Abstract
Although the full aetiology of autism spectrum disorder (ASD) is unknown, familial and twin studies demonstrate high heritability of 60–90%, indicating a predominant role of genetics in the development of the disorder. The genetic architecture of ASD consists of a complex array of rare and common variants of all classes of genetic variation usually acting additively to augment individual risk. The relative contribution of heredity in ASD persists despite selective pressures against the classic autistic phenotype; a phenomenon thought to be explained, in part, by the incidence of spontaneous (or de novo) mutations. Notably, environmental exposures attributed as salient risk factors for ASD may play a causal role in the emergence of deleterious de novo variations, with several ASD-associated agents having significant mutagenic potential. To explore this hypothesis, this review article assesses published epidemiological data with evidence derived from assays of mutagenicity, both in vivo and in vitro, to determine the likely role such agents may play in augmenting the genetic liability in ASD. Broadly, these exposures were observed to elicit genomic alterations through one or a combination of: (1) direct interaction with genetic material; (2) impaired DNA repair; or (3) oxidative DNA damage. However, the direct contribution of these factors to the ASD phenotype cannot be determined without further analysis. The development of comprehensive prospective birth cohorts in combination with genome sequencing is essential to forming a causal, mechanistic account of de novo mutations in ASD that links exposure, genotypic alterations, and phenotypic consequences.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) is a pervasive neurodevelopmental condition estimated to affect ~1–1.5% of the global population [1, 2]. The behavioural phenotype of the disorder is characterised by early-onset dysfunction in social-communicative reciprocity and behavioural inflexibility [3], resulting in clinically significant impairment across a range of interpersonal, academic, and occupational contexts. Although age-related gains in adaptive functioning may attenuate symptomology in a subset of cases over time [4], the core deficits associated with childhood ASD minimally remit across the lifespan [5].
A strong contribution of heritable factors in the aetiology of ASD is supported by disproportionately increased risk of onset among first-degree relatives of probands [6, 7] and monozygotic twin concordance exceeding 60–90% [8, 9]. The genetic architecture of the disorder consists of a complex array of both rare (e.g., copy-number and single nucleotide variants, chromosomal abnormalities) [10] and common single nucleotide polymorphisms [2] acting additively to augment individual ASD risk (Fig. 1). The relative contribution of these mutations to the aetiology of the disorder is estimated at 2.5–15% and 12–52% [11, 12], respectively, with more recent evidence supporting the role of tandem repeat variations as additional, and incredibly salient, components of the ASD genotype (see section Elevated genomic sensitivity to exposure-induced mutagenesis). In several cases of syndromic ASD, a single genetic mutation seems sufficient to induce symptom onset [13], indicating that the disorder phenotype coincides disruption in important loss-of-function intolerant genes.
Both heritable and non-heritable factors can independently and reciprocally influence the development of ASD symptomatology. Up to 5–15% of ASD probands possess risk-associated de novo mutations, indicating the significance of non-familial genetic variability in determining disorder risk. The mutagenic/genotoxic potential of non-heritable factors associated with ASD suggests that these toxicants may play a role in the elicitation of spontaneous mutations. Figure created with BioRender.com.
The contribution of heredity in ASD persists despite strong selective pressures against the deleterious genetic events associated with disorder onset. Interestingly, the reproductive challenges faced by individuals with ASD are not physical, but instead mostly social [14, 15]. Fecundity is consequently reduced among affected individuals [16], limiting the potential transmission of putative risk variants to offspring in subsequent generations. Despite this, the prevalence of ASD has demonstrated stability or increases over time, with non-heritable aetiological factors insufficiently compensating for loss of high-risk genetic variants from the reproductive gene pool [17, 18].
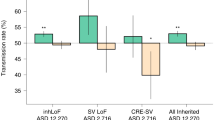
Preservation of the genetic liability of ASD despite reduced transmission of risk variants has been theorised to occur, in part, due to spontaneous de novo mutations [12, 19] (see Table 1). Depending on the study, which includes the complexity of the ASD phenotype, it is estimated that between 5 and 15% of ASD probands carry de novo mutations presumed to be involved in the disease [12], with a greater burden of de novo risk observed in simplex families without prior history of neurodevelopmental concerns [20]. Spontaneous genetic events are therefore more likely to represent important contributory factors to sporadic cases of ASD. Notably, although the rate of genomic de novo mutations is approximately equal between affected and unaffected familial trios, transmission of pathogenic mutations in important loss-of-function intolerant genes and gene pathways is observably higher among ASD probands [21,22,23], thereby consituting a critical feature of disorder aetiology.
Environmental exposures that are classically attributed as salient risk factors for ASD and other neurodevelopmental disorders could represent a catalyst for deleterious de novo variation, with several disorder-associated agents having significant mutagenic and genotoxic potential [24]. However, although the neurotoxicity and teratogenicity conferred by these toxicants is well established [25], their potential role in the genesis of de novo mutations of relevance to ASD has received little attention. Toxicogenomic analyses suggest disorder-associated exposures may perturb known ASD susceptibility genes through mutagenetic chemical-gene interactions [26], however a paucity of evidence limits the current conceptualisation of this relationship to very few exposures (see Table 2). Moreover, in the last decade only one article has addressed the likely contribution of such agents to the de novo burden of ASD [24], pre-dating advances in next generation sequencing which have since contributed to a more comprehensive understanding of the significance of de novo variants to ASD. This review combines epidemiological data with evidence derived from assays of mutagenicity (i.e., in vivo and in vitro) to assess the plausibility of environmental exposures as sources of de novo ASD-associated genetic events. Elucidating the role of these agents in eliciting mutations will assist to delineate the basis of aetiological risk associated with non-familial forms of ASD. It may also encourage primary health interventions aimed at reducing the negative impact of environmental exposures on ASD risk.
Mutagenicity and genotoxicity of early-exposure to established ASD toxicants
Although there is no support for a causal role for any single environmental risk factor in the onset of ASD, several pre-conceptual and prenatal environmental agents have been associated with increased risk of ASD and ASD-like symptoms. The most widely cited sources of environmental risk include exposure to toxins and heavy metals in utero [27], birth and maternal residency proximal to sources of ambient air pollution [28], prescription [29, 30] and illicit [31, 32] drug use during pregnancy, and parental health factors including age [33], maternal obesity and diabetes [34, 35], and pre- and peri-gestational vitamin status [36,37,38].
Very few direct assessments of the potential role of exposures in eliciting de novo mutations in ASD exist. However, significant observations have surfaced from cohort studies as to how parental age may contribute to spontaneous mutations associated with the disorder. Advanced paternal [39,40,41] and, more recently, maternal age [12, 33] at the time of conception have been demonstrated to confer an elevated risk of ASD development. To this end, age-associated accumulation of gametal DNA damage and failure of intrinsic repair mechanisms to excise acquired errors act as potent inducers of ASD-associated de novo risk. It is hypothesised that lifelong spermatogenesis affords recurrent opportunities for DNA damage and mis-repair in the generation of mature sperm [42], whereas prolonged meiotic arrest enhances the likelihood of damage-induced lesions accruing in the genetic structures of primary oocytes [43]. It is important to note that de novo risk persists beyond insemination, with rapid mitotic events during early embryogenesis elevating the risk of DNA damage and mutational events. Further, there is mounting evidence of differential mutational rates and mechanisms in postzygotic somatic mosaicism [44, 45], which is a current area of much-needed additional research.
Table 2 provides an extensive audit of current environmental factors identified via epidemiological studies as salient influencers of ASD risk. Although less thoroughly researched than the association with parental age, many exhibit evidence of mutagenicity and genotoxicity in human and relevant mammalian models. Broadly, these exposures can be observed to contribute to genomic alterations through one of three potential modes of action: (1) direct interaction with genetic material, both at the nucleic and chromosomal level; (2) interference with endogenous DNA repair; and (3) indirect DNA damage elicited through exposure-induced oxidative stress (see Fig. 2). Each of these processes has the capability to elicit genotypic abnormalities in biological systems and are, therefore, important to consider in relation to the de novo mutational burden of ASD.
ASD-associated toxicants (e.g., herbicides, heavy metals) can induce de novo mutations in parental germline cells which may be transmitted to offspring in the subsequent generation. For example, agent-induced double stranded breaks (DSBs) and impaired BRCA1-directed homologous recombination (HR) DNA damage response (DDR) can elicit de novo mutations and hamper their repair. Offspring may also acquire agent-induced mutations at later stages of development, resulting in somatic mosaicism. Gene(s) impacted by these processes can lead to aberrant neural development and functioning, contributing to the onset of ASD. Figure created with BioRender.com.
Direct interaction with genetic material
Several environmental agents exert their mutagenic and/or genotoxic potential by altering the DNA structure. Such mutations are primarily the consequence of toxicant-induced double-stranded breaks (DSBs): lesions to the DNA duplex which result in loss of stability and integrity in both strands of the nucleic acid helix. DSB repair is readily enacted by endogenous systems of non-homologous end-joining (NHEJ; i.e., ligation of two strands of damaged DNA) and homologous recombination (HR; i.e., template-dependent repair) [46], however aberrations may occur if the capacity for repair is exceeded by the degree of damage, or if repair-directed ligation results in erroneous rearrangement of DNA fragments [47]. The structural abnormalities that arise from un- or mis-repaired DSBs act greatly contribute to genomic instability, conferring risk for an array of genomic anomalies including nucleotide insertions and deletions (collectively referred to as indels) [48], gross chromosomal rearrangements [49], and fragmentation of chromosomes or chromatids [50], all of which can be sustained through multiple cycles of mitotic division. If such damage were to occur pre-gestationally in maternal or paternal gametes, or post-conceptually in early stages of embryogenesis, it is plausible that this could lead to presentation of novel genomic variation in offspring that is absent in either parental genome [51].
