Abstract
For over a century, psychiatric disorders have been defined by expert opinion and clinical observation. The modern DSM has relied on a consensus of experts to define categorical syndromes based on clusters of symptoms and signs, and, to some extent, external validators, such as longitudinal course and response to treatment. In the absence of an established etiology, psychiatry has struggled to validate these descriptive syndromes, and to define the boundaries between disorders and between normal and pathologic variation. Recent advances in genomic research, coupled with large-scale collaborative efforts like the Psychiatric Genomics Consortium, have identified hundreds of common and rare genetic variations that contribute to a range of neuropsychiatric disorders. At the same time, they have begun to address deeper questions about the structure and classification of mental disorders: To what extent do genetic findings support or challenge our clinical nosology? Are there genetic boundaries between psychiatric and neurologic illness? Do the data support a boundary between disorder and normal variation? Is it possible to envision a nosology based on genetically informed disease mechanisms? This review provides an overview of conceptual issues and genetic findings that bear on the relationships among and boundaries between psychiatric disorders and other conditions. We highlight implications for the evolving classification of psychopathology and the challenges for clinical translation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Change history
14 March 2018
In the Table 1 legend, the reference numbers and symbols were not correctly presented in the footnotes. The corrected footnotes are presented below.
References
American Psychiatric Association. Diagnostic and statistical manual of mental disorders: DSM-5. 5th edn. Arlington, VA: American Psychiatric Association; 2013., xliv, p. 947.
Rudin E. Studien uber Vererbung und entstehung geistiger Storungen. I. Zur vererbung und neuentstehung der Dementia praecox (Studies on the inheritance and origin of mental illness. I. The problem of the inheritance and primary origin of dementia praecox). Monographien aus dem Gesamtgebiet der Neurologie und Psychiatrie, Number 12. Berlin: Springer; 1916.
Luxenburger H. Vorlaufiger Bericht uder psychiatrische Serienuntersuchungen und Zwillingen. Z Gesamt Neurol Psychiatr. 1928;116:297–326.
Heston LL. Psychiatric disorder in foster home reared children of schizophrenic mothers. Br J Psychiatry. 1966;112:819–25.
Kendler KS, Eaves LJ. Psychiatric genetics, review of psychiatry, Vol 24. Arlington, VA: American Psychiatric Publishing, Inc.; 2005.
Kendler KS. Twin studies of psychiatric illness: an update. Arch Gen Psychiatry. 2001;58:1005–14.
Kendler KS, Prescott CA, Myers J, Neale MC. The structure of genetic and environmental risk factors for common psychiatric and substance use disorders in men and women. Arch Gen Psychiatry. 2003;60:929–37.
Kety SS. The significance of genetic factors in the etiology of schizophrenia: results from the national study of adoptees in Denmark. J Psychiatr Res. 1987;21:423–9.
Sprich S, Biederman J, Crawford MH, Mundy E, Faraone SV. Adoptive and biological families of children and adolescents with ADHD. J Am Acad Child Adolesc Psychiatry. 2000;39:1432–7.
Verhulst B, Neale MC, Kendler KS. The heritability of alcohol use disorders: a meta-analysis of twin and adoption studies. Psychol Med. 2015;45:1061–72.
Kendler KS, Aggen SH, Knudsen GP, Roysamb E, Neale MC, Reichborn-Kjennerud T. The structure of genetic and environmental risk factors for syndromal and subsyndromal common DSM-IV axis I and all axis II disorders. Am J Psychiatry. 2011;168:29–39.
Purcell SM, Wray NR, Stone JL, Visscher PM, O’Donovan MC, et al. International Schizophrenia Consortium Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460:748–52.
Stefansson H, Ophoff RA, Steinberg S, Andreassen OA, Cichon S, Rujescu D, et al. Common variants conferring risk of schizophrenia. Nature. 2009;460:744–7.
Stefansson H, Rujescu D, Cichon S, Pietilainen OP, Ingason A, Steinberg S, et al. Large recurrent microdeletions associated with schizophrenia. Nature. 2008;455:232–6.
International Schizophrenia Consortium. Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature. 2008;455:237–41.
Psychiatric GWAS Consortium Coordinating Committee. Genomewide association studies: history, rationale, and prospects for psychiatric disorders. Am J Psychiatry. 2009;166:540–56.
Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, et al. 10 Years of GWAS Discovery: biology, function, and translation. Am J Hum Genet. 2017;101:5–22.
Schizophrenia Working Group of the Psychiatric Genomics C. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Wray NR, Lee SH, Mehta D, Vinkhuyzen AA, Dudbridge F, Middeldorp CM. Research review: Polygenic methods and their application to psychiatric traits. J Child Psychol Psychiatry. 2014;55:1068–87.
Otowa T, Hek K, Lee M, Byrne EM, Mirza SS, Nivard MG, et al. Meta-analysis of genome-wide association studies of anxiety disorders. Mol Psychiatry. 2016;21:1485.
Duncan LE, Ratanatharathorn A, Aiello AE, Almli LM, Amstadter AB, Ashley-Koch AE et al. Largest GWAS of PTSD (N = 20 070) yields genetic overlap with schizophrenia and sex differences in heritability. Mol Psychiatry. 2017 Apr 25. doi:10.1038/mp.2017.77. [Epub ahead of print].
Cross-Disorder Group of the Psychiatric Genomics C, Lee SH, Ripke S, Neale BM, Faraone SV, Purcell SM, et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat Genet. 2013;45:984–94.
Charney AW, Ruderfer DM, Stahl EA, Moran JL, Chambert K, Belliveau RA, et al. Evidence for genetic heterogeneity between clinical subtypes of bipolar disorder. Transl Psychiatry. 2017;7:e993.
CNV Schizophrenia Working Groups of the Psychiatric Genomics Consortium. Contribution of copy number variants to schizophrenia from a genome-wide study of 41,321 subjects. Nat Genet. 2017;49:27–35.
Genovese G, Fromer M, Stahl EA, Ruderfer DM, Chambert K, Landen M, et al. Increased burden of ultra-rare protein-altering variants among 4877 individuals with schizophrenia. Nat Neurosci. 2016;19:1433–41.
Deciphering Developmental Disorders S. Prevalence and architecture of de novo mutations in developmental disorders. Nature. 2017;542:433–8.
Willsey AJ, Fernandez TV, Yu D, King RA, Dietrich A, Xing J, et al. De novo coding variants are strongly associated with Tourette disorder. Neuron. 2017;94:486–99. e489
Vorstman JA, Parr JR, Moreno-De-Luca D, Anney RJ, Nurnberger JI Jr., Hallmayer JF. Autism genetics: opportunities and challenges for clinical translation. Nat Rev Genet. 2017;18:362–76.
de la Torre-Ubieta L, Won H, Stein JL, Geschwind DH. Advancing the understanding of autism disease mechanisms through genetics. Nat Med. 2016;22:345–61.
Singh T, Kurki MI, Curtis D, Purcell SM, Crooks L, McRae J, et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat Neurosci. 2016;19:571–7.
Solovieff N, Cotsapas C, Lee PH, Purcell SM, Smoller JW. Pleiotropy in complex traits: challenges and strategies. Nat Rev Genet. 2013;14:483–95.
Malhotra D, Sebat J. CNVs: harbingers of a rare variant revolution in psychiatric genetics. Cell. 2012;148:1223–41.
Tesli M, Espeseth T, Bettella F, Mattingsdal M, Aas M, Melle I, et al. Polygenic risk score and the psychosis continuum model. Acta Psychiatr Scand. 2014;130:311–7.
Hatzimanolis A, Bhatnagar P, Moes A, Wang R, Roussos P, Bitsios P, et al. Common genetic variation and schizophrenia polygenic risk influence neurocognitive performance in young adulthood. Am J Med Genet B Neuropsychiatr Genet. 2015;168:392–401.
McIntosh AM, Gow A, Luciano M, Davies G, Liewald DC, Harris SE, et al. Polygenic risk for schizophrenia is associated with cognitive change between childhood and old age. Biol Psychiatry. 2013;73:938–43.
Roussos P, Giakoumaki SG, Zouraraki C, Fiullard JF, Karagiorga V-E, Tsapakis E-M, et al. The relationship of common risk variants and polygenic risk for schizophrenia to sensorimotor gating. Biol Psychiatry. 2016;79:988–96.
Kauppi K, Westlye LT, Tesli M, Bettella F, Brandt CL, Mattingsdal M, et al. Polygenic risk for schizophrenia associated with working memory-related prefrontal brain activation in patients with schizophrenia and healthy controls. Schizophr Bull. 2015;41:736–43.
Riglin L, Collishaw S, Richards A, Thapar AK, Maughan B, O’Donovan MC, et al. Schizophrenia risk alleles and neurodevelopmental outcomes in childhood: a population-based cohort study. Lancet Psychiatry. 2017;4:57–62.
Cross Disorder Group of the Psychiatric GWAS Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–9.
Nivard MG, Gage SH, Hottenga JJ, van Beijsterveldt CE, Abdellaoui A, Bartels M, et al. Genetic overlap between schizophrenia and developmental psychopathology: longitudinal and multivariate polygenic risk prediction of common psychiatric traits during development. Schizophr Bull. 2017;43:1197–207.
Jones HJ, Stergiakouli E, Tansey KE, Hubbard L, Heron J, Cannon M, et al. Phenotypic manifestation of genetic risk for schizophrenia during adolescence in the general population. JAMA Psychiatry. 2016;73:221–8.
Lo MT, Hinds DA, Tung JY, Franz C, Fan CC, Wang Y, et al. Genome-wide analyses for personality traits identify six genomic loci and show correlations with psychiatric disorders. Nat Genet. 2017;49:152–6.
Zheng J, Erzurumluoglu AM, Elsworth BL, Kemp JP, Howe L, Haycock PC, et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics. 2017;33:272–9.
Network and Pathway Analysis Subgroup of Psychiatric Genomics Consortium. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat Neurosci. 2015;18:199–209.
Sekar A, Bialas AR, de Rivera H, Davis A, Hammond TR, Kamitaki N, et al. Schizophrenia risk from complex variation of complement component 4. Nature. 2016;530:177–83.
Devor A, Andreassen OA, Wang Y, Maki-Marttunen T, Smeland OB, Fan CC, et al. Genetic evidence for role of integration of fast and slow neurotransmission in schizophrenia. Mol Psychiatry. 2017;22:792–801.
Gandal MJ, Leppa V, Won H, Parikshak NN, Geschwind DH. The road to precision psychiatry: translating genetics into disease mechanisms. Nat Neurosci. 2016;19:1397–407.
Breen G, Li Q, Roth BL, O’Donnell P, Didriksen M, Dolmetsch R, et al. Translating genome-wide association findings into new therapeutics for psychiatry. Nat Neurosci. 2016;19:1392–6.
Kathiresan S. Developing medicines that mimic the natural successes of the human genome: lessons from NPC1L1, HMGCR, PCSK9, APOC3, and CETP. J Am Coll Cardiol. 2015;65:1562–6.
Lahey BB, Applegate B, Hakes JK, Zald DH, Hariri AR, Rathouz PJ. Is there a general factor of prevalent psychopathology during adulthood? J Abnorm Psychol. 2012;121:971–7.
Kerekes N, Brandstrom S, Lundstrom S, Rastam M, Nilsson T, Anckarsater H. ADHD, autism spectrum disorder, temperament, and character: phenotypical associations and etiology in a Swedish childhood twin study. Compr Psychiatry. 2013;54:1140–7.
Plomin R, Haworth CM, Davis OS. Common disorders are quantitative traits. Nat Rev Genet. 2009;10:872–8.
Larsson H, Anckarsater H, Rastam M, Chang Z, Lichtenstein P. Childhood attention-deficit hyperactivity disorder as an extreme of a continuous trait: a quantitative genetic study of 8500 twin pairs. J Child Psychol Psychiatry. 2012;53:73–80.
Clarke TK, Lupton MK, Fernandez-Pujals AM, Starr J, Davies G, Cox S, et al. Common polygenic risk for autism spectrum disorder (ASD) is associated with cognitive ability in the general population. Mol Psychiatry. 2016;21:419–25.
Groen-Blokhuis MM, Middeldorp CM, Kan KJ, Abdellaoui A, van Beijsterveldt CE, Ehli EA, et al. Attention-deficit/hyperactivity disorder polygenic risk scores predict attention problems in a population-based sample of children. J Am Acad Child Adolesc Psychiatry. 2014;53:1123–9. e1126
Robinson EB, St Pourcain B, Anttila V, Kosmicki JA, Bulik-Sullivan B, Grove J, et al. Genetic risk for autism spectrum disorders and neuropsychiatric variation in the general population. Nat Genet. 2016;48:552–5.
