Abstract
Machine and deep learning methods, today’s core of artificial intelligence, have been applied with increasing success and impact in many commercial and research settings. They are powerful tools for large scale data analysis, prediction and classification, especially in very data-rich environments (“big data”), and have started to find their way into medical applications. Here we will first give an overview of machine learning methods, with a focus on deep and recurrent neural networks, their relation to statistics, and the core principles behind them. We will then discuss and review directions along which (deep) neural networks can be, or already have been, applied in the context of psychiatry, and will try to delineate their future potential in this area. We will also comment on an emerging area that so far has been much less well explored: by embedding semantically interpretable computational models of brain dynamics or behavior into a statistical machine learning context, insights into dysfunction beyond mere prediction and classification may be gained. Especially this marriage of computational models with statistical inference may offer insights into neural and behavioral mechanisms that could open completely novel avenues for psychiatric treatment.
Similar content being viewed by others
Introduction
In recent years, artificial neural networks (NN) have become a huge success story in artificial intelligence (AI) research, achieving human to super-human performance in many domains in which more traditional AI approaches, much based on symbolic information processing and logical inference [1,2,3,4], had failed or progressed slowly for many decades. These include areas like visual object, pattern, and scene recognition [5,6,7], natural language processing [8, 9], video game playing [10], to cognitively challenging board games like Go [11] or goal-directed planning tasks [12]. Some of these achievements are particularly impressing: In Go, for instance, the combinatorial explosion of potential moves is much more severe than in chess; hence proficient Go playing relies much more on heuristic strategies rather than brute force scanning of potential trajectories of board constellations [11]. Some of these heuristic rules, accumulated by humans over centuries, it appears were rediscovered by NN-based algorithms within days without any human input other than the rules of the game [11]. Not surprisingly, therefore, NN-based algorithms have found their way into many everyday products, industrial and medical applications that require recognition of visual objects or scenes, spoken or written language, or prediction of outcomes, future events, or subject characteristics based on sensory or other types of input data [8].
A hallmark feature of NN-based systems is that they can learn and adapt: They consist of (1) a network architecture which describes the “anatomical” layout of the system and how its processing units, the artificial ‘neurons’, are wired (Fig. 1); (2) a loss or optimization function which specifies the overall goals of the learning process, and (3) a “training algorithm” which iteratively changes parameters of the NN, like the connection strengths between units, such that the target function is ultimately optimized based on the inputs the NN receives. The idea of artificial NNs as a mathematical formalization of nervous system activity for the purpose of computation reaches back at least to work of McCulloch and Pitts [13] and Alan Turing [14] in the forties of the last century. Later in the fifties and early sixties Frank Rosenblatt, an American psychologist, devised the “Perceptron” as a simple adaptive model of perception [15]. The perceptron consists of a sensory surface (or sheet of input neurons; Fig. 1a) connected to one or more output units which are supposed to assign the input pattern to one of several classes, e.g., for the purpose of recognizing handwritten letters. To achieve correct classification, or mapping of input patterns to output classes, Rosenblatt formulated a learning rule which iteratively adapts the connection weights between the input and output units upon each presentation of a training sample such that the actual outputs move closer and closer to the desired outputs. Formally, the learning rule acts to minimize the sum of squared deviations between actual and desired outputs. Classification, i.e., assignment of input patterns (like bundles of symptoms) to output categories (like a medical diagnosis), remains one of the major objectives of NN applications.
The perceptron and linear classification problems. a The perceptron, devised by Frank Rosenblatt [15], was one of the first feed-forward neural networks (NNs). It consists of a layer of input units (at which inputs or “predictors” xi are presented) which project to one (or several) output unit(s) with connection weights wi. The output unit forms a weighted sum of the inputs and compares it to a threshold, as indicated: If the weighted sum is larger, the input is assigned to one class, otherwise to the other class—as illustrated for a two-dimensional feature space with vector points color-coded according to class membership. Several output units would allow to represent multi-class problems. Formally, the weighted sum is a linear combination of the input features, and hence the surface separating the classes will always be a line, plane, or hyper-plane for a perceptron. b Graph shows a noisy version of the “XOR-problem”, with the strict logical version represented as a Boolean “truth table” on the right. In this case, the classes are such that they cannot be separated by a linear classifier (a line as indicated) as represented by the perceptron. c Adding one “hidden unit” in between the input layer and the output unit of the perceptron solves the XOR problem. Numbers in the graph denote the connection weights and activation thresholds: Due to its higher threshold (θ = 1.5), the hidden unit only activates once both input units are on (x1 = x2 = 1) and turns off the output unit by its large inhibitory weight. Thus, this NN implements the truth table on the left
In the early days, and continuing well into the nineties, NN research met strong and prominent skepticism to outright opposition within the areas of AI and cognitive science [1, 16]. This was due in part to the “cognitive revolution” [17, 18] and early success of formal, logic-based programming languages, which favored systems which explicitly manipulated strings of symbols through sets of syntactical (grammatical) rules [4]. NN research was particularly curbed by the book “Perceptrons” by Marvin Minsky, one of the “godfathers” of AI, and Seymour Papert [19], who identified and worked out severe computational limitations of the perceptron (with a prominent example illustrated in Fig. 1b). Later [20] it was recognized that these limitations could be overcome by inserting one to several so-called “hidden layers” of units between the input and the output stage (Fig. 1c). In fact, a famous and remarkable mathematical theorem (due to Cybenko [21], and Funahashi [22]), called the “universal approximation theorem”, states that with just one layer of hidden units a feedforward NN could essentially achieve any desired mapping between sets of input and output patterns. While this is just a statement about the expressive power of such networks, a famous learning rule which enabled training NNs across several such hidden layers, dubbed “back-propagation” and popularized by David Rumelhart, Geoffrey Hinton, and Ronald Williams [20], contributed to the ‘second wave’ of NN research in the eighties [23]. The idea is that any input pattern is first propagated forward through the network to the output stage, where the actual output is compared to the desired (teacher) output, and an error signal proportional to the mismatch between these two is then propagated back through the network for adjusting the weights between each pair of layers.
However, training a network across several hidden layers in practice proved to be very hard, mainly for a problem now known as the one of “vanishing or exploding gradients” [24,25,26]. As the error signal is back-propagated, it tends to either die out or blow up exponentially across successive network layers, for simple mathematical reasons. How to prevent this is actually still a hot topic in NN research [27,28,29]. This fact about training, and the above mentioned theorems assuring us that more than one hidden layer is—formally speaking—not actually needed, may have hampered NN research exploring NN architectures with very many hidden layers. It was again the group by Geoffrey Hinton [30, 31] who helped triggering the “third wave” of NN research by showing how training large structures could be achieved via layer-by-layer pre-training (Fig. 2d). NNs with very many hidden layers are what are called “deep NNs” these days (Fig. 2b), and efficient procedures for pre-training and initializing single layer connections, the strong rise in affordable parallel (mainly GPU-based) computing facilities, and the availability of large amounts of data (“big data”), is what makes training such networks (i.e., deep learning) efficient and feasible. There are also other reasons to their success, like their propensity to build up more and more elaborated representations across successive layers, as discussed below.
Different neural network (NN) architectures and activation functions. a “NN representation” of a multivariate General Linear Model (GLM): In a GLM, each output is computed from a weighted sum of the input features (or predictors, regressors), as highlighted for the first unit, plus some bias term w0i. A GLM may thus be seen as a simple NN with linear activation function. Most statistical regression and classification models could be brought into this NN form. b In a true NN, each unit still computes a weighted sum of the inputs from the previous layer and adds a bias term, but the unit’s activation function ϕ is nonlinear, often a monotonically increasing sigmoid function as indicated by the blue curves. There is no formal definition of the term “deep”, but often networks with 4 or more hidden layers are considered deep, while networks with just one (as in Fig. 1c) or two are termed “shallow”. Activity is propagated forward from the input layer through all hidden layers until it reaches the output stage. c In a recurrent NN (RNN), in contrast to a pure feedforward NN (FNN), also feedback connections between units are allowed. Although this may seem like a minor addition to the FNN architecture, formally RNNs become dynamical systems that can recognize temporally extended input sequences (like spoken or written text), can have internal states, or autonomously produce output sequences. d Auto-encoder NN (AE-NN) often used for pre-training (initializing) single layers of deep NNs (as in b). An AE-NN is supposed to reproduce as output (using decoding weights Wdecode) its own input pattern (projected through encoding weights Wencode), thereby forming a lower-dimensional internal representation of the input within its hidden layer
Overview over neural network architectures and training procedures
NNs come in a variety of different designs and architectures derived from different principles, or conceived for different purposes. Most commonly, feedforward NNs (FNNs) are employed in current applications, that is networks where activity is propagated uni-directionally layer-by-layer from the input up to the output stage, with no feedback connections within or between layers (Figs. 2a, b and 3). In the simplest case, several layers may be connected in series in an all-to-all manner, i.e., with connections from each preceding layer unit to all units of the subsequent layer (Fig. 2b), called a multilayer perceptron (MLP). One way of pre-training MLPs is auto-encoders (AE) [31, 32], networks which build a compressed representation of the data by reconstructing the input at the output layer from a “down-sampled” version at an intermediate layer with (much) fewer units (Fig. 2d). Several such AEs may be stacked on top of each other, each receiving as input the compressed representation from the previous auto-encoder, until finally via several perceptron-type layers, or back-propagation through all layers, the mapping onto the ultimately desired output is achieved [32]. This way, efficient feature representations of the input space are iteratively constructed through the set of auto-encoding stages. Convolutional deep NNs (CNN, Fig. 3; reviewed in [5, 33, 34]) were originally designed in analogy with the primate visual system [5] and exploit spatial invariances in the data. They extract different feature maps from the input, each represented by a set of units with different “receptive fields”, i.e., tuned to a different spot of input space, but sharing the same set of weights. This “weight sharing” is an efficient principle to dramatically reduce the number of to-be-trained parameters, while at the same time allowing the system to recognize the same patterns across different spatial locations. Several such convolutional layers are usually alternated with dimensionality reduction (“pooling”) and nonlinear transformation stages in common CNN architectures. CNNs pretty much represent the state-of-the-art for visual object and scene recognition [6, 7].
