Abstract
Pulmonary enteric adenocarcinoma is a rare non-small cell lung cancer subtype. It is poorly characterized and cannot be distinguished from metastatic colorectal or upper gastrointestinal adenocarcinomas by means of routine pathological methods. As DNA methylation patterns are known to be highly tissue specific, we aimed to develop a methylation-based algorithm to differentiate these entities. To this end, genome-wide methylation profiles of 600 primary pulmonary, colorectal, and upper gastrointestinal adenocarcinomas obtained from The Cancer Genome Atlas and the Gene Expression Omnibus database were used as a reference cohort to train a machine learning algorithm. The resulting classifier correctly classified all samples from a validation cohort consisting of 680 primary pulmonary, colorectal and upper gastrointestinal adenocarcinomas, demonstrating the ability of the algorithm to reliably distinguish these three entities. We then analyzed methylation data of 15 pulmonary enteric adenocarcinomas as well as four pulmonary metastases and four primary colorectal adenocarcinomas with the algorithm. All 15 pulmonary enteric adenocarcinomas were reliably classified as primary pulmonary tumors and all four metastases as well as all four primary colorectal cancer samples were identified as colorectal adenocarcinomas. In a t-distributed stochastic neighbor embedding analysis, the pulmonary enteric adenocarcinoma samples did not form a separate methylation subclass but rather diffusely intermixed with other pulmonary cancers. Additional characterization of the pulmonary enteric adenocarcinoma series using fluorescence in situ hybridization, next-generation sequencing and copy number analysis revealed KRAS mutations in nine of 15 samples (60%) and a high number of structural chromosomal changes. Except for an unusually high rate of chromosome 20 gain (67%), the molecular data was mostly reminiscent of standard pulmonary adenocarcinomas. In conclusion, we provide sound evidence of the pulmonary origin of pulmonary enteric adenocarcinomas and in addition provide a publicly available machine learning-based algorithm to reliably distinguish these tumors from metastatic colorectal cancer.
Similar content being viewed by others
Introduction
The differentiation of a primary tumor and a metastatic lesion originating from a distant primary site is crucial in cancer diagnosis as it has tremendous effects on the individual treatment and prognosis. Over the last decades, there have been great advances in this field mainly relying on immunohistochemistry [1]. However, in some cases it is still very difficult or even impossible to distinguish primary and metastatic tumors. This is particularly true for the differentiation of pulmonary enteric adenocarcinoma and metastatic colorectal carcinoma to the lung.
The first description of pulmonary enteric adenocarcinoma dates back to 1991 [2]. However, pulmonary enteric adenocarcinomas were not included in the World Health Organization classification of lung tumors until 2015 due to the difficulty of the diagnosis [3, 4]. By definition, these tumors show an enteric morphology in >50% of the tumor mass. This pattern can consist of glandular, papillary, cribriform or solid growth, lined by tall-columnar tumor cells with luminal necrosis or prominent nuclear debris [5]. Furthermore, pulmonary enteric adenocarcinomas express at least one immunohistochemical marker typical for enteric differentiation (CDX2, CK20 or MUC2). The diagnosis of pulmonary enteric adenocarcinoma is further complicated by the fact that TTF-1 expression is frequently lost in non-small cell lung cancer with mucinous or enteric differentiation. Therefore, CK7 was considered to be a helpful marker to distinguish pulmonary enteric adenocarcinomas from metastatic colorectal cancer [6, 7]. However, there are several reports on CK7 negative pulmonary enteric adenocarcinomas and CK7 expression has been described in up to 17% of colorectal cancers [8,9,10,11]. A recent study determined an intermediate sensitivity (71%) and specificity (82%) for the combinatorial use of CK7 and CDX2 for pulmonary enteric adenocarcinoma identification [12]. Furthermore, apart from metastatic colorectal cancer, metastases from malignancies arising from the upper gastrointestinal tract, such as stomach or esophageal adenocarcinoma, often have to be considered as another differential diagnosis. This further complicates the diagnosis of pulmonary enteric adenocarcinoma, as CK7 expression is even more common in these tumors than in colorectal adenocarcinomas. In most instances, this disqualifies CK7 as a reliable marker to exclude a gastrointestinal primary. In addition, pulmonary enteric adenocarcinomas were reported to frequently lack Surfactant and Napsin A expression, two markers that are commonly used to demonstrate the pulmonary origin of an adenocarcinoma [13]. More recently, SATB2 and β-catenin immunohistochemistry have been described as a putative helpful diagnostic markers in this setting, correctly identifying six of seven investigated pulmonary enteric adenocarcinomas [14]. Apart from immunohistochemistry, pulmonary enteric adenocarcinomas are poorly characterized with only small case studies mainly focusing on single genetic alterations [2, 3, 10, 13, 15,16,17].