ASD-associated toxicants including chlorpyrifos, benzene, diesel particulate matter, and vinyl chloride induce various cellular phenotypes of genomic instability indicative of DSB DNA damage. Chlorpyrifos is an organophosphate congener widely applied in residential pest control, the use of which has been shown to possibly increase the incidence of ASD among prenatally exposed children [52]. Critically, chlorpyrifos have been shown to elicit DSBs and genetic rearrangements in the well-established ASD susceptibility gene KMT2A [53], a transcriptional coactivator gene repeatedly reported to harbour loss-of-function de novo variants among individuals with ASD [12, 54,55,56,57]. Conversely, benzene is an organic chemical compound widely applied in the industrial manufacture of plastics, and is present in petrochemicals including gasoline. Occupational exposure to benzene and benzene metabolites has been repeatedly associated with DSBs in intracellular nuclei [58], and confers abnormal morphology [59] and segregation of sex chromosomes in mature sperm [60]. Chromosomal instability elicited via DSBs increases the incidence of missegregation through disrupted genomic and protein integrity during mitotic division [61], potentiating the uneven distribution of genetic material in daughter cells. X/Y chromosome aneuploidy is frequently observed in clinical ASD cohorts [62], hence atypical inheritance of sex-linked genes (e.g., FMR1, NLGN3/4) orchestrated through toxicant-induced DSBs and DNA missegregation in parental gametes is a feasible mechanism of ASD risk.
Although potentially less deleterious than entire chromosomal losses or gains, micronuclei are important structural anomalies indicative of genomic instability and DSB mis-repair in cells exposed to harmful genotoxins [63]. The incidence of micronuclei and related extra-nuclear structures positively correlate with occupational exposure to diesel particulate matter among mechanics [64] and vinyl chloride in thermoplastic manufacturing facilities [65], both of which are considered salient toxicants in the aetiology of ASD [66, 67]. These extra-nuclear bodies can originate through the enveloping of acentric chromosome and chromatid fragments formed as consequence of asymmetrical DSB repair [47, 63]. Compartmentalised fragments of damaged DNA within these structures are vulnerable to acquisition of mutations through repeated cycles of defective replication and impaired recruitment of damage response pathways [68], and can reincorporate into the primary nucleus following breakdown of the nuclear membrane during mitosis. Daughters of micronucleated parent cells are therefore likely to harbour an array of genomic rearrangements including indels, chromosomal translocations, and copy-number variation in gene-coding sequences [61], all of which represent core features of the molecular architecture of ASD.
Interference of endogenous DNA repair responses
Toxicants capable of eliciting damage to the DNA structure are ubiquitous in the environment, however the harmful consequences of these are largely mitigated through rapid-activation of DNA damage response (DDR) pathways. DDRs are responsible for the detection and amelioration of aberrations in the genomic structure, and if necessary, can initiate apoptotic events to prevent transmission of lesioned genetic materials through DNA replication and mitotic division [69]. Several DDR pathways are perturbed through exposure to ASD-associated genotoxins, limiting the repair system’s ability to rectify acquired mutations and, by proxy, their affiliated biological consequences. Specifically, these include: the aforementioned NHEJ and HR pathways, which are responsible for extruding acquired DSBs; mismatch repair (MMR), which is necessary to amend errors in base-to-base alignment in replicated DNA; and base- (BER) and nucleotide excision repair (NER), which are enacted to correct single-base and bulky DNA lesions, respectively [46]. In many cases, mutations arise through exposure-induced delay or impaired recruitment of the machinery required to initiate DDR, hampering the overall DNA repair efficiency. The consequence of ineffective or inhibited repair responses by these systems are likely to coincide a vast array of genetic anomalies, ranging from single base-pairs to entire gene-encoding sequences, conferring significant and arguably the most pronounced risk to the acquisition of deleterious disorder-associated mutations.
The most common mode of DDR interference exerted by ASD-associated exposures is suppression of genes whose products are necessary to detect, excise, or amend lesions in the nucleotide structure. Several genes encoding proteins responsible for the identification of genomic anomalies are markedly downregulated in the NER (e.g., XPC) [69], BER (e.g., APEX1) [70], MMR (e.g., MSH3, MSH6) [69], and NHEJ (e.g., KU70/80) [71] pathways following exposure to ASD toxicants, suggesting these exposures may impair the ability of DDR mechanisms to effectively respond to damage in the DNA structure or sequence. Activation of the DDR pathways hinges on the capacity of the cell to detect aberrations in the genetic code, hence interference in the expression of these may cause delay or complete arrest in the damage-directed repair response. Although less frequently observed, genes encoding products required to effectively excise detected aberrations also demonstrate dysregulation following exposure to disorder-associated environmental agents, specifically within the NER response to induced DNA adducts (e.g., XPG) [72]. In such instances, acquired mutations remain unextruded despite appropriate activation of the DDR, precluding the repair response and permitting DNA damage and, thereby, de novo variation to persist.
Critically, exposure-induced dysregulation of genes required in the assembly and incision of corrected DNA sequences can be observed in both the HR (e.g., RAD51) [73, 74] and NHEJ (e.g., XRCC4) [71] DSB response pathways, suggesting ASD-associated toxicants are not only capable of inducing DSB lesions, but also inhibiting their effective repair. This represents a salient risk for the acquisition of de novo mutations, as un- or mis-repaired DSBs exert potent mutagenic potential as previously discussed (see section Direct interaction with genetic material). Remarkably, suppressed expression of HR repair genes following exposure to valproate, an anticonvulsant drug associated with elevated ASD risk [75], has been linked to idiosyncratic deficits in social-communicative behaviours in mammalian models [74], suggestive of a potentially causative relationship between impaired DDR and development of the disorder phenotype. Although it is plausible that this relationship could be mediated, at least in part, by accumulation of genetic aberrations through failure to appropriately repair acquired lesions, experimental confirmation is required.
Relative to HR, NHEJ DSB repair confers greater risk of acquiring structural anomalies through its ability to ligate strands of damaged DNA irrespective of homology. The unspecific nature of this mechanism allows tethering of mismatched or significantly mutilated termini, increasing the likelihood of gross chromosomal rearrangements or loss of genetic material [68]. To enact HR following lesion acquisition, an intact sister chromatid is required to act as a template for replication and synthesis of the damaged genetic material, resulting in high-fidelity repair. Initiation of HR is largely dependent on the breast cancer type 1 susceptibility protein, BRCA1, which can elicit DNA resection to inhibit NHEJ and trigger homology-directed repair [76]. Susceptibility to mutagenic events in early embryogenesis necessitates accurate DDR, hence HR is implemented as the preferred mechanism for DSB repair at this stage of development [77]. Exposure to ASD toxicants including valproate and phthalate, the latter a common plasticiser, downregulate BRCA1 in mammalian ovarian cells [69, 73] thereby reducing availability of the BRCA1 protein for the purposes of DNA resection. This increases reliance of the developing cellular system on the error-prone NHEJ DDR to ameliorate acquired DSBs, elevating the risk of, and frequency at which, the foetus will sustain de novo mutations.
Oxidative DNA damage
Generation of oxidative stress and reactive oxidative species (ROS) are important components of cellular metabolism, however, can become genotoxic when produced en masse in response to harmful environmental toxicants. The capacity for this otherwise non-pathogenic process to elicit DNA damage is due to the highly reactive hydroxyl radical (•OH). •OH forms double bonds with bases in the nucleic structure and abstracts hydrogen atoms from both the carbon-hydrogen bonds of 2’-deoxyribose and the methyl groups of thymine nucleotides [78]. This can elicit several genomic changes, including base-specific modifications, DNA-DNA and DNA-protein crosslinks, and both single- and DSBs [79]. Several classes of ASD-associated exposures elevate •OH ROS production and redox cycling across a range of mammalian and human tissue, suggestive of potentially ubiquitous consequences to the biological system.
Occupational and residential exposures are among the primary sources of oxidative DNA damage for several key ASD-associated toxicants. Prolonged contact with ROS-inducing agents increases the intracellular burden of oxidation, permitting accrual of genotoxic and cytotoxic lesions as consequence of persistent generation of •OH radicals. Among those aetiologically relevant exposures, the contaminants individuals are likely to interact with chronically – either through workplace activities or residential pollutants—include oxidative chemicals used in the production of synthetic materials (e.g., phthalates, styrene) [80,81,82], components of petroleum exhaust (e.g., 1,3-butadiene) [83], and toxic chemicals such as lead [84, 85] and mercury [86] utilised in industrial manufacturing processes. Of note, the oxidative DNA damage associative of chronic inhalation of 1,3-butadiene has been demonstrated to selectively target HRAS, a high-confidence ASD susceptibility gene, resulting in elevated point mutation accrual in affected cells [87]. Although typically assessed through urinary markers of oxidative DNA damage (i.e., 8-hydroxy-2′-deoxyguanosine, or 8-OHdG) [83, 88] or conveniently acquired biologicals (e.g., blood samples, buccal cells) [80, 89], occupational exposure to chemicals including phthalates [82] and styrene [90] have been directly observed to induce DNA aberration in gametes, supporting the capacity for ROS-induced lesions to localise to reproductive tissue. If viable, these mutated cells may result in genetic anomalies in subsequent offspring, thereby affording a mechanism for elicitation of non-familial genomic variability.
To counteract the pathological consequences of environmentally-induced oxidative stress, the biological system relies on a complex array of endogenous and exogeneous antioxidant defence mechanisms. These include metabolic antioxidant enzymes (e.g., glutathione peroxidase, catalase, superoxide dismutase), non-enzymatic proteins (e.g., lactoferrin), and scavengers of free radicals (e.g., iron) [91, 92]. Collectively, these constituents protect intracellular DNA and the broader cellular system from the damaging effects of excessive oxidative stress. It is therefore unsurprising that antioxidant deficiencies or interruption of their endogenous functions can indirectly contribute to the acquisition of genomic lesions if sustained over time. Maternal multivitamin supplementation during pregnancy has been shown to both reduce the incidence of ASD onset [37, 93] and buffer against oxidative DNA damage [94, 95], whereas ASD toxicants such as formaldehyde, a gaseous chemical used in the production of building materials, and flame retardant constituents including polybrominated diphenyl ethers disrupt antioxidant status in both gametal [96,97,98] and non-reproductive cells [99], potentiating oxidative imbalances and ROS-induced DNA damage in afflicted tissues.
