Abstract
The molecular pathomechanisms of major depressive disorder (MDD) are still not completely understood. Here, we follow the hypothesis, that mitochondria dysfunction which is inevitably associated with bioenergetic disbalance is a risk factor that contributes to the susceptibility of an individual to develop MDD. Thus, we investigated molecular mechanisms related to mitochondrial function in induced neuronal progenitor cells (NPCs) which were reprogrammed from fibroblasts of eight MDD patients and eight non-depressed controls. We found significantly lower maximal respiration rates, altered cytosolic basal calcium levels, and smaller soma size in NPCs derived from MDD patients. These findings are partially consistent with our earlier observations in MDD patient-derived fibroblasts. Furthermore, we differentiated MDD and control NPCs into iPS-neurons and analyzed their passive biophysical and active electrophysiological properties to investigate whether neuronal function can be related to altered mitochondrial activity and bioenergetics. Interestingly, MDD patient-derived iPS-neurons showed significantly lower membrane capacitance, a less hyperpolarized membrane potential, increased Na+ current density and increased spontaneous electrical activity. Our findings indicate that functional differences evident in fibroblasts derived from MDD patients are partially present after reprogramming to induced-NPCs, could relate to altered function of iPS-neurons and thus might be associated with the aetiology of major depressive disorder.
Similar content being viewed by others
Introduction
Depressive syndrome is a debilitating and severe mental disorder, affecting about 5 million people per year in Germany and 350 million people worldwide (World Health Organization, 2016). The molecular mechanisms underlying the cause and progress of this complex disease are still not completely understood. Hypotheses argue for a combination of neurobiological factors which affect cellular function and neuronal communication, thereby increasing the risk for MDD, in conjunction with environmental (psychosocial) stress. The interaction of all factors has been shown to be associated with the onset and severity of depressive episodes [1]. Involved molecular mechanisms include genetically inherited and acquired neurobiological issues such as the dysregulation of monoaminergic, glutamatergic and GABAergic neurotransmission as well as reduced neuroplasticity and neurogenesis as a consequence of impaired BDNF signaling (reviewed in [2]). Furthermore, mitochondrial dysfunction associated with reduced bioenergetic capability is considered an important risk factor for MDD as well as other psychiatric disorders [3,4,5], by constraining proper cellular function and rendering the cells vulnerable to stress, especially during increased metabolic demand. Mitochondrial disorders that present with MDD as well as mitochondrial abnormalities and involved pathomechanisms in patients living with depression are reported in [6,7,8,9,10] and reviewed in [3, 5, 11, 12].
Especially brain neurons rely heavily on mitochondrial energy provision to maintain membrane excitability and operate neurotransmission, as well as to sustain plasticity [13]. However, MDD is increasingly viewed as a systemic disease with somatic manifestations outside the brain [10, 14]. Literature already reports mitochondria-related effects in fibroblasts [6, 10], muscle- [9], peripheral mononuclear blood cells [7], and platelets [8, 15] of depressed patients, suggesting that mitochondria-related pathomechanisms associated with MDD can be identified and studied in neuronal and also in peripheral non-neuronal cells.
To challenge the hypothesis that mitochondria dysfunction contributes to the aetiology of the disease, we aimed to identify and characterize molecular pathomechanisms related to mitochondrial function and bioenergetic imbalance in a human cellular model of major depression [16,17,18,19]. We used skin fibroblasts from eight MDD patients and eight non-depressed controls [6] to reprogram and differentiate these cell lines to neural progenitor cells (NPCs) and neurons (iPS-neurons), shifting our cellular model from non-neuronal to neuronal cells. The advent of induced pluripotent stem cell (iPSC) technology [20, 21] opens a new avenue for studying basic biological and pathological mechanisms [17, 22,23,24]. The generation of disease- and patient-specific iPSCs and their differentiation into specific cell types helps unravelling the molecular mechanisms, which underlie the etiology of complex diseases such as MDD [16, 17]. As a measure of mitochondrial function in MDD and non-depressed control NPCs, we assessed the performance of the oxidative phosphorylating system (OXPHOS) by analyzing the oxygen consumption rates (OCRs) in different respiratory states and the mitochondrial membrane potential (MMP) as an indicator for the bioenergetic state. Moreover, the cellular ATP content was analyzed as the primary energetic outcome of mitochondrial function. The cytosolic Ca2+ level provided important information about cellular Ca2+ homeostasis, since Ca2+ ions serve as important signalling molecules regulating most of mitochondrial and cellular functions, ranging from electron transport chain (ETC) to apoptosis. Interestingly, the reprogrammed NPCs presented functional alterations, which might be associated with bioenergetic dysfunction of mitochondria in MDD patient cells and were in line with our findings obtained in the founder fibroblasts [6].
Moreover, to characterize electrophysiological properties of neurons derived from MDD patients and to identify molecular phenotypes of mitochondrial impairment in these cells, we differentiated the NPCs into iPS-neurons. iPS-neurons were analyzed by means of whole-cell voltage- and current-clamp recordings. We found differences in the membrane capacitance, resting membrane potential, Na+ current density and spontaneous activity of MDD iPS-neurons when compared with cells derived from non-depressed controls. In summary, we identified alterations in bioenergetic parameters in NPCs, and found altered electrophysiological properties in iPS-neurons of patients suffering from depression, which might be related to reduced bioenergetics. Moreover, our findings indicate that functional differences evident in fibroblasts from MDD patients [6] are partially present after reprogramming and differentiation to induced-NPCs, thereby affecting functional properties of iPS-neurons. We discuss that the underlying mechanisms might be associated with the aetiology of major depressive disorder (MDD). Mitochondrial alterations or altered energy metabolism in iPSC-derived neural lineages has already been studied [25,26,27,28,29,30,31], but to the best of our knowledge, this is the first report addressing altered energy metabolism in iPSC-derived neural lineages in the context of major depression.
Methods and materials
Skin biopsies and primary human fibroblast cultivation
Skin biopsies were conducted by the Department of Dermatology, University Hospital of Regensburg, Regensburg, Germany. All participants gave written informed consent, and all study procedures were approved by the ethics committee of the University of Regensburg (ref: 13-101-0271). Human fibroblasts were obtained and cultivated as previously described [6]. Until use, cells were stored in a robotic storage system (SmartFreezer®, Angelantoni) in the Central Biobank Regensburg at the vapor phase of liquid nitrogen and standardized conditions. All cell lines (fibroblasts, iPSCs, NPCs) were routinely tested for mycoplasma contamination.