As it is currently impossible to distinguish pulmonary enteric adenocarcinomas from metastatic gastrointestinal adenocarcinomas based on conventional histomorphology and immunohistochemistry, gastrointestinal malignancies have to be ruled out by careful clinical and radiological examination. This is a complex, time-consuming, and stressful procedure for the patients and delays the initiation of optimal therapy. Therefore, there is an urgent need to identify reliable methods to differentiate pulmonary enteric adenocarcinomas from metastatic gastrointestinal cancer.
DNA methylation patterns have been shown to be very tissue specific [18, 19]. This led to the recent development of a methylation-based classification of brain tumors and novel algorithms to identify primary sites of cancers of unknown primary [20, 21]. Along these lines, we hypothesized that DNA methylation may also be able to distinguish pulmonary enteric adenocarcinomas from metastatic gastrointestinal cancer. Therefore, we used publicly available methylation data of primary pulmonary, colorectal as well as esophageal and gastric (upper gastrointestinal) adenocarcinomas to develop a machine learning-based algorithm that reliably distinguishes pulmonary enteric adenocarcinomas from metastatic gastrointestinal cancers. Furthermore, we investigated if there is molecular evidence of a separate pulmonary enteric adenocarcinoma methylation class among lung tumors and characterized our cohort for molecular alterations.
Material and methods
Patients and samples
IDAT files containing raw methylation data were obtained from The Cancer Genome Atlas and the Gene Expression Omnibus database [22]. After excluding duplicates and normal tissue samples, The Cancer Genome Atlas dataset consisted of 455 pulmonary, 391 colorectal, and 375 upper gastrointestinal adenocarcinoma specimens. The Gene Expression Omnibus dataset included 27 pulmonary (GSE83842 and GSE94785) and 32 colorectal adenocarcinoma samples (GSE77954, GSE77965, GSE98990 and GSE75546) [23,24,25,26,27].
Reference cohort: All samples were randomly numbered using the sample function from the R base package. The first 200 pulmonary, 200 colorectal, and 200 upper gastrointestinal adenocarcinoma specimens were assigned to a reference cohort (n = 600) that was used for generation of the machine learning model.
Validation cohort: The remaining samples were used as a validation cohort (n = 680) to validate the model generated using the reference cohort.
Test cases: Formalin-fixed paraffin embedded tissue samples were obtained from the archives of the Institute of Pathology of the Charité–University Hospital Berlin. We included six resection specimens and eight biopsy samples that met the morphological, immunohistochemical, and clinical criteria for pulmonary enteric adenocarcinoma. We purposely included biopsy samples as the vast majority of lung malignancies are diagnosed based on biopsy tissue and it is especially challenging to distinguish pulmonary enteric adenocarcinoma from metastatic colorectal cancer in this setting due to the limited amount of tissue available. However, it is crucial to differentiate between primary and metastatic disease prior to surgery as this has direct impact on the surgical technique [28]. The Cancer Genome Atlas dataset included one tumor that met the criteria for pulmonary enteric adenocarcinoma (TCGA-55-A4DF). This sample was also included in the test cases, so a total of 15 pulmonary enteric adenocarcinomas were investigated.