Germine L, Robinson EB, Smoller JW, Calkins ME, Moore TM, Hakonarson H, et al. Association between polygenic risk for schizophrenia, neurocognition and social cognition across development. Transl Psychiatry. 2016;6:e924.
Insel T, Cuthbert B, Garvey M, Heinssen R, Pine DS, Quinn K, et al. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am J Psychiatry. 2010;167:748–51.
Insel TR Director’s Blog: Transforming diagnosis. https://www.nimh.nih.gov/about/directors/thomas-insel/blog/2013/transforming-diagnosis.shtml. 2013.
Hyman SE. The diagnosis of mental disorders: the problem of reification. Annu Rev Clin Psychol. 2010;6:12.11–25.
Kendler KS. Levels of explanation in psychiatric and substance use disorders: implications for the development of an etiologically based nosology. Mol Psychiatry. 2012;17:11–21.
Duncan L, Yilmaz Z, Gaspar H, Walters R, Goldstein J, Anttila V, et al. Significant locus and metabolic genetic correlations revealed in genome-wide association study of anorexia nervosa. Am J Psychiatry. 2017;174:850–8.
Robins E, Guze S. Establishment of diagnostic validity in psychiatric illness: its application to schizophrenia. Am J Psychiatry. 1970;126:983–7.
Kendler KS. Toward a scientific psychiatric nosology. Strengths and limitations. Arch Gen Psychiatry. 1990;47:969–73.
Tsuang M, Faraone S, Lyons M. Identification of the phenotype in psychiatric genetics. Eur Arch Psychiatry Clin Neurosci. 1993;243:131–42.
Kendler KS. Reflections on the relationship between psychiatric genetics and psychiatric nosology. Am J Psychiatry. 2006;163:1138–46.
Kendler KS. An historical framework for psychiatric nosology. Psychol Med. 2009;39:1935–41.
Scassellati C, Bonvicini C, Faraone SV, Gennarelli M. Biomarkers and attention-deficit/hyperactivity disorder: a systematic review and meta-analyses. J Am Acad Child Adolesc Psychiatry. 2012;51:1003–19. e1020
Okser S, Pahikkala T, Airola A, Salakoski T, Ripatti S, Aittokallio T. Regularized machine learning in the genetic prediction of complex traits. PLoS Genet. 2014;10:e1004754.
Wray NR, Yang J, Hayes BJ, Price AL, Goddard ME, Visscher PM. Pitfalls of predicting complex traits from SNPs. Nat Rev Genet. 2013;14:507–15.
Levinson DF, Mostafavi S, Milaneschi Y, Rivera M, Ripke S, Wray NR, et al. Genetic studies of major depressive disorder: why are there no genome-wide association study findings and what can we do about it? Biol Psychiatry. 2014;76:510–2.
Holland D, Wang Y, Thompson WK, Schork A, Chen CH, Lo MT, et al. Estimating effect sizes and expected replication probabilities from GWAS summary statistics. Front Genet. 2016;7:15.
Zuk O, Schaffner SF, Samocha K, Do R, Hechter E, Kathiresan S, et al. Searching for missing heritability: designing rare variant association studies. Proc Natl Acad Sci USA. 2014;111:E455–464.
Moutsianas L, Agarwala V, Fuchsberger C, Flannick J, Rivas MA, Gaulton KJ, et al. The power of gene-based rare variant methods to detect disease-associated variation and test hypotheses about complex disease. PLoS Genet. 2015;11:e1005165.
Sullivan PF, Agrawal A, Bulik CM, Andreassen OA, Borglum A, Breen G et al. Psychiatric genomics: an update and an agenda. BiorXiv. 2017. https://doi.org/10.1101/115600.
Boyle EA, Li YI, Pritchard JK. An expanded view of complex traits: from polygenic to omnigenic. Cell. 2017;169:1177–86.
Anttila V, Bulik-Sullivan B, Finucane H, Bras J, Duncan L, Escott-Price V et al. Analysis of shared heritability in common disorders of the brain. BioRXiv. 2017. https://doi.org/10.1101/048991.
Smoller JW. The use of electronic health records for psychiatric phenotyping and genomics. Am J Med Genet B Neuropsychiatr Genet. 2017 May 30. doi:10.1002/ajmg.b.32548. [Epub ahead of print].
Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J, et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12:e1001779.
