Abstract
Although most women with luminal breast cancer do well on endocrine therapy alone, some will develop fatal recurrence thereby necessitating the need to prospectively determine those for whom additional cytotoxic therapy will be beneficial. Categorical combinations of immunohistochemical measures of ER, PR, HER2, and KI67 are traditionally used to classify patients into luminal A-like and B-like subtypes for chemotherapeutic reasons, but this may lead to the loss of prognostically relevant information. Here, we compared the prognostic value of quantitative measures of these markers, combined in the IHC4-score, to categorical combinations in subtypes. Using image analysis-based scores for all four markers, we computed the IHC4-score for 2498 patients with luminal breast cancer from two European study populations. We defined subtypes (A-like (ER + and PR + : and HER2- and low KI67) and B-like (ER + and/or PR + : and HER2 + or high KI67)) by combining binary categories of these markers. Hazard ratios and 95% confidence intervals for associations with 10-year breast cancer-specific survival were estimated in Cox proportional-hazard models. We accounted for clinical prognostic factors, including grade, tumor size, lymph-nodal involvement, and age, by using the PREDICT-score. Overall, Subtypes [hazard ratio (95% confidence interval) B-like vs. A-like = 1.64 (1.25–2.14); P-value < 0.001] and IHC4-score [hazard ratio (95% confidence interval)/1 standard deviation = 1.32 (1.20–1.44); P-value < 0.001] were prognostic in univariable models. However, IHC4-score [hazard ratio (95% confidence interval)/1 standard deviation = 1.24 (1.11–1.37); P-value < 0.001; likelihood ratio chi-square (LRχ2) = 12.5] provided more prognostic information than Subtype [hazard ratio (95% confidence interval) B-like vs. A-like = 1.38 (1.02–1.88); P-value = 0.04; LRχ2 = 4.3] in multivariable models. Further, higher values of the IHC4-score were associated with worse prognosis, regardless of subtype (P-heterogeneity = 0.97). These findings enhance the value of the IHC4-score as an adjunct to clinical prognostication tools for aiding chemotherapy decision-making in luminal breast cancer patients, irrespective of subtype.
Similar content being viewed by others
Introduction
Breast cancer is the most common malignancy and the leading cause of cancer-related mortality among women worldwide [1]. With the advent and increasing uptake of screening programs, the incidence of early stage, hormone receptor positive (HR + )/luminal-like [estrogen receptor positive (ER + ) and/or progesterone receptor positive (PR + )] breast cancer has continued to rise [2, 3]. Accounting for almost 70% of all cases, luminal-like tumors comprise the majority of breast cancers in Western populations [4]. These tumors are notably heterogeneous, encompassing subtypes with distinct molecular profiles and clinical outcomes [5,6,7,8]. Based on gene expression profiling, two main subtypes of luminal-like tumors have been identified. Denoted as luminal A and B, these subtypes are differentiated by their relative expressions of hormone and proliferation-related genes and, in a subset of luminal B tumors, by the amplification of the human epidermal growth factor receptor 2 (HER2/neu) gene [5, 9].
Although most women with luminal-like disease do well on endocrine therapy alone, some of them develop fatal recurrence thereby necessitating the need for additional cytotoxic therapy. Owing to the debilitating side-effects associated with chemotherapy, the need to prospectively distinguish women for whom its addition will be beneficial from those for whom this may not be needed remains a challenge in translational breast cancer research [10]. Several prognostic tools have been developed to address this, including those that rely on standard clinical prognostic factors [11,12,13] and others, such as the IHC4-score, that are based on immunohistochemical measures on ER, PR, HER2, and KI67 [13]. In addition, some international guidelines have endorsed the use of immunohistochemistry-based (i.e., luminal A-like and B-like) subtypes together with multiparameter molecular tests [14,15,16,17] to aid chemotherapy decision-making [18,19,20]. However, because standard immunohistochemistry-based luminal A-like/B-like subtype definition is based on dichotomous categories of the individual immunohistochemical markers, this may lead to the loss of prognostically relevant information. Nonetheless, it remains unclear whether quantitative measures of ER, PR, HER2, and KI67 provide more prognostic information than categorial combinations in breast cancer subtypes.
Multiparameter molecular testing is one method of quantification of hormone and proliferation-related genes that has been shown to be prognostic in breast cancer [14,15,16,17], but this is expensive, and it remains unclear whether it improves the prognostication of clinical tools such as PREDICT. PREDICT [11, 12] is a popular, breast cancer prognostication, and treatment benefit tool that remains the only breast cancer prognostication tool to be endorsed by the American Joint Committee on Cancer to date. Like traditional clinical prognostic factors in PREDICT, immunohistochemical markers are cheap to perform, widely available, and are typically assessed as part of the routine workup for most breast cancer patients. It has previously been shown that combined visual assessments of ER, PR, HER2, and KI67 in the IHC4-score provided additional prognostic information to standard clinicopathological factors and contained comparative prognostic information to the 21-gene (Oncotype DX) panel test [13].