Generation of control and MDD patient iPSCs
iPSCs were generated using the episomal protocol described by [21]. Briefly, 5 × 105 fibroblasts of passage 4 or less were electroporated with 600 ng of each the episomal vectors pCXB-EBNA1, pCE-hsk, pCE-hUL, pCE-hOCT3/4 and pCE-mp53DD (Addgene plasmid #41857, # 41814, # 41855, # 41813, and # 41856, gift from Shinya Yamanaka) using the Amaxa Nucleofactor (Lonza). The cells were then cultured in TeSR-E7 medium for 3–4 weeks on Matrigel-coated dishes (Corning) until the colonies appeared. iPSC colonies were manually picked and cultured using mTeSR1 medium on Matrigel (StemCell Technologies). The generated iPSC cultures were analyzed for pluripotency by the PluriTest® method, a patented bioinformatics assay for the quality assessment of iPSCs, which was adjusted for next-generation sequencing (NGS) data as published by [32].
iPSC differentiation to NPCs and neuron differentiation
NPCs were derived from iPSCs using a protocol described in [33]. Briefly, NPCs were expanded in expansion medium containing 50% Neurobasal medium, 50% Advanced DMEM/F12, and neural induction supplement, (all from Life Technologies), and subsequently dissociated with Accutase (Life Technologies) and maintained on Geltrex-coated dishes at a density of 1 × 105 cells per cm2. To obtain mature neurons, NPCs from passages 5 to 10 were plated onto poly-L-ornithine/laminin (Sigma-Aldrich) coated glass coverslips and differentiated in Neurobasal medium supplemented for 21 days with 1% B27, 0.5% GlutaMax, 0.5% non-essential amino acids, 0.5% Culture One (Thermo Fisher Scientific), 200 nM ascorbic acid (Carl Roth), 20 ng/ml BDNF, 20 ng/ml GDNF (both PeproTech), 1 mM dibutyryl-cAMP (Stemcell), 4 μg/ml laminin (Sigma) and 50 U/ml Penicillin, 50 μg/ml Streptomycin (Thermo Fisher Scientific).
Immunofluorescence
Human fibroblasts, iPSCs, NPCs and induced neurons grown on glass coverslips (Menzel Gläser), were fixed in 4% paraformaldehyde (Carl Roth) and blocked by 10% normal goat serum in PBST (PBS + 0.5% Triton X-100) for 20 min. Cells were then incubated with primary antibodies in PBST (PBS + 0.1% Triton X-100, 2% normal goat serum) overnight at 4 °C, following 1 h incubation at RT with corresponding Alexa 488/Cy3/Cy5 coupled secondary antibodies (1:1000, Thermo Fisher Scientific). Cell nuclei were stained with DAPI. Fibroblasts were labelled with anti-SMA (mouse; 1:250, ab7817, Abcam), whereas iPSCs were immunostained using anti-NANOG (rabbit; 1:250, ab21624, Abcam) antibodies. NPCs were labeled with anti-PAX6 (mouse; 1:10; deposited to the DSHB by Kawakami, A.) and anti-SOX2, ab97959 antibodies (rabbit; 1:1000, ab97959, Abcam). For immunostaining of the induced neurons following antibodies were used: anti-Tuj 1 (mouse; 1:2000, G7121, Promega), anti-MAP2 (chicken; 1:5000, ab5392, Abcam), anti-synaptophysin (rabbit; 1:500, ab52636, Abcam), anti-VGLUT1 (rabbit; 1:1000, ab180188, Abcam), and anti-NEUN (rabbit; 1:500, ab177487, Abcam). Coverslips were fixed onto glass object slides (Menzel Gläser) using Fluorescence Mounting Medium (Dako) and images were acquired using a Zeiss Observer Z.1 microscope (Zeiss). For quantitative analysis of immunofluorescence labelling, all nuclei (DAPI) of NPCs of three visual fields (inspected with a 40X objective) were counted and the % of cells, that were/were not stained/double stained with PAX6 and SOX2 were calculated.
qPCR analysis of expression of neuronal markers
Total RNA was isolated from iPSCs and from induced neurons after 21 DIV using RNA Plus Kit (Macherey-Nagel) according to the manufacturer’s instructions. First strand cDNA synthesis from 1 µg of total RNA was performed with QuantiTect Reverse Transcription Kit (Qiagen). Quantitative RT-PCR experiments were performed with Rotor-Gene-Q machine (Qiagen) using the 1× Takyon SYBR Master Mix (Eurogentec) and intron-spanning primers specific for different neural subtypes, listed in Supplementary Table 1. Measurements were completed in triplicate and results were analyzed with a Rotor-Gene-Q software version 2.3 (Qiagen). Relative expression levels of each target gene were calculated using the comparative Ct method and HPRT1 as a housekeeping gene for normalization. Gene expression of various neural subtypes was represented as heat map without statistical analysis.
Analysis of mitochondrial respiration
Analysis of mitochondrial respiration was performed using Seahorse XFp Flux analyzer with a Seahorse XFp Mito Stress Test Kit (Agilent Technologies) according to manufacturer´s recommendations. Briefly, 8 × 104 NPCs were grown in XFp eight-well miniplates at 37 °C, humidified air and 5% CO2. Oxygen consumption rate (OCR) was measured with sequential injection of 1 µM Oligomycin, 1 µM FCCP, and each 0.5 µM Rotenone/Antimycin A (Biomol). After the measurements, nuclei were stained with DAPI and counted in order to to normalize respiration data.
Luminescent assay for ATP content
1 × 106 cells were pelleted in a 1.5 ml Eppendorf cup and stored at −20 °C. ATP content was measured by CellTiter-Glo®Cell Viability Kit (Promega). CellTiter-Glo®Reagent containing CellTiter®Substrate and CellTiter®Buffer was thawed on ice. The ATP standard curve ranged between concentrations of 10 µM, 1 µM, 100 nM, 10 nM and 1 nM ATP in 1x PBS. Cell pellets were resuspended in 500 µl PBS, heated at 100 °C for 2 min and kept on ice afterwards. Triplicates of 50 µl per sample and each standard were applied to a black 96-well-plate together with 50 µl of CellTiter-Glo®Reagent. The absorption was measured with the VarioScan at an integration time of 1 s. The RLU generated by the SkanIT Software was used to calculate the actual ATP concentrations using the ATP standard curve. ATP concentrations were normalized to µg/mL protein using a BCA assay (Thermo Fisher Scientific).
Mitochondrial membrane potential (JC-1)
Mitochondrial membrane potential was measured in the NPCs as described in [6, 34]. Briefly, 2.8 × 106 NPCs were grown overnight on glass coverslips (diameter 25 mm; Menzel Gläser) and were loaded with 1 µM of JC-1/Pluronic in OptiMEM (Thermo Fisher Scientific) for 30 min at 37 °C (humidified air and 5% CO2). For every subject, we manually selected and circled 20 cells (as regions of interest, ROIs) in each of 10 visual fields at three different inspection days using FIJI/ImageJ [34, 35], adding up to a total of n = 600 cells. Mitochondrial membrane potential was compared as ratio of red over green fluorescence intensity in the ROIs. Code availability: The macro used for background subtraction and analysis will be provided on request.
