Abstract
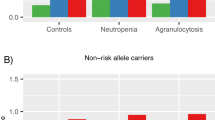
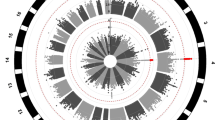
Individuals of African ancestry in the United States and Europe are at increased risk of developing schizophrenia and have poorer clinical outcomes. The antipsychotic clozapine, the only licensed medication for treatment-resistant schizophrenia, is under-prescribed and has high rates of discontinuation in individuals of African ancestry, due in part to increased rates of neutropenia. The genetic basis of lower neutrophil levels in those of African ancestry has not previously been investigated in the context of clozapine treatment. We sought to identify risk alleles in the first genome-wide association study of neutrophil levels during clozapine treatment, in 552 individuals with treatment-resistant schizophrenia and robustly inferred African genetic ancestry. Two genome-wide significant loci were associated with low neutrophil counts during clozapine treatment. The most significantly associated locus was driven by rs2814778 (β = −0.9, P = 4.21 × 10−21), a known regulatory variant in the atypical chemokine receptor 1 (ACKR1) gene. Individuals homozygous for the C allele at rs2814778 were significantly more likely to develop neutropenia and have to stop clozapine treatment (OR = 20.4, P = 3.44 × 10−7). This genotype, also termed “Duffy-null”, has previously been shown to be associated with lower neutrophil levels in those of African ancestry. Our results indicate the relevance of the rs2814778 genotype for those taking clozapine and its potential as a pharmacogenetic test, dependent on the outcome of additional safety studies, to assist decision making in the initiation and on-going management of clozapine treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Cantor-Graae E, Selten JP. Schizophrenia and migration: a meta-analysis and review. Am J Psychiatry. 2005;162:12–24.
McGrath J, Saha S, Chant D, Welham J. Schizophrenia: a concise overview of incidence, prevalence, and mortality. Epidemiol Rev. 2008;30:67–76.
Fearon P, Kirkbride JB, Morgan C, Dazzan P, Morgan K, Lloyd T, et al. Incidence of schizophrenia and other psychoses in ethnic minority groups: results from the MRC AESOP Study. Psychol Med. 2006;36:1541–50.
Bresnahan M, Begg MD, Brown A, Schaefer C, Sohler N, Insel B, et al. Race and risk of schizophrenia in a US birth cohort: another example of health disparity? Int J Epidemiol. 2007;36:751–8.
Brugha T, Jenkins R, Bebbington P, Meltzer H, Lewis G, Farrell M. Risk factors and the prevalence of neurosis and psychosis in ethnic groups in Great Britain. Soc Psychiatry Psychiatr Epidemiol. 2004;39:939–46.
Boydell J, van Os J, McKenzie K, Allardyce J, Goel R, McCreadie RG, et al. Incidence of schizophrenia in ethnic minorities in London: ecological study into interactions with environment. BMJ. 2001;323:1336–8.
Bhui K, Stansfeld S, Hull S, Priebe S, Mole F, Feder G. Ethnic variations in pathways to and use of specialist mental health services in the UK. Systematic review. Br J Psychiatry. 2003;182:105–16.
Morgan C, Fearon P, Lappin J, Heslin M, Donoghue K, Lomas B, et al. Ethnicity and long-term course and outcome of psychotic disorders in a UK sample: the AESOP-10 study. Br J Psychiatry. 2017;211:88–94.
Kane J, Honigfeld G, Singer J, Meltzer H. Clozapine for the treatment-resistant schizophrenic. A double-blind comparison with chlorpromazine. Arch Gen Psychiatry. 1988;45:789–96.
Leucht S, Corves C, Arbter D, Engel RR, Li C, Davis JM. Second-generation versus first-generation antipsychotic drugs for schizophrenia: a meta-analysis. Lancet. 2009;373:31–41.
Tiihonen J, Mittendorfer-Rutz E, Majak M, Mehtala J, Hoti F, Jedenius E, et al. Real-world effectiveness of antipsychotic treatments in a nationwide cohort of 29823 patients with schizophrenia. JAMA Psychiatry. 2017;74:686–93.
Kuno E, Rothbard AB. Racial disparities in antipsychotic prescription patterns for patients with schizophrenia. Am J Psychiatry. 2002;159:567–72.
Kelly DL, Dixon LB, Kreyenbuhl JA, Medoff D, Lehman AF, Love RC, et al. Clozapine utilization and outcomes by race in a public mental health system: 1994-2000. J Clin Psychiatry. 2006;67:1404–11.