Precision Medicine Initiative (PMI) Working Group. The Precision Medicine Initiative Cohort Program—Building a Research Foundation for 21st Century Medicine September 17, 2015. https://www.nih.gov/sites/default/files/research-training/initiatives/pmi/pmi-working-group-report-20150917-2.pdf.
Bush WS, Oetjens MT, Crawford DC. Unravelling the human genome-phenome relationship using phenome-wide association studies. Nat Rev Genet. 2016;17:129–45.
Riglin L, Collishaw S, Thapar AK, Dalsgaard S, Langley K, Smith GD, et al. Association of Genetic Risk Variants With Attention-Deficit/Hyperactivity Disorder Trajectories in the General Population. JAMA Psychiatry. 2016;73:1285–92.
Gulsuner S, Walsh T, Watts AC, Lee MK, Thornton AM, Casadei S, et al. Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network. Cell. 2013;154:518–29.
Willsey AJ, Sanders SJ, Li M, Dong S, Tebbenkamp AT, Muhle RA, et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell. 2013;155:997–1007.
Niarchou M, Zammit S, Lewis G. The Avon Longitudinal Study of Parents and Children (ALSPAC) birth cohort as a resource for studying psychopathology in childhood and adolescence: a summary of findings for depression and psychosis. Soc Psychiatry Psychiatr Epidemiol. 2015;50:1017–27.
Mors O, Perto GP, Mortensen PB. The Danish Psychiatric Central Research Register. Scand J Public Health. 2011;39(7 Suppl):54–57.
Lee PH, Baker JT, Holmes AJ, Jahanshad N, Ge T, Jung JY, et al. Partitioning heritability analysis reveals a shared genetic basis of brain anatomy and schizophrenia. Mol Psychiatry. 2016;21:1680–9.
Bearden CE, Thompson PM. Emerging global initiatives in neurogenetics: the enhancing neuroimaging genetics through meta-analysis (ENIGMA) consortium. Neuron. 2017;94:232–6.
Cuthbert BN. Research domain criteria: toward future psychiatric nosologies. Dialogues Clin Neurosci. 2015;17:89–97.
Evans DM, Davey Smith G. Mendelian randomization: new applications in the coming age of hypothesis-free causality. Annu Rev Genom Hum Genet. 2015;16:327–50.
Emdin CA, Khera AV, Natarajan P, Klarin D, Zekavat SM, Hsiao AJ, et al. Genetic association of waist-to-hip ratio with cardiometabolic traits, type 2 diabetes, and coronary heart disease. JAMA. 2017;317:626–34.
Voight BF, Peloso GM, Orho-Melander M, Frikke-Schmidt R, Barbalic M, Jensen MK, et al. Plasma HDL cholesterol and risk of myocardial infarction: a mendelian randomisation study. Lancet. 2012;380:572–80.
Vaucher J, Keating BJ, Lasserre AM, Gan W, Lyall DM, Ward J et al. Cannabis use and risk of schizophrenia: a Mendelian randomization study. Mol Psychiatry. 2017 Jan 24. doi:10.1038/mp.2016.252. [Epub ahead of print].
Pickrell JK, Berisa T, Liu JZ, Segurel L, Tung JY, Hinds DA. Detection and interpretation of shared genetic influences on 42 human traits. Nat Genet. 2016;48:709–17.
Schaefer GB, Mendelsohn NJ, Professional P, Guidelines C. Clinical genetics evaluation in identifying the etiology of autism spectrum disorders: 2013 guideline revisions. Genet Med. 2013;15:399–407.
Samocha KE, Robinson EB, Sanders SJ, Stevens C, Sabo A, McGrath LM, et al. A framework for the interpretation of de novo mutation in human disease. Nat Genet. 2014;46:944–50.
Robinson EB, Samocha KE, Kosmicki JA, McGrath L, Neale BM, Perlis RH, et al. Autism spectrum disorder severity reflects the average contribution of de novo and familial influences. Proc Natl Acad Sci USA. 2014;111:15161–5.
Zhu X, Need AC, Petrovski S, Goldstein DB. One gene, many neuropsychiatric disorders: lessons from Mendelian diseases. Nat Neurosci. 2014;17:773–81.
Lichtenstein P, Carlstrom E, Rastam M, Gillberg C, Anckarsater H. The genetics of autism spectrum disorders and related neuropsychiatric disorders in childhood. Am J Psychiatry. 2010;167:1357–63.
Polderman TJ, Benyamin B, de Leeuw CA, Sullivan PF, van Bochoven A, Visscher PM, et al. Meta-analysis of the heritability of human traits based on fifty years of twin studies. Nat Genet. 2015;47:702–9.
Song J, Bergen SE, Kuja-Halkola R, Larsson H, Landen M, Lichtenstein P. Bipolar disorder and its relation to major psychiatric disorders: a family-based study in the Swedish population. Bipolar Disord. 2015;17:184–93.
Cardno AG, Rijsdijk FV, Sham PC, Murray RM, McGuffin P. A twin study of genetic relationships between psychotic symptoms. Am J Psychiatry. 2002;159:539–45.
McGuffin P, Rijsdijk F, Andrew M, Sham P, Katz R, Cardno A. The heritability of bipolar affective disorder and the genetic relationship to unipolar depression. Arch Gen Psychiatry. 2003;60:497–502.
Bulik CM, Thornton LM, Root TL, Pisetsky EM, Lichtenstein P, Pedersen NL. Understanding the relation between anorexia nervosa and bulimia nervosa in a Swedish national twin sample. Biol Psychiatry. 2010;67:71–77.
Mathews CA, Grados MA. Familiality of Tourette syndrome, obsessive-compulsive disorder, and attention-deficit/hyperactivity disorder: heritability analysis in a large sib-pair sample. J Am Acad Child Adolesc Psychiatry. 2011;50:46–54.
Cederlof M, Thornton LM, Baker J, Lichtenstein P, Larsson H, Ruck C, et al. Etiological overlap between obsessive-compulsive disorder and anorexia nervosa: a longitudinal cohort, multigenerational family and twin study. World Psychiatry. 2015;14:333–8.
Tick B, Bolton P, Happe F, Rutter M, Rijsdijk F. Heritability of autism spectrum disorders: a meta-analysis of twin studies. J Child Psychol Psychiatry. 2016;57:585–95.
Sartor CE, Grant JD, Lynskey MT, McCutcheon VV, Waldron M, Statham DJ, et al. Common heritable contributions to low-risk trauma, high-risk trauma, posttraumatic stress disorder, and major depression. Arch General Psychiatry. 2012;69:293–9.
Stein MB, Jang KL, Taylor S, Vernon PA, Livesley WJ. Genetic and environmental influences on trauma exposure and posttraumatic stress disorder symptoms: a twin study. Am J Psychiatry. 2002;159:1675–81.
Acknowledgements
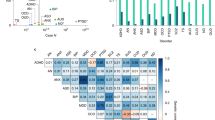
Supported in part by NIH awards K24MH094614 (JWS), the Research Council of Norway (223273) and KG Jebsen Stiftelsen (OAA). Dr. Smoller is a Tepper Family MGH Research Scholar. “All of Us” is a service mark of the U.S. Department of Health and Human Services. The authors thank Nicholas Merriam for assistance with producing the figure.
Author contributions
Dr. Smoller is an unpaid member of the Scientific Advisory Board of PsyBrain Inc. and the Bipolar/Depression Research Community Advisory Panel of 23andMe. Dr. Andreassen has received speaker’s honorarium from Lundbeck. The remaining authors have no disclosures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: Modifications have been made to the Table 1 legend. Full information regarding corrections made can be found in the erratum/correction article for this article.
Rights and permissions
About this article
Cite this article
Smoller, J.W., Andreassen, O.A., Edenberg, H.J. et al. Psychiatric genetics and the structure of psychopathology. Mol Psychiatry 24, 409–420 (2019). https://doi.org/10.1038/s41380-017-0010-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-017-0010-4
This article is cited by
-
Characterizing the phenotypic and genetic structure of psychopathology in UK Biobank
Nature Mental Health (2024)
-
Understanding Environmental Exposures and ADHD: a Pathway Forward
Prevention Science (2024)
-
The Promise and Challenges of Integrating Biological and Prevention Sciences: A Community-Engaged Model for the Next Generation of Translational Research
Prevention Science (2024)
-
Association between Autism Spectrum Disorder (ASD) and vision problems. A systematic review and meta-analysis
Molecular Psychiatry (2023)
-
The need for evidence-based updating of ICD and DSM models of psychotic and mood disorders
Molecular Psychiatry (2023)