Imaging of cytosolic Ca2+ (Fura-2/AM)
Cytosolic Ca2+ levels in NPCs were measured as previously described in [36]. Briefly, 2.8 × 106 NPCs were grown on glass coverslips in six-well plate and loaded with 2 µM Fura-2/AM and Pluronic F127 in OptiMEM (Thermo Fisher Scientific) for 30 min at 37 °C. Cytosolic Ca2+ level in NPCs was assessed by measuring the fluorescence at 510 nm after excitation at 340 or 380 nm using Observer Z.1 inverted microscope (Zeiss). Cells were selected as ROIs in the visual field using the FIJI/ImageJ [35] as described for the procedure used to assess the MMP. Code availability: The macro used for background subtraction and analysis will be provided on request.
Electrophysiology
Whole-cell patch-clamp recordings were performed on induced neurons during their 4th week of differentiation. A total of 333 induced neurons (162 Control-Neurons and 171 MDD-Neurons) were recorded and analyzed. The extracellular solution was composed of 140 mM NaCl, 5 mM KCl, 2 mM CaCl2, 1 mM MgCl2, 10 mM HEPES, pH 7.3. Micropipettes were made of borosilicate glass (Science Products) by means of a horizontal pipette puller (Zeitz Instruments) and fire-polished to obtain a series resistance of 3–5 MΩ. Micropipettes were filled with intracellular solution (140 mM KCl, 1 mM MgCl2, 0.1 mM CaCl2, 5 mM EGTA, 10 mM HEPES). Recordings were made using a Heka Electronic EPC-10 amplifier (HEKA Electronic). The liquid-liquid junction potential was calculated to be 4 mV (LJP calculator of the pClamp software suite, Axon Instruments) and was corrected in the amplifier settings of the Patchmaster software (HEKA Electronic). The series resistance was assessed but not compensated. The resting membrane potential and capacitance were recorded directly after reaching the whole-cell configuration. For voltage-clamp recordings, membrane potential was held at −80 mV and depolarized in steps of 10 mV to evoke voltage-activated Na+- and K+-channels. In current-clamp mode, manually adjusted currents in the range of −2 to −240 pA (dependent on the resting membrane potential and the series resistance), were injected to hyperpolarize the membrane potential of neurons to about −80 mV. The cells were then depolarized by 20 steps with a 2–20 pA increment to reach the threshold and evoke action potentials (APs). Spontaneous APs were recorded at a holding potential of about −45 to −50 mV. All patch-clamp recordings were carried out at room temperature. Data were analyzed using Patchmaster v2 × 90.03 (HEKA Electronic). Cells with resting membrane potential of 0 mV and above were excluded from the analysis. The experimenter was blind to the group parameter (MDD or control) during the experiment and for the analysis of the electrophysiological data to avoid any biased treatment of the data. Data were then unblinded for statistical analysis.
Statistical analysis
Graphical depiction and statistical analysis were conducted with Graph Pad Prism 8.0.2 (GraphPad Software). To exclude any bias, cell lines were renamed and anonymized before starting the differentiation protocol and experiments. For all experiments, except patch-clamp recordings, the means of two to three technical replicates were calculated and two to three biological replicates were averaged. Measurements were conducted pairwise allowing direct comparison of MDD vs. controls. Statistical outliers were detected and eliminated using ROUT-Method. Normal distribution was implied because of high sample size. Variance homogeneity was evaluated before performing statistical analysis, cases of variance inhomogeneity are mentioned, and statistical analysis was therefore adjusted adequately. Results of spontaneous activity were compared by Fisher’s exact test. The resting membrane potential showed no variance homogeneity, thus the Welch-test was applied. All other results showed variance homogeneity and were compared using unpaired Student’s t Test. Results are presented as mean ± SEM, unless otherwise stated. p value limit for statistical significance is set to ≤0.05.
Unlike the variables subjected to standard averaging and mean comparison procedure, ratio of the fluorescence signals yielded a complex data structure with multiple cells nested in visual fields. Therefore, multilevel regression analyses were conducted using the statistical software R (Team RC, 2021), to account for the nested data structure. Alterations in the variables between pre- and posttests were analyzed using linear mixed regression models using the package “lme4” [37]. Type-I-Error probabilities for the regression coefficients were calculated using the package “lmerTest” [38]. The model accounted for random intercepts of patients and images, which thus defined the nested structure of the data. The fixed factor was defined by the group (i.e., patient vs. control). Tests were run two-tailed. A detailed description of the model is given by [39].
Results
Based on our findings of altered bioenergetic properties and mitochondrial function in primary dermal fibroblasts of patients suffering from MDD [6], we were interested in the identification and characterization of MDD-associated pathomechanisms in neural cells. Thus, we used primary skin fibroblasts characterized in our earlier study [6] (information on study participants is given in the Supplementary Table 2) and reprogrammed them to iPSCs by transient episomal transduction according to the Yamanaka protocol (Fig. 1A) [20, 21]. Pluripotency of iPSCs was validated by PluriTest analysis, a bioinformatics assay for the quality assessment of iPSCs by transcription profiling based on NGS data [32] obtained from the respective iPSC clones. The PluriTest analysis revealed that 13 of the iPSC clones used in this study could be considered pluripotent. Three others were very close to the empirical threshold (pluripotency >1424; novelty <2.5) (Fig. 1B and Table 1). The iPSC colonies were differentiated to early NPCs within 7 days after culture medium was changed from mTeSR to neural induction medium. Further maturation of NPCs was allowed for 5 passages before cells were stained for neural progenitor markers SOX2 and PAX6 [40] (Fig. 1C). Table 2 presents the percentage of cells positively labelled with the respective antibody and indicates that the majority of the cells co-expresses both NPC markers and can be regarded as neural progenitors. The three cell lines that failed the PluriTest analysis, showed prominent expression of the neuronal progenitor markers PAX6 and SOX2 as well, thus we decided to not exclude these cell lines from our study. To seek for functional differences between NPCs from MDD and non-depressed control subjects, we analyzed bioenergetic properties and mitochondrial function.