Clinicopathological details regarding the included cases are summarized in Supplementary Table S1. All pulmonary enteric adenocarcinoma patients underwent endoscopic examination as well as computed tomography (CT) of the head, chest, and the abdomen. Six patients (PEAD 1, PEAD 3, PEAD 5, PEAD 8, PEAD 10, and PEAD 11) also had positron emission tomography computed tomography (PET/CT) performed. There were no indications for an extrapulmonary primary tumor. For validation purpose, we also analyzed four cases of pulmonary metastases of colorectal cancers (MCC 1–4) as well as four primary colorectal adenocarcinoma (CRAD 1–4) samples.
Histological reevaluation and immunohistochemistry
To evaluate if the pulmonary enteric adenocarcinoma samples could also be differentiated from metastatic colorectal cancer (“primary pulmonary” vs. “metastatic”) by means of conventional histomorphology and immunohistochemistry, histology was reevaluated by five senior pathologists blinded to clinical information.
Immunohistochemical staining for ALK, CDX2, CK7, CK20, and TTF-1 was performed on the Leica BOND-MAX and Ventana BenchMark XT automated slide stainer according to the manufacturer’s instructions. Antibodies and their according manufacturers as well as concentrations are shown in Supplementary Table S2.
Fluorescence in situ hybridization
Fluorescence in situ hybridization was done according to previously described protocols using the Vysis LSI ALK dual color probe (Abbott Molecular, USA), the MET/CEP7 dual color probe (Abbott Molecular, USA) and the RF POSEIDON ROS1 Break probe (Kreatech, The Netherlands) [29]. For each sample, signals were evaluated in at least 50 non-overlapping tumor cells. Cases were classified as ALK or ROS1 positive if at least 15% of all evaluated tumor cell nuclei displayed break apart signal patterns [30]. Cases with ≥5 MET signals per tumor cell or a MET/CEP7 ratio ≥2 were regarded as MET positive [31, 32].
DNA extraction
Semi-automated DNA extraction from formalin-fixed paraffin embedded samples was performed according the manufacturer’s instructions (Maxwell RSC FFPE Plus DNA Purification Kit, Custom, AX4920, Promega, USA).
Next-generation sequencing
Next-generation sequencing was performed using the Ion AmpliSeq Colon Lung Cancer Research Panel v2 according to manufacturer’s instructions and as described previously [33]. This panel covers 22 relevant genes associated with lung and colorectal cancer (KRAS, EGFR, BRAF, PIK3CA, AKT1, ERBB2, PTEN, NRAS, STK11, MAP2K1, ALK, DDR2, CTNNB1, MET, TP53, SMAD4, FBX7, FGFR3, NOTCH1, ERBB4, FGFR1, and FGFR2). For all samples the optimal amount of 10 ng DNA input was used.
Methylation analysis
The Infinium HD FFPE DNA Restore Kit was used for DNA restoration and methylation analysis was performed using the Illumina Infinium MethylationEPIC BeadChip, each according to protocols supplied by the manufacturer.
Raw and unprocessed methylation data for pulmonary enteric adenocarcinoma and metastatic colorectal cancer samples is available at the Gene Expression Omnibus repository under the accession number GSE116699.
Copy number analysis
Copy number analysis was based on raw methylation array data. Individual genome-wide copy number plots were generated using the conumee package [34]. Copy number plots were screened for focal amplifications and deletions, defined as a negative or positive deviation of the mean line of more than 0.4. The region of interest was visualized using the Integrative Genomics Viewer (Broad Institute, Version 2.4.13) to confirm the aberration and to identify potentially cancer-relevant genes at these loci.
Summary copy number plots showing the rate of chromosomal aberrations over multiple samples were generated using a customized version of the conumee package by D.S. The summary copy number plots for all pulmonary enteric adenocarcinoma specimens were then compared to primary lung adenocarcinoma and colorectal adenocarcinoma summary copy number plots. A chromosomal aberration was considered as characteristic for an entity if it occurred in at least 50% of samples while being present in less than 25% of the comparison entity.
Statistical analysis
Statistical analysis was performed using RStudio version 1.1.444 based on the statistical language R version 3.4.2 [35, 36].