De novo risk persists beyond embryogenesis
Spontaneous genetic mutations can be acquired at any stage of the lifespan [44, 45], and although those preceding or in early embryogenesis elicit a greater number of cells harbouring genetic anomalies (see Fig. 2), exposure-induced mutations in later periods of pregnancy or childhood may perturb neurodevelopment if sustained in temporally-critical brain-expressed genes. Mutations elicited in the post-zygotic period are restricted to specific subsets of dividing somatic cells, resulting in genetically dissimilar assemblages of tissue within the biological system [100]. This phenomenon, referred to as somatic mosaicism, permits variable expression of genes across clusters of daughter cells with diverse parental lineages, creating irregularity in the availability and integrity of the encoded protein [101]. If harboured in neural tissue prior to scheduled periods of brain development, disruption of genes central to processes of proliferation, synaptogenesis, or synaptic pruning may result in aberrant or stunted neurological growth; a cornerstone feature of the disorder phenotype (see Fig. 2). The deleterious effects of localised genetic lesions in ASD have been evidenced in histopathological studies of post-mortem brain tissue, with atypical segments of neocortical architecture in the frontal and temporal regions arising as consequence of somatic mutations in a subset of ASD patients [102, 103].
The incidence of de novo mutations and their role in the underlying pathology of ASD beyond early development has received little research attention, however the mutational capacity of environmental factors represents a source of somatic mosaicism likely to influence neural maturation throughout childhood. Although in isolation these are unlikely to elicit symptomatic presentations of ASD or ASD-like behaviours, such mutations may play an influential role in augmenting the severity and persistence of symptoms for at-risk individuals, hence contributing to the functional impairment coinciding clinical diagnosis.
Elevated genomic sensitivity to exposure-induced mutagenesis
Elevated genomic instability among ASD probands may enhance sensitivity to exposure-induced mutagenesis, increasing opportunity for mutation accrual and, by extension, de novo variation. Tandem repeat DNA motifs are highly liable genomic constituents prone to spontaneous somatic mutations [104], with increasing numbers of motif iterations (i.e., expansions) elevating the likelihood of novel genetic variation [105]. Constituting ~6% of the human genome, these units of repetitive DNA are known to contribute to molecular dysfunction across several complex clinical phenotypes, with recent analyses implicating repetitive DNA variants in the aetiology of ASD. Genome-wide interrogation of rare tandem repeat expansions has suggested that up to 2.6% of ASD risk may be explained by tandem expansions of repeat sequences enriched in exonic and splice sites across the genome, and often correlating with fragile sites [106]. In a separate study of a subset of the samples analysed by Trost and colleagues [106], the role of tandem repeat alterations in ASD was validated [107], further signalling that tandem repeats represent a significant component of the genetic aetiology of ASD. Critically, ASD-associated tandem repeat expansions were determined to be further expansions of large, inherited motifs, suggesting a transmission bias of these genomically unstable units among ASD probands [106, 108]. This preferential inheritance of mutationally liable DNA may therefore augment susceptibility to the mutagenic actions of environmental toxicants, likely increasing the incidence of putative de novo events across predominantly protein-coding and alternative transcription sites (we use the term putative due to the complexities in defining what is considered de novo in dynamic regions of the genome such as tandem repeats). The phenotypic consequences of such mutational events would vary depending on the putative gene affected, however given the elevated burden of de novo variation in loss-of-function intolerant regions among sporadic cases of ASD (see Table 1), the effects on clinical presentation are likely to be pronounced.
Environmental agents and epigenetics
Harmful exposures in the pre- and post-pregnancy period may elicit non-mutational epigenetic changes to the expression of genes central to early development [109]. Epigenetic modifications provoke transformations in the conformational arrangement of the nuclear structure and modulate intracellular activity, regulating the capacity for molecular machinery to transcribe and translate encoded genetic materials without directly affecting the underlying DNA. Several toxicants implicated in the aetiology of ASD are known to disrupt epigenetic mechanisms, particularly processes of DNA methylation [110], across developmentally sensitive stages of neural differentiation, proliferation, and migration [111], indicating that these non-genomic gene-environment interactions are important determinants in the susceptibility profile of ASD. Research suggests, however, that the role of the epigenome in perturbing processes central to the pathophysiology of ASD may extend beyond dysregulated gene expression. Studies concerning the mechanisms of tumorigenesis, a pathology largely underpinned by mutation accrual and disrupted gene regulation, have discerned that aberrant functioning in key epigenetic regulators may be causative of de novo genetic events [112]. For example, methylation of cytosine residues in CpG dinucleotides has been associated with repair-resistant T:G nucleotide mismatches, elevating base-level mutation rates at CpG sites relative to other dinucleotides across the genome [113]. Furthermore, accumulation and persistence of de novo events has been demonstrated to coincide promoter hypermethylation of key cell signalling and DDR-associated genes [112], thereby contributing to de novo mutations through both elicitation and impaired rectification of novel DNA variants. Evidence implicating DNA methylation as a target of ASD-associated toxicants has been reviewed elsewhere [110], and provides compelling support for the physiological consequences these agents exert on the epigenome, suggestive of its potential role in elicitation of de novo variation.
Limitations and future directions
The parallels between epidemiological trends and mutagenicity following exposure to ASD toxicants leads to intriguing hypotheses regarding the role of environmental agents in eliciting de novo mutations. Nonetheless, discerning causality in relation to the incidence of sporadic ASD remains challenging given the current state of the available research literature and therefore our review should support hypothesis building.
A key limitation in characterising the gene-environment relationship underscoring the de novo burden of ASD is the disparity in approaches from which heritable and non-heritable aetiological evidence is derived. On the one hand, identification of environmental agents that are likely to influence ASD development has depended on comparing the rates of ASD between exposed versus unexposed children from population cohorts (see Table 2). Since family-based designs are typically not employed within such studies, data is lacking both on the rate of de novo mutations in these samples and therefore the potential causative role of environmental exposures on de novo mutations. Clinic-referred samples, on the other hand, have afforded great insights into the contribution of rare, de novo mutations to the aetiology of ASD, but have rarely surveyed environmental risk as a modifier of this contribution, with the exception of parental age. As consequence, there is no informative estimate as to the proportion of de novo mutations among ASD probands which develop in response to environmental toxicant exposure. Moreover, although evidence supports a contribution of de novo mutational load in ASD at the level of specific genes and pathways, there is no overwhelming support for this effect genome-wide [21]. Thus, although we consider this line of inquiry to be of significant aetiological interest, we acknowledge the inherent limitations in trying to bridge these literatures.
A further limitation for the field is the absence of longitudinal evidence - human or otherwise - which simultaneously considers acquisition of non-familial genetic variation and ASD outcomes following pre- or post-conception toxicant exposure. The prognostic trajectory of ASD and other neurodevelopmental conditions has led to increasing interest and activity in establishing prospective birth cohorts to monitor environmental correlates of childhood disorder phenotypes (e.g., the Children of Nurses’ Health Study [66], Table 2). This has permitted the detrimental effects of developmental exposures to be observed in relation to symptomatic onset in later life, enhancing understanding of the non-heritable aetiology of ASD. Nevertheless, incorporation of the biological techniques necessary to detect and trace mutational events among such cohorts is limited, if not entirely absent. As such, this review has drawn upon broader evidence of exposures’ mutagenicity from in vivo and in vitro methodologies. Although the literature reviewed provides tantalizing insights into the ways in which environmental exposures might be linked to de novo mutational events, disorder-specific causation is lacking. To establish the role of disorder-relevant genotoxins in enhancing the de novo burden of ASD, both the genotypic and phenotypic consequences of parental/prenatal exposures must be assessed prospectively from birth in broader population cohorts. This will not only permit identification of novel genetic variation resulting from harmful toxicants, but allow the corollaries of acquired mutations on neurodevelopment to be traced across sensitive periods of childhood.
Identification of spontaneous de novo mutations relies on availability of the genetic information from each of the mother, father, and offspring, allowing intergenerational divergence in the DNA sequence to be distinguished from inherited allelic compositions [12]. Detection and assessment of acquired genomic abnormalities has traditionally relied on cytogenetic assays which, although capable of discerning significant aberrations in the chromosomal structure [101], cannot capture less conspicuous mutations known to contribute to the genetic aetiology of ASD. Current conceptualisations of the mutagenicity and genotoxicity intrinsic to several environmental toxicants has therefore been limited to their capacity to elicit large-scale malformations in the genomic structure, neglecting the subtler effects these exposures may exert on the genetic sequence. With the development of next generation sequencing (NGS) methods, however, the ability to identify nucleic acid deviations in as little as a single base pair in offspring relative to parental genomes is achievable, accurate, and increasingly affordable. NGS involves comprehensive assessment of the nucleic arrangement of either the entire genome (i.e., whole genome sequencing) or specifically gene-encoding segments of DNA (i.e., whole exome sequencing), allowing variation in the genomic code to be detected and compared between individual genomes. Such techniques have provided important insights into the rate of spontaneous genetic variation in cases of sporadic ASD, demonstrating the importance of genotypic variation to ASD development [19]. The advances in technology which spurred the advent of NGS have significantly reduced the costs associated with performing high-throughput sequencing, hence it is feasible to apply these to existing and emerging birth cohorts to assess the origin and frequency of de novo mutations among infants diagnosed with ASD in childhood. Such data would provide crucial insight into the genuine risks environmental toxicants pose to accentuating the genetic aetiology associated with ASD, enhancing our understanding of the mechanisms by which these agents contribute to the ASD phenotype.
Concerns for the harmful effects of prenatal exposure to environmental toxicants has led to a maternal bias in epidemiological studies assessing trends in ASD onset. Although such concerns are founded on well-established evidence for the disruptive consequences of exogenous agents on foetal development in utero [25], this focus has arguably undermined the significance of pre-conception paternal exposures. Relative to oocytes, spermatocytes appear to possess heightened sensitivity to genotoxic agents within the environment, putting male gametes at greater risk of acquiring and transmitting genomic lesions to the subsequent generation [91]. As previously discussed, recurrent spermatogenesis across the lifespan elevates the risk of defective DNA replication and repair in male relative to female gametes [43]. Should the testicular tissue from which spermatozoa divide acquire environmentally-induced genomic damage, as has been evinced following exposure to several known ASD toxicants (see Table 2), it is likely that resultant sperm cells will harbour genetic abnormalities. Germline susceptibility to environmental agents may be additionally mediated by the intrinsic ability of cells to effectively employ DDR following harmful exposures, the capacity for which is significantly compromised in sperm [114]. During spermatogenesis, the nuclear chromatin within the cell becomes highly condensed to improve motility for insemination. As consequence, the capacity to effectively excise and extrude genomic aberrations is limited, hence mutations in the genomic sequence may persist to fertilisation. Although there is evidence to support the capacity of the fused oocyte to perform post-fertilisation DDR of spermatic DNA, the frequency of de novo mutations of paternal origin in atypically developing offspring [33] suggests this repair response is imperfect. Given such sensitivity to mutagenic agents, assessing the incidence of ASD among offspring of fathers exposed to toxicants is essential to characterising the role of paternally acquired mutations in enhancing onset risk, further characterising the aetiological basis and more specifically the de novo origin of the disorder.
Notwithstanding these limitations, understanding the mutagenic mechanisms of environmental toxicants and, by proxy, their role in the elicitation of de novo risk variants, may inform public health initiatives to combat increasing rates of ASD within the community. Mutagenic exposures such as parental drug use and pre- and post-conceptual health factors offer targetable and inexpensive intervention opportunities which may assist to circumvent disorder-associated mutational events. For example, maternal folate and vitamin D supplementation prior to and during gestation has been shown to significantly reduce the rate of ASD among offspring [37]. In contrast, administration of substances such as paracetamol [29, 115] and antidepressant medications [30] during pregnancy is associated with increased risk of disorder onset in the subsequent generation. Addressing these modifiable factors may prove to be effective combatants to the de novo burden of ASD, warranting further research in the interest of public health. To this end, the advent and increased accessibility of novel molecular tools such as induced pluripotent stem cell (iPSCs)-derived neuronal lines has equipped researchers to evaluate agent-induced mutagenesis in clinically relevant tissue [116], facilitating the evaluation of the current hypothesis and its utility in informing primary intervention strategies. Recent advances in developing evidence-based list of genes relevant to autism and the neurobehavioral characteristics associated with it [117] will also enable more specific hypothesis testing in model systems, as well as epidemiological studies.
Concluding remarks
The paradox of stable (or increasing) ASD rates in the general population despite reduced fecundity provides a compelling rationale for the contribution of de novo mutations to the genetic aetiology of ASD [12]. The potential for non-heritable factors such as environmental toxicants to elicit de novo events may afford an explanation for the underlying mutational trigger, leading to novel pathogenic mutationsin specific disorder-associated gene(s), thereby contributing to symptomatic onset. At both the pre- and post-gestational period, harmful environmental agents may induce genomic lesions through a myriad of genotoxic mechanisms, summarised within this review into direct, repair inhibition, and oxidative DNA damage induction. Acquired mutations in parental gametes or the developing embryo may potentiate the disorder phenotype if localised to genes salient in processes of general development or specific neurodevelopmental pathways, however this risk may persist throughout maturation through the phenomenon of somatic mosaicism. The mutagenic potential of ASD toxicants may be further potentiated by intrinsic genomic instability among ASD probands, with recent identification of rare tandem repeat expansions among affected individuals likely increasing mutational liability to environmental toxins. Whereas iPSC derived neuronal lines might now be used to test toxin exposure on mutation rates [116], determining the specific contribution of these environmentally-induced DNA alterations to ASD is difficult, as there is a paucity of population-based, longitudinal evidence necessary to draw conclusive links between exposure, genotypic responses, and phenotypic consequences. In addition, neglect for the critical influence of paternal exposures on offspring outcomes is evident in available epidemiological surveys of ASD trends, creating a maternally-biased view of the contribution of environment to the disorder phenotype. The development of comprehensive prospective birth cohorts in tandem with increasing fidelity and accessibility of high-throughput sequencing offers unprecedented opportunities to trace the effects and outcomes of developmental genotoxin exposure. This will deepen our understanding of the complex gene-environment relationships underpinning the aetiology of ASD.
References
Lord C, Brugha TS, Charman T, Cusack J, Dumas G, Frazier T, et al. Autism spectrum disorder. Nat Rev Dis Prim. 2020;6:5.
Grove J, Ripke S, Als TD, Mattheisen M, Walters RK, Won H, et al. Identification of common genetic risk variants for autism spectrum disorder. Nat Genet. 2019;51:431–44.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders (DSM-5). American Psychiatric Association Publishing: Washington, DC, 2013.
Farley M, Cottle KJ, Bilder D, Viskochil J, Coon H, McMahon W. Mid-life social outcomes for a population-based sample of adults with ASD. Autism Res. 2018;11:142–52.
Baxter AJ, Brugha TS, Erskine HE, Scheurer RW, Vos T, Scott JG. The epidemiology and global burden of autism spectrum disorders. Psychol Med. 2015;45:601–13.
Hansen SN, Schendel DE, Francis RW, Windham GC, Bresnahan M, Levine SZ, et al. Recurrence risk of autism in siblings and cousins: a multinational, population-based study. J Am Acad Child Adolesc Psychiatry. 2019;58:866–75.
Sandin S, Lichtenstein P, Kuja-Halkola R, Larsson H, Hultman CM, Reichenberg A. The familial risk of autism. JAMA. 2014;311:1770–7.
Castelbaum L, Sylvester CM, Zhang Y, Yu Q, Constantino JN. On the nature of monozygotic twin concordance and discordance for autistic trait severity: a quantitative analysis. Behav Genet. 2019;50:263–72.
Tick B, Bolton P, Happé F, Rutter M, Rijsdijk F. Heritability of autism spectrum disorders: a meta-analysis of twin studies. J Child Psychol Psychiatry. 2016;57:585–95.
Kenny EM, Cormican P, Furlong S, Heron E, Kenny G, Fahey C, et al. Excess of rare novel loss-of-function variants in synaptic genes in schizophrenia and autism spectrum disorders. Mol Psychiatry. 2014;19:872–9.
Gaugler T, Klei L, Sanders SJ, Bodea CA, Goldberg AP, Lee AB, et al. Most genetic risk for autism resides with common variation. Nat Gen. 2014;46:881–5.
Iossifov I, O’Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature. 2014;515:216–21.
Fernandez BA, Scherer SW. Syndromic autism spectrum disorders: moving from a clinically defined to a molecularly defined approach. Dialogues Clin Neurosci. 2017;19:353–71.
Pelphrey KA, Shultz S, Hudac CM, Vander Wyk BC. Research review: Constraining heterogeneity: the social brain and its development in autism spectrum disorder. J Child Psychol Psychiatry. 2011;52:631–44.
Keller MC, Miller G. Resolving the paradox of common, harmful, heritable mental disorders: which evolutionary genetic models work best? Behav Brain Sci. 2006;29:385–404.
Power RA, Kyaga S, Uher R, MacCabe JH, Långström N, Landen M, et al. Fecundity of patients with schizophrenia, autism, bipolar disorder, depression, anorexia nervosa, or substance abuse vs their unaffected siblings. JAMA Psychiatry. 2013;70:22–30.
Nevison CD. A comparison of temporal trends in United States autism prevalence to trends in suspected environmental factors. Environ Health. 2014;13:73.
Taylor MJ, Rosenqvist MA, Larsson H, Gillberg C, D’Onofrio BM, Lichtenstein P, et al. Etiology of autism spectrum disorders and autistic traits over time. JAMA Psychiatry. 2020;77:936–43.
Sanders SJ, Murtha MT, Gupta AR, Murdoch JD, Raubeson MJ, Willsey AJ, et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature. 2012;485:237–41.
Leppa Virpi M, Kravitz Stephanie N, Martin Christa L, Andrieux J, Le Caignec C, Martin-Coignard D, et al. Rare inherited and de novo CNVs reveal complex contributions to ASD risk in multiplex families. Am J Hum Genet. 2016;99:540–54.
Pinto D, Delaby E, Merico D, Barbosa M, Merikangas A, Klei L, et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am J Hum Genet. 2014;94:677–94.
Marshall CR, Noor A, Vincent JB, Lionel AC, Feuk L, Skaug J, et al. Structural variation of chromosomes in autism spectrum disorder. Am J Hum Genet. 2008;82:477–88.
Sebat J, Lakshmi B, Malhotra D, Troge J, Lese-Martin C, Walsh T, et al. Strong association of de novo copy number mutations with autism. Science. 2007;316:445–9.
Kinney DK, Barch DH, Chayka B, Napoleon S, Munir KM. Environmental risk factors for autism: do they help cause de novo genetic mutations that contribute to the disorder? Med Hypotheses. 2010;74:102–6.
Bölte S, Girdler S, Marschik PB. The contribution of environmental exposure to the etiology of autism spectrum disorder. Cell Mol Life Sci. 2019;76:1275–97.
Carter CJ, Blizard RA. Autism genes are selectively targeted by environmental pollutants including pesticides, heavy metals, bisphenol A, phthalates and many others in food, cosmetics or household products. Neurochem Int. 2016;101:83–109.
von Ehrenstein OS, Aralis H, Cockburn M, Ritz B. In utero exposure to toxic air pollutants and risk of childhood autism. Epidemiology. 2014;25:851–8.
Kalkbrenner AE, Windham GC, Zheng C, McConnell R, Lee Nora L, Schauer JJ, et al. Air toxics in relation to autism diagnosis, phenotype, and severity in a U.S. family-based study. Environ Health Perspect. 2018;126:037004.
Masarwa R, Levine H, Gorelik E, Reif S, Perlman A, Matok I. Prenatal exposure to acetaminophen and risk for attention deficit hyperactivity disorder and autistic spectrum disorder: a systematic review, meta-analysis, and meta-regression analysis of cohort studies. Am J Epidemiol. 2018;187:1817–27.
Rai D, Lee BK, Dalman C, Golding J, Lewis G, Magnusson C. Parental depression, maternal antidepressant use during pregnancy, and risk of autism spectrum disorders: population based case-control study. BMJ. 2013;346:f2059.
Guennewig B, Bitar M, Obiorah I, Hanks J, O’Brien EA, Kaczorowski DC, et al. THC exposure of human iPSC neurons impacts genes associated with neuropsychiatric disorders. Transl Psychiatry. 2018;8:89–89.
Ornoy A, Weinstein-Fudim L, Ergaz Z. Prenatal factors associated with autism spectrum disorder (ASD). Reprod Toxicol. 2015;56:155–69.
Wu S, Wu F, Ding Y, Hou J, Bi J, Zhang Z. Advanced parental age and autism risk in children: a systematic review and meta-analysis. Acta Psychiatr Scand. 2017;135:29–41.
Li M, Fallin MD, Riley A, Landa R, Walker SO, Silverstein M, et al. The association of maternal obesity and diabetes with autism and other developmental disabilities. Pediatrics. 2016;137:e20152206.