A Patient-derived dermal fibroblasts were reprogrammed into iPSCs and subsequently differentiated into neurons. Representative images of fibroblasts, iPSCs, NPCs and iPS-neurons expressing corresponding cell specific markers. Scale bar indicates 20 µm. B Characterization of 16 pluripotent cell lines (black dots) from this study using PluriTest. Kernel density estimations for iPSCs (yellow to red), and non-pluripotent stem cells (shades of blue/gray) are shown. Black dashed lines indicate the empirical thresholds for novelty and pluripotency. High quality cells should be located in the upper left quadrant, with a pluripotency score above the threshold and a novelty score below the threshold. It can be seen that the empirical thresholds detect high quality pluripotent cells from this study, but that this population still contains some cells with reduced pluripotency. C Representative images of NPCs differentiated from control and MDD patients show that majority of the cells are expressing typical neural progenitor cell markers PAX6 and SOX2. Scale bar indicates 20 µm.
Mitochondrial membrane potential
The cationic and lipophilic fluorescent dye JC-1 accumulates in the mitochondrial membrane to an extent, which is dependent on the strength of the electric field. In negatively charged (i.e., highly energized) mitochondria, JC-1 molecules accumulate and form red fluorescing aggregates, while the fluorescence changes to green, when the dye molecules disaggregate into monomers in response to dissipation of the transmembrane potential. The ratio of the fluorescence signals emitted by the two states of JC-1 is then analyzed as a measure of the MMP. We found that NPCs from MDD patients presented a lower, however a non-significantly different red/green ratio when compared to the control NPCs from non-depressed subjects. This can be judged as an indication to a more depolarized MMP in patient cells (MDD 2.171 ± 0.16 vs. Cntrl 2.417 ± 0.114, mean ± SEM, p = 0.148, t = 21.291, df = 14.058, linear mixed regression) (Fig. 2A).
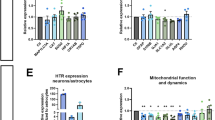
A Mitochondrial membrane potential of NPCs. Red/green (JC-1 aggregate/monomer) ratios of control (CNTRL) and MDD NPCs. Dot plots show mean red/green ratios ± SEM, control n = 8, MDD n = 8. Difference is not significant. B, C Oxygen consumption rates (OCR) of control and MDD NPCs. B Representative OCR measurement during the Mito Stress Test for MDD and control NPCs. C Maximal respiration is significantly different as analyzed by two-tailed t-test (indicated by *). Considering our hypothesis of reduced oxygen consumption in MDD cells, a one-tailed t-test indicate significant difference of all parameters except spare-respiratory capacity (indicated by #). Dot plots show normalized mean OCR values ± SEM, Control n = 8, MDD n = 8. D Basal Ca2+ calcium levels were significantly altered in MDD NPCs. Shown are the Fura-2 340 nm/380 nm fluorescence ratios of non-depressed control and MDD NPC lines. Dots show mean ratios (340 nm/380 nm, ratios ± SEM). Control n = 8, MDD n = 8. E ATP levels were not significantly different between MDD and CNTRL NPCs. Dots show normalized mean RLU values ± SEM. F MDD NPCs are significantly smaller than CNTRLs. Size is analyzed by assessing area (pixels) of Fura-2-loaded cells. Significant differences between MDD and non-depressive controls are indicated with *.
Mitochondrial oxidative phosphorylation system (OXPHOS)—Respirometry
The consumption of molecular oxygen by accepting the electrons delivered by the ETC at complex IV can be used as a readout for the function and performance of the OXPHOS. Figure 2B depicts a representative Seahorse measurement using the Mito Stress Kit reporting the OCRs (OCR) in various respiratory states. Although all assessed parameters were lower in patients suffering from depression, we found a significant difference between the groups in the maximal respiration only ([pmol/min/1000 cells] MDD 9.74 ± 0.85 vs. Cntrl 12.19 ± 1.41, p = 0.038, t = 2,551, df = 7, paired t-test, two-tailed) (Fig. 2C). The basal respiration, as well as ATP-related oxygen consumption, non-mitochondrial OCR and proton leak were not significantly lower in MDD NPCs compared to healthy controls. However, based on our earlier results on the reduced respiration of fibroblasts from patients suffering from depression, we would expect/assume a reduced respiration in NPCs as well. Following this hypothesis, we can justify to use a one-tailed test instead, leading to significant differences between the MDD and control groups in the other respiratory parameters (Table 3).
Cytosolic Ca2+ levels, cellular ATP-content, and cell size
As an additional measure for metabolic/bioenergetic activity and mitochondrial function, we compared the cytosolic Ca2+ levels in MDD and control NPCs by Fura-2 Ca2+ imaging via linear multilevel regression models, accounting for the nesting of multiple measurements within cell lines. We detected a small but significant difference in cytosolic Ca2+ levels between MDD and control NPCs (Fura-2 emission ratio: MDD 0.574 ± 0.008, Cntrl 0.558 ± 0.005, mean ± SEM, t = 2.247, df = 13.889, p = 0.041), consistent with the important function of Ca2+ as a signaling molecule and mediator in mitochondrial and cellular physiology (Fig. 2D). Although we found reduced respiration and altered Ca2+ levels in MDD NPCs, the cellular ATP content was not different between the two groups ([nM/µg/ml] Cntrl 19.15 ± 4.11, MDD 18.62 ± 4.22, mean ± SEM, p = 0.69, paired t-test, two-tailed) (Fig. 2E). This observation points to a compensatory mechanism in the NPCs, which compensates for a possibly reduced energy production according to a reduced OXPHOS. Although we could not detect a deficit in cellular ATP levels in the MDD NPCs, this group presented a significantly smaller cell size, as assessed by pixel counting of the fluorescent area during Fura-2 imaging ([pixel] MDD 168.1 ± 1.21, Cntrl 191.3 ± 1.29, mean ± SEM, p = 0.0001, paired t-test, two-tailed) (Fig. 2F).
Functional characteristics of iPS-neurons
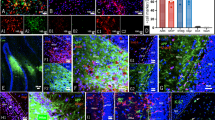
Next, we were interested whether differential functional phenotypes can also be associated with iPS-neurons from MDD or control subjects. To this end, we differentiated NPC lines from eight MDD patients and matching controls to iPS-neurons. iPS-neurons showed clear bipolar or multipolar neuronal morphology (Fig. 3A) and expressed the typical neuronal markers class III beta-tubulin, MAP2 and NEUN (Fig. 3B). Moreover, they were immunopositive for the specific synaptic proteins VGLUT1 and SYP at DIV 21 (Fig. 3C, D). A qPCR analysis of neuronal marker expression in 5 pairs of iPS-neuron lines indicated the expression of MAP2, GAD1, and GRIN1, whereas SLC6A4, TH, and CHAT showed only low expression (Fig. 3E), indicating the differentiation of NPCs to functional neurons capable of glutamatergic and GABAergic signaling. The Ct values for NPCs and iPS-neurons are presented in the Supplementary Table 3. To functionally characterize and compare electrophysiological properties of iPS-neurons, we performed whole-cell voltage-clamp and current-clamp recordings of iPS-neurons during their 4th week of differentiation and analyzed their passive biophysical as well as their active neuronal properties. Figure 4A–D shows exemplary whole-cell recordings of a typical iPS-neuron. In total, we recorded 333 neurons derived and differentiated from the eight MDD patients and eight non-depressed controls. We found that the membrane capacitance of iPS-neurons from MDD patients, which was compensated as C-slow by the inbuilt capacitance compensation algorithm of the EPC 10 amplifier, was significantly smaller compared with non-depressed control neurons ([pF] MDD 15.75 ± 0.76 vs. Cntrl 18.12 ± 0.9, p = 0.044, t = 2.022, df = 328, t-test, unpaired) (Fig. 5A). Since C-slow is associated with the size of the membrane surface, the capacitance can be regarded as a measure for cell size. Interestingly, our data indicate that iPS-neurons from MDD patients are smaller than from non-depressed controls. This finding is consistent with the reduced cell size in MDD NPCs (Fig. 2F). The series resistance was not different between the two groups ([MΩ] MDD 23.09 ± 1.035 vs. Cntrl 24.79 ± 1.11, p = 0.26, t = 1.121, df = 318, t-test, unpaired) (Fig. 5B). Interestingly, we found that the resting membrane potential measured directly after establishing the whole-cell configuration was different between the groups: MDD iPS-neurons had a significantly lower (i.e., less hyperpolarized) resting membrane potential than iPS-neurons derived from the control subjects ([mV] MDD –23.68 ± 0.84 vs. Cntrl −26.68 ± 1.02, p = 0.024, t = 2.275, df = 312.3, t-test, unpaired, with Welch’s correction (Fig. 5C). In current-clamp mode, the neurons were able to fire APs when the membrane potential was adjusted to approximately −80 mV by current injection and then depolarized by modulating the injected current. The mean current injected to adjust the membrane potential to −80 mV was not different between MDD and non-depressed control iPS-neurons ([pA] MDD −84.08 ± 5.214 vs. Cntrl –81.14 ± 5.29, p = 0.69, t = 0.40, df = 299, t-test, unpaired) (Fig. 5D). In voltage-clamp experiments, voltage-activated sodium inward currents and potassium outward currents were induced by stepwise depolarization of the membrane potential starting at a Vhold of −80 mV. To consider the between-group difference in membrane capacitance as an indicator for different cell size, we analyzed the current density (pA/pF) by relating the Na+ and K+ current amplitudes to the measured membrane capacitance and found a higher INa+ current density in MDD iPS-neurons ([pA/pF] MDD −47.4 ± 2.9 vs. Cntrl −35.2 ± 1.9, p = 0.0006, t = 3.465, df = 291, t-test, unpaired) (Fig. 5E). The amplitudes of the potassium (IK+) outward currents at +20 mV were not different between MDD and non-depressed iPS-neurons ([pA/pF] MDD −40.8 ± 2.2 vs. Cntrl −41.9 ± 2.3, p = 0.7151, t = 0.3653, df = 307, t-test, unpaired) (Fig. 5G). Assessing the amplitudes of INa+ at 0 mV to minimize effects of intrinsic space clamp problems in large neurons, we again detected significantly larger INa+ current density in MDD iPS-neurons ([pA/pF] MDD −35.15 ± 2.3 vs. Cntrl −27.09 ± 1.4, p = 0.0001, t = 11.71, df = 135, t-test, unpaired) (Fig. 5F). In addition, we characterized spontaneous electrical activity of cultured iPS-neurons by current-clamp recording and adjusted the resting membrane potential to about −50 mV by direct current injection. Under this condition, we found that a significantly higher proportion of MDD-derived iPS-neurons (35/163 = 21.74%), were spontaneously active and generated APs when compared with cells reprogrammed from non-depressed controls (18/162 = 11.11%) indicating an increased activity of MDD neurons (p = 0.016, Fisher’s exact test) (Fig. 5H).
A Neurons differentiated from NPCs show clear bipolar or -multipolar neuronal morphology. B Representative images of iPS-neurons expressing typical neuronal markers such as class III beta-tubulin (TUJ1), microtubule associated protein 2 (MAP2), and neuronal nuclear protein (NEUN). Scale bar indicates 20 µm. C, D Formation of glutamatergic synapses in iPS-neurons after 3–4 weeks in the culture. VGLUT1 and synaptophysin (SYP) immunostaining visualizes synaptic vesicle clusters (green, white arrows). Scale bar indicates 10 µm. E Heat map representation of relative gene expression by cycle threshold of RT-PCR reactions. Changes of expression of induced neurons after 21 days of neural differentiation relative to corresponding iPSCs are represented as downregulation (blue), no change (white) or upregulation (red).
A Exemplary differential interference contrast photomicrograph (400×) of a cultured iPSC-derived neuron (DIV 21) from a MDD patient. A patch-pipette is attached to the soma of the cell. B Whole-cell recording in the current-clamp mode allows recording of the membrane potential. Injection of a supra-threshold depolarizing current induced action potentials. C Whole-cell recording in the voltage-clamp mode showed voltage-activated transient Na+-inward (INa+) and delayed K+-outward currents (IK+) following a pulse protocol stepping from Vhold = −80 mV to more depolarized potentials (−70 to +40 mV). An exemplary current/voltage relationship for voltage-activated Na+ (INa+, dark gray filled circles) and K+ currents (IK+, light gray filled circles) is presented in (D).
A Analysis of the membrane capacitance as a measure of the size of the recorded cell revealed a significantly smaller mean value in cells derived from MDD patients. B Series resistance of iPS-neurons measured in whole-cell voltage-clamp recording. No between-group difference could be detected. C The resting membrane potential measured in the current-clamp mode was less hyperpolarized in MDD iPS-derived neurons. D The injected current (in current-clamp experiments) to drive the membrane potential to −80 mV is not different between MDD and CNTRL iPS-neurons. E Maximum INa+ current density and current density at 0 mV (F) is significantly higher in MDD iPS-neurons, whereas IK+ current density at +20 mV shows no between-groups difference (G). H The fraction of spontaneously active iPS-neurons, which fire action potentials at the adjusted membrane potential of −50 mV in a time frame of 30 s, was significantly higher in the MDD iPS-neuron group.
Discussion
In order to identify and characterize molecular pathomechanisms associated with major depression, we established a human cellular model, which is based on dermal fibroblasts derived from skin biopsies of MDD patients and non-depressed controls [6] and then reprogrammed and differentiated these fibroblasts to iPSCs, NPCs and iPSC-derived neurons.
A reduced value in the pluripotency parameter might be due to contaminating re-differentiated cells (fibroblasts) in the iPSC samples used for the Pluritest. Considering the high proportion of SOX2 and PAX6 positive cells, we decided not to exclude these iPSC lines from our study.
Analysis of the mitochondrial function of reprogrammed human NPCs revealed a reduced function of the OXPHOS in MDD NPCs. These findings indicate a significant mitochondrial alteration, which was already evident in the founder cells (i.e., the fibroblasts) of patients suffering from depression [6]. Other functional parameters of the bioenergetic status of the NPCs, such as MMP or cellular ATP content were not found to be significantly different between the MDD and the control group. A partially altered functional phenotype might in part be explained by an altered metabolic dependence of NPCs. Whereas fibroblasts rely highly on OXPHOS for ATP production, NPCs gain a significant part of their energy via the glycolytic pathway [26, 41]. Thus, potential deficits in their mitochondrial OXPHOS and ETC activity might be compensated by the predominant generation of energy through glycolytic metabolism.
The MDD patients were asked to participate in our study and to donate a skin biopsy at the end of their inpatient stay [6]. At this time, the patients received antidepressant medication and were nearly in remission. Importantly, after several cell divisions of fibroblasts in vitro, the confounding variability in these samples on the basis of the subjects’ medication use, should be eliminated [10]. Thus, the alterations in mitochondria function we have found in our study might be associated rather with a trait than with the state of depression. The etiology of depression is associated with genetic (polygenic) [42,43,44], as well as with various environmental risk factors such as stressful life events directly influencing physiological processes and mental health of an individual, immediately and in later life. These threats are suggested to affect gene-environment interactions and epigenetics by altering chromatin thereby leading to altered molecular patterns observed in depression [45,46,47,48,49]. We observed mitochondrial alterations in fibroblasts of depressed patients [6] but also in iPSC-derived NPCs reprogrammed from these fibroblasts. So far, our observations can not answer the question, whether this functional phenotype relates to a genetic or epigenetic basis. Although much of the epigenetic memory is erased during the reprogramming process, iPSCs retain a transcriptional memory of the original cells upon reprogramming [50,51,52]. However, analysis of potential risk variants in the genome of our patient’s cells and the investigation of molecular patterns (e.g., DNA methylation and microRNAs) is an important next step in our search for molecular pathomechanisms of depression.
Today, depression is increasingly viewed as a systemic disease with somatic manifestations outside the brain. Following this hypothesis, we have detected mitochondrial alterations in fibroblasts [6] and in reprogrammed NPCs of depressed patients. In order to search for neuron-specific disease-related phenotypes and to deepen our understanding of the pathomechanisms involved in depression, we differentiated the NPCs of depressed and non-depressed individuals to iPS-neurons and performed electrophysiological characterization of the biophysical properties by means of whole-cell patch-clamp recording. We recorded from the iPS-neurons after 21 days of differentiation and found that the resting membrane potential was significantly less negative in iPS-neurons derived from MDD patients compared to non-depressed controls. Moreover, consistent with a significantly reduced soma size of NPCs, MDD neurons showed a smaller membrane capacitance, which is indicative for a smaller size or altered geometry of the neuron [53,54,55]. Intriguingly, iPS-neurons from depressed patients showed a higher Na+ current density and an increased fraction of spontaneously active cells in vitro. In general, a lower resting membrane potential as well as a smaller cell size could be a result of reduced energy availability in consequence of mitochondrial dysfunction [56], or due to a delayed neural development, since a hyperpolarized resting membrane potential as well as cell size is associated with neuronal maturation [57, 58]. The increased Na+ current density and network activity analyzed as the fraction of spontaneously active neurons when recorded in current-clamp mode is an intriguing observation, which needs deeper investigation. Altered neurite growth and morphology of iPS-neurons derived from depressed patients which were resistant to serotonin reuptake inhibitors have recently been demonstrated and were associated with a reduced expression of the Protocadherin alpha genes in these patients [16]. How this altered activity in iPS-neurons in vitro relates to interregional network activity and functional connectivity dynamics in vivo [59, 60], is not known, but an interesting and important objective for future investigations.
Limitations
Although we paired MDD patients with non-depressed control subjects of the same sex and a similar age, the study is limited by a high interpersonal variability based on the ages of the subjects within the group, differing lifestyles (e.g., sportive activity) and genetic as well as physiological differences. Moreover, the mental health status of a non-depressed control is based on the self-reported absence of any history of depression in his/her former life. We can not control for a putative load of genetic or environmental risk factors which might have accumulated in the controls. In this line of argumentation, we need to analyze nuclear and mitochondrial DNA for genetic risk factors and further elaborate how much of possible epigenetic patterns could be transmitted from fibroblasts to NPCs and iPS-neurons.
We noticed that the neurons showed quite depolarized RMPs compared to other reports [57]. Since such a low RMP renders voltage-gated sodium channels mostly steady-state inactivated, we adjusted the membrane potential to about −50 mV by current injection to provide a permissive condition/environment in which AP firing may occur. A depolarized (low) RMP points to a possibly immature developmental state of the induced neurons [57]. In general, neuronal development and electrophysiological properties depend also on the differentiation protocol and substrate (feeder layer, presence of glial cells). It is difficult to decide at which point in time neurons grown in vitro can be considered fully mature. Although we found that after 21 days of differentiation, the cells showed the typical almond shaped morphology with an extended meshwork of neurites, generated typical APs with overshoots and showed expression of neuronal and synaptic markers, we will use older cultures for future investigations to increase the maturity of our cellular model. However, we observed functional differences between MDD and control neurons after 21 days post differentiation indicating a disease-related phenotype at this rather early state, although depression is not considered a classical neurodevelopmental disorder.
The direct assessment of the energy metabolism and respirometry of iPS-neurons was not feasible in the present study because of the limited growth area and geometry of the XFp Seahorse culture plates, which did not support uniform distribution of iPS-neurons. An adapted experimental setting will help to overcome this problem in future and already ongoing studies to address the important question of bioenergetics in iPS-neurons derived from patients suffering from MDD.
In addition, the question of the effect of chronic exposure of patient-derived cells to stress-induced hormones [24] and their potentially diminished cellular resistance will be investigated in detail in the future by our group.
Conclusions
We suggest that NPCs and neurons derived from iPSCs of MDD patients can be used to identify and characterize molecular and cellular pathomechanisms of the disease. Our findings indicate that disease-specific alterations observed in fibroblasts are also present in induced-NPCs and might affect the physiology of differentiated iPS-neurons after reprogramming and differentiation. This approach may lead to new insights into the molecular biology and pathophysiology of depression as well as to the discovery of new drug targets and successful therapies.
References
Klengel T, Binder EB. Gene-environment interactions in major depressive disorder. Can J Psychiatry. 2013;58:76–83.
Hasler G. Pathophysiology of depression: do we have any solid evidence of interest to clinicians? World Psychiatry. 2010;9:155–61.
Manji H, Kato T, Di Prospero NA, Ness S, Beal MF, Krams M, et al. Impaired mitochondrial function in psychiatric disorders. Nat Rev Neurosci. 2012;13:293–307.
Gardner A, Boles RG. Beyond the serotonin hypothesis: mitochondria, inflammation and neurodegeneration in major depression and affective spectrum disorders. Prog Neuropsychopharmacol Biol Psychiatry. 2011;35:730–43.
Klinedinst NJ, Regenold WT. A mitochondrial bioenergetic basis of depression. J Bioenerg Biomembr. 2015;47:155–71.
Kuffner K, Triebelhorn J, Meindl K, Benner C, Manook A, Sudria-Lopez D, et al. Major Depressive Disorder is Associated with Impaired Mitochondrial Function in Skin Fibroblasts. Cells. 2020;9:884.
Karabatsiakis A, Bock C, Salinas-Manrique J, Kolassa S, Calzia E, Dietrich DE, et al. Mitochondrial respiration in peripheral blood mononuclear cells correlates with depressive subsymptoms and severity of major depression. Transl Psychiatry. 2014;4:e397.
Hroudova J, Fisar Z, Kitzlerova E, Zverova M, Raboch J. Mitochondrial respiration in blood platelets of depressive patients. Mitochondrion. 2013;13:795–800.
Gardner A, Johansson A, Wibom R, Nennesmo I, von Dobeln U, Hagenfeldt L, et al. Alterations of mitochondrial function and correlations with personality traits in selected major depressive disorder patients. J Affect Disord. 2003;76:55–68.
Garbett KA, Vereczkei A, Kalman S, Wang L, Korade Z, Shelton RC, et al. Fibroblasts from patients with major depressive disorder show distinct transcriptional response to metabolic stressors. Transl Psychiatry. 2015;5:e523.
Anglin RE, Garside SL, Tarnopolsky MA, Mazurek MF, Rosebush PI. The psychiatric manifestations of mitochondrial disorders: a case and review of the literature. J Clin Psychiatry. 2012;73:506–12.
Allen J, Romay-Tallon R, Brymer KJ, Caruncho HJ, Kalynchuk LE. Mitochondria and Mood: Mitochondrial Dysfunction as a Key Player in the Manifestation of Depression. Front Neurosci. 2018;12:386.
Picard M, McEwen BS. Mitochondria impact brain function and cognition. Proc Natl Acad Sci USA. 2014;111:7–8.
Verhoeven JE, Revesz D, Wolkowitz OM, Penninx BW. Cellular aging in depression: Permanent imprint or reversible process?: An overview of the current evidence, mechanistic pathways, and targets for interventions. Bioessays. 2014;36:968–78.
Sjovall F, Ehinger JK, Marelsson SE, Morota S, Frostner EA, Uchino H, et al. Mitochondrial respiration in human viable platelets-methodology and influence of gender, age and storage. Mitochondrion. 2013;13:7–14.
Vadodaria KC, Ji Y, Skime M, Paquola AC, Nelson T, Hall-Flavin D, et al. Altered serotonergic circuitry in SSRI-resistant major depressive disorder patient-derived neurons. Mol Psychiatry. 2019;24:808–18.
Soliman MA, Aboharb F, Zeltner N, Studer L. Pluripotent stem cells in neuropsychiatric disorders. Mol Psychiatry. 2017;22:1241–9.
Vadodaria KC, Mertens J, Paquola A, Bardy C, Li X, Jappelli R, et al. Generation of functional human serotonergic neurons from fibroblasts. Mol Psychiatry. 2016;21:49–61.
Lu J, Zhong X, Liu H, Hao L, Huang CT, Sherafat MA, et al. Generation of serotonin neurons from human pluripotent stem cells. Nat Biotechnol. 2016;34:89–94.
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–72.
Okita K, Matsumura Y, Sato Y, Okada A, Morizane A, Okamoto S, et al. A more efficient method to generate integration-free human iPS cells. Nat Methods. 2011;8:409–12.
Inoue H, Nagata N, Kurokawa H, Yamanaka S. iPS cells: a game changer for future medicine. EMBO J. 2014;33:409–17.
Takahashi K, Yamanaka S. Induced pluripotent stem cells in medicine and biology. Development. 2013;140:2457–61.
Heard KJ, Shokhirev MN, Becronis C, Fredlender C, Zahid N, Le AT, et al. Chronic cortisol differentially impacts stem cell-derived astrocytes from major depressive disorder patients. Transl Psychiatry. 2021;11:608.
Consortium HDi. Bioenergetic deficits in Huntington’s disease iPSC-derived neural cells and rescue with glycolytic metabolites. Hum Mol Genet. 2020;29:1757–71.
Lorenz C, Lesimple P, Bukowiecki R, Zink A, Inak G, Mlody B, et al. Human iPSC-Derived Neural Progenitors Are an Effective Drug Discovery Model for Neurological mtDNA Disorders. Cell Stem Cell. 2017;20:659–74 e659.
Mertens J, Wang QW, Kim Y, Yu DX, Pham S, Yang B, et al. Differential responses to lithium in hyperexcitable neurons from patients with bipolar disorder. Nature. 2015;527:95–99.
McNeill RV, Ziegler GC, Radtke F, Nieberler M, Lesch KP, Kittel-Schneider S. Mental health dished up-the use of iPSC models in neuropsychiatric research. J Neural Transm (Vienna). 2020;127:1547–68.
Srivastava R, Faust T, Ramos A, Ishizuka K, Sawa A. Dynamic Changes of the Mitochondria in Psychiatric Illnesses: New Mechanistic Insights From Human Neuronal Models. Biol Psychiatry. 2018;83:751–60.
Gimenez-Palomo A, Dodd S, Anmella G, Carvalho AF, Scaini G, Quevedo J, et al. The Role of Mitochondria in Mood Disorders: From Physiology to Pathophysiology and to Treatment. Front Psychiatry. 2021;12:546801.
Osete JR, Akkouh IA, de Assis DR, Szabo A, Frei E, Hughes T, et al. Lithium increases mitochondrial respiration in iPSC-derived neural precursor cells from lithium responders. Mol Psychiatry. 2021;26:6789–805.