Whiskey E, Olofinjana O, Taylor D. The importance of the recognition of benign ethnic neutropenia in black patients during treatment with clozapine: case reports and database study. J Psychopharmacol. 2011;25:842–5.
Davis MC, Fuller MA, Strauss ME, Konicki PE, Jaskiw GE. Discontinuation of clozapine: a 15-year naturalistic retrospective study of 320 patients. Acta Psychiatr Scand. 2014;130:30–39.
Moeller FG, Chen YW, Steinberg JL, Petty F, Ripper GW, Shah N, et al. Risk factors for clozapine discontinuation among 805 patients in the VA hospital system. Ann Clin Psychiatry. 1995;7:167–73.
Munro J, O’Sullivan D, Andrews C, Arana A, Mortimer A, Kerwin R. Active monitoring of 12,760 clozapine recipients in the UK and Ireland. Beyond pharmacovigilance. Br J Psychiatry. 1999;175:576–80.
Kelly DL, Kreyenbuhl J, Dixon L, Love RC, Medoff D, Conley RR. Clozapine underutilization and discontinuation in African Americans due to leucopenia. Schizophr Bull. 2007;33:1221–4.
Myles N, Myles H, Xia S, Large M, Kisely S, Galletly C. et al. Meta-analysis examining the epidemiology of clozapine-associated neutropenia. Acta Psychiatr Scand. 2018;138:101–9.
Saito T, Ikeda M, Mushiroda T, Ozeki T, Kondo K, Shimasaki A, et al. Pharmacogenomic study of clozapine-induced agranulocytosis/granulocytopenia in a Japanese population. Biol Psychiatry. 2016;80:636–42.
Goldstein JI, Fredrik Jarskog L, Hilliard C, Alfirevic A, Duncan L, Fourches D, et al. Clozapine-induced agranulocytosis is associated with rare HLA-DQB1 and HLA-B alleles. Nat Commun. 2014;5:4757.
Legge SE, Hamshere ML, Ripke S, Pardinas AF, Goldstein JI, Rees E. et al. Genome-wide common and rare variant analysis provides novel insights into clozapine-associated neutropenia. Mol Psychiatry. 2017;22:1502–8.
Gibson C, Berliner N. How we evaluate and treat neutropenia in adults. Blood. 2014;124:1251–8. quiz 1378
Pardinas AF, Holmans P, Pocklington AJ, Escott-Price V, Ripke S, Carrera N. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat Genet. 2018;50:381–9.
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Anderson CA, Pettersson FH, Clarke GM, Cardon LR, Morris AP, Zondervan KT. Data quality control in genetic case-control association studies. Nat Protoc. 2010;5:1564–73.
McCarthy S, Das S, Kretzschmar W, Delaneau O, Wood AR, Teumer A, et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet. 2016;48:1279–83.
Das S, Forer L, Schonherr S, Sidore C, Locke AE, Kwong A, et al. Next-generation genotype imputation service and methods. Nat Genet. 2016;48:1284–7.
Advanced Research Computing @ Cardiff (ARCCA). Introduction to RAVEN. https://www.cardiff.ac.uk/advanced-researchcomputing/about-us/our-supercomputers (accessed 29 March 2016).
van Leeuwen EM, Kanterakis A, Deelen P, Kattenberg MV, Genome of the Netherlands Consortium, Slagboom PE, et al. Population-specific genotype imputations using minimac or IMPUTE2. Nat Protoc. 2015;10:1285–96.
Huang J, Howie B, McCarthy S, Memari Y, Walter K, Min JL, et al. Improved imputation of low-frequency and rare variants using the UK10K haplotype reference panel. Nat Commun. 2015;6:8111.
Phillips C, Salas A, Sanchez JJ, Fondevila M, Gomez-Tato A, Alvarez-Dios J, et al. Inferring ancestral origin using a single multiplex assay of ancestry-informative marker SNPs. Forensic Sci Int Genet. 2007;1:273–80.
Bulbul O, Filoglu G, Zorlu T, Altuncul H, Freire-Aradas A, Sochtig J, et al. Inference of biogeographical ancestry across central regions of Eurasia. Int J Leg Med. 2016;130:73–79.
Shriver MD, Parra EJ, Dios S, Bonilla C, Norton H, Jovel C, et al. Skin pigmentation, biogeographical ancestry and admixture mapping. Hum Genet. 2003;112:387–99.