Owing to the limitations of visual scoring, particularly for KI67 [21, 22], automated methods have been suggested as potential alternatives. We have previously demonstrated the independent prognostic value of automated scores for ER, PR, HER2, and KI67 separately, but it remains unclear whether combining these in the IHC4-score algorithm will provide additional prognostic information to immunohistochemistry-based subtypes or other clinical prognostic factors. Further, although patients with luminal A-like breast cancer generally have better clinical outcomes than those with luminal B-like disease [5, 9, 16], it is unknown if the dynamic range of the IHC4-score could be leveraged to further stratify these patients into clinically relevant subgroups for treatment decision-making.
Our primary aim in this study was, therefore, to investigate the comparative prognostic performance of image analysis-based, quantitative, measures of ER, PR, HER2, and KI67, combined in the IHC4-score, vs. categorical combinations of these markers in luminal (A-like/B-like) breast cancer subtype. As a secondary aim, we evaluated the prognostic significance of the image analysis-based IHC4-score in relation to clinical prognostic factors, combined in the clinical treatment score (C-score) and PREDICT-score.
Materials and Methods
Study population
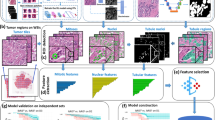
The current analysis included 2498 patients with luminal-like invasive breast cancer from two study populations from Poland (N = 558) and the United Kingdom (N = 1940). The analysis comprised of women with luminal-like, i.e., ER + and/or PR + tumors and for whom we also had complete data on image analysis-based scores for ER, PR, HER2, and KI67 (Fig. 1). These scores were obtained by digital image analysis of tissue microarrays and analyzed as part of other projects [23,24,25] within the Breast Cancer Association Consortium. Details of both study populations have been previously described [26, 27], but in brief: The Polish Breast Cancer Study is a population-based study in Poland that enrolled women 20–74 years with histologically or cytologically confirmed breast cancer at five participating hospitals in Warsaw and Lodz over a three-year period between 2000 and 2003 [27]. The Study of Epidemiology and Risk Factors in Cancer Heredity (SEARCH) is a population-based study that began in the UK in 1996 [26]. Patients were ascertained through the Eastern Cancer Registration and Information Center and included women < 55 years of age diagnosed with invasive breast cancer between 1991 and mid-1996 who were alive at the start of the study and those < 70 years who were diagnosed from mid-1996 onwards.
Consort diagram indicating the total number of patients included in this analysis. The patients were women with clinically determined invasive luminal-like breast cancer from two study populations in Europe (the Polish Breast Cancer Study (PBCS) in Poland and the Study of Epidemiology and Risk Factors in Cancer Heredity (SEARCH) in the United Kingdom) with complete information on image analysis-based scores of ER, PR, HER2, and KI67, as well as other relevant clinicopathological and follow-up data
Data on relevant clinicopathological characteristics, including ER, PR, HER2, histologic grade, tumor size, nodal involvement, endocrine therapy, and systemic therapy were obtained from clinical records. Patients were followed up from recruitment for the development of outcomes of interest, i.e., breast cancer-specific deaths. Among the patients included in this analysis, a total of 316 breast cancer-specific deaths (N = 255 and 61, for the SEARCH and Polish breast cancer studies, respectively) occurred over a median follow-up period of 7.05 years (8.01 and 5.0 years for the SEARCH and Polish study populations, respectively). In both studies, deaths were ascertained through linkage to registries, as well as by curating clinical records. Ethical approvals were obtained from local ethics committees and all participants provided written informed consent.
Immunostaining and scoring of tissue microarrays for ER, PR, HER2, and KI67
Staining for all four markers was performed in the respective study groups by using standard laboratory techniques (Supplementary Table 1). Tissue microarray sections for ER and PR were stained by using mouse monoclonal antibodies 6F11/2 (Novocastra) and PR636 (Dako) clones, respectively, while tissue microarrays for HER2 and KI67 were stained using Herceptest kit K5207 (Dako) and MIB-1 (Dako), respectively. Dichotomous categories (positive and negative) of ER, PR, and HER2 were obtained from clinical records. ER and PR were scored using the Allred scoring method and values > 2 ( > 10% positive cells) were considered positive. For HER2, 3 + on immunohistochemistry or HER2 amplification on fluorescent in situ hybridization were HER2 + . Quantitative measures on these markers were generated by using digital pathology image analysis performed in two institutions in the UK: the Cancer Research Institute in Cambridge and the Institute of Cancer Research (ICR) in London. ER, PR, and HER2 were scored in Cambridge while KI67 was scored at the ICR. Both institutions used the Ariol machine (Leica Biosystems, Newcastle UK) for scoring. Ariol has functionality for the automatic separation of malignant and non-malignant cells based on their shape and size characteristics and, by using color deconvolution, it can detect (3–3'-diaminobenzidine) positive and negative (hematoxylin) staining malignant epithelial cells. Details of the optimized Ariol algorithms and protocols that were used for the scoring of each of these four markers have been previously described [23, 24]. In brief, for ER, PR, and KI67 nuclear staining, the Ariol system was tuned to distinguish between malignant and non-malignant cells and to count positively and negatively staining malignant cells. Based on the number of positive and number of negative tumor nuclei presented by the machine, the percentage of cells stained (0–100%) was calculated as the ratio of positive nuclei to the sum of positive and negative nuclei per tissue core. For HER2, the US Food and Drug Administration-approved Herceptest score [28] (0, 1 + , 2 + , 3 + ), generated to American Society of Clinical Oncology/College of American Pathologists guidelines was calculated by the system. As previously reported [23, 24], we observed good agreement with standardized pathologists scores for ER (observed agreement = 90%; kappa = 0.76), PR (observed agreement = 84%; kappa = 0.66), HER2 (observed agreement = 90%, kappa = 0.69), and KI67 (observed agreement = 87%; kappa = 0.64).