Convolutional neural network (CNN). A CNN consists of four distinct processing stages: An input image (left) as, e.g., obtained from sMRI, is first processed by multiple feature maps, shown as black, blue and red neural sheets (layers) here, with examples of corresponding features illustrated on the left. Each feature unit has its own spatial receptive field, illustrated by the projections from the input image to the feature units, and different units from the same feature map share the same set of weights (formally, they perform a convolution of the image, indicated by the star (*), with a filter defined by the weights, hence the term CNN). The sum of weighted inputs to each feature unit is then passed through a nonlinear activation function (e.g., a rectified linear unit [ReLU] as illustrated). In a third step, each map is “down-sampled” through a pooling operation (e.g., by taking the maximum of the outputs from a set of neighboring units). These three processing stages may be repeated multiple times (making the network deep), until finally a set of fully connected layers is used to predict the probability for a given class such as a psychiatric disorder. The “Colin_27_T1_seg_MNI.nii” anatomical template available on http://brainmap.org/ale/ was used to create the input image
From a statistical point of view, FNNs are complex nonlinear regression or classification devices, in contrast to the commonplace linear regression models or classifiers implemented in statistical packages. In a typical regression problem, the model is trained with a set of pairs of input (regressor or predictor) patterns and output (target or response) patterns. By training one means that parameters of the system, like the connection (or regression) weights, are adapted such as to minimize (resp. maximize) a loss function like the mean sum of squared errors [MSE] between desired and actual outputs, or the likelihood function (Fig. 4). While in a regression approach the outputs are usually continuously valued (i.e., real numbers), in classification we are dealing with categorical outputs, and the goal is commonly to adapt parameters such that the probability of observing the correct class given an instance from that class is maximized (cf. Fig. 1). Linear models have dominated statistics for so many decades because they have a number of highly desirable properties: The associated loss or likelihood functions can usually be solved fast, analytically, and efficiently in just one step, and these solutions are unique and represent a global optimum (Fig. 4a). Furthermore, model fitting works with relatively small sample sizes, and statistical testing on these models is well understood. In contrast, NN regressors or classifiers need to be solved numerically, with time-consuming iterative schemes like back-propagation. Usually they have complicated loss functions with very many local minima (Fig. 4b) and other intricacies that make finding a global optimum nearly impossible [33, 35]. Moreover, they require large amounts of data for training their many parameters [36, 37]. In general, further measures, called regularization approaches, are necessary to prevent over-fitting which would come with poor generalization to new observations [36, 37]. Yet, when sufficient data are available, NNs unlike linear statistical models can discover complex patterns in the data, can form highly predictive nonlinear feature combinations, and can build abstract high-level representations of the data. That is where their unique power lies, and what makes them superior to more traditional statistical approaches if computational and data resources permit.
Training neural networks (NNs). For training, NNs are presented with a set of paired inputs (xi) and outputs (yi), as in any typical regression or classification problem. The NN (and a GLM or other statistical models as well, cf. Fig. 2a), can be understood as computing a prediction function \(\widehat {y_i}\)=fw(xi) of its inputs given its parameters w, and the goal of training is to optimize the parameters w of this function such that some loss or error function Err(w) is minimized (or optimized, more generally). One of the most common loss functions is “squared-error-loss”, the sum or mean (MSE) of the squared differences between truly observed and predicted outcomes. To this, often a “regularization term”, e.g., the sum of the squared weights, is added that can be regulated by some meta-parameter λ to prevent over-fitting to the data. a A “convex optimization problem” with a unique global minimum in the error function, as is the case, e.g., for the GLM. b A highly non-convex optimization problem, with numerous local minima and different regions of slope, as typical for highly nonlinear models like deep NNs. One common numerical scheme to “solve” this problem is “gradient descent”, where one iteratively updates parameter estimates \({\hat{\mathbf w}}_{n}\) moving against the gradient of the error function Err(w) with some learning rate γ. This procedure would, ideally, slide into the next minimum, but more elaborated schemes like stochastic gradient descent, adaptive-rate or second-order methods (which take second derivatives into account) are necessary to alleviate at least some of the problems
There are also NN architectures with feedback (“recurrent”) connections within or between layers, called recurrent neural networks (RNNs; Fig. 2c). Although, at first, including recurrent connections may seem like a minor change of architecture, mathematically this makes RNNs dynamical systems which can exhibit a full new range of behaviors. Just like FNNs are powerful nonlinear regressors or classifiers, RNNs are, in essence, nonlinear time series models. RNNs are even harder to train than FNNs [35, 38], but once trained, they can be run forward in time to produce predictions of future outcomes or states. One of the most powerful and state-of-the-art RNNs is called “Long Short-Term Memory” (LSTM), as it can efficiently solve problems with very long temporal dependencies [25]. Deep RNNs like LSTM networks [12, 25, 39] are a major vehicle these days for language and text processing tasks, including automatic sentence completion [8, 9] or topic inference models [40], or for modeling consumer behavior [41]. Deep RNNs can form complex internal models of the external world that enable sophisticated planning and problem solving [12, 42].
We confer the interested reader to Box 1 for more technical details on how to train neural networks, and how they relate to statistics.
Deep networks in psychiatric research and clinical practice
Statistical and machine learning (ML) techniques for nonlinear regression and classification, like support vector machines (SVM) and kernel methods [43], or shallow neural networks, have long been in place in psychiatry and neuroscience (see [44,45,46,47] for review). However, deep learning (DL) algorithms, on which this short review will focus, often outperform these earlier ML techniques by considerable margins [6, 48]. It is not yet fully understood why this is so. Part of the reason may be that deep neural networks (DNNs) can infer suitable high-level representations without much domain-specific knowledge and prior feature construction [49]. Recent advances in pre-training and transfer-training procedures also enabled to navigate their complex optimization landscapes more efficiently [33, 50]. Moreover, there may be fundamental computational reasons: For instance, the compositional capabilities and the space of “computable functions” grows much faster for deep than for shallow NNs with the number of parameters [49, 51].
Given the ability of DNNs to learn abstract representations from raw data [5, 49, 52] and their success in image and speech recognition [6, 9], DL methods have promptly found their way into (bio-)medical research and health care [53,54,55,56]. Big companies like IBM and Google are already harnessing DL and related algorithms to guide personalized medicine, e.g., IBM’s Watson (although not strictly a DNN) or Google’s DeepMind Health. DNNs are especially advancing medical fields which largely depend on image analysis such as tumor detection and segmentation [57]. This has raised hopes that DNNs may likewise assist in tackling open issues in psychiatry, such as reliable diagnostic decisions, predicting risk and disease trajectories to facilitate early and preemptive interventions, indicating the most effective personalized treatments, or discovering potential new drugs.
Diagnosis and prognosis based on neuroimaging data
So far, most studies employing DL in psychiatry have focused on diagnostics [56]. Computer aided diagnostic tools which classify mental illness could assist clinicians in forming more reliable, unbiased, and standardized diagnostic decisions across sites in less time. In general, however, diagnostic classification based on neuroimaging data is not an easy endeavor. A wealth of studies has looked into neuro-functional and structural abnormalities which discriminate psychiatric disease from health (mainly) on the basis of mass univariate statistics. One take-home from these studies is that alterations are often rather subtle and reliably detected only between groups [58], not at an individual level. Features and their statistical relationships (or their compositional structure) required to accurately classify single individuals are therefore likely to be more complex, potentially not even discernible within a single imaging modality [59]. On the other hand, cross-modal feature combinations and interactions are expected to be even harder to detect, as they may only materialize at very high (abstract) levels of analysis [60].
Deep NNs are particularly suited for these challenges as they efficiently capture higher-order statistical relationships [8, 33, 49], and thus learn to extract features with far less parameters than shallow architectures [49]. This is due to their multi-layered design, where highly complex and intricate nonlinear relations among input features could be extracted and represented by layers further up in the processing hierarchy. By rather seamlessly integrating complementary data sets obtained from multiple imaging modalities such as functional magnetic resonance imaging (fMRI), structural MRI (sMRI), and positron emission tomography (PET) (Fig. 5), DL-based systems could provide clinicians with valuable insights otherwise not immediately accessible. Moreover, their ability to directly work on raw neuroimaging data [61, 62], rather than on hand-selected and pre-selected features, could remove tedious and error-prone data preprocessing stages in the future.
Illustration of multi-modal integration in DNNs (inspired by Fig. 8 in Calhoun and Sui [59]). While lower layers of a DNN may represent modality-specific properties, higher layers may learn to represent complex feature combinations from different modalities (left). Right: In data space, similar to the XOR problem (Fig. 1b), data from a single modality may not easily allow to discriminate two different disease states, while a nonlinear combination from both modalities would
Accordingly, DNNs have shown convincing first results in classifying psychiatric disorders. Most studies have focused on diagnosing dementia [54, 63,64,65,66,67,68,69,70] (see [56] for older studies) and attention deficit hyperactivity disorder [71,72,73,74,75,76], most likely due to the accessibility of moderately large publically available neuroimaging data sets (e.g. ADNI, OASIS, and ADHD-200 databases). For these, often balanced accuracy levels well above 90% have been achieved [77,78,79,80] (see also [56] for an overview). Notably, a few of these studies also investigated the ability to predict disease trajectories such as the conversion from mild cognitive impairment (MCI) to Alzheimer’s disease (AD) [70] (see [81] for review), which is essential to detect disease at an early stage and prevent its progression. Studies classifying other mental disorders such as schizophrenia [60, 82,83,84,85,86], autism [87,88,89], Parkinson’s disease [80], depression [90], substance abuse disorder [91], and epilepsy [92, 93], are slowly accumulating as well.
ML algorithms fed with multimodal data, allowing them to harvest predictive inter-relationships among data types [59, 94, 95] (Fig. 5), also consistently outperform unimodal data in diagnostic decisions [84, 96,97,98]. Psychiatric symptoms are most likely a result of multiple etiological processes spanning many levels of computation in the nervous system [99]. Multimodal data, as e.g., obtained from neuroimaging and genomics, potentially provides complementary information on etiological mechanisms, such as insights into how genes shape structure, and how structure in turn implements function. While also more “traditional” classifiers like SVMs or discriminant analysis could be, and have been [100, 101], fed with features from multiple modalities, particularly informative and predictive cross-modal links may form specifically at deeper levels of complexity (cf. Fig. 5). Consistent with this idea, DNNs have been found to outperform shallow architectures when rendering diagnoses on the basis of multimodal data [69, 70, 84, 95]. As a concrete example, Lu and Popuri [70] used DNNs to fuse features obtained from sMRI, related to gray matter volume at different spatial scales, and fluorodeoxyglucose PET (FDG-PET) for assessing mean glucose metabolism, to predict progression to AD. Feature representations were first learned independently via stacked AEs (unsupervised pre-training), and then fused at a later stage with a DNN which took as input these lower-level representations and provided the probabilities for the two classes as output (see Fig. 5). The performance increases obtained in this study by merging modalities compared to single-modality DNNs may still seem relatively modest (<4%). The full potential of multi-modal DNNs may only unfold when larger sample sizes become available for which these architectures are most suited. Nevertheless, these studies highlight how algorithms which leverage the joint information available from multiple data sources may be helpful for arriving at a more complete characterization of the disease [59], especially since we often lack strong hypotheses on how data from different modalities may be related, such that strongly data-driven methods like DNNs may be of particular value.
However, based on the number of studies conducted so far, it is too early to say how factors such as type of disorder, DNN architecture and the specific input provided, or data modality affect classification performance. What can be said, however, is that deep architectures are able to achieve performance levels at least comparable to shallow ones [56], which is encouraging given that at times the latter already outperform experienced clinicians [102], and that sample sizes in neuroimaging are yet limited.
Predictions based on mobile phone data and large data bases
Rather than looking into (neuro)-biological data which are currently limited in terms of sample size, AI—specifically DL architectures—may prove particularly powerful in areas in which we already possess large and ever growing data sets such as electronic health records (EHRs), social media platforms, and ecological momentary assessments (EMA). DNNs have recently been successfully employed to predict medical diagnoses based on EHRs [103, 104], and could mine social media platforms, like “Reddit” or “Twitter”, for posts indicative of mental illness [66, 105].
Arguably the highest potential for AI may lie in identifying structure in data obtained from wearable devices like mobile phones and other sensors. Modern mobile-based sensor technologies, in principle, offer extraordinary possibilities to (passively) collect vast amounts of data in temporally highly resolved, ecologically valid, and yet unobtrusive settings. As mobile phones are by now with us almost the entire day, prepared for collecting and sensing a wide range of mental health dependent variables, the information we need for tracking mental well-being may, in principle, already be available to large degree. However, the sheer amount of collectable data, the challenges of fusing different modalities and sources, and the non-trivial temporal dependencies within them, call for learning algorithms which are extremely powerful and efficient in particular for time series data.
Features which could, in principle, be extracted from mobile phone usage and sensors, such as movement patterns and indicators of social interactions, derived, e.g., from GPS, calls, and text messages, have already proven to be predictive of mental health status [106,107,108,109,110,111]. For instance, deep architectures applied to smartphone data could successfully predict mental health related variables such as sleep quality or stress from physical activity [112,113,114]. They have also been used to monitor PD based on motor movements [115, 116], or to detect depressive states based on typing dynamics [90]. In this latter example, the authors collected meta-data related to typing duration, speed, and acceleration, and were able to accurately (>90%) classify depressive states in bipolar patients assessed weekly through the Hamilton Depression Rating Scale. Given sufficient typing sessions for training, their DNN even achieved high individual-subject-level predictions on single typing sessions, illustrating how these approaches may be harvested for personalized therapy. Particularly noteworthy in this context are also efforts of tracking dynamics and predicting upcoming (future) mental states. Suhara et al. [117] forecast severe depressive states based on individual histories of mood, behavioral logs, and sleep information using a LSTM architecture. This highlights how networks which are capable of learning long-term temporal dependencies from smartphone data could be used to predict future pathological mental states or risks thereof (Fig. 6 illustrates a processing pipeline for this type of approach). It is highly likely that such forecasting will improve if we find efficient ways to utilize the entire information available from sensor and user data, e.g., by integrating physiological, motor, environmental, and social information.
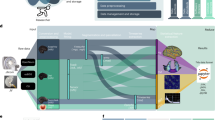
Schematic workflow for the potential application of RNNs in the context of mobile devices and sensors. Sensor readings and other meta-data from wearables and smartphones (box-1) may be used to extract mental health-related features in social, physical, physiological, and medical domains (box-2). The RNN could be trained to learn the temporal dependencies within and among these features (box-3). Based on these, it can perform ahead-predictions of, e.g., the onset of specific symptoms (or the risk thereof) and feed this information back to the patient in order to simply raise awareness, provide advice (e.g., to consult a doctor soon), or to suggest behavioral interventions (box-4). The illustration of sensor glasses in box 1 was inspired by Google Glasses
The advancement of technologies that assist in predicting state trajectories, including symptom onset or risk thereof, brings up unprecedented opportunities for affordable targeted interventions at early stages, or possibilities to evaluate treatments. As in the case of social media blogs, features which predict risk of mental illness or symptom onset could be used for specific feedbacks and interventions, inviting users to seek expert advice, follow practical exercises and treatments, or simply raise awareness [118]. Combining algorithms with transfer learning could further help to efficiently pre-train such models on a wide pool of user data, while fine-tuning could help to adapt treatments to the specific needs of individuals. Thus, possibilities in mobile applications seem endless, and RNN related architectures will likely play a crucial role. On the down side, such applications which process so rich, detailed, and sensitive personal data, obviously also come with profound ethical and security issues [119, 120]. Such data could potentially be exploited by insurers, lawyers and employers to form long-term judgments which cut an individual’s access to services, jobs, and benefits, with substantial implications for their personal lives. Perhaps even worse, these data could be misused for manipulating individuals and political processes as recently evidenced in the case about Cambridge Analytica. How to efficiently deal with such issues is currently an open problem.
Open issues and further challenges for AI in psychiatry
Low sample size issues and multi-modal integration
A major caveat with DL-based classification is that data in this domain are usually very high-dimensional relative to the typical sample sizes collected in neuroscientific studies. For a single imaging modality, current sample sizes are on the order of 102–104, which is modest relative to the number of voxels in the image (and thus the number of input parameters), leave alone multi-modal data for which the situation is much worse. In contrast, areas in which DNNs typically excel other ML methods and shallow networks, such as image recognition or speech analysis, consist of data bases with n > 106 [6, 121]. Training models with very many parameters on small sample sizes poses a severe challenge to finding solutions that will generalize well to the population [122] (termed “curse of dimensionality” or “p»n problems”, see, e.g., Ch. 18 in Hastie et al. [36]). A real breakthrough in (multi-modal) image classification and feature extraction will therefore possibly have to wait until larger samples, currently under way (e.g., [123]), have been collected and made available to machine-learners [124].
Until then, alternative strategies for dealing with this problem will have to be explored, such as strong regularization schemes (see “Overview over neural network architectures and training procedures” section), applying sparse architectures, reducing input dimensionality by, e.g., prior feature selection or dimensionality reduction techniques, or exploiting transfer learning [125]. For instance, specific types of deep architectures achieve good generalization performance by their “sparse” design. CNNs for example, which are particularly suited for processing raw imaging data since their architecture inherently recognizes translational and other invariances [5, 126], reduce the number of parameters through “weight sharing” (same set of weights for many units copied across space), their local connectivity, and pooling of unit responses [5] (see Fig. 3). Evidence indicates that CNNs outperform shallow architectures when predicting mental health related aspects from raw imaging data [62, 77, 79], while reaching at least comparable performance levels as shallow architectures when these were provided with pre-designed features [62, 80]. Other studies have reduced the dimensionality of the input data through prior feature selection. While such approaches may be less suitable for clinical applications, they yield agreeable results for small samples of n < 100 (e.g., [83, 87, 127]).
Alternatively, one may increase the effective “n” by transferring knowledge gained with one data set (or task) to another (termed transfer learning) [125]. The idea behind this approach is that since representations learned by DNNs develop from more general, data set unspecific features in early layers, to highly specific features in final network layers close to the output stage [50], one can use different (albeit sufficiently similar) data sets for training the network on the more general features, and then perform fine-tuning of selected layers on the actual data set in question. In the simplest case, this could mean pre-training a network on a given sample collected at one site, and fine-tuning it on another target sample from a different site [77]. In the case of models which are to make predictions on a single individual, for instance for reliably predicting seizure onset based on a limited amount of subject-specific EEG recordings, data from other epileptic patients have been applied to first learn general seizure representations, and then these pre-trained NNs were used as initialization for parameter fine-tuning on the subject-level [93]. Notably, since pre-training is commonly performed on unlabeled data first (e.g., through auto-encoders, Fig. 2d, cf [57].), i.e., is unsupervised and thus less bound to a particular target output [30, 128], transfer learning is not restricted to data obtained from the same type of sample and/or imaging modality. For example, Gupta et al. [129] pre-trained a CNN for dementia classification based on sMRI images through a sparse autoencoder (cf. Figure 2d) on natural images. Surprisingly, three-way classification results on AD, MCI, and HC were superior when pre-training was performed on random patches of natural rather than on structural MRI images (see also [130] for a similar procedure on speech data and posttraumatic stress disorder classification). The idea behind this is that these distinct data sets share basic structural statistics, and extracting patterns from one data set can therefore be leveraged for the other (see also [67, 79, 131, 132]).
From a different angle, training on multiple data sets obtained from different sites may actually be necessary to improve generalization to the overall population. It ensures that models do not learn to exploit site-specific nuisance variables predictive of, but not relevant to, a disorder, such as treatment or medication effects which could be related to differences in site-specific health care, medical supply, or other sample-specific properties [133].
Redefining diagnostic schemes
Psychotherapeutic and pharmacological treatments indicated through “conventional” diagnosis often fail to show effect, leaving a considerable proportion of psychiatric patients low or non-responsive. In part, this may be attributed to the fact that our current diagnostic systems base psychiatric diagnoses on the observation and duration of a list of descriptive and static symptoms, rather than considering functional deficits, etiological mechanisms, or dynamical aspects [134] of the disease. As a result, the same psychiatric diagnosis may refer to a very heterogeneous set of individuals with quite different functional deficits and in need of specifically tailored therapeutic interventions [135, 136]. In essence, current psychiatric diagnoses may often just be a coarse indicator of, but may not accurately capture, the underlying neural and psychological problem. This poses an inadvertent constraint on any prediction algorithm which is designed to imitate the diagnostician: In the currently most common supervised DL approaches, the quality of the expert-provided diagnostic labels used for training defines an upper bound on the system’s performance. If AI is to help guide personalized medicine, it needs to go beyond the mere prediction of symptoms by current diagnostic schemes, but rather has to help refining our diagnoses and their neurobiological underpinnings.
Several avenues are conceivable along which AI can assist in this effort. For one, the strength of DNNs in learning intricate relationships from data on their own, without much prior input from the researchers, could be exploited to extract novel biomarkers which may account for much more variation in an illness-related phenotype and signify well targeted interventions and treatments. Representations learned by different layers in a DNN hierarchy have been shown to sometimes yield interpretable features which are specifically altered in a given disorder [60, 83, 137]. Thus, although trained with traditional diagnostic labels, the extracted feature maps, and their specific differences and commonalities across the psychiatric spectrum, may help to refine nosological schemes. Another possible way forward is to omit diagnostic labels altogether and rather train the network to predict directly future outcomes like relapse times, hallucinatory episodes, mood assessments, performance in different functional domains as included in RDoC [135, 138], problems with family or at work, and others [90, 117, 139]. Alternatively, one could use unsupervised approaches for identifying novel, more natural and predictive demarcations in feature spaces spanned by symptom assessments, RDoC-type testing, imaging data, and other relevant information [140]. One such method is deep (stacked) AEs which are forced to build compact yet highly informative representations of the data by projecting them down into much lower-dimensional manifolds at deep layers, from which the inputs are ought to be reconstructed as faithfully as possible [32] (see Fig. 2d for the basic principle). Specific constraints on the AE architecture and optimization process may ensure that the deep-layer representations have certain desired properties and are interpretable.
As mentioned earlier, multi-modal architectures may be particularly important to gain insights into how different levels of analysis, such as genes, molecules, and networks, are linked. Since we currently largely lack concrete hypotheses on the precise interaction of these different levels, data-driven approaches uncovering higher order statistical dependencies are essential. In this regard, a potential limitation is that DNNs are often criticized for being a “black box”, with their inner workings not well understood. However, approaches for opening the black box are emerging (see next section), thus also addressing questions about accountability. Even if it is unclear how exactly a DNN achieved its performance, the trained NN of course remains a valuable tool for prediction. Even misclassifications can prove valuable in informing the user about potentially important contingencies that may have been missed, like that a drug could have yet unknown alternative targets [141], or—the other way around—that a patient may share more biological commonalities than expected with a disorder s/he was not originally been diagnosed with [133].
Other issues related to the interpretability of DNN models will be discussed in the next section.
Outlook: adding meaning to deep networks—tools for mechanistic insight
NN models, although originally inspired by processes in the nervous system (e.g., [142,143,144]), primarily serve as regression and classification tools in machine learning: They are applied to predict yet unobserved properties (like personality traits), category labels (like clinical diagnoses), or future trajectories (like prognoses of disease development) of human subjects or sensory objects. RNNs [12], or FNNs coupled with, e.g., reinforcement learning methods [10], also serve as active AI agents to carry out actions upon received histories of inputs. However, trained NN models are hard to interpret semantically in a specific clinical setting (but see [145, 146]), and are not commonly used yet as tools to gain insight into neural, psychiatric and behavioral mechanisms. Mechanistic insight beyond “mere” prediction may enable to design novel treatments and to identify optimal leveraging points for clinical interventions, and it may help to better connect neurobiological and pharmacological processes to their behavioral and psychiatric consequences.
There are two fundamental ways we could fill neural network models with “meaning”:
First, we could move them closer to biological reality. In fact, biophysical neural network models, with spiking neurons driven by ionic conductances and structural features derived from anatomical data, have a long tradition of their own in computational neuroscience (e.g., [143, 147, 148]). But only quite recently attempts have been started to infer such models more systematically through a loss function or statistically principled means from experimental data [149,150,151,152,153,154,155,156]. Such models, once trained, can often reproduce and predict physiological observations in quantitative detail [157, 158]. The trouble with such models is that they are much more complicated in structure and function than artificial NN models, and hence their training is even more tedious and computationally demanding than those of deep networks, without so far offering an apparent advantage from a pure data analysis point of view. As a potential way forward, biophysically derived mean-field models which “summarize” the behavior of larger populations of neurons [159,160,161,162,163] could provide an intermediate step for parameter estimation.
Another development to be mentioned in this context is computational models of behavior, like reinforcement learning models, which are statistically inferred from the data in a maximum likelihood or Bayesian sense [152, 164,165,166]. Like biophysical networks, these models are also phrased in terms of semantically directly interpretable quantities, in this case action values, choice probabilities, rewards, and the like. While these models allow for mechanistic insight and computationally based assessment of behavior [167], on their own they lack the computationally expressive power of DNNs (often they are linear, for instance), and are not designed to be general-purpose data-analytical tools that could, e.g., easily incorporate other data modalities. They could be coupled to DNNs, however [10, 11].
Second, instead of constructing biologically directly interpretable models, we could try to interpret the unit-activities and layer-activities in trained NNs in a biological and/or psychological context. For FNNs, several approaches have been established recently for probing and visualizing the representations constructed from the input data by successive hidden layers [168,169,170,171]. These approaches are sometimes similar to what in vivo electrophysiologists do, using specifically designed input stimuli to map out the “receptive fields” of hidden units [171]. This may yield interesting insights into the higher-level features the deep NN uses for predicting its targets. Indeed, CNNs trained on natural images have been shown to learn biologically plausible representations such as present in the ventral processing stream [145, 146, 172].
For RNNs, on the other hand, a natural framework for interpreting the activity evolution in these networks and relating them to brain processes is provided by dynamical systems theory [37, 162, 173, 174]. Mathematically, a RNN is a dynamical system, and as such will exhibit a range of phenomena like attractor states, oscillations (limit cycles), or chaos, that are found in neurophysiological activity as well, and that have long been assumed to be the underpinning of neural computation [174, 175]. In fact, most computational neuroscientists view neural information processing as fundamentally being implemented in terms of dynamical systems phenomena (for instance, working memory contents as attractor states [176], or decision making as noisy transitions between competing attractor states [177, 178]). RNNs may allow to extract this computationally relevant network dynamics directly from physiological recordings [173], even though their units and connection weights are mere abstractions of biophysical neurons and synapses.
In summary, although in past years NNs have been mainly used as sophisticated nonlinear tools for classification, regression, and prediction, a very exciting development is to employ them to also to gain insight into physiological, computational, and cognitive mechanisms.
References
Fodor JA, Pylyshyn ZW. Connectionism and cognitive architecture: a critical analysis. Cognition. 1988;28:3–71.
Minsky M, Papert SA. Artificial intelligence. Eugene: University of Oregan Press; 1972.
Minsky M. Semantic information processing. Cambridge: MIT Press; 1968.
Newell A, Simon HA. Human problem solving, vol. 104. Englewood Cliffs, NJ: Prentice-Hall; 1972.
LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436–44.
Krizhevsky A, Sutskever I, Hinton GE. Imagenet classification with deep convolutional neural networks. Proceedings of the Advances in Neural Information Processing Systems. 2012;1097–105.
Farabet C, Couprie C, Najman L, LeCun Y. Learning hierarchical features for scene labeling. IEEE Trans Pattern Anal Mach Intell. 2013;35:1915–29.
Schmidhuber J. Deep learning in neural networks: an overview. Neural Netw. 2015;61:85–117.
Graves A, Mohamed A-r, Hinton G. Speech recognition with deep recurrent neural networks. 2013 IEEE International Conference on Acoustics, Speech and Signal Processing. 2013;6645–9.
Mnih V, Kavukcuoglu K, Silver D, Rusu AA, Veness J, Bellemare MG, et al. Human-level control through deep reinforcement learning. Nature. 2015;518:529–33.
Silver D, Schrittwieser J, Simonyan K, Antonoglou I, Huang A, Guez A, et al. Mastering the game of go without human knowledge. Nature. 2017;550:354.
Graves A, Wayne G, Reynolds M, Harley T, Danihelka I, Grabska-Barwińska A, et al. Hybrid computing using a neural network with dynamic external memory. Nature. 2016;538:471.
McCulloch WS, Pitts W. A logical calculus of the ideas immanent in nervous activity. Bull Math Biophys. 1943;5:115–33.
Turing AM. Intelligent machinery, a heretical theory. National Physical Lab. Report. 1948.
Rosenblatt F. The perceptron: a probabilistic model for information storage and organization in the brain. Psychol Rev. 1958;65:386.
Gallistel CR, Gibbon J. The symbolic foundations of conditioned behavior. Hove: Psychology Press; 2002.
Miller GA. The magical number seven, plus or minus two: some limits on our capacity for processing information. Psychol Rev. 1956;63:81.
Miller GA, Galanter E, Pribram KH. Plans and the structure of behavior. New York: Adams Bannister Cox; 1986.
Minsky M, Papert S. Perceptrons: an introduction to computational geometry. Cambridge, MA: MIT Press; 1969.
Rumelhart DE, Hinton G, Williams RJ. Learning internal representations by error propagation. Parallel distributed processing: exploration in the microstructure of cognition, vol. 1. Cambridge, MA: MIT Press; 1986. p. 318–62.
Cybenko G. Approximation by superpositions of a sigmoidal function. Math Control Signals Syst. 1989;2:303–14.
Funahashi K-I. On the approximate realization of continuous mappings by neural networks. Neural Netw. 1989;2:183–92.
Rumelhart DE, McClelland JL Parallel distribution processing: exploration in the microstructure of cognition. Cambridge, MA: MIT Press; 1986.
Hochreiter S. Untersuchungen zu dynamischen neuronalen Netzen. Diploma, Tech Univ München. 1991;91:1.
Hochreiter S, Schmidhuber J. Long short-term memory. Neural Comput. 1997;9:1735–80.
Bengio Y, Simard P, Frasconi P. Learning long-term dependencies with gradient descent is difficult. IEEE Trans Neural Netw. 1994;5:157–66.
Le QV, Jaitly N, Hinton GE. A simple way to initialize recurrent networks of rectified linear units. arXiv preprint arXiv:150400941 2015:1–9.
Arjovsky M, Shah A, Bengio Y. Unitary evolution recurrent neural networks. Proceedings of the International Conference on Machine Learning. 2016;1120–8.
Neyshabur B, Wu Y, Salakhutdinov RR, Srebro N. Path-normalized optimization of recurrent neural networks with relu activations. Proceedings of the Advances in Neural Information Processing Systems. 2016;3477–85.
Hinton GE, Osindero S, Teh Y-W. A fast learning algorithm for deep belief nets. Neural Comput. 2006;18:1527–54.
Hinton GE, Salakhutdinov RR. Reducing the dimensionality of data with neural networks. Science. 2006;313:504–7.
Bengio Y, Lamblin P, Popovici D, Larochelle H. Greedy layer-wise training of deep networks. Proceedings of the Advances in Neural Information Processing Systems. 2007;153–60.
Goodfellow I, Bengio Y, Courville A, Bengio Y. Deep learning, vol. 1. Cambridge: MIT Press; 2016.
Liu W, Wang Z, Liu X, Zeng N, Liu Y, Alsaadi FE. A survey of deep neural network architectures and their applications. Neurocomputing. 2017;234:11–26.
Martens J, Sutskever I. Learning recurrent neural networks with hessian-free optimization. Proceedings of the Proceedings of the 28th International Conference on Machine Learning (ICML-11). Citeseer; 2011;1033–40.
Hastie T, Tibshirani R, Friedman JH. The elements of statistical learning: data mining, inference, and prediction. 2 edn. New York: Springer; 2009.
Durstewitz D. Advanced data analysis in neuroscience: integrating statistical and computational models. New York: Springer; 2017.
Pascanu R, Mikolov T, Bengio Y. On the difficulty of training recurrent neural networks. Proceedings of the International Conference on Machine Learning. 2013;1310–8.
Sak H, Senior A, Beaufays F. Long short-term memory recurrent neural network architectures for large scale acoustic modeling. Proceedings of the Fifteenth Annual Conference of the International Speech Communication Association. 2014.
Zaheer M, Kottur S, Ahmed A, Moura J, Smola A. Canopy fast sampling with cover trees. Proceedings of the International Conference on Machine Learning. 2017;3977–86.
Lang T, Rettenmeier M. Understanding consumer behavior with recurrent neural networks. Proceedings of the International Workshop on Machine Learning Methods for Recommender Systems. 2017.
Graves A, Bellemare MG, Menick J, Munos R, Kavukcuoglu K. Automated curriculum learning for neural networks. arXiv preprint arXiv:170403003 2017.
Schölkopf B, Smola AJ. Learning with kernels: support vector machines, regularization, optimization, and beyond. Cambridge: MIT Press; 2002.
Orru G, Pettersson-Yeo W, Marquand AF, Sartori G, Mechelli A. Using support vector machine to identify imaging biomarkers of neurological and psychiatric disease: a critical review. Neurosci Biobehav Rev. 2012;36:1140–52.
Wolfers T, Buitelaar JK, Beckmann CF, Franke B, Marquand AF. From estimating activation locality to predicting disorder: a review of pattern recognition for neuroimaging-based psychiatric diagnostics. Neurosci Biobehav Rev. 2015;57:328–49.
Woo C-W, Chang LJ, Lindquist MA, Wager TD. Building better biomarkers: brain models in translational neuroimaging. Nat Neurosci. 2017;20:365.
Bzdok D, Meyer-Lindenberg A. Machine learning for precision psychiatry: opportunities and challenges. Biol Psychiatry. 2018;3:223–30.
Le QV. Building high-level features using large scale unsupervised learning. Proceedings of the IEEE International Conference on Acoustics, Speech and Signal Processing. 2013;8595–8.
Bengio Y. Learning deep architectures for AI. Found trends® Mach Learn. 2009;2:1–127.
Yosinski J, Clune J, Bengio Y, Lipson H. How transferable are features in deep neural networks? Proceedings of the Advances in Neural Information Processing Systems. 2014;3320–8.
Montufar GF, Pascanu R, Cho K, Bengio Y. On the number of linear regions of deep neural networks. Proceedings of the Advances in Neural Information Processing Systems. 2014;2924–32.
Bengio Y, Goodfellow IJ, Courville A. Deep learning. Nature. 2015;521:436–44.
Brosch T, Tam R. Initiative AsDN. Manifold learning of brain MRIs by deep learning. Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer; 2013. p. 633–40.
Li H, Habes M, Fan Y. Deep ordinal ranking for multi-category diagnosis of Alzheimer’s disease using hippocampal MRIdata. arXiv preprint arXiv:170901599; 2017.
Mamoshina P, Vieira A, Putin E, Zhavoronkov A. Applications of deep learning in biomedicine. Mol Pharm. 2016;13:1445–54.
Vieira S, Pinaya WH, Mechelli A. Using deep learning to investigate the neuroimaging correlates of psychiatric and neurological disorders: methods and applications. Neurosci Biobehav Rev. 2017;74:58–75.
Shen D, Wu G, Suk H-I. Deep learning in medical image analysis. Annu Rev Biomed Eng. 2017;19:221–48.
Arbabshirani MR, Plis S, Sui J, Calhoun VD. Single subject prediction of brain disorders in neuroimaging: promises and pitfalls. Neuroimage. 2017;145:137–65.
Calhoun VD, Sui J. Multimodal fusion of brain imaging data: a key to finding the missing link (s) in complex mental illness. Biol Psychiatry. 2016;1:230–44.
Plis SM, Hjelm DR, Salakhutdinov R, Allen EA, Bockholt HJ, Long JD et al. Deep learning for neuroimaging: a validation study. Front Neurosci. 2014; 8:1–11.
Jang H, Plis SM, Calhoun VD, Lee J-H. Task-specific feature extraction and classification of fMRI volumes using a deep neural network initialized with a deep belief network: evaluation using sensorimotor tasks. Neuroimage. 2017;145:314–28.
Cole JH, Poudel RPK, Tsagkrasoulis D, Caan MWA, Steves C, Spector TD, et al. Predicting brain age with deep learning from raw imaging data results in a reliable and heritable biomarker. Neuroimage. 2017;163:115–24.
Liu M, Zhang J, Adeli E, Shen D. Deep multi-task multi-channel learning for joint classification and regression of brain status. Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer; 2017. p. 3–11.
Ambastha AK, Leong TY. A deep learning approach to neuroanatomical characterisation of Alzheimer’s disease. Stud Health Technol Inform. 2017;245:1249.
Khajehnejad M, Saatlou FH, Mohammadzade H. Alzheimer’s disease early diagnosis using manifold-based semi-supervised learning. Brain Sci. 2017;7:109.
Shickel B, Heesacker M, Benton S, Rashidi P. HashtagHealthcare: from tweets to mental health journals using deep transfer learning. arXiv preprint arXiv:170801372; 2017:1–10.
Wang S, Shen Y, Chen W, Xiao T, Hu J. Automatic recognition of mild cognitive impairment from MRI images using expedited convolutional neural networks. Proceedings of the International Conference on Artificial Neural Networks. Springer; 2017. p. 373–80.
Suk H-I, Lee S-W, Shen D. Deep ensemble learning of sparse regression models for brain disease diagnosis. Med Image Anal. 2017;37:101–13.
Shi J, Zheng X, Li Y, Zhang Q, Ying S. Multimodal neuroimaging feature learning with multimodal stacked deep polynomial networks for diagnosis of Alzheimer’s disease. IEEE J Biomed Health Inform. 2018;22:173–83.
Lu D, Popuri K, Ding GW, Balachandar R, Beg MF. Multimodal and multiscale deep neural networks for the early diagnosis of Alzheimer’s disease using structural MR and FDG-PET images. Sci Rep. 2018;8:5697.
Deshpande G, Wang P, Rangaprakash D, Wilamowski B. Fully connected cascade artificial neural network architecture for attention deficit hyperactivity disorder classification from functional magnetic resonance imaging data. IEEE Trans Cybern. 2015;45:2668–79.
Han X, Zhong Y, He L, Philip SY, Zhang L. The unsupervised hierarchical convolutional sparse auto-encoder for neuroimaging data classification. International Conference on Brain Informatics and Health. Springer; 2015, p. 156–166.
Hao AJ, He BL, Yin CH. Discrimination of ADHD children based on Deep Bayesian Network. Journal IET International Conference on Biomedical Image and Signal Processing; 2015.
Kuang D, He L. Classification on ADHD with deep learning. Proceedings of the International Conference on Cloud Computing and Big Data. 2014;27–32.
Kuang D, Guo X, An X, Zhao Y, He L. Discrimination of ADHD based on fMRI data with deep belief network. Proceedings of the International Conference on Intelligent Computing. Springer; 2014. p. 225–32.
Zou L, Zheng J, Miao C, Mckeown MJ, Wang ZJ. 3D CNN based automatic diagnosis of attention deficit hyperactivity disorder using functional and structural MRI. IEEE Access. 2017;5:23626–36.
Hosseini-Asl E, Ghazal M, Mahmoud A, Aslantas A, Shalaby AM, Casanova MF, et al. Alzheimer’s disease diagnostics by a 3D deeply supervised adaptable convolutional network. Front Biosci. 2018;23:584–96.
Sarraf S, Tofighi G. DeepAD: Alzheimer′ s Disease classification via deep convolutional neural networks using MRI and fMRI. bioRxiv 2016;070441.
Payan A, Montana G. Predicting Alzheimer’s disease: a neuroimaging study with 3D convolutional neural networks. arXiv preprint arXiv:150202506 2015:1–9.
Choi H, Ha S, Im HJ, Paek SH, Lee DS. Refining diagnosis of Parkinson’s disease with deep learning-based interpretation of dopamine transporter imaging. NeuroImage. 2017;16:586–94.
Liu X, Chen K, Wu T, Weidman D, Lure F, Li J. Use of multi-modality imaging and artificial intelligence for diagnosis and prognosis of early stages of alzheimer’s disease. Transl Res. 2018; 194:56–67.
Dakka J, Bashivan P, Gheiratmand M, Rish I, Jha S, Greiner R. Learning neural markers of Schizophrenia disorder using recurrent neural networks. arXiv preprint arXiv:171200512; 2017:1–6.
Kim J, Calhoun VD, Shim E, Lee J-H. Deep neural network with weight sparsity control and pre-training extracts hierarchical features and enhances classification performance: evidence from whole-brain resting-state functional connectivity patterns of schizophrenia. Neuroimage. 2016;124:127–46.
Ulloa A, Plis S, Calhoun V. Improving classification rate of Schizophrenia using a multimodal multi-layer perceptron model with structural and functional MR. arXiv preprint arXiv:180404591; 2018:1–9.
Zeng L-L, Wang H, Hu P, Yang B, Pu W, Shen H, et al. Multi-site diagnostic classification of Schizophrenia using discriminant deep learning with functional connectivity MRI. EBioMedicine. 2018;30:74–85.
Yan W, Plis S, Calhoun VD, Liu S, Jiang R, Jiang T-Z, et al. Discriminating schizophrenia from normal controls using resting state functional network connectivity: a deep neural network and layer-wise relevance propagation method. Proceedings of the 27th International Workshop on Machine Learning for Signal Processing, 2017;1–6.
Guo X, Dominick KC, Minai AA, Li H, Erickson CA, Lu LJ. Diagnosing autism spectrum disorder from brain resting-state functional connectivity patterns using a deep neural network with a novel feature selection method. Front Neurosci. 2017;11:460.
Heinsfeld AS, Franco AR, Craddock RC, Buchweitz A, Meneguzzi F. Identification of autism spectrum disorder using deep learning and the ABIDE dataset. NeuroImage. 2018;17:16–23.
Heinsfeld AS. Identification of autism disorder through functional MRI and deep learning. Pontifícia Universidade Católica do Rio Grande do Sul; 2017; 17:16–23.
Cao B, Zheng L, Zhang C, Yu PS, Piscitello A, Zulueta J, et al. DeepMood: modeling mobile phone typing dynamics for mood detection. Proceedings of the 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. ACM; 2017. p. 747–55.
Wang SH, Lv YD, Sui Y, Liu S, Wang SJ, Zhang YD. Alcoholism detection by data augmentation and convolutional neural network with stochastic pooling. J Med Syst. 2017;42:2.
Munsell BC, Wee C-Y, Keller SS, Weber B, Elger C, da Silva LAT, et al. Evaluation of machine learning algorithms for treatment outcome prediction in patients with epilepsy based on structural connectome data. Neuroimage. 2015;118:219–30.
Thodoroff P, Pineau J, Lim A. Learning robust features using deep learning for automatic seizure detection. Proceedings of the Machine Learning for Healthcare Conference. 2016;178–90.
Pearlson GD, Calhoun VD, Liu J. An introductory review of parallel independent component analysis (p-ICA) and a guide to applying p-ICA to genetic data and imaging phenotypes to identify disease-associated biological pathways and systems in common complex disorders. Front Genet. 2015;6:276.
Liu S, Liu S, Cai W, Che H, Pujol S, Kikinis R, et al. Multimodal neuroimaging feature learning for multiclass diagnosis of Alzheimer’s disease. IEEE Trans Biomed Eng. 2015;62:1132–40.
Dai D, Wang J, Hua J, He H. Classification of ADHD children through multimodal magnetic resonance imaging. Front Syst Neurosci. 2012;6:63.