Xiang AH, Wang X, Martinez MP, Walthall JC, Curry ES, Page K, et al. Association of maternal diabetes with autism in offspring. JAMA. 2015;313:1425–34.
Chen J, Xin K, Wei J, Zhang K, Xiao H. Lower maternal serum 25(OH) D in first trimester associated with higher autism risk in Chinese offspring. J Psychosom Res. 2016;89:98–101.
Levine SZ, Kodesh A, Viktorin A, Smith L, Uher R, Reichenberg A, et al. Association of maternal use of folic acid and multivitamin supplements in the periods before and during pregnancy with the risk of autism spectrum disorder in offspring. JAMA Psychiatry. 2018;75:176–84.
Raghavan R, Riley AW, Volk H, Caruso D, Hironaka L, Sices L, et al. Maternal multivitamin intake, plasma folate and vitamin B12 levels and autism spectrum disorder risk in offspring. Paediatr Perinat Epidemiol. 2018;32:100–11.
Kong A, Frigge ML, Masson G, Besenbacher S, Sulem P, Magnusson G, et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature. 2012;488:471–5.
Yuen RKC, Merico D, Cao H, Pellecchia G, Alipanahi B, Thiruvahindrapuram B, et al. Genome-wide characteristics of de novo mutations in autism. NPJ Genom Med. 2016;1:160271–10.
Michaelson JJ, Shi Y, Gujral M, Zheng H, Malhotra D, Jin X, et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell. 2012;151:1431–42.
Crow JF. The origins, patterns and implications of human spontaneous mutation. Nat Rev Genet. 2000;1:40–47.
Wong WSW, Solomon BD, Bodian DL, Kothiyal P, Eley G, Huddleston KC, et al. New observations on maternal age effect on germline de novo mutations. Nat Commun. 2016;7:10486.
Lim ET, Uddin M, De Rubeis S, Chan Y, Kamumbu AS, Zhang X, et al. Rates, distribution and implications of postzygotic mosaic mutations in autism spectrum disorder. Nat Neurosci. 2017;20:1217–24.
Bae T, Tomasini L, Mariani J, Zhou B, Roychowdhury T, Franjic D, et al. Different mutational rates and mechanisms in human cells at pregastrulation and neurogenesis. Science. 2018;359:550–5.
Chatterjee N, Walker GC. Mechanisms of DNA damage, repair, and mutagenesis. Environ Mol Mutagen. 2017;58:235–63.
Janssen A, Medema RH. Genetic instability: tipping the balance. Oncogene. 2013;32:4459–70.
Sankaranarayanan K, Taleei R, Rahmanian S, Nikjoo H. Ionizing radiation and genetic risks. XVII. Formation mechanisms underlying naturally occurring DNA deletions in the human genome and their potential relevance for bridging the gap between induced DNA double-strand breaks and deletions in irradiated germ cells. Mutat Res Rev Mutat Res. 2013;753:114–30.
Varga T, Aplan PD. Chromosomal aberrations induced by double strand DNA breaks. DNA Repair. 2005;4:1038–46.
Kaye JA, Melo JA, Cheung SK, Vaze MB, Haber JE, Toczyski DP. DNA breaks promote genomic instability by impeding proper chromosome segregation. Curr Biol. 2004;14:2096–106.
Buchanan JA, Scherer SW. Contemplating effects of genomic structural variation. Genet Med. 2008;10:639–47.
Shelton JF, Geraghty EM, Tancredi DJ, Delwiche LD, Schmidt RJ, Ritz B, et al. Neurodevelopmental disorders and prenatal residential proximity to agricultural pesticides: the CHARGE study. Environ Health Perspect. 2014;122:1103–9.
Lu C, Liu X, Liu C, Wang J, Li C, Liu Q, et al. Chlorpyrifos induces MLL translocations through caspase 3-dependent genomic instability and topoisomerase II inhibition in human fetal liver hematopoietic stem cells. Toxicol Sci. 2015;147:588–606.
Yuen R, Merico D, Bookman M,JLH, Thiruvahindrapuram B, Patel RV, et al. Whole genome sequencing resource identifies 18 new candidate genes for autism spectrum disorder. Nat Neurosci. 2017;20:602–11.
De Rubeis S, He X, Goldberg AP, Poultney CS, Samocha K, Cicek AE, et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature. 2014;515:209–15.
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature. 2015;519:223–8.
Satterstrom FK, Kosmicki JA, Wang J, Breen MS, De Rubeis S, An JY, et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell. 2020;180:568–84.e523.
Costa-Amaral IC, Carvalho LVB, Santos MVC, Valente D, Pereira AC, Figueiredo VO, et al. Environmental assessment and evaluation of oxidative stress and genotoxicity biomarkers related to chronic occupational exposure to benzene. Int J Environ Res Public Health. 2019;16:2240.
Marchetti F, Eskenazi B, Weldon RH, Li G, Zhang L, Rappaport SM, et al. Occupational exposure to benzene and chromosomal structural aberrations in the sperm of Chinese men. Environ Health Perspect. 2012;120:229–34.
Ji Z, Weldon RH, Marchetti F, Chen H, Li G, Xing C, et al. Comparison of aneuploidies of chromosomes 21, X, and Y in the blood lymphocytes and sperm of workers exposed to benzene. Environ Mol Mutagen. 2012;53:218–26.
Ly P, Brunner SF, Shoshani O, Kim DH, Lan W, Pyntikova T, et al. Chromosome segregation errors generate a diverse spectrum of simple and complex genomic rearrangements. Nat Genet. 2019;51:705–15.
Tartaglia NR, Wilson R, Miller JS, Rafalko J, Cordeiro L, Davis S, et al. Autism spectrum disorder in males with sex chromosome aneuploidy: XXY/Klinefelter Syndrome, XYY, and XXYY. J Dev Behav Pediatr. 2017;38:197–207.
Fenech M, Kirsch-Volders M, Natarajan AT, Surralles J, Crott JW, Parry J, et al. Molecular mechanisms of micronucleus, nucleoplasmic bridge and nuclear bud formation in mammalian and human cells. Mutagenesis. 2011;26:125–32.
León-Mejía G, Luna-Rodríguez I, Trindade C, Oliveros-Ortíz L, Anaya-Romero M, Luna-Carrascal J, et al. Cytotoxic and genotoxic effects in mechanics occupationally exposed to diesel engine exhaust. Ecotoxicol Environ Saf. 2019;171:264–73.
Kumar AK, Balachandar V, Arun M, Ahamed SAKM, Kumar SS, Balamuralikrishnan B. et al. A comprehensive analysis of plausible genotoxic covariates among workers of a polyvinyl chloride plant exposed to vinyl chloride monomer. Arch Environ Contam Toxicol. 2013;64:652–8.
Roberts AL, Lyall K, Hart Jaime E, Laden F, Just Allan C, Bobb, et al. Perinatal air pollutant exposures and autism spectrum disorder in the Children of Nurses’ Health Study II participants. Environ Health Perspect. 2013;121:978–84.
Windham GC, Zhang L, Gunier R, Croen LA, Grether JK. Autism spectrum disorders in relation to distribution of hazardous air pollutants in the San Francisco Bay Area. Environ Health Perspect. 2006;114:1438–44.
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y, et al. DNA breaks and chromosome pulverization from errors in mitosis. Nature. 2012;482:53–58.
Liu X, Craig Z. R. Environmentally relevant exposure to dibutyl phthalate disrupts DNA damage repair gene expression in the mouse ovary. Biol Reprod. 2019;101:854–67.
Attia SM, Harisa GI, Hassan MH, Bakheet SA. Beryllium chloride-induced oxidative DNA damage and alteration in the expression patterns of DNA repair-related genes. Mutagenesis. 2013;28:555–9.
Sanie-Jahromi F, Saadat M. Effects of electromagnetic field, cisplatin and morphine on cytotoxicity and expression levels of DNA repair genes. Mol Biol Rep. 2018;45:807–14.
Yang G, Ibuki Y. Cigarette sidestream smoke delays nucleotide excision repair: inhibited accumulation of repair proteins at DNA lesions. Carcinogenesis. 2017;39:56–65.
Lamparter C, Winn LM. Tissue-specific effects of valproic acid on DNA repair genes and apoptosis in postimplantation mouse embryos. Toxicol Sci. 2014;141:59–67.
Servadio M, Manduca A, Melancia F, Leboffe L, Schiavi S, Campolongo P, et al. Impaired repair of DNA damage is associated with autistic-like traits in rats prenatally exposed to valproic acid. Eur Neuropsychopharmacol. 2018;28:85–96.
Christensen J, Grønborg TK, Sørensen MJ, Schendel D, Parner ET, Pedersen LH, et al. Prenatal valproate exposure and risk of autism spectrum disorders and childhood autism. JAMA. 2013;309:1696–703.
Daley JM, Sung P. 53BP1, BRCA1, and the choice between recombination and end joining at DNA double-strand breaks. Mol Cell Biol. 2014;34:1380–88.
Chiruvella KK, Sebastian R, Sharma S, Karande AA, Choudhary B, Raghavan SC. Time-dependent predominance of nonhomologous DNA end-joining pathways during embryonic development in mice. J Mol Biol. 2012;417:197–211.
Cooke MS, Evans MD, Dizdaroglu M, Lunec J. Oxidative DNA damage: mechanisms, mutation, and disease. FASEB J. 2003;17:1195–214.
Dobrakowski M, Pawlas N, Kasperczyk A, Kozłowska A, Olewińska E, Machoń-Grecka A, et al. Oxidative DNA damage and oxidative stress in lead-exposed workers. Hum Exp Toxicol. 2016;36:744–54.
Cavallo D, Tranfo G, Ursini CL, Fresegna AM, Ciervo A, Maiello R, et al. Biomarkers of early genotoxicity and oxidative stress for occupational risk assessment of exposure to styrene in the fibreglass reinforced plastic industry. Toxicol Lett. 2018;298:53–59.
Gurdemir G, Erkekoglu P, Balci A, Sur U, Ozkemahli G, Tutkun E, et al. Oxidative stress parameters, selenium levels, DNA damage, and phthalate levels in plastic workers. J Environ Pathol Toxicol Oncol. 2019;38:253–70.