Methylation data was processed using the minfi package based on a workflow suggested by Maksimovic et al. [37, 38]. As a first step we filtered for samples with poor overall quality, defined as a mean detection p value < 0.05. Normalization was done using the functional normalization algorithm (funnorm) [39]. Next, we excluded CpG sites that (a) are located on the sex chromosomes, (b) are associated with single nucleotide polymorphisms (SNPs), (c) have previously been reported as cross-reactive, (d) are not covered by both the Illumina Infinium Human Methylation 450 K BeadChip and the Infinium MethylationEPIC BeadChip or (e) had a detection p value > 0.01 in more than 10% of samples [40]. For further analysis, M-values were generated using the shiftBetas function from the Harman package with shiftBy = 1e−04 to avoid infinite M-values [41]. The 10,000 CpG sites with the highest standard deviation across all samples from the reference cohort were selected for further analysis.
T-distributed stochastic neighbor embedding analyses were generated using the Rtsne package with default parameters, 2000 iterations and a perplexity of 20 [42].
Random forest models were generated using the caret package with standard parameters [43]. The optimal ‘mtry’ value was determined using the trainControl function from the caret package and was set at 140.
The generated random forest classifier was then included in a simple R script that can predict the tissue type (pulmonary, colorectal or upper gastrointestinal) solely based on raw IDAT files as well as a sample annotation sheet. In case of missing values for required CpG sites, the missing data is replaced with the mean M-value across all samples analyzed in this study to minimize any possible confounding effect on the classification.
Results
Pulmonary enteric adenocarcinomas cannot be reliably distinguished from metastatic colorectal cancer by histomorphological and immunohistochemical investigation
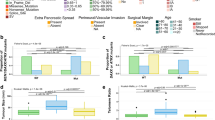
All pulmonary enteric adenocarcinoma cases of the series were reinvestigated for immunohistochemical marker expression (Table 1 and Fig. 1). As expected, all pulmonary enteric adenocarcinoma samples (15 of 15, 100%) were positive for CDX2, while eight of 15 tumors (53%) also expressed CK20. CK7 expression was present in eleven of 15 pulmonary enteric adenocarcinomas (73%), while only two of 15 pulmonary enteric adenocarcinomas (13%) showed focal immunoreactivity against TTF-1. Four of four metastatic colorectal cancers (100%) and four of four primary colorectal specimens (100%) expressed CDX2 and CK20, but one of four metastatic colorectal cancer samples (13%) was also positive for CK7.
All pulmonary enteric adenocarcinomas and metastatic colorectal cancer samples were investigated by five senior pathologists that were asked to identify the tissue origin of the tumors based on morphology and immunoprofile. The results from their classification are displayed in Table 2. All pulmonary enteric adenocarcinoma samples were falsely classified as a metastatic lesion by at least one pathologist. Additionally, there were also discordant results regarding the classification of the metastatic colorectal cancer specimens. Particularly, one colorectal metastasis was classified as a tumor with pulmonary origin by all investigators.
Molecular characterization of pulmonary enteric adenocarcinomas
KRAS mutations were the most frequent alterations and were observed in nine of 15 samples (60%), followed by TP53 mutations in five of 15 samples (33%). There were no cases with ALK, ROS1 or MET alterations. Detailed results for molecular characterization are summarized in Table 3.
A summary copy number profile of the entire pulmonary enteric adenocarcinoma cohort is displayed in Fig. 2. An overlay of this graph with the summary copy number profiles of primary pulmonary and colorectal adenocarcinomas is depicted in Supplementary Figure S1. In total, pulmonary enteric adenocarcinomas demonstrated high rates of chromosomal aberrations. The most frequent alterations were partial loss of chromosome 3p (66%) and chromosome 1q (53%) as well as gain of chromosome 1p (53%) and chromosome 20q (53%). The detailed regions are listed in Supplementary Table 2. The general copy number profiles of pulmonary enteric adenocarcinomas were reminiscent of primary pulmonary cancers with comparable rates of e.g. gain of chromosome 1p or loss of chromosome 13q. The exception to this was a high frequency (53%) of gain of chromosome 20q that is not typically seen in pulmonary adenocarcinomas (about 20%) but is a frequent event in colorectal cancer (about 65%). Focal amplifications (e.g. ERBB2, MYC) or homozygous deletions (e.g. CDKN2A/B) are also summarized in Table 3. An exemplary copy number plot of one pulmonary enteric adenocarcinoma sample with focal MYC and ERBB2 amplification is displayed in Supplementary Figure S2.