Li JZ, Absher DM, Tang H, Southwick AM, Casto AM, Ramachandran S, et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science. 2008;319:1100–4.
Tishkoff SA, Kidd KK. Implications of biogeography of human populations for ‘race’ and medicine. Nat Genet. 2004;36:S21–27.
Bryc K, Durand EY, Macpherson JM, Reich D, Mountain JL. The genetic ancestry of African Americans, Latinos, and European Americans across the United States. Am J Hum Genet. 2015;96:37–53.
Avena S, Via M, Ziv E, Perez-Stable EJ, Gignoux CR, Dejean C, et al. Heterogeneity in genetic admixture across different regions of Argentina. PLoS One. 2012;7:e34695.
Graffelman J. Exploring diallelic genetic markers: the HardyWeinberg Package. J Stat Softw. 2015;64:1–23.
Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82.
Yang J, Zaitlen NA, Goddard ME, Visscher PM, Price AL. Advantages and pitfalls in the application of mixed-model association methods. Nat Genet. 2014;46:100–6.
Yang J, Ferreira T, Morris AP, Medland SE, Genetic Investigation of ANthropometric Traits (GIANT) Consortium, DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium, et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat Genet. 2012;44:361–75.
Zheng X, Shen J, Cox C, Wakefield JC, Ehm MG, Nelson MR, et al. HIBAG--HLA genotype imputation with attribute bagging. Pharm J. 2014;14:192–200.
Levin AM, Adrianto I, Datta I, Iannuzzi MC, Trudeau S, Li J, et al. Association of HLA-DRB1 with sarcoidosis susceptibility and progression in African Americans. Am J Respir Cell Mol Biol. 2015;53:206–16.
Conomos MP, Miller MB, Thornton TA. Robust inference of population structure for ancestry prediction and correction of stratification in the presence of relatedness. Genet Epidemiol. 2015;39:276–93.
Reich D, Nalls MA, Kao WH, Akylbekova EL, Tandon A, Patterson N, et al. Reduced neutrophil count in people of African descent is due to a regulatory variant in the Duffy antigen receptor for chemokines gene. PLoS Genet. 2009;5:e1000360.
Rajagopal S. Clozapine, agranulocytosis, and benign ethnic neutropenia. Postgrad Med J. 2005;81:545–6.
Manu P, Sarvaiya N, Rogozea LM, Kane JM, Correll CU. Benign ethnic neutropenia and clozapine use: a systematic review of the evidence and treatment recommendations. J Clin Psychiatry. 2016;77:e909–916.
Haddy TB, Rana SR, Castro O. Benign ethnic neutropenia: what is a normal absolute neutrophil count? J Lab Clin Med. 1999;133:15–22.
Thobakgale CF, Ndung’u T. Neutrophil counts in persons of African origin. Curr Opin Hematol. 2014;21:50–57.
Reiner AP, Lettre G, Nalls MA, Ganesh SK, Mathias R, Austin MA, et al. Genome-wide association study of white blood cell count in 16,388 African Americans: the continental origins and genetic epidemiology network (COGENT). PLoS Genet. 2011;7:e1002108.
Nalls MA, Wilson JG, Patterson NJ, Tandon A, Zmuda JM, Huntsman S, et al. Admixture mapping of white cell count: genetic locus responsible for lower white blood cell count in the Health ABC and Jackson Heart studies. Am J Hum Genet. 2008;82:81–87.
The Charge Consortium Hematology Working Group. Meta-analysis of rare and common exome chip variants identifies S1PR4 and other loci influencing blood cell traits. Nat Genet. 2016;48:867–76.
Davis MB, Walens A, Hire R, Mumin K, Brown AM, Ford D, et al. Distinct transcript isoforms of the atypical chemokine receptor 1 (ACKR1)/Duffy antigen receptor for chemokines (DARC) gene are expressed in lymphoblasts and altered isoform levels are associated with genetic ancestry and the Duffy-null allele. PLoS One. 2015;10:e0140098.
Pierron D, Heiske M, Razafindrazaka H, Pereda-Loth V, Sanchez J, Alva O, et al. Strong selection during the last millennium for African ancestry in the admixed population of Madagascar. Nat Commun. 2018;9:932.
Duchene J, Novitzky-Basso I, Thiriot A, Casanova-Acebes M, Bianchini M, Etheridge SL, et al. Atypical chemokine receptor 1 on nucleated erythroid cells regulates hematopoiesis. Nat Immunol. 2017;18:753–61.