Huffman AM, Wu H, Rosati A, Rahil T, Sites CK, Whitcomb BW, et al. Associations of urinary phthalate metabolites and lipid peroxidation with sperm mitochondrial DNA copy number and deletions. Environ Res. 2018;163:10–15.
Arayasiri M, Mahidol C, Navasumrit P, Autrup H, Ruchirawat M. Biomonitoring of benzene and 1,3-butadiene exposure and early biological effects in traffic policemen. Sci Total Environ. 2010;408:4855–62.
Chinde S, Kumari M, Devi KR, Murty US, Rahman MF, Kumari SI, et al. Assessment of genotoxic effects of lead in occupationally exposed workers. Environ Sci Pollut Res. 2014;21:11469–80.
Jasso-Pineda Y, Díaz-Barriga F, Calderón J, Yáñez L, Carrizales L, Pérez-Maldonado IN. DNA damage and decreased DNA repair in peripheral blood mononuclear cells in individuals exposed to arsenic and lead in a mining site. Biol Trace Elem Res. 2012;146:141–9.
Cruz-Esquivel Á, Marrugo-Negrete J, Calao-Ramos C. Genetic damage in human populations at mining sites in the upper basin of the San Jorge River, Colombia. Environ Sci Pollut Res. 2019;26:10961–71.
Sills RC, Hong HL, Boorman GA, Devereux TR, Melnick RL. Point mutations of K-ras and H-ras genes in forestomach neoplasms from control B6C3F1 mice and following exposure to 1,3-butadiene, isoprene or chloroprene for up to 2-years. Chem Biol Interact. 2001;135-136:373–86.
Kim SS, Meeker JD, Keil AP, Aung MT, Bommarito PA, Cantonwine DE, et al. Exposure to 17 trace metals in pregnancy and associations with urinary oxidative stress biomarkers. Environ Res. 2019;179:108854.
Tucker JD, Sorensen KJ, Ruder AM, McKernan LT, Forrester CL, Butler MA. Cytogenetic analysis of an exposed-referent study: perchloroethylene-exposed dry cleaners compared to unexposed laundry workers. Environ Health. 2011;10:16.
Migliore L, Colognato R, Naccarati A, Bergamaschi E. Relationship between genotoxicity biomarkers in somatic and germ cells: findings from a biomonitoring study. Mutagenesis. 2006;21:149–52.
García-Rodríguez A, Gosálvez J, Agarwal A, Roy R, Johnston S. DNA damage and repair in human reproductive cells. Int J Mol Sci. 2018;20:31.
Poljsak B, Šuput D, Milisav I. Achieving the balance between ROS and antioxidants: when to use the synthetic antioxidants. Oxid Med Cell Longev. 2013;2013:956792.
Tan M, Yang T, Zhu J, Li Q, Lai X, Li Y, et al. Maternal folic acid and micronutrient supplementation is associated with vitamin levels and symptoms in children with autism spectrum disorders. Reprod Toxicol. 2020;91:109–15.
Haq SH, AlAfaleq NO, Johari RA. Vitamin D treatment reverses the induced oxidative stress damage to DNA. Pak J Biol Sci. 2019;22:8–14.
Polidoro L, Properzi G, Marampon F, Gravina GL, Festuccia C, Di Cesare E, et al. Vitamin D protects human endothelial cells from H2O2 oxidant injury through the Mek/Erk-Sirt1 axis activation. J Cardiovasc Transl Res. 2013;6:221–31.
Razi M, Malekinejad H, Sayrafi R, Hosseinchi MR, Feyzi S, Moshtagion SM, et al. Adverse effects of long-time exposure to formaldehyde vapour on testicular tissue and sperm parameters in rats. Vet Res Forum. 2013;4:213–9.
Wang H-X, Wang X-Y, Zhou D-X, Zheng L-R, Zhang J, Huo Y-W, et al. Effects of low-dose, long-term formaldehyde exposure on the structure and functions of the ovary in rats. Toxicol Ind Health. 2012;29:609–15.
Li X, Zhu Y, Zhang C, Liu J, Zhou G, Jing L, et al. BDE-209 induces male reproductive toxicity via cell cycle arrest and apoptosis mediated by DNA damage response signaling pathways. Environ Pollut. 2019;255:113097.
Manuguerra S, Espinosa Ruiz C, Santulli A, Messina CM. Sub-lethal doses of polybrominated diphenyl ethers, in vitro, promote oxidative stress and modulate molecular markers related to cell cycle, antioxidant balance and cellular energy management. Int J Environ Res Public Health. 2019;16:588.
Acuna-Hidalgo R, et al. Post-zygotic point mutations are an underrecognized source of de novo genomic variation. Am J Hum Genet. 2015;97:67–74.
D’Gama AM, Walsh CA. Somatic mosaicism and neurodevelopmental disease. Nat Neurosci. 2018;21:1504–14.
Stoner R, Chow ML, Boyle MP, Sunkin SM, Mouton PR, Roy S, et al. Patches of disorganization in the neocortex of children with autism. N Engl J Med. 2014;370:1209–19.
Casanova MF, El-Baz AS, Kamat SS, Dombroski BA, Khalifa F, Elnakib A, et al. Focal cortical dysplasias in autism spectrum disorders. Acta Neuropathol Commun. 2013;1:67.
Hannan AJ. Tandem repeats mediating genetic plasticity in health and disease. Nat Rev Genet. 2018;19:286–98.
López Castel A, Cleary JD, Pearson CE. Repeat instability as the basis for human diseases and as a potential target for therapy. Nat Rev Mol Cell Biol. 2010;11:165–70.
Trost B, Engchuan W, Nguyen CM, Thiruvahindrapuram B, Dolzhenko E, Backstrom I, et al. Genome-wide detection of tandem DNA repeats that are expanded in autism. Nature. 2020;586:80–86.
Mitra I, Huang B, Nima M, Ma N, Lamkin M, Yanicky R, et al. Genome-wide patterns of de novo tandem repeat mutations and their contribution to autism spectrum disorders. Nature. 2021;589:246–50.
Hannan AJ. Repeat DNA expands our understanding of autism spectrum disorder. Nature. 2021;589:200–2.
Vogel Ciernia A, LaSalle J. The landscape of DNA methylation amid a perfect storm of autism aetiologies. Nat Rev Neurosci. 2016;17:411–23.
Keil KP, Lein PJ. DNA methylation: a mechanism linking environmental chemical exposures to risk of autism spectrum disorders? Environ Epigenet. 2016;2:dvv012.
Tran NQV, Miyake K. Neurodevelopmental disorders and environmental toxicants: epigenetics as an underlying mechanism. Int J Genomics. 2017;2017:7526592.
You JS, Jones PA. Cancer genetics and epigenetics: two sides of the same coin? Cancer Cell. 2012;22:9–20.
Acuna-Hidalgo R, Veltman JA, Hoischen A. New insights into the generation and role of de novo mutations in health and disease. Genome Biol. 2016;17:241.
Smith TB, Dun MD, Smith ND, Curry BJ, Connaughton HS, Aitken RJ. The presence of a truncated base excision repair pathway in human spermatozoa that is mediated by OGG1. J Cell Sci. 2013;126:1488–97.
Ji Y, Azuine RE, Zhang Y, Hou W, Hong X, Wang G, et al. Association of cord plasma biomarkers of in utero acetaminophen exposure with risk of attention-deficit/hyperactivity disorder and autism spectrum disorder in childhood. JAMA Psychiatry. 2020;77:180–9.
Sahin M, Sur M. Genes, circuits, and precision therapies for autism and related neurodevelopmental disorders. Science. 2015;350:aab3897.
Schaaf CP, Betancur C, Yuen RKC, Parr JR, Skuse DH, Gallagher L, et al. A framework for an evidence-based gene list relevant to autism spectrum disorder. Nat Rev Genet. 2020;21:367–76.
Pinto D, Pagnamenta AT, Klei L, Anney R, Merico D, Regan R, et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature. 2010;466:368–72.
Levy D, Ronemus M, Yamrom B, Lee Y-h, Leotta A, Kendall J, et al. Rare de novo and transmitted copy-number variation in autistic spectrum disorders. Neuron. 2011;70:886–97.
O’Roak BJ, Deriziotis P, Lee C, Vives L, Schwartz JJ, Girirajan S, et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat Genet. 2011;43:585–89.
Sanders SJ, Ercan-Sencicek AG, Hus V, Luo R, Murtha MT, Moreno-De-Luca D, et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron. 2011;70:863–85.
Iossifov I, Ronemus M, Levy D, Wang Z, Hakker I, Rosenbaum J, et al. De novo gene disruptions in children on the autistic spectrum. Neuron. 2012;74:285–99.
Y-h Jiang, Yuen Ryan KC, Jin X, Wang M, Chen N, Wu X, et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. A J Hum Genet. 2013;93:249–63.
Dong S, Walker Michael F, Carriero Nicholas J, DiCola M, Willsey AJ, Ye Adam Y, et al. De novo insertions and deletions of predominantly paternal origin are associated with autism spectrum disorder. Cell Rep. 2014;9:16–23.
O’Roak BJ, Stessman HA, Boyle EA, Witherspoon KT, Martin B, Lee C, et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nat Commun. 2014;5:5595.
Krumm N, Turner TN, Baker C, Vives L, Mohajeri K, Witherspoon K, et al. Excess of rare, inherited truncating mutations in autism. Nat Genet. 2015;47:582–8.
Sanders STJ, He X, Willsey AJ, Ercan-Sencicek AG, Samocha Kaitlin E, Cicek AE, et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron. 2015;87:1215–33.
Turner TN, Coe BP, Dickel DE, Hoekzema K, Nelson BJ, Zody MC, et al. Genomic patterns of de novo mutation in simplex autism. Cell. 2017;171:710–22.e712.
An J-Y, Lin K, Zhu L, Werling DM, Dong S, Brand H, et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder. Science. 2018;362:eaat6576.
Gi M, Fujioka M, Kakehashi A, Okuno T, Masumura K, Nohmi T, et al. In vivo positive mutagenicity of 1,4-dioxane and quantitative analysis of its mutagenicity and carcinogenicity in rats. Arch Toxicol. 2018;92:3207–21.
Itoh S, Hattori C. In vivo genotoxicity of 1,4-dioxane evaluated by liver and bone marrow micronucleus tests and Pig-a assay in rats. Mutat Res Genet Toxicol Environ Mutagen. 2019;837:8–14.