Zhang D, Wang Y, Zhou L, Yuan H, Shen D. Multimodal classification of Alzheimer’s disease and mild cognitive impairment. Neuroimage. 2011;55:856–67.
Yang J, Yin Y, Zhang Z, Long J, Dong J, Zhang Y, et al. Predictive brain networks for major depression in a semi-multimodal fusion hierarchical feature reduction framework. Neurosci Lett. 2018;665:163–9.
Schumann G, Binder EB, Holte A, de Kloet ER, Oedegaard KJ, Robbins TW, et al. Stratified medicine for mental disorders. Eur Neuropsychopharmacol. 2014;24:5–50.
Sui J, Qi S, van Erp TGM, Bustillo J, Jiang R, Lin D, et al. Multimodal neuromarkers in schizophrenia via cognition-guided MRI fusion. Nat Commun. 2018;9:3028.
Sui J, Adali T, Yu Q, Calhoun VD. A review of multivariate methods for multimodal fusion of brain imaging data. J Neurosci Methods. 2012;204:68–81.
Kloppel S, Stonnington CM, Barnes J, Chen F, Chu C, Good CD, et al. Accuracy of dementia diagnosis: a direct comparison between radiologists and a computerized method. Brain. 2008;131(Pt 11):2969–74.
Miotto R, Li L, Kidd BA, Dudley JT. Deep patient: an unsupervised representation to predict the future of patients from the electronic health records. Sci Rep. 2016;6:26094.
Tran T, Kavuluru R. Predicting mental conditions based on “history of present illness” in psychiatric notes with deep neural networks. J Biomed Inform. 2017;75:S138–S148.
Gkotsis G, Oellrich A, Velupillai S, Liakata M, Hubbard TJ, Dobson RJ, et al. Characterisation of mental health conditions in social media using informed deep learning. Sci Rep. 2017;7:45141.
Thayer JF, Ahs F, Fredrikson M, Sollers JJ 3rd, Wager TD. A meta-analysis of heart rate variability and neuroimaging studies: implications for heart rate variability as a marker of stress and health. Neurosci Biobehav Rev. 2012;36:747–56.
Holt-Lunstad J, Smith TB, Layton JB. Social relationships and mortality risk: a meta-analytic review. PLoS Med. 2010;7:e1000316.
Taylor CB, Sallis JF, Needle R. The relation of physical activity and exercise to mental health. Public Health Rep. 1985;100:195.
Canzian L, Musolesi M. Trajectories of depression: unobtrusive monitoring of depressive states by means of smartphone mobility traces analysis. Proceedings of the ACM International Joint Conference on Pervasive and Ubiquitous Computing. ACM; 2015. p. 1293–304.
Mehrotra A, Hendley R, Musolesi M. Towards multi-modal anticipatory monitoring of depressive states through the analysis of human-smartphone interaction. Proceedings of the ACM International Joint Conference on Pervasive and Ubiquitous Computing. ACM; 2016. p. 1132–8.
Abdullah S, Matthews M, Frank E, Doherty G, Gay G, Choudhury T. Automatic detection of social rhythms in bipolar disorder. J Am Med Inform Assoc. 2016;23:538–43.
Mikelsons G, Smith M, Mehrotra A, Musolesi M. Towards deep learning models for psychological state prediction using smartphone data: challenges and opportunities. arXiv preprint arXiv:171106350; 2017.
Sathyanarayana A, Joty S, Fernandez-Luque L, Ofli F, Srivastava J, Elmagarmid A et al. Sleep quality prediction from wearable data using deep learning. JMIR mHealth and uHealth 2016; 4:e125.
Aung MH, Matthews M, Choudhury T. Sensing behavioral symptoms of mental health and delivering personalized interventions using mobile technologies. Depress Anxiety 2017;34:603–9.
Stamate C, Magoulas GD, Küppers S, Nomikou E, Daskalopoulos I, Luchini MU et al. Deep learning Parkinson's from smartphone data. Proceedings of the International Conference on Pervasive Computing and Communications. 2017;1-40.
Stamate C, Magoulas G, Kueppers S, Nomikou E, Daskalopoulos I, Jha A, et al. The cloudUPDRS app: a medical device for the clinical assessment of Parkinson’s Disease. Pervasive Mob Comput. 2018;43:146–66.
Suhara Y, Xu Y, Pentland AS. Deepmood: forecasting depressed mood based on self-reported histories via recurrent neural networks. Proceedings of the 26th International Conference on World Wide Web. 2017;715–24.
Donker T, Petrie K, Proudfoot J, Clarke J, Birch M-R, Christensen H. Smartphones for smarter delivery of mental health programs: a systematic review. J Med Internet Res 2013; 15:e247.
Dehling T, Gao F, Schneider S, Sunyaev A. Exploring the far side of mobile health: information security and privacy of mobile health apps on iOS and android. JMIR Mhealth Uhealth. 2015;3:e8.
Marzano L, Bardill A, Fields B, Herd K, Veale D, Grey N, et al. The application of mHealth to mental health: opportunities and challenges. Lancet Psychiatry. 2015;2:942–8.
He K, Zhang X, Ren S, Sun J. Delving deep into rectifiers: surpassing human-level performance on imagenet classification. Proceedings of the International Conference on Computer Vision. 2015;1026–34.
Whelan R, Garavan H. When optimism hurts: inflated predictions in psychiatric neuroimaging. Biol Psychiatry. 2014;75:746–8.
Collins FS, Varmus H. A new initiative on precision medicine. New Engl J Med. 2015;372:793–5.
Bzdok D, Yeo BT. Inference in the age of big data: future perspectives on neuroscience. Neuroimage. 2017;155:549–64.
Caruana R. Multitask learning. Learning to learn. Springer; 1998, p. 95–133.
Zhou Y, Song S, Cheung N-M. On classification of distorted images with deep convolutional neural networks. Proceedings of the International Conference on Acoustics, Speech and Signal Processing. 2017;1213–7.
Suk H-I, Lee S-W, Shen D. Initiative AsDN. Latent feature representation with stacked auto-encoder for AD/MCI diagnosis. Brain Struct Funct. 2015;220:841–59.
Raina R, Battle A, Lee H, Packer B, Ng AY. Self-taught learning: transfer learning from unlabeled data. Proceedings of the 24th International Conference on Machine learning. ACM; 2007. 759–66.
Gupta A, Ayhan M, Maida A. Natural image bases to represent neuroimaging data. Proceedings of the International Conference on Machine Learning. 2013;987–94.
Banerjee D, Islam K, Mei G, Xiao L, Zhang G, Xu R et al. A deep transfer learning approach for improved post-traumatic stress disorder diagnosis. Proceedings of the International Conference on Data Mining. 2017;11–20.
Li F, Tran L, Thung K-H, Ji S, Shen D, Li J. Robust deep learning for improved classification of AD/MCI patients. Proceedings of the International Workshop on Machine Learning in Medical Imaging. Springer; 2014. p. 240–7.
Tan X, Liu Y, Li Y, Wang P, Zeng X, Yan F, et al. Localized instance fusion of MRI data of Alzheimer’s disease for classification based on instance transfer ensemble learning. Biomed Eng Online. 2018;17:49.
Yahata N, Morimoto J, Hashimoto R, Lisi G, Shibata K, Kawakubo Y, et al. A small number of abnormal brain connections predicts adult autism spectrum disorder. Nat Commun. 2016;7:11254.
Nelson B, McGorry PD, Wichers M, Wigman JT, Hartmann JA. Moving from static to dynamic models of the onset of mental disorder: a review. JAMA Psychiatry. 2017;74:528–34.
Insel T, Cuthbert B, Garvey M, Heinssen R, Pine DS, Quinn K et al. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am Psychiatric Assoc. 2010;167:748–51.
Haro JM, Ayuso‐Mateos JL, Bitter I, Demotes‐Mainard J, Leboyer M, Lewis SW, et al. ROAMER: roadmap for mental health research in Europe. Int J Methods Psychiatr Res. 2014;23(S1):1–14.
Suk H-I, Lee S-W, Shen D. Initiative AsDN. Hierarchical feature representation and multimodal fusion with deep learning for AD/MCI diagnosis. Neuroimage. 2014;101:569–82.
Insel TR. The NIMH Research Domain Criteria (RDoC) Project: precision medicine for psychiatry. Am J Psychiatry. 2014;171:395–7.
Rios A, Kavuluru R. Ordinal convolutional neural networks for predicting RDoC positive valence psychiatric symptom severity scores. J Biomed Inform. 2017;75s:S85–s93.
Mehrotra A, Musolesi M. Using autoencoders to automatically extract mobility features for predicting depressive states. Proc ACM Interact Mob Wearable Ubiquitous Technol. 2018;2:1–20.
Aliper A, Plis S, Artemov A, Ulloa A, Mamoshina P, Zhavoronkov A. Deep learning applications for predicting pharmacological properties of drugs and drug repurposing using transcriptomic data. Mol Pharm. 2016;13:2524–30.
Hertz J, Krogh A, Palmer RG. Introduction to the theory of neural computation. Addison-Wesley/Addison Wesley Longman; 1991.
Dayan P, Abbott LF. Theoretical neuroscience, vol. 806. Cambridge, MA: MIT Press; 2001.
Hassabis D, Kumaran D, Summerfield C, Botvinick M. Neuroscience-inspired artificial intelligence. Neuron. 2017;95:245–58.
Kriegeskorte N. Deep neural networks: a new framework for modeling biological vision and brain information processing. Annu Rev Vision Sci. 2015;1:417–46.
Yamins DL, DiCarlo JJ. Using goal-driven deep learning models to understand sensory cortex. Nat Neurosci. 2016;19:356–65.
Durstewitz D, Seamans JK, Sejnowski TJ. Dopamine-mediated stabilization of delay-period activity in a network model of prefrontal cortex. J Neurophysiol. 2000;83:1733–50.
Koch C, Segev I. Methods in neuronal modeling: from ions to networks. MIT Press; 1998.
Druckmann S, Banitt Y, Gidon AA, Schürmann F, Markram H, Segev I. A novel multiple objective optimization framework for constraining conductance-based neuron models by experimental data. Front Neurosci. 2007;1:1.
Fisher D, Olasagasti I, Tank DW, Aksay ER, Goldman MS. A modeling framework for deriving the structural and functional architecture of a short-term memory microcircuit. Neuron. 2013;79:987–1000.
Hertäg L, Hass J, Golovko T, Durstewitz D. An approximation to the adaptive exponential integrate-and-fire neuron model allows fast and predictive fitting to physiological data. Front Comput Neurosci. 2012;6:62.
Durstewitz D, Koppe G, Toutounji H. Computational models as statistical tools. Curr Opin Behav Sci. 2016;11:93–99.
Speiser A, Yan J, Archer EW, Buesing L, Turaga SC, Macke JH. Fast amortized inference of neural activity from calcium imaging data with variational autoencoders. Proceedings of the Advances in Neural Information Processing Systems. 2017;4027–37.
Nonnenmacher M, Turaga SC, Macke JH. Extracting low-dimensional dynamics from multiple large-scale neural population recordings by learning to predict correlations. Proceedings of the Advances in Neural Information Processing Systems. 2017;5706–16.
Lueckmann J-M, Goncalves PJ, Bassetto G, Öcal K, Nonnenmacher M, Macke JH. Flexible statistical inference for mechanistic models of neural dynamics. Proceedings of the Advances in Neural Information Processing Systems. 2017;1289–99.
Putzky P, Franzen F, Bassetto G, Macke JH. A Bayesian model for identifying hierarchically organised states in neural population activity. Proceedings of the Advances in Neural Information Processing Systems. 2014;3095–103.
Markram H, Muller E, Ramaswamy S, Reimann MW, Abdellah M, Sanchez CA, et al. Reconstruction and simulation of neocortical microcircuitry. Cell. 2015;163:456–92.
Hass J, Hertäg L, Durstewitz D. A detailed data-driven network model of prefrontal cortex reproduces key features of in vivo activity. PLoS Comput Biol. 2016;12:e1004930.
Hertäg L, Durstewitz D, Brunel N. Analytical approximations of the firing rate of an adaptive exponential integrate-and-fire neuron in the presence of synaptic noise. Front Comput Neurosci. 2014;8:116.
Brunel N, Wang XJ. Effects of neuromodulation in a cortical network model of object working memory dominated by recurrent inhibition. J Comput Neurosci. 2001;11:63–85.
Amit DJ, Brunel N. Model of global spontaneous activity and local structured activity during delay periods in the cerebral cortex. Cereb Cortex. 1997;7:237–52.
Breakspear M. Dynamic models of large-scale brain activity. Nat Neurosci. 2017;20:340.
Stephan KE, Iglesias S, Heinzle J, Diaconescu AO. Translational perspectives for computational neuroimaging. Neuron. 2015;87:716–32.
Huys QJM, Maia TV, Frank MJ. Computational psychiatry as a bridge from neuroscience to clinical applications. Nat Neurosci. 2016;19:404.
Mathys CD, Lomakina EI, Daunizeau J, Iglesias S, Brodersen KH, Friston KJ, et al. Uncertainty in perception and the Hierarchical Gaussian Filter. Front Hum Neurosci. 2014;8:825.
Koppe G, Mallien AS, Berger S, Bartsch D, Gass P, Vollmayr B, et al. CACNA1C gene regulates behavioral strategies in operant rule learning. PLoS Biol. 2017;15:e2000936.
Stephan KE, Schlagenhauf F, Huys QJM, Raman S, Aponte EA, Brodersen KH, et al. Computational neuroimaging strategies for single patient predictions. Neuroimage. 2017;145(Pt B):180–99.
Finnegan A, Song JS. Maximum entropy methods for extracting the learned features of deep neural networks. PLoS Comput Biol. 2017;13:e1005836.
Yosinski J, Clune J, Nguyen A, Fuchs T, Lipson H. Understanding neural networks through deep visualization. arXiv preprint arXiv:150606579 2015:1–12.
Kietzmann TC, McClure P, Kriegeskorte N. Deep neural networks in computational neuroscience. bioRxiv 2017: 133504.
Erhan D, Bengio Y, Courville A, Vincent P. Visualizing higher-layer features of a deep network. Univ Montr. 2009;1341:1.
Güçlü U, van Gerven MA. Deep neural networks reveal a gradient in the complexity of neural representations across the ventral stream. J Neurosci. 2015;35:10005–14.
Durstewitz D. A state space approach for piecewise-linear recurrent neural networks for identifying computational dynamics from neural measurements. PLoS Comput Biol. 2017;13:e1005542.
Durstewitz D, Huys QJM, Koppe G. Psychiatric illnesses as disorders of network dynamics. arXiv preprint arXiv:180906303 2018:1–24.
Wilson HR. Spikes, decisions, and actions: the dynamical foundations of neurosciences. 1999.
Durstewitz D, Seamans JK, Sejnowski TJ. Neurocomputational models of working memory. Nat Neurosci. 2000;3(11s):1184.
Wang X-J. Probabilistic decision making by slow reverberation in cortical circuits. Neuron. 2002;36:955–68.
Albantakis L, Deco G. The encoding of alternatives in multiple-choice decision making. Proc Natl Acad Sci. 2009;106:10308–13.
Neal RM. Bayesian learning for neural networks, vol. 118. New York: Springer; 2012.
Haykin SS. Kalman filtering and neural networks. Wiley Online Library; 2001.
Kingma DP, Welling M. Auto-encoding variational bayes. Proceedings of the 2nd International Conference on Learning Representations (ICLR); 2014.
Werbos PJ. Backpropagation through time: what it does and how to do it. Proc IEEE. 1990;78:1550–60.
Acknowledgements
DD was supported by grants from the German Federal Ministry of Education and Research (BMBF) within the e:Med program (01ZX1311A [SP7] and 01ZX1314G [SP10]), and by the German Science Foundation (DFG; Du 354/8-2). AML was supported by the BMBF (01ZX1314G [SP4]) and by the European Union’s Seventh Framework Program (FP7, grant 602805 and 602450).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
AML has received consultant fees from Boehringer Ingelheim, Elsevier, Walt Disney Pictures, Brainsway, Lundbeck Int. Neuroscience Foundation, Sumitomo Dainippon Pharma Co., Academic Medical Center of the University of Amsterdam, Synapsis Foundation-Alzheimer Research Switzerland, IBS Center for Synaptic Brain Dysfunction, Blueprint Partnership, University of Cambridge, Dt. Zentrum für Neurodegenerative Erkrankungen, Universität Zürich, L.E.K. Consulting, ICARE Schizophrenia, Science Advances, and has received fees for lectures, interviews and travels from Lundbeck International Foundation, Paul-Martini-Stiftung, Lilly Deutschland, Atheneum, Fama Public Relations, Institut d’investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), Jansen-Cilag, Hertie Stiftung, Bodelschwingh-Klinik, Pfizer, Atheneum, Universität Freiburg, Schizophrenia Academy, Hong Kong Society of Biological Psychiatry, Fama Public Relations, Spanish Society of Psychiatry, Reunions I Ciencia S.L., Brain Center Rudolf Magnus UMC Utrecht. The other authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Durstewitz, D., Koppe, G. & Meyer-Lindenberg, A. Deep neural networks in psychiatry. Mol Psychiatry 24, 1583–1598 (2019). https://doi.org/10.1038/s41380-019-0365-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-019-0365-9
This article is cited by
-
Robust and replicable functional brain signatures of 22q11.2 deletion syndrome and associated psychosis: a deep neural network-based multi-cohort study
Molecular Psychiatry (2024)
-
Unraveling minds in the digital era: a review on mapping mental health disorders through machine learning techniques using online social media
Social Network Analysis and Mining (2024)
-
Transdiagnostic subgroups of cognitive impairment in early affective and psychotic illness
Neuropsychopharmacology (2024)
-
Applications of artificial intelligence−machine learning for detection of stress: a critical overview
Molecular Psychiatry (2023)
-
On the Analyses of Medical Images Using Traditional Machine Learning Techniques and Convolutional Neural Networks
Archives of Computational Methods in Engineering (2023)