Permanyer M, Bosnjak B, Forster R. Dual role for atypical chemokine receptor 1 in myeloid cell hematopoiesis and distribution. Cell Mol Immunol. 2018;15:399–401.
Richardson CM, Davis EA, Vyas GR, DiPaula BA, McMahon RP, Kelly DL. Evaluation of the safety of clozapine use in patients with benign neutropenia. J Clin Psychiatry. 2016;77:e1454–e1459.
Howes RE, Patil AP, Piel FB, Nyangiri OA, Kabaria CW, Gething PW, et al. The global distribution of the Duffy blood group. Nat Commun. 2011;2:266.
Meyer S, Vollmert C, Trost N, Bronnimann C, Gottschalk J, Buser A, et al. High-throughput Kell, Kidd, and Duffy matrix-assisted laser desorption/ionization, time-of-flight mass spectrometry-based blood group genotyping of 4000 donors shows close to full concordance with serotyping and detects new alleles. Transfusion. 2014;54:3198–207.
Lopez GH, Morrison J, Condon JA, Wilson B, Martin JR, Liew YW, et al. Duffy blood group phenotype-genotype correlations using high-resolution melting analysis PCR and microarray reveal complex cases including a new null FY*A allele: the role for sequencing in genotyping algorithms. Vox Sang. 2015;109:296–303.
Hoher G, Fiegenbaum M, Almeida S. Molecular basis of the Duffy blood group system. Blood Transfus. 2018;16:93–100.
Rios M, Chaudhuri A, Mallinson G, Sausais L, Gomensoro-Garcia AE, Hannon J, et al. New genotypes in Fy(a-b-) individuals: nonsense mutations (Trp to stop) in the coding sequence of either FY A or FY B. Brit J Haematol. 2000;108:448–54.
Langhi DM Jr., Bordin JO. Duffy blood group and malaria. Hematology. 2006;11:389–98.
Liu YD, Zhang B, Kuang H, Korakavi G, Lu LY, Yu XC. Zinc finger protein 618 regulates the function of UHRF2 (ubiquitin-like with PHD and ring finger domains 2) as a specific 5-hydroxymethylcytosine reader. J Biol Chem. 2016;291:13679–88.
Acknowledgements
This project was supported by Medical Research Council (MRC) Centre (MR/L010305/1), Program (G0800509) and Project ("STRATA", MR/L011794/1) grants to Cardiff University. The project has received funding from the European Union’s Seventh Framework Programme for research, technological development and demonstration under grant agreement no. 279227 (CRESTAR Consortium; http://www.crestar-project.eu/). This publication reflects only the authors’ views and the European Union is not liable for any use that may be made of the information contained therein. We acknowledge Leyden Delta and Magna Laboratories, UK, for supporting the CLOZUK2 sample collection, anonymisation and data preparation (Andy Walker and Anouschka Colson). We acknowledge deCODE genetics (Hreinn Stefansson and colleagues) for genotyping of the CLOZUK2 sample. We acknowledge the MRC Centre laboratory staff (particularly Lucinda Hopkins, Lesley Bates and Catherine Bresner) at Cardiff University for laboratory sample management and Wayne Lawrence and Mark Einon at Cardiff University for support with the use and setup of computational infrastructures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
DAC is a full-time employee and stockholder of Eli Lilly and Company. MH, JAJ and KJ are full-time employees of Leyden Delta B.V. The remaining authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Legge, S.E., Pardiñas, A.F., Helthuis, M. et al. A genome-wide association study in individuals of African ancestry reveals the importance of the Duffy-null genotype in the assessment of clozapine-related neutropenia. Mol Psychiatry 24, 328–337 (2019). https://doi.org/10.1038/s41380-018-0335-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-018-0335-7
This article is cited by
-
A genome-wide association study of neutrophil count in individuals associated to an African continental ancestry group facilitates studies of malaria pathogenesis
Human Genomics (2024)
-
Clinical associations with a polygenic predisposition to benign lower white blood cell counts
Nature Communications (2024)
-
The lived experience of clozapine discontinuation in patients and carers following suspected clozapine-induced neutropenia
BMC Psychiatry (2023)
-
Mediation and longitudinal analysis to interpret the association between clozapine pharmacokinetics, pharmacogenomics, and absolute neutrophil count
Schizophrenia (2023)
-
Genome-wide association analyses of symptom severity among clozapine-treated patients with schizophrenia spectrum disorders
Translational Psychiatry (2022)