Sanchez AB, Garcia CCM, Freitas FP, Batista GL, Lopes FS, Carvalho VH, et al. DNA adduct formation in the lungs and brain of rats exposed to low concentrations of [13C2]-acetaldehyde. Chem Res Toxicol. 2018;31:332–9.
Zhang X, Duan H, Gao F, Li Y, Huang C, Niu Y, et al. Increased micronucleus, nucleoplasmic bridge, and nuclear bud frequencies in the peripheral blood lymphocytes of diesel engine exhaust-exposed workers. Toxicol Sci. 2014;143:408–17.
Liu B, Song L, Zhang L, Wu M, Wang L, Cao Z, et al. Prenatal aluminum exposure is associated with increased newborn mitochondrial DNA copy number. Environ Pollut. 2019;252:330–5.
D’Souza SP, Vijayalaxmi KK, Naik P. Assessment of genotoxicity of aluminium acetate in bone marrow, male germ cells and fetal liver cells of Swiss albino mice. Mutat Res Genet Toxicol Environ Mutagen. 2014;766:16–22.
Kopp B, Zalko D, Audebert M. Genotoxicity of 11 heavy metals detected as food contaminants in two human cell lines. Environ Mol Mutagen. 2018;59:202–10.
Demir E, Qin T, Li Y, Zhang Y, Guo X, Ingle T, et al. Cytotoxicity and genotoxicity of cadmium oxide nanoparticles evaluated using in vitro assays. Mutat Res Genet Toxicol Environ Mutagen. 2020;850-851:503149.
Lehti V, Hinkka-Yli-Salomäki S, Cheslack-Postava K, Gissler M, Brown AS, Sourander A. The risk of childhood autism among second-generation migrants in Finland: a case–control study. BMC Pediatr. 2013;13:171.
Nguyen DT, Nguyen HH, Nguyen TD, Nguyen TTH, Nakano K, Maejima K, et al. Whole genome sequencing of a Vietnamese family from a dioxin contamination hotspot reveals novel variants in the son with undiagnosed intellectual disability. Int J Environ Res Public Health. 2018;15:2629.
Ton ND, Nakagawa H, Ha NH, Duong NT, Nhung VP, Hien LTT, et al. Whole genome sequencing and mutation rate analysis of trios with paternal dioxin exposure. Hum Mutat. 2018;39:1384–92.
Lan A, Kalimian M, Amram B, Kofman O. Prenatal chlorpyrifos leads to autism-like deficits in C57Bl6/J mice. Environ Health. 2017;16:43.
Mužinić V, Ramić S, želježić D. Chromosome missegregation and aneuploidy induction in human peripheral blood lymphocytes in vitro by low concentrations of chlorpyrifos, imidacloprid and α-cypermethrin. Environ Mol Mutagen. 2019;60:72–84.
Salazar-Arredondo E, Solís-Heredia MdJ, Rojas-García E, Hernández-Ochoa I, Quintanilla-Vega B. Sperm chromatin alteration and DNA damage by methyl-parathion, chlorpyrifos and diazinon and their oxon metabolites in human spermatozoa. Reprod Toxicol. 2008;25:455–60.
Neale BM, Kou Y, Liu L, Ma’ayan A, Samocha KE, Sabo A, et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature. 2012;485:242–5.
Yang F, Chen J, Miao M-H, Yuan W, Li L, Liang H, et al. Risk of autism spectrum disorder in offspring following paternal use of selective serotonin reuptake inhibitors before conception: a population-based cohort study. BMJ Open. 2017;7:e016368.
Battal D, Aktas A, Sungur MA, Kadioglu E, Derici Eker E, Sahin NO, et al. In vivo genotoxicity assessment of sertraline by using alkaline comet assay and the cytokinesis-block micronucleus assay. Basic Clin Pharm Toxicol. 2013;113:339–46.
Alzahrani HA. Sister chromatid exchanges and sperm abnormalities produced by antidepressant drug fluoxetine in mouse treated in vivo. Eur Rev Med Pharm Sci. 2012;16:2154–61.
Attia SM, Bakheet SA. Citalopram at the recommended human doses after long-term treatment is genotoxic for male germ cell. Food Chem Toxicol. 2013;53:281–5.
de Freitas TAB, Palazzo RP, de Andrade FM, Reichert CL, Pechansky F, Kessler F, et al. Genomic instability in human lymphocytes from male users of crack cocaine. Int J Environ Res Public Health. 2014;11:10003–15.
Yujra VQ, Moretti EG, Claudio SR, Silva MJD. Oliveira Fd, Oshima CTF et al. Genotoxicity and mutagenicity induced by acute crack cocaine exposure in mice. Drug Chem Toxicol. 2016;39:388–91.
Lewis L, Borowa-Mazgaj B, de Conti A, Chappell GA, Luo Y-S, Bodnar W, et al. Population-based analysis of DNA damage and epigenetic effects of 1,3-butadiene in the mouse. Chem Res Toxicol. 2019;32:887–98.
Jaiswal PK, Srivastava S, Gupta J, Thakur IS. Dibenzofuran induces oxidative stress, disruption of trans-mitochondrial membrane potential (ΔΨm) and G1 arrest in human hepatoma cell line. Toxicol Lett. 2012;214:137–44.
Gwinn MR, Johns DO, Bateson TF, Guyton KZ. A review of the genotoxicity of 1,2-dichloroethane (EDC). Mutat Res Rev Mutat Res. 2011;727:42–53.
Kalkbrenner AE, Daniels JL, Chen J-C, Poole C, Emch M, Morrissey J. Perinatal exposure to hazardous air pollutants and autism spectrum disorders at age 8. Epidemiology. 2010;21:631–41.
Talbott EO, Marshall LP, Rager JR, Arena VC, Sharma RK, Stacy SL. Air toxics and the risk of autism spectrum disorder: the results of a population based case–control study in southwestern Pennsylvania. Environ Health. 2015;14:80.
Everatt R, Slapšytė G, Mierauskienė J, Dedonytė V, Bakienė L. Biomonitoring study of dry cleaning workers using cytogenetic tests and the comet assay. J Occup Environl Hyg. 2013;10:609–21.
Toyooka T, Yanagiba Y, Ibuki Y, Wang R-S. Trichloroethylene exposure results in the phosphorylation of histone H2AX in a human hepatic cell line through cytochrome P450 2E1-mediated oxidative stress. J Appl Toxicol. 2018;38:1224–32.
Loch-Caruso R, Hassan I, Harris SM, Kumar A, Bjork F, Lash LH. Trichloroethylene exposure in mid-pregnancy decreased fetal weight and increased placental markers of oxidative stress in rats. Reprod Toxicol. 2019;83:38–45.
Fahmy MA, Hassan NHA, Farghaly AA, Hassan EES. Studies on the genotoxic effect of beryllium chloride and the possible protective role of selenium/vitamins A, C and E. Mutat Res Genet Toxicol Environ Mutagen. 2008;652:103–11.
Muller CD, Garcia SC, Brucker N, Goethel G, Sauer E, Lacerda LM, et al. Occupational risk assessment of exposure to metals in chrome plating workers. Drug Chem Toxicol. 2020;27:1–8.
Fu S-C, Liu J-M, Lee K-I, Tang F-C, Fang K-M, Yang C-Y, et al. Cr(VI) induces ROS-mediated mitochondrial-dependent apoptosis in neuronal cells via the activation of Akt/ERK/AMPK signaling pathway. Toxicol Vitr. 2020;65:104795.
Dickerson AS, Rahbar MH, Han I, Bakian AV, Bilder DA, Harrington RA, et al. Autism spectrum disorder prevalence and proximity to industrial facilities releasing arsenic, lead or mercury. Sci Total Environ. 2015;536:245–51.
Hill DS, Cabrera R, Wallis Schultz D, Zhu H, Lu W, Finnell RH, et al. Autism-like behavior and epigenetic changes associated with autism as consequences of in utero exposure to environmental pollutants in a mouse model. Behav Neurol. 2015;2015:426263.
Hossain S, Bhowmick S, Jahan S, Rozario L, Sarkar M, Islam S, et al. Maternal lead exposure decreases the levels of brain development and cognition-related proteins with concomitant upsurges of oxidative stress, inflammatory response and apoptosis in the offspring rats. NeuroToxicology. 2016;56:150–58.
Abudayyak M, Guzel E, Özhan G. Nickel oxide nanoparticles are highly toxic to SH-SY5Y neuronal cells. Neurochem Int. 2017;108:7–14.
Lou J, Jin L, Wu N, Tan Y, Song Y, Gao M, et al. DNA damage and oxidative stress in human B lymphoblastoid cells after combined exposure to hexavalent chromium and nickel compounds. Food Chem Toxicol. 2013;55:533–40.
Ji WD, Chen JK, Lu JC, Wu ZL, Yi F, Feng SM. Alterations of FHIT gene and P16 gene in nickel transformed human bronchial epithelial cells. Biomed Environ Sci. 2006;19:277–84.
Mohamed AA-R, Abdellatief SA, Khater SI, Ali H, Al-Gabri NA. Fenpropathrin induces testicular damage, apoptosis, and genomic DNA damage in adult rats: Protective role of camel milk. Ecotoxicol Environ Saf. 2019;181:548–58.
Martínez M-A, Lopez-Torres B, Rodríguez J-L, Martínez M, Maximiliano J-E, Martínez-Larrañaga M-R, et al. Toxicologic evidence of developmental neurotoxicity of type II pyrethroids cyfluthrin and alpha-cypermethrin in SH-SY5Y cells. Food Chem Toxicol. 2020;137:111173.
Braun JM, Kalkbrenner AE, Just AC, Yolton K, Calafat AM, Sjödin A, et al. Gestational exposure to endocrine-disrupting chemicals and reciprocal social, repetitive, and stereotypic behaviors in 4- and 5-year-old children: the HOME study. Environ Health Perspect. 2014;122:513–20.
Dong D, Yu J, Wu Y, Fu N, Villela NA, Yang P. Maternal diabetes triggers DNA damage and DNA damage response in neurulation stage embryos through oxidative stress. Biochem Biophys Res Commun. 2015;467:407–12.
Schmidt RJ, Iosif A-M, Guerrero Angel E, Ozonoff S. Association of maternal prenatal vitamin use with risk for autism spectrum disorder recurrence in young siblings. JAMA Psychiatry. 2019;76:391–8.
Wiegersma AM, Dalman C, Lee BK, Karlsson H, Gardner RM. Association of prenatal maternal anemia with neurodevelopmental disorders. JAMA Psychiatry. 2019;76:1–12.
Toblli JE, Cao G, Oliveri L, Angerosa M. Effects of iron deficiency anemia and its treatment with iron polymaltose complex in pregnant rats, their fetuses and placentas: oxidative stress markers and pregnancy outcome. Placenta. 2012;33:81–87.
Zaka-Ur-Rab Z, Adnan M, Ahmad SM, Islam N. Effect of oral iron on markers of oxidative stress and antioxidant status in children with iron deficiency anaemia. J Clin Diagn Res. 2016;10:SC13–19.
Jetten MJA, Gaj S, Ruiz-Aracama A, de Kok TM, van Delft JHM, Lommen A, et al. ‘Omics analysis of low dose acetaminophen intake demonstrates novel response pathways in humans. Toxicol Appl Pharm. 2012;259:320–8.
Smarr MM, Kannan K, Chen Z, Kim S, Buck Louis GM. Male urinary paracetamol and semen quality. Andrology. 2017;5:1082–8.
Narita M, Oyabu A, Imura Y, Kamada N, Yokoyama T, Tano K, et al. Nonexploratory movement and behavioral alterations in a thalidomide or valproic acid-induced autism model rat. Neurosci Res. 2010;66:2–6.
Lee CJJ, Gonçalves LL, Wells PG. Embryopathic effects of thalidomide and its hydrolysis products in rabbit embryo culture: evidence for a prostaglandin H synthase (PHS)-dependent, reactive oxygen species (ROS)-mediated mechanism. FASEB J. 2011;25:2468–83.
Wani TH, Chakrabarty A, Shibata N, Yamazaki H, Guengerich FP, Chowdhury G. The dihydroxy metabolite of the teratogen thalidomide causes oxidative DNA damage. Chem Res Toxicol. 2017;30:1622–8.
Costa S, Costa C, Madureira J, Valdiglesias V, Teixeira-Gomes A, Guedes de Pinho P, et al. Occupational exposure to formaldehyde and early biomarkers of cancer risk, immunotoxicity and susceptibility. Environ Res. 2019;179:108740.
U.S. Environmental protection agency. toxicological review of quinoline. In: Agency USEP (ed). Washington, DC; 2001. CAS No. 91-22-5.
Zhang W, Liu Y, An Z, Huang D, Qi Y, Zhang Y. Mediating effect of ROS on mtDNA damage and low ATP content induced by arsenic trioxide in mouse oocytes. Toxicol Vitr. 2011;25:979–84.
Govindaraju M, Shekar HS, Sateesha SB, Vasudeva Raju P, Sambasiva Rao KR, Rao KSJ, et al. Copper interactions with DNA of chromatin and its role in neurodegenerative disorders. J Pharm Anal. 2013;3:354–9.
Alimba CG, Dhillon V, Bakare AA, Fenech M. Genotoxicity and cytotoxicity of chromium, copper, manganese and lead, and their mixture in WIL2-NS human B lymphoblastoid cells is enhanced by folate depletion. Mutat Res Genet Toxicol Environ Mutagen. 2016;798–799:35–47.
Stringari J, Nunes AK, Franco JL, Bohrer D, Garcia SC, Dafre AL, et al. Prenatal methylmercury exposure hampers glutathione antioxidant system ontogenesis and causes long-lasting oxidative stress in the mouse brain. Toxicol Appl Pharm. 2008;227:147–54.
Robitaille S, Mailloux RJ, Chan HM. Methylmercury alters glutathione homeostasis by inhibiting glutaredoxin 1 and enhancing glutathione biosynthesis in cultured human astrocytoma cells. Toxicol Lett. 2016;256:1–10.
Ni M, Li X, Yin Z, Jiang H, Sidoryk-Węgrzynowicz M, Milatovic D, et al. Methylmercury induces acute oxidative stress, altering Nrf2 protein level in primary microglial cells. Toxicol Sci. 2010;116:590–603.
Kašuba V, Milić M, Rozgaj R, Kopjar N, Mladinić M, Žunec S, et al. Effects of low doses of glyphosate on DNA damage, cell proliferation and oxidative stress in the HepG2 cell line. Environ Sci Pollut Res. 2017;24:19267–81.
Zhang J-W, Xu D-Q, Feng X-Z. The toxic effects and possible mechanisms of glyphosate on mouse oocytes. Chemosphere. 2019;237:124435.
Roberts EM, English PB, Grether JK, Windham GC, Somberg L, Wolff C. Maternal residence near agricultural pesticide applications and autism spectrum disorders among children in the California Central Valley. Environ Health Perspect. 2007;115:1482–9.
Sebastian R, Raghavan SC. Induction of DNA damage and erroneous repair can explain genomic instability caused by endosulfan. Carcinogenesis. 2016;37:929–40.
Xu D, Liang D, Guo Y, Sun Y. Endosulfan causes the alterations of DNA damage response through ATM-p53 signaling pathway in human leukemia cells. Environ Pollut. 2018;238:1048–55.
Ponsonby A-L, Symeonides C, Saffery R, Mueller JF, O’Hely M, Sly PD, et al. Prenatal phthalate exposure, oxidative stress-related genetic vulnerability and early life neurodevelopment: a birth cohort study. NeuroToxicology. 2020;80:20–28.
Lee W, Cho J-H, Lee Y, Lee S, Kim DH, Ha S, et al. Dibutyl phthalate impairs neural progenitor cell proliferation and hippocampal neurogenesis. Food Chem Toxicol. 2019;129:239–48.
Goodrich AJ, Volk HE, Tancredi DJ, McConnell R, Lurmann FW, Hansen RL, et al. Joint effects of prenatal air pollutant exposure and maternal folic acid supplementation on risk of autism spectrum disorder. Autism Res. 2018;11:69–80.
Wang T-C, Song Y-S, Wang H, Zhang J, Yu S-F, Gu Y-E, et al. Oxidative DNA damage and global DNA hypomethylation are related to folate deficiency in chromate manufacturing workers. J Hazard Mater. 2012;213-214:440–6.
Unnikrishnan A, Prychitko TM, Patel HV, Chowdhury ME, Pilling AB, Ventrella-Lucente LF, et al. Folate deficiency regulates expression of DNA polymerase β in response to oxidative stress. Free Radic Biol Med. 2011;50:270–80.
Magnusson C, Lundberg M, Lee BK, Rai D, Karlsson H, Gardner R, et al. Maternal vitamin D deficiency and the risk of autism spectrum disorders: population-based study. BJPsych Open. 2016;2:170–2.
O’Callaghan-Gordo C, Kogevinas M, Fthenou E, Pedersen M, Espinosa A, Chalkiadaki G, et al. Vitamin D insufficient levels during pregnancy and micronuclei frequency in peripheral blood T lymphocytes mothers and newborns (Rhea cohort, Crete). Clin Nutr. 2017;36:1029–35.
Kopjar N, Fuchs N, Žunec S, Mikolić A, Micek V, Kozina G, et al. DNA damaging effects, oxidative stress responses and cholinesterase activity in blood and brain of wistar rats exposed to Δ(9)-tetrahydrocannabinol. Molecules. 2019;24:1560.
Russo C, Ferk F, Mišík M, Ropek N, Nersesyan A, Mejri D, et al. Low doses of widely consumed cannabinoids (cannabidiol and cannabidivarin) cause DNA damage and chromosomal aberrations in human-derived cells. Arch Toxicol. 2019;93:179–88.
Rubenstein E, Young JC, Croen LA, DiGuiseppi C, Dowling NF, Lee L-C, et al. Brief report: maternal opioid prescription from preconception through pregnancy and the odds of autism spectrum disorder and autism features in children. J Autism Dev Disord. 2019;49:376–82.
Rezaei M, Khodaei F, Sayah Bargard M, Abasinia M. Long-term methadone intake and genotoxicity in addicted patients. Jundishapur J Nat Pharm Prod. 2015;10:e17008.
Akhigbe R, Ajayi A. Testicular toxicity following chronic codeine administration is via oxidative DNA damage and up-regulation of NO/TNF-α and caspase 3 activities. PloS ONE. 2020;15:e0224052.
Zhao H, Wang Q, Yan T, Zhang Y, Xu H-J, Yu H-P, et al. Maternal valproic acid exposure leads to neurogenesis defects and autism-like behaviors in non-human primates. Transl Psychiatry. 2019;9:267.
Hamzawy MA, El-Ghandour YB, Abdel-Aziem SH, Ali ZH. Leptin and camel milk abate oxidative stress status, genotoxicity induced in valproic acid rat model of autism. Int J Immunopathol Pharm. 2018;32:2058738418785514.
Acknowledgements
MAB is supported by a Senior Research Fellowship (SRF B) from the National Health and Medical Research Council of Australia. ZH is supported by a research grant funded by the National Health and Medical Research Council of Australia (APP1146644). KP is supported by an Australian Government Research Training Program (RTP) Scholarship. SWS is the GlaxoSmithKline Chair in Genome Sciences at the Hospital for Sick Children and University of Toronto.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pugsley, K., Scherer, S.W., Bellgrove, M.A. et al. Environmental exposures associated with elevated risk for autism spectrum disorder may augment the burden of deleterious de novo mutations among probands. Mol Psychiatry 27, 710–730 (2022). https://doi.org/10.1038/s41380-021-01142-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-021-01142-w
This article is cited by
-
Autism spectrum disorders and the gastrointestinal tract: insights into mechanisms and clinical relevance
Nature Reviews Gastroenterology & Hepatology (2024)
-
A Comprehensive Review on Plant-Based Medications and Chemical Approaches for Autism Spectrum Disorders (ASDs) Psychopharmacotherapy
Indian Journal of Microbiology (2024)
-
Neuroimaging genetics approaches to identify new biomarkers for the early diagnosis of autism spectrum disorder
Molecular Psychiatry (2023)
-
Bisphenol A contamination in aquatic environments: a review of sources, environmental concerns, and microbial remediation
Environmental Monitoring and Assessment (2023)
-
Attitudes among parents of persons with autism spectrum disorder towards information about genetic risk and future health
European Journal of Human Genetics (2022)