Subtype definition based on binary categories of ER, PR, HER2, and KI67
Subtypes were defined according to the published St Gallen criteria [18] as follows: Luminal A-like: tumors that homogeneously expressed ER and PR (i.e., ER + and PR + ) in addition to being HER2– and low proliferating (image analysis-based KI67 ≤ 12%). We have previously reported a cutoff point of 12% for image analysis-based KI67, which corresponded to a visual score of 25%, to provide the best discrimination in terms of survival in this population [25], hence its adoption here. The luminal B-like subtype comprised tumors that were: (a) ER + and/or PR + and high proliferating (image analysis-based KI67 > 12%); (b) ER + and/or PR + and HER2 + .
Quantitative IHC4-score generation
The average score for ER, PR, HER2, and KI67 across the total number of cores per patient was taken as the patient’s score on each marker. IHC4-scores were generated using the published algorithm [13]:
The ER10 variable was calculated by dividing the ER% score for each patient by a factor of 10 to generate a range of values 0–10. In the original algorithm, the ER10 variable was generated by dividing the H-score (30–300) by a factor of 30 to give values ranging from 1–10. All other components of the IHC4-score were the same as in the published algorithm [13].
Clinical prognostic factors
We accounted for standard clinical prognostic factors, including age at diagnosis, tumor size, histologic grade, and number of lymph nodes involved using two methods. The first was based on the C-score reported by Cuzick et al. [13]:
Where N is the number of nodes (0, 1–3, 4 + ), T is tumor size ( ≤ 1 cm, > 1 to ≤ 2 cm, > 2 to ≤ 3 cm, > 3 cm), Gr is grade (1–3) and Age being the patients age at diagnosis ( < 65, ≥ 65 years). Data on specific treatment regimen received by each patient were not available; as such, the aromatase inhibitor (AI) vs. tamoxifen component was not computed.
The second was based on the parameters used for the PREDICT prognostication model:
We did not have information on mode of detection, therefore, could not compute the screen-detected vs. interval (or non-screen-detected) component of the PREDICT-score. Notably, organized mammography screening was not available in Poland during the study. All other components were as in the original equation.
Statistical analysis
Participant’s ages were categorized as < 35, 35–50, 50–65, and > 65 years. Chi-square, for categorical variables, and non-parametric Kruskal–Wallis tests, for continuous variables, were used to assess the frequencies of tumor clinicopathological characteristics (including age at diagnosis, histologic grade, stage, morphology, size, lymph-nodal status, and treatment), overall and by study population. Histograms and box plots were used to assess the distribution of the IHC4-score, overall and by study population. In univariable Cox proportional hazard regression models, we assessed the associations between subtype (B-like vs. A-like) and continuous measures of the IHC4-score, C-score, and PREDICT-score with 10-year breast cancer-specific survival. Additionally, for subtypes (luminal A-like and B-like) and quartiles (Q1–Q4) of the IHC4-score, we examined 10-year breast cancer-specific survival in Kaplan–Meier survival curves. For the IHC4-score this analysis was further stratified by nodal status (i.e., node-negative and node-positive). In multivariable Cox proportional models, we adjusted for study population (in combined analysis), treatment, and other standard clinical factors, including age at diagnosis, tumor size, histologic grade, and lymph-nodal involvement. These features were combined by means of the C-score and PREDICT-score. The relative contributions of the C-score and PREDICT-score to a prognostic model were determined by assessing the change in likelihood ratio chi-square (∆LRχ2) when either of these was removed from the full model. We used loglikelihood and LRχ2 values to compare model fit between prognostic scores. All analyses were performed overall and following stratification by study population. In subtype-specific analysis, we examined the prognostic value of the IHC4-score within each of the luminal-like breast cancer subtypes. To determine whether automated IHC4-score can be used to further stratify luminal-like breast cancer patients into prognostically relevant subgroups, we dichotomized the IHC4-score at the mean + 1 standard deviation threshold and examined associations with 10-year breast cancer-specific survival in Kaplan–Meier curves and in multivariable Cox proportional hazard models. Violations of the proportionality assumption of the Cox model were assessed by modeling the predictors as time-varying covariates. As part of sensitivity analysis, we redefined luminal-like breast cancer subtype by using a cut-off point of ≥ 1% on ER and PR [20] and examined the prognostic value of the IHC4-score in the resulting luminal A-like and B-like subtypes. Owing to the low prevalence of chemotherapy in this population (~7%), we could not perform analyses stratified by chemotherapy; instead, the few women who received chemotherapy (N = 178) were excluded from the survival analysis. All tests were two-sided, and analyses were conducted using Stata statistical software version 14.1 (StataCorp, Lakeway Drive, TX, USA).
Results
Description of study population
As shown in Table 1, the majority (86%) of the patients were between the ages of 35 and 65 years at diagnosis, with women from the SEARCH study being younger than those from the Polish study on average (P-value < 0.001). Overall, and in both study populations, most of the tumors (82%) were of intermediate or low histologic grade and fewer (18%) were high grade. Similarly, most (~97%) of the tumors from both study populations were stage I and II. Small ( < 2 cm) and intermediate (2–5 cm) size tumors were predominant in both studies (98% and 97% for SEARCH and Polish studies, respectively). The majority (70%) of the tumors were invasive ductal carcinomas; however, the Polish study had a substantially higher frequency of ‘other’’ non-ductal or lobular invasive carcinomas than the SEARCH study (28% vs. 7%; P-value < 0.001). A higher proportion (61%) of the patients had node-negative than positive (39%) disease, which was slightly fewer in the Polish (56%) than the SEARCH (62%) study population. Only 9% of the patients were HER2 + and this did not differ by study population (P-value = 0.12).
Dynamic range of image analysis-based scores for immunohistochemistry markers in the IHC4-score
Image analysis produced quantitative scores (0–100%) for each of the three nuclear markers (Supplementary Fig. 1) with median (standard deviation) scores of 62% (34), 57% (38), and 9% (11) for ER, PR, and KI67, respectively. When combined with data on HER2 in the IHC4 algorithm, these markers produced an IHC4-score with a dynamic range of −148 to 289 (mean = 33, standard deviation = 65; Supplementary Fig. 2A). The distribution of the IHC4-score differed by study population, with patients from the Polish study generally having higher values than those from the SEARCH study population (Supplementary Fig. 2B).
Associations between Subtype, IHC4-score, C-score, and PREDICT-score with 10-year breast cancer-specific survival
In Kaplan–Meier curves (Fig. 2) and in univariable models (Table 2), Subtype (luminal B-like vs. A-like) [hazard ratio (95% confidence interval) = 1.64 (1.25–2.14); P-value < 0.001] and IHC4-score [hazard ratio (95% confidence interval)/1 standard deviation = 1.32 (1.20–1.44); P-value < 0.001] were significantly associated with survival overall. However, the IHC4-score (LRχ2 = 40.1) provided more prognostic information than subtype (LRχ2 = 23.4). A similar pattern of association was seen in both the Polish and SEARCH study populations (Table 2).
Both the C-score [hazard ratio (95% confidence interval)/1 standard deviation = 1.78 (1.67–1.90); P-value = < 0.001], and PREDICT-score [hazard ratio (95% confidence interval)/1 standard deviation = 2.34 (2.07–2.65); P-value < 0.001] were associated with 10-year breast cancer-specific survival, with the model based on PREDICT-score (LRχ2 = 178.5) fitting better than that based on the C-score (LRχ2 = 168.2). In addition, PREDICT-score provided more prognostic information [∆LRχ2 = 9.0; P-value = 0.002] than the C-score [∆LRχ2 = 5.6; P-value = 0.02] in this population, hence it was used as the adjustment factor in multivariable models.
The IHC4-score remained significantly associated with survival, overall [hazard ratio (95% confidence interval)/1 standard deviation = 1.24 (1.11–1.37); P-value< 0.001] and in both the Polish [hazard ratio (95% confidence interval)/1 standard deviation = 1.46 (1.17–1.74); P-value 0.002] and SEARCH [hazard ratio (95% confidence interval)/SD = 1.19 (1.05–1.33); P-value = 0.007] study populations after adjusting for PREDICT-score (Table 2). Further, when we performed analyses stratified by lymph-nodal involvement, higher values of the IHC4-score were associated with worse breast cancer-specific survival in women with node-negative (Fig. 3a; logrank P-value = 0.002) and node-positive (Fig. 3b; logrank P-value = 0.002) disease.
Association between IHC4-score and 10-year breast cancer-specific survival within luminal A-like and B-like subtypes of breast cancer
Overall, the IHC4-score was associated with survival in both luminal A-like [hazard ratio (95% confidence interval)/1 standard deviation = 1.20 (0.98–1.43); P-value = 0.07) and luminal B-like [hazard ratio (95% confidence interval)/1 standard deviation = 1.22 (1.03–1.41); P-value = 0.01] subtypes (P-value for heterogeneity = 0.97). Although the hazard ratio estimate was slightly attenuated in luminal A-like [hazard ratio (95% confidence interval)/1 standard deviation = 1.10 (0.88, 1.33); P-value = 0.37)] subtype defined using 1% threshold for ER and PR, the estimates remained essentially the same for luminal B-like tumors [hazard ratio (95% confidence interval)/1 standard deviation = 1.22 (1.03–1.41); P-value = 0.02].
We observed an overlap in the distribution of the IHC4-score between luminal A-like and B-like subtypes, overall and in both study populations (Fig. 4). Thus, by comparing women with IHC4-score above the mean + 1 standard deviation (denoted as high IHC4-score) with those that had scores below this threshold (low IHC4-score), we observed those with high IHC4-score to have significantly worse survival outcomes than those with low scores, overall and in both study populations (Fig. 4). Following adjustment for the PREDICT-score in Cox proportional hazard models, we observed differences in high vs. low IHC-score categories in the luminal B-like subtype overall and in both the Polish and SEARCH study populations (Table 3). However, differences did not attain statistical significance in the luminal A-like subtype in the Polish study, which is likely due to the limited number of events (number (deaths/cases) = 7/99 and 5/41 in the low and high IHC4-score categories, respectively) in this sub-population (Table 3).
Distribution of the IHC4-score in luminal A-like and B-like subtypes of breast cancer and Kaplan–Meier survival curves for the associations between subtypes of luminal-like breast cancer stratified by levels of IHC4-score, overall (a and b), and among patients in the Polish (c and d) and SEARCH (e and f) study populations
Discussion
In the current analysis, we combined quantitative scores of ER, PR, HER2, and KI67 in the IHC4-score and compared its prognostic performance with categorical combinations defining A-like and B-like subtypes of luminal-like breast cancer. We also investigated the prognostic value of the IHC4-score in relation to other clinical prognostic factors, combined in the C-score and PREDICT-score. Our findings show that the IHC4-score provided more prognostic information than immunohistochemistry-based subtyping of luminal-like breast cancer. Additionally, the IHC4-score was associated with survival in both the luminal A-like and B-like subtypes after adjusting for the PREDICT-score, which provided more prognostic information than the C-score in this study population. Our findings also suggest that the dynamic range of the IHC4-score can be leveraged to provide prognostic information in both node-negative and node-positive disease and to further stratify women with luminal A-like or B-like breast cancer into subgroups with different prognoses.
Successive St. Gallen panels [18, 20, 29] have endorsed the use of immunohistochemical markers for the surrogate definition of the A-like and B-like subtypes of luminal-like breast cancer for deciding systemic therapy options. Based on current guidelines [20], most patients with the luminal B-like subtype receive chemotherapy in addition to standard endocrine treatment. Conversely, endocrine therapy is the mainstay of treatment for luminal A-like disease, except for a subset of patients for whom the addition of chemotherapy may be warranted. Some indications for cytotoxic therapy in luminal A-like patients include high 21-gene recurrence score and high-risk status on the 70-gene panel [18, 20]. However, dichotomization of ER, PR, HER2, and KI67 for subtype definition may be associated with the loss of prognostically relevant information. Moreover, intratumor heterogeneity may lead to discordant classifications of breast cancer subtypes since conventional subtyping approaches assign patients into discrete categories based on the topographical region of the tumor that has been sampled [30]. By combining quantitative data on all four markers, the IHC4-score has a dynamic range that can allow for the evaluation of dose–response relationships with survival thereby avoiding the pitfalls associated with assigning patients into discrete categories [13, 31].
We have previously demonstrated the prognostic value of automated scores on the individual immunohistochemical markers that are currently used to define breast cancer subtypes [23, 25]. However, the prognostic significance of combining automated scores on all four markers has not been previously studied. In this analysis, we showed that a combined score of these markers is significantly associated with 10-year breast cancer-specific survival even after adjustment for the PREDICT-score. This finding is particularly relevant given that, unlike expression-based assays, the IHC4-score and PREDICT-score are based on routinely determined clinicopathological and immunohistochemical parameters in clinical practice thereby making them potentially available to many patients with luminal-like breast cancer. Although multiparameter molecular tests [14,15,16,17] also quantify the expression of several genes, including those related to ER, PR, HER2, and proliferation, these are expensive, not widely available, and it remains unclear whether they provide additional prognostic information to PREDICT.
Overall, the range of clinical applications of the IHC4-score is still evolving [32,33,34]. One previous study documented its capacity to distinguish breast cancer patients with intermediate Nottingham Prognostic Index into subgroups with low and high-risk of recurrence [32, 33]. Our findings from the current study provide clues into other potential uses of the IHC4-score in clinical practice. For instance, the finding of significant associations with survival for the IHC4-score and PREDICT-score in luminal A-like and B-like patients suggest that chemotherapy recommendations should be based on its predicted absolute benefit regardless of immunohistochemistry-based subtype. In addition, the overlap in the distribution of the IHC4-score between A-like and B-like tumors that we observed may be indicative of the need to leverage quantitative information on immunohistochemical markers to provide additional prognostic information beyond what is contained in immunohistochemistry-based subtypes.
Despite its potential benefits, the widespread adoption of the IHC4-score may be affected by concerns regarding its analytical validity. There is the perception that immunohistochemical methods lack reproducibility and suffer from variable degrees of between-laboratory discordance. However, Dodson et al. [35] showed in a recent multi-institutional analytical validity study that risk of recurrence estimates with the IHC4 + C-score were tolerant of variations in staining and scoring across different laboratories. Moreover, several international efforts have led to the publication of guidelines that will help to enhance the validity of assays performed in laboratories across the globe [36,37,38].
An important strength of this study was that we used a digital image analysis-based approach for the centralized scoring of all four immunohistochemical markers, which yielded quantitative scores on ER, PR, and KI67. Although visual scoring by a trained expert can guarantee accurate discrimination between epithelial and stromal cells and between malignant and benign epithelial cells, this method is labor intensive and suffers from varying degrees of intra- and inter-observer discordance [21, 22]. In contrast, image analysis-based methods are high-throughput, highly reproducible, and show good agreement with pathologist’s-based scores [23, 24, 39,40,41,42,43,44]. Previous studies looking at the IHC4-score have focused on its prognostic performance in luminal-like breast cancer as a homogeneous entity and none has evaluated its prognostic value in relation to PREDICT-score. To the best of our knowledge, ours is the first study to specifically investigate the prognostic value of the IHC4-score in relation to subtypes of luminal-like breast cancer, as well as the PREDICT-score. Furthermore, our analysis involved patients from two study populations for whom the IHC4-score had never been applied, which allowed us to compare the results across populations.
In terms of limitations, despite their promise as alternatives to visual scoring, concerns exist regarding the accuracy of image analysis-based methods in discriminating between malignant and benign epithelial cells. We have previously documented the underestimation of hazard ratio estimates when comparing image analysis with pathologist’s-based scores [23, 24]. This was due to the attenuation of the performance of the image analysis algorithm in the presence of mixed cell populations. However, we utilized tissue microarrays for this study, which may limit the impact of mixed cell populations on our results since cores on tissue microarrays are typically enriched for tumor cells. Also, at the time most of our patients were recruited, ER + and PR + tumors were defined based on Allred score of > 2, corresponding to a proportion score of > 10%. However, this threshold has evolved over time, with current recommendations stipulating a cutoff point of ≥ 1% [20]. Nonetheless, many studies still utilize the 10% threshold. Moreover, when we redefined subtypes based on the 1% threshold as part of sensitivity analysis our results remained essentially the same.
This study was not designed to assess the predictive value of the IHC4-score for chemotherapy response. In view of results from a few studies showing poor chemotherapy response in high-risk luminal A-like tumors [45], an important area of future research will be the determination of the predictive value of the IHC4-score for chemotherapy response in patients with luminal A-like disease. Interestingly, recent findings suggest that the Magee Equation [46], another inexpensive tool that is based on ER, PR, HER2, and KI67 in addition to the Nottingham score and tumor size, can be used to predict pathologic response to neoadjuvant chemotherapy in ER + /HER2-negative/equivocal breast cancer [47]. The PREDICT-score also provides information on estimated treatment benefit for both ER + and ER- breast cancer patients. However, both the Magee Equation and PREDICT-score are based on visual assessments of all four immunohistochemical markers and it remains unclear whether the incorporation of automated measures can help refine the discriminatory accuracy of both tools, particularly in women with equivocal scores.
In conclusion, findings from this study showed that quantitative measures of ER, PR, HER2, and KI67, combined in the IHC4-score, provided more prognostic information than categorical combinations in immunohistochemistry-based subtypes of luminal-like breast cancer. In addition, the IHC4-score was associated with 10-year breast cancer-specific survival in patients with both luminal A-like and B-like tumors even after accounting for PREDICT-score, which was the strongest prognostic factor in this population. Taken together, these findings support the view that the IHC4-score can be used as an inexpensive adjunct to other clinical prognostication tools to aid treatment decision-making in patients with luminal-like breast cancer, irrespective of subtype. Given the prognostic strength of the PREDICT-score that we observed, further studies will be needed to determine whether combining the IHC4-score and PREDICT-score will provide superior prognostic information than PREDICT-score plus HER2 and KI67.
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–86.
Li CI, Daling JR, Malone KE. Incidence of invasive breast cancer by hormone receptor status from 1992 to 1998. J Clin Oncol. 2003;21:28–34.
Rosenberg PS, Barker KA, Anderson WF. Estrogen receptor status and the future burden of invasive and in situ breast cancers in the United States. J Natl Cancer Inst. 2015;107:djv159.
Howlader N, Altekruse SF, Li CI, Chen VW, Clarke CA, Ries LAG, et al. US incidence of breast cancer subtypes defined by joint hormone receptor and HER2 Status. J Natl Cancer Inst. 2014;106:dju055.
Sørlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA. 2001;98:10869–74.
Ciriello G, Sinha R, Hoadley KA, Jacobsen AS, Reva B, Perou CM, et al. The molecular diversity of Luminal A breast tumors. Breast Cancer Res Treat. 2013;141:409–20.
Howell SJ. Advances in the treatment of luminal breast cancer. Curr Opin Obstet Gynecol. 2013;25:49–54.
Ignatiadis M, Sotiriou C. Luminal breast cancer: from biology to treatment. Nat Rev Clin Oncol. 2013;10:494–506.
Sørlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA. 2003;100:8418–23.
Dowsett M, Goldhirsch A, Hayes DF, Senn H-J, Wood W, Viale G. International web-based consultation on priorities for translational breast cancer research. Breast Cancer Res. 2007;9:1–7.
Wishart GC, Bajdik CD, Dicks E, Provenzano E, Schmidt MK, Sherman M, et al. PREDICT Plus: development and validation of a prognostic model for early breast cancer that includes HER2. Br J Cancer. 2012;107:800–7.
Candido dos Reis FJ, Wishart GC, Dicks EM, Greenberg D, Rashbass J, Schmidt MK, et al. An updated PREDICT breast cancer prognostication and treatment benefit prediction model with independent validation. Breast Cancer Res. 2017;19:58.
Cuzick J, Dowsett M, Pineda S, Wale C, Salter J, Quinn E, et al. Prognostic value of a combined estrogen receptor, progesterone receptor, Ki-67, and human epidermal growth factor receptor 2 immunohistochemical score and comparison with the Genomic Health recurrence score in early breast cancer. J Clin Oncol. 2011;29:4273–8.
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, et al. A Multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004;351:2817–26.
Albain KS, Barlow WE, Shak S, Hortobagyi GN, Livingston RB, Yeh IT, et al. Prognostic and predictive value of the 21-gene recurrence score assay in postmenopausal women with node-positive, oestrogen-receptor-positive breast cancer on chemotherapy: a retrospective analysis of a randomised trial. Lancet Oncol. 2010;11:55–65.
Nielsen TO, Parker JS, Leung S, Voduc D, Ebbert M, Vickery T, et al. A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor–positive breast cancer. Clin Cancer Res. 2010;16:5222–32.
Wallden B, Storhoff J, Nielsen T, Dowidar N, Schaper C, Ferree S, et al. Development and verification of the PAM50-based Prosigna breast cancer gene signature assay. BMC Med Genom. 2015;8:54.
Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thürlimann B, et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol. 2013;24:2206–23.
Senkus E, Kyriakides S, Penault-Llorca F, Poortmans P, Thompson A, Zackrisson S, et al. Primary breast cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2013;26:v8–v30.
Coates AS, Members P, Winer EP, Members P, Goldhirsch A, Members P, et al. Tailoring therapies—improving the management of early breast cancer: St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2015. Ann Oncol. 2015;26:1533–46.
Mikami Y, Ueno T, Yoshimura K, Tsuda H, Kurosumi M, Masuda S, et al. Interobserver concordance of Ki67 labeling index in breast cancer: Japan Breast Cancer Research Group Ki67 ring study. Cancer Sci. 2013;104:1539–43.
Polley M-YC, SCY Leung, McShane LM, Gao D, Hugh JC, Mastropasqua MG, et al. An international Ki67 reproducibility study. J Natl Cancer Inst. 2013;105:1897–906.
Howat WJ, Blows FM, Provenzano E, Brook MN, Morris L, Gazinska P, et al. Performance of automated scoring of ER, PR, HER2, CK5/6 and EGFR in breast cancer tissue microarrays in the Breast Cancer Association Consortium. J Pathol Clin Res. 2014;1:18–32.
Abubakar M, Howat WJ, Daley F, Zabaglo L, McDuffus LA, Blows F, et al. High‐throughput automated scoring of Ki67 in breast cancer tissue microarrays from the Breast Cancer Association Consortium. J Pathol Clin Res. 2016;2:138–53.
Abubakar M, Orr N, Daley F, Coulson P, Ali HR, Blows F, et al. Prognostic value of automated KI67 scoring in breast cancer: a centralised evaluation of 8088 patients from 10 study groups. Breast Cancer Res. 2016;18:104.
Lesueur F, Pharoah PD, Laing S, Ahmed S, Jordan C, Smith PL, et al. Allelic association of the human homologue of the mouse modifier Ptprj with breast cancer. Hum Mol Genet. 2005;14:2349–56.
García-Closas M, Egan KM, Newcomb PA, Brinton LA, Titus-Ernstoff L, Chanock S, et al. Polymorphisms in DNA double-strand break repair genes and risk of breast cancer: two population-based studies in USA and Poland, and meta-analyses. Hum Genet. 2006;119:376–88.
Mayr D, Heim S, Werhan C, Zeindl-Eberhart E, Kirchner T. Comprehensive immunohistochemical analysis of Her-2/neu oncoprotein overexpression in breast cancer: HercepTest™ (Dako) for manual testing and Her-2/neuTest 4B5 (Ventana) for Ventana BenchMark automatic staining system with correlation to results of fluorescence in situ hybridization (FISH). Virchows Arch. 2009;454:241–8.
Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thürlimann B, Senn H-J, et al. Strategies for subtypes—dealing with the diversity of breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22:1736–47.
López-Knowles E, Gao Q, Cheang MCU, Morden J, Parker J, Martin L-A, et al. Heterogeneity in global gene expression profiles between biopsy specimens taken peri-surgically from primary ER-positive breast carcinomas. Breast Cancer Res. 2016;18:39.
Bartlett JMS, Christiansen J, Gustavson M, Rimm DL, Piper T, van de Velde CJ, et al. Validation of the IHC4 breast cancer prognostic algorithm using multiple approaches on the multinational TEAM clinical trial. Arch Pathol Lab Med. 2016;140:66–74.
Yeo B, Zabaglo L, Hills M, Dodson A, Smith I, Dowsett M. Clinical utility of the IHC4+C score in oestrogen receptor-positive early breast cancer: a prospective decision impact study. Br J Cancer. 2015;113:390–5.
Barton S, Zabaglo L, A’Hern R, Turner N, Ferguson T, O’Neill S, et al. Assessment of the contribution of the IHC4+C score to decision making in clinical practice in early breast cancer. Br J Cancer. 2012;106:1760–5.
Lakhanpal R, Sestak I, Shadbolt B, Bennett GM, Brown M, Phillips T, et al. IHC4 score plus clinical treatment score predicts locoregional recurrence in early breast cancer. Breast . 2016;29:147–52.
Dodson A, Zabaglo L, Yeo B, Miller K, Smith I, Dowsett M. Risk of recurrence estimates with IHC4+C are tolerant of variations in staining and scoring: an analytical validity study. J Clin Pathol. 2016;69:128–35.