DNA methylation profiling for pulmonary enteric adenocarcinoma classification
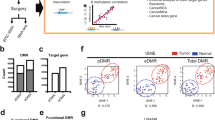
Unsupervised t-distributed stochastic neighbor embedding of the whole dataset (n = 1303) revealed three main clusters, representing primary pulmonary, colorectal, and upper gastrointestinal adenocarcinoma samples (Fig. 3). There were some overlaps among the pulmonary and upper gastrointestinal cancer clusters as well as the colorectal and upper gastrointestinal clusters. However, no pulmonary adenocarcinoma cases grouped with the colorectal cancer cluster or vice versa. The 15 pulmonary enteric adenocarcinoma samples reliably grouped with the primary pulmonary adenocarcinomas and the four metastatic colorectal cancer samples clustered with the primary colorectal adenocarcinomas. Both pulmonary enteric adenocarcinomas and colorectal metastases did not form a separate methylation cluster but rather intermixed with primary pulmonary or colorectal samples, respectively. With a proportion of 77% (10 of 14 cases with histomorphological data available), mucinous lung adenocarcinomas were overrepresented among the non-small cell lung cancer cases that accumulated in the upper gastrointestinal adenocarcinoma cluster. The rate of mucinous adenocarcinomas in the main lung cancer cluster was 6% (27 of 454 cases with histomorphological data available), indicating that the mucinous differentiation results in a closer epigenetic relation to upper gastrointestinal adenocarcinomas. Additionally, upper gastrointestinal cancers associated with the Epstein-Barr virus (EBV) formed a separate subgroup (Supplementary Figure S4).
Unsupervised t-distributed stochastic neighbor embedding for all cases (n = 1303). The tissue origin is annotated by color. Distinct clusters representing pulmonary adenocarcinomas, colorectal adenocarcinomas and upper gastrointestinal adenocarcinomas can be observed with small overlaps between the pulmonary and the upper gastrointestinal adenocarcinoma cluster as well as the colorectal and the upper gastrointestinal adenocarcinoma cluster. All pulmonary enteric adenocarcinomas and metastatic colorectal cancers are assigned to the correct clusters
We then separated the publicly available cases into a reference (n = 600) and a validation cohort (n = 680). The reference cohort was used to train the machine learning algorithm to identify the correct tumor type. When applied to the validation cohort, the resulting classifier correctly classified all specimens as pulmonary, colorectal or upper gastrointestinal carcinomas. This also included the cases that fell into divergent clusters in the t-distributed stochastic neighbor embedding plot. We then applied this system to the test cases. All 15 pulmonary enteric adenocarcinoma samples were classified as carcinomas originating from the lung and all four metastatic colorectal cancer specimens as colorectal tumors.
The resulting algorithm was integrated in an R Script that is publicly available at https://github.com/aennecken/PEAD. Using raw IDAT files and a sample annotation file, the algorithm can be applied to single or multiple samples. Results are given in the form of an HTML document, including the final prediction and additional information, such as the proportion of votes for the individual diagnosis. Samples are classified as pulmonary, colorectal or upper gastrointestinal tumors. The diagnosis is given if the proportion of votes for one of the three diagnoses is at least 50%. If neither diagnosis reaches 50% of votes, an alternative primary site should be excluded. An example report for the 14 pulmonary enteric adenocarcinoma samples and the four metastatic colorectal cancer samples investigated in this study is available with the online full text version of this paper (Supplementary File S1).
Mean decrease in accuracy was used to assess the importance of the individual CpG sites for random forest decision making. The 100 CpG probes with the highest mean decrease in accuracy are listed in Supplementary Table S4. The most frequent genes that were associated with these CpG probes were CACNