylation and ANKRD37 overexpression, and cg26741686 hypermethylation favored ANKRD37 overexpression, and ANKRD37 overexpression reduced hippocampal volume. The pairwise relationships of rs1053218 with hippocampal volume, rs1053218 with cg26741686 methylation, cg26741686 methylation with ANKRD37 expression, and ANKRD37 expression with hippocampal volume could be replicated in an independent healthy young (n = 443) dataset and observed in elderly people (n = 194), and were more significant in patients with late-onset Alzheimer’s disease (n = 76). This study revealed a novel causal molecular association mechanism of ANKRD37 with human hippocampal volume, which may facilitate the design of prevention and treatment strategies for hippocampal impairment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
Custom code that supports the findings of this study is available from the corresponding author upon request.
References
Lisman J, Buzsáki G, Eichenbaum H, Nadel L, Ranganath C, Redish AD. Viewpoints: how the hippocampus contributes to memory, navigation and cognition. Nat Neurosci. 2017;20:1434–47.
Kim EJ, Pellman B, Kim JJ. Stress effects on the hippocampus: a critical review. Learn Mem. 2015;22:411–6.
Moodley K, Chan D. The hippocampus in neurodegenerative disease. Front Neurol Neurosci. 2014;34:95–108.
Chatzikonstantinou A. Epilepsy and the hippocampus. Front Neurol Neurosci. 2014;34:121–42.
MacQueen G, Frodl T. The hippocampus in major depression: evidence for the convergence of the bench and bedside in psychiatric research? Mol Psychiatry. 2011;16:252–64.
Roeske MJ, Konradi C, Heckers S, Lewis AS. Hippocampal volume and hippocampal neuron density, number and size in schizophrenia: a systematic review and meta-analysis of postmortem studies. Mol Psychiatry. 2021;26:3524–35.
Halliday G. Pathology and hippocampal atrophy in Alzheimer’s disease. Lancet Neurol. 2017;16:862–4.
Bayram E, Caldwell JZ, Banks SJ. Current understanding of magnetic resonance imaging biomarkers and memory in Alzheimer’s disease. Alzheimers Dement (N Y). 2018;4:395–413.
Henneman WJ, Sluimer JD, Barnes J, van der Flier WM, Sluimer IC, Fox NC, et al. Hippocampal atrophy rates in Alzheimer disease: added value over whole brain volume measures. Neurology. 2009;72:999–1007.
Jack CR Jr, Petersen RC, Xu Y, O’Brien PC, Smith GE, Ivnik RJ, et al. Rates of hippocampal atrophy correlate with change in clinical status in aging and AD. Neurology. 2000;55:484–90.
Stein JL, Medland SE, Vasquez AA, Hibar DP, Senstad RE, Winkler AM, et al. Identification of common variants associated with human hippocampal and intracranial volumes. Nat Genet. 2012;44:552–61.
Hibar DP, Stein JL, Renteria ME, Arias-Vasquez A, Desrivières S, Jahanshad N, et al. Common genetic variants influence human subcortical brain structures. Nature. 2015;520:224–9.
Elliott LT, Sharp K, Alfaro-Almagro F, Shi S, Miller KL, Douaud G, et al. Genome-wide association studies of brain imaging phenotypes in UK Biobank. Nature. 2018;562:210–6.
Alexander RP, Fang G, Rozowsky J, Snyder M, Gerstein MB. Annotating non-coding regions of the genome. Nat Rev Genet. 2010;11:559–71.
Jasinska AJ, Zelaya I, Service SK, Peterson CB, Cantor RM, Choi OW, et al. Genetic variation and gene expression across multiple tissues and developmental stages in a nonhuman primate. Nat Genet. 2017;49:1714–21.
Kepa A, Martinez Medina L, Erk S, Srivastava DP, Fernandes A, Toro R, et al. Associations of the intellectual disability gene MYT1L with helix–loop–helix gene expression, hippocampus volume and hippocampus activation during memory retrieval. Neuropsychopharmacology. 2017;42:2516–26.
Jia T, Chu C, Liu Y, van Dongen J, Papastergios E, Armstrong NJ, et al. Epigenome-wide meta-analysis of blood DNA methylation and its association with subcortical volumes: findings from the ENIGMA Epigenetics Working Group. Mol Psychiatry. 2021;26:3884–95.
McRae AF, Powell JE, Henders AK, Bowdler L, Hemani G, Shah S, et al. Contribution of genetic variation to transgenerational inheritance of DNA methylation. Genome Biol. 2014;15:1–10.
Gutierrez-Arcelus M, Lappalainen T, Montgomery SB, Buil A, Ongen H, Yurovsky A, et al. Passive and active DNA methylation and the interplay with genetic variation in gene regulation. Elife. 2013;2:e00523.
Wu H, Zhang Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell. 2014;156:45–68.
Blake LE, Roux J, Hernando-Herraez I, Banovich NE, Perez RG, Hsiao CJ, et al. A comparison of gene expression and DNA methylation patterns across tissues and species. Genome Res. 2020;30:250–62.
Zhu Z, Zhang F, Hu H, Bakshi A, Robinson MR, Powell JE, et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet. 2016;48:481–7.
Giambartolomei C, Vukcevic D, Schadt EE, Franke L, Hingorani AD, Wallace C, et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014;10:e1004383.
Millstein J, Zhang B, Zhu J, Schadt EE. Disentangling molecular relationships with a causal inference test. BMC Genet. 2009;10:1–15.
Delaneau O, Ongen H, Brown AA, Fort A, Panousis NI, Dermitzakis ET. A complete tool set for molecular QTL discovery and analysis. Nat Commun. 2017;8:15452.
Davies MN, Volta M, Pidsley R, Lunnon K, Dixit A, Lovestone S, et al. Functional annotation of the human brain methylome identifies tissue-specific epigenetic variation across brain and blood. Genome Biol. 2012;13:R43.
Montano C, Taub MA, Jaffe A, Briem E, Feinberg JI, Trygvadottir R, et al. Association of DNA methylation differences with schizophrenia in an epigenome-wide association study. JAMA Psychiatry. 2016;73:506–14.
Schulz H, Ruppert AK, Herms S, Wolf C, Mirza-Schreiber N, Stegle O, et al. Genome-wide mapping of genetic determinants influencing DNA methylation and gene expression in human hippocampus. Nat Commun. 2017;8:1511.
Burgess S, Davey Smith G, Davies NM, Dudbridge F, Gill D, Glymour MM, et al. Guidelines for performing Mendelian randomization investigations. Wellcome Open Res. 2019;4:186.
Schumann G, Loth E, Banaschewski T, Barbot A, Barker G, Büchel C, et al. The IMAGEN study: reinforcement-related behaviour in normal brain function and psychopathology. Mol Psychiatry. 2010;15:1128–39.
Diedenhofen B, Musch J. cocor: a comprehensive solution for the statistical comparison of correlations. PLoS One. 2015;10:e0121945.
Smith SM, Douaud G, Chen W, Hanayik T, Alfaro-Almagro F, Sharp K, et al. An expanded set of genome-wide association studies of brain imaging phenotypes in UK Biobank. Nat Neurosci. 2021;24:737–45.
Bork P. Hundreds of ankyrin‐like repeats in functionally diverse proteins: mobile modules that cross phyla horizontally? Proteins. 1993;17:363–74.
Benita Y, Kikuchi H, Smith AD, Zhang MQ, Chung DC, Xavier RJ, et al. An integrative genomics approach identifies Hypoxia Inducible Factor-1 (HIF-1)-target genes that form the core response to hypoxia. Nucleic Acids Res. 2009;37:4587–602.
Galbraith MD, Allen MA, Bensard CL, Wang X, Schwinn MK, Qin B, et al. HIF1A employs CDK8-mediator to stimulate RNAPII elongation in response to hypoxia. Cell. 2013;153:1327–39.
Majmundar AJ, Wong WJ, Simon MC. Hypoxia-inducible factors and the response to hypoxic stress. Mol Cell. 2013;40:294–309.
Zhang X, Le W. Pathological role of hypoxia in Alzheimer’s disease. Exp Neurol. 2010;223:299–303.
Sun X, He G, Qing H, Zhou W, Dobie F, Cai F, et al. Hypoxia facilitates Alzheimer’s disease pathogenesis by up-regulating BACE1 gene expression. Proc Natl Acad Sci U S A. 2006;103:18727–32.
Lana D, Ugolini F, Giovannini MG. An Overview on the Differential Interplay Among Neurons–Astrocytes–Microglia in CA1 and CA3 Hippocampus in Hypoxia/Ischemia. Front Cell Neurosci. 2020;14:585833.
Yasukochi Y, Shin S, Wakabayashi H, Maeda T. Transcriptomic changes in young Japanese males after exposure to acute hypobaric hypoxia. Front Genet. 2020;11:559074.
Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012;13:484–92.
Neri F, Rapelli S, Krepelova A, Incarnato D, Parlato C, Basile G, et al. Intragenic DNA methylation prevents spurious transcription initiation. Nature. 2017;543:72–77.
Weinberg DN, Papillon-Cavanagh S, Chen H, Yue Y, Chen X, Rajagopalan KN, et al. The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape. Nature. 2019;573:281–6.
Wu H, Coskun V, Tao J, Xie W, Ge W, Yoshikawa K, et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Nature. 2010;329:444–8.
Acknowledgements
This work was partly supported by the National Key Research and Development Program of China (Grant No. 2018YFC1314301), National Natural Science Foundation of China (Grant No. 82001797), Tianjin Applied Basic Research Diversified Investment Foundation (Grant No. 21JCYBJC01360), Tianjin Health Technology Project (Grant No. TJWJ2021QN002), Science&Technology Development Fund of Tianjin Education Commission for Higher Education (2019KJ195), National Key Research and Development Program of China (Grant No. 2017YFA0604401 and 2017YFA0504102), National Natural Science Foundation of China (Grant No. 82030053, 32070675 and 81801687), Natural Science Foundation of Tianjin City (Grant No. 19JCJQJC63600 for MJL and 18JCJQJC48200 for XW), Tianjin Key Medical Discipline (Specialty) Construction Project (Grant No. TJYXZDXK-001A). Further support was received by GS from the Horizon 2020 funded ERC Advanced Grant “STRATIFY” (Brain network-based stratification of reinforcement-related disorders) (695313), the National Institute of Health (NIH) (R01DA049238, A decentralized macro and micro gene-by-environment interaction analysis of substance use behavior and its brain biomarkers), the Human Brain Project (SGA3; 945539), and the Chinese National High-end Foreign Expert Recruitment Plan. Further support was provided by grants from the ANR (AAPG2019-GeBra), the Eranet Neuron (ANR-18-NEUR00002-01- ADORe). Healthy elderly controls and LOAD cases data collection and sharing for this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
Author information
Authors and Affiliations
Consortia
Contributions
JX, MJL, XW, and CY designed the study. JX, XX, YD, MJL, XW, and CY wrote the manuscript. JX, QL, XX and ZS analysed the data. All authors critically reviewed the manuscript. XS, NL, YH, XS, YH, WQ and SZ were the principal investigators. TB, HF, AG, PG, AH, RB, JM, EA, FN, TP, LP, SH, HW, PS and GS acquired the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, J., Xia, X., Li, Q. et al. A causal association of ANKRD37 with human hippocampal volume. Mol Psychiatry 27, 4432–4445 (2022). https://doi.org/10.1038/s41380-022-01800-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01800-7