Wolff AC, Hammond MEH, Schwartz JN, Hagerty KL, Allred DC, Cote RJ, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. Arch Pathol Lab Med. 2007;131:18–43.
Hammond MEH, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer (unabridged version). Arch Pathol Lab Med. 2010;134:e48–e72.
Dowsett M, Nielsen TO, A’Hern R, Bartlett J, Coombes RC, Cuzick J, et al. Assessment of Ki67 in breast cancer: recommendations from the International Ki67 in Breast Cancer working group. J Natl Cancer Inst. 2011;103:1656–64.
Bolton KL, Garcia-Closas M, Pfeiffer RM, Duggan MA, Howat WJ, Hewitt SM, et al. Assessment of automated image analysis of breast cancer tissue microarrays for epidemiologic studies. Cancer Epidemiol Biomark Prev. 2010;19:992–9.
Faratian D, Kay C, Robson T, Campbell FM, Grant M, Rea D, et al. Automated image analysis for high-throughput quantitative detection of ER and PR expression levels in large-scale clinical studies: The TEAM Trial Experience. Histopathology. 2009;55:587–93.
Ali HR, Irwin M, Morris L, Dawson SJ, Blows FM, Provenzano E, et al. Astronomical algorithms for automated analysis of tissue protein expression in breast cancer. Br J Cancer. 2013;108:602–12.
Turbin D, Leung S, Cheang MU, Kennecke H, Montgomery K, McKinney S, et al. Automated quantitative analysis of estrogen receptor expression in breast carcinoma does not differ from expert pathologist scoring: a tissue microarray study of 3,484 cases. Breast Cancer Res Treat. 2008;110:417–26.
Gudlaugsson E, Skaland I, Janssen EAM, Smaaland R, Shao Z, Malpica A, et al. Comparison of the effect of different techniques for measurement of Ki67 proliferation on reproducibility and prognosis prediction accuracy in breast cancer. Histopathology. 2012;61:1134–44.
Konsti J, Lundin M, Joensuu H, Lehtimäki T, Sihto H, Holli K, et al. Development and evaluation of a virtual microscopy application for automated assessment of Ki-67 expression in breast cancer. BMC Clin Pathol. 2011;11:3.
Nielsen TO, Jensen M-B, Burugu S, Gao D, Jørgensen CLT, Balslev E, et al. High-risk premenopausal Luminal A breast cancer patients derive no benefit from adjuvant cyclophosphamide-based chemotherapy: results from the DBCG77B clinical trial. Clin Cancer Res. 2017;23:946–53.
Klein ME, Dabbs DJ, Shuai Y, Brufsky AM, Jankowitz R, Puhalla SL, et al. Prediction of the Oncotype DX recurrence score: use of pathology-generated equations derived by linear regression analysis. Mod Pathol. 2013;26:658–64.
Farrugia DJ, Landmann A, Zhu L, Diego EJ, Johnson RR, Bonaventura M, et al. Magee Equation 3 predicts pathologic response to neoadjuvant systemic chemotherapy in estrogen receptor positive, HER2 negative/equivocal breast tumors. Mod Pathol. 2017;30:1078–85.
Acknowledgements
MD acknowledges support from the Royal Marsden NIHR Biomedical Research Center. MA acknowledges funding support from CRUK and the Institute of Cancer Research (ICR), London at the time part of this work was conducted at the ICR.
Funding
The Polish Breast Cancer Study was funded by Intramural Research Funds of the National Cancer Institute, Department of Health and Human Services, USA. The Study of Epidemiology and Risk Factors in Cancer Heredity is funded by a program grant from Cancer Research UK (CRUK) (C490/A10124. C490/A16561) and supported by the UK National Institute for Health Research (NIHR) Biomedical Research Center at the University of Cambridge. Part of this work was supported by the European Community’s Seventh Framework Program under grant agreement number 223175 (grant number HEALTH-F2–2009223175) (COGS).
Author contributions
MA, MGC, MD, PDP conceived the study design and analytical concept. MA performed the statistical analysis and drafted the manuscript. MGC, MD, and PDP contributed to the interpretation of results and the critical revision of the manuscript. JF, JL, HRA, FB, CC, DE, MS, MA, MGC, PDP contributed to data generation. All authors participated in the revision of the manuscript and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abubakar, M., Figueroa, J., Ali, H.R. et al. Combined quantitative measures of ER, PR, HER2, and KI67 provide more prognostic information than categorical combinations in luminal breast cancer. Mod Pathol 32, 1244–1256 (2019). https://doi.org/10.1038/s41379-019-0270-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-019-0270-4
This article is cited by
-
Ki67 increase after core needle biopsy associated with worse disease outcome in HER2-negative breast cancer patients
Scientific Reports (2023)
-
The role of interleukin‑7 serum level as biological marker in breast cancer: a cross‑sectional, observational, and analytical study
World Journal of Surgical Oncology (2022)
-
Single cell imaging-based chromatin biomarkers for tumor progression
Scientific Reports (2021)
-
Impact of breast cancer risk factors on clinically relevant prognostic biomarkers for primary breast cancer
Breast Cancer Research and Treatment (2021)
-
Management of early breast cancer during the COVID-19 pandemic in Brazil
Breast Cancer Research and Treatment (2020)