Schulze M, Hoja S, Winner B, Winkler J, Edenhofer F, Riemenschneider MJ. Model Testing of PluriTest with Next-Generation Sequencing Data. Stem Cells Dev. 2016;25:569–71.
Yan Y, Shin S, Jha BS, Liu Q, Sheng J, Li F, et al. Efficient and rapid derivation of primitive neural stem cells and generation of brain subtype neurons from human pluripotent stem cells. Stem Cells Transl Med. 2013;2:862–70.
Milenkovic VM, Slim D, Bader S, Koch V, Heinl ES, Alvarez-Carbonell D, et al. CRISPR-Cas9 Mediated TSPO Gene Knockout alters Respiration and Cellular Metabolism in Human Primary Microglia Cells. Int J Mol Sci. 2019;20:3359.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Bader S, Wolf L, Milenkovic VM, Gruber M, Nothdurfter C, Rupprecht R, et al. Differential effects of TSPO ligands on mitochondrial function in mouse microglia cells. Psychoneuroendocrinology. 2019;106:65–76.
Bates D, Machler M, Bolker BM, Walker SC. Fitting Linear Mixed-Effects Models Using lme4. J Stat Softw. 2015;67:1–48.
Kuznetsova A, Brockhoff PB, Christensen RHB. lmerTest Package: Tests in Linear Mixed Effects Models. J Stat Softw. 2017;82:1–26.
Hilbert S, Stadler M, Lindl A, Naumann F, Bühner M. Analyzing longitudinal intervention studies with linear mixed models. Test Psychometrics Methodol Appl Psychol. 2019;26:101–19.
Zhang X, Huang CT, Chen J, Pankratz MT, Xi J, Li J, et al. Pax6 is a human neuroectoderm cell fate determinant. Cell Stem Cell. 2010;7:90–100.
Zheng X, Boyer L, Jin M, Mertens J, Kim Y, Ma L, et al. Metabolic reprogramming during neuronal differentiation from aerobic glycolysis to neuronal oxidative phosphorylation. Elife 2016;5:e13374.
Shadrina M, Bondarenko EA, Slominsky PA. Genetics Factors in Major Depression Disease. Front Psychiatry. 2018;9:334.
Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018;50:668–81.
Gratten J, Wray NR, Keller MC, Visscher PM. Large-scale genomics unveils the genetic architecture of psychiatric disorders. Nat Neurosci. 2014;17:782–90.
Sun H, Kennedy PJ, Nestler EJ. Epigenetics of the depressed brain: role of histone acetylation and methylation. Neuropsychopharmacology. 2013;38:124–37.
Nestler EJ. Epigenetic mechanisms of depression. JAMA Psychiatry. 2014;71:454–6.
Uchida S, Yamagata H, Seki T, Watanabe Y. Epigenetic mechanisms of major depression: targeting neuronal plasticity. Psychiatry Clin Neurosci. 2018;72:212–27.
Barbu MC, Shen X, Walker RM, Howard DM, Evans KL, Whalley HC, et al. Epigenetic prediction of major depressive disorder. Mol Psychiatry. 2021;26:5112–23.
Lin E, Tsai SJ. Epigenetics and Depression: An Update. Psychiatry Investig. 2019;16:654–61.
Doi A, Park IH, Wen B, Murakami P, Aryee MJ, Irizarry R, et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat Genet. 2009;41:1350–3.
Lister R, Pelizzola M, Kida YS, Hawkins RD, Nery JR, Hon G, et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature. 2011;471:68–73.
Ohi Y, Qin H, Hong C, Blouin L, Polo JM, Guo T, et al. Incomplete DNA methylation underlies a transcriptional memory of somatic cells in human iPS cells. Nat Cell Biol. 2011;13:541–9.
Lindau M, Neher E. Patch-clamp techniques for time-resolved capacitance measurements in single cells. Pflug Arch. 1988;411:137–46.
Hernández-Balaguera E, Vara H, Polo JL. Identification of Capacitance Distribution in Neuronal Membranes from a Fractional-Order Electrical Circuit and Whole-Cell Patch-Clamped Cells. J Electrochem Soc. 2018;165:G3104–G3111.
Kim MH, von Gersdorff H. Extending the realm of membrane capacitance measurements to nerve terminals with complex morphologies. J Physiol. 2010;588:2011–2.
Vaarmann A, Mandel M, Zeb A, Wareski P, Liiv J, Kuum M, et al. Mitochondrial biogenesis is required for axonal growth. Development. 2016;143:1981–92.
Pre D, Nestor MW, Sproul AA, Jacob S, Koppensteiner P, Chinchalongporn V, et al. A time course analysis of the electrophysiological properties of neurons differentiated from human induced pluripotent stem cells (iPSCs). PLoS ONE. 2014;9:e103418.
Fang D, Qing Y, Yan S, Chen D, Yan SS. Development and Dynamic Regulation of Mitochondrial Network in Human Midbrain Dopaminergic Neurons Differentiated from iPSCs. Stem Cell Rep. 2016;7:678–92.
Helm K, Viol K, Weiger TM, Tass PA, Grefkes C, Del Monte D, et al. Neuronal connectivity in major depressive disorder: a systematic review. Neuropsychiatr Dis Treat. 2018;14:2715–37.
Drysdale AT, Grosenick L, Downar J, Dunlop K, Mansouri F, Meng Y, et al. Resting-state connectivity biomarkers define neurophysiological subtypes of depression. Nat Med. 2017;23:28–38.
Acknowledgements
The authors would like to thank Richard Warth for providing access to the Seahorse device.
Funding
The work has been supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) project number 422182557 to CHW, and GRK2174 to CHW, KK, JT, and IC), the Bavarian State Ministry of Science and the Arts (Bavarian Research Networks ForIPS and ForInter, grants to MJR), and the BMBF (Research Grant No. 01EE1401B to TCB and RR). KD is fellowship of the “Else-Kröner-Fresenius-Stiftung. Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
CHW, JT and KK conceived and designed the experiments. JT, IC, KK, SB, TJ, KM, VMM and CHW performed the experiments and/or contributed to data acquisition. JT, IC, KK, SB, KM, TRH, MJR, SH, VMM and CHW analyzed and interpreted the data. CHW and JT wrote the original manuscript. CHW, JT and RR critically revised and edited the manuscript. KD, MB, CN, AM, TCB, CB and RR provided resources, and CHW was responsible for general supervision. All authors approved the final version of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Triebelhorn, J., Cardon, I., Kuffner, K. et al. Induced neural progenitor cells and iPS-neurons from major depressive disorder patients show altered bioenergetics and electrophysiological properties. Mol Psychiatry (2022). https://doi.org/10.1038/s41380-022-01660-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41380-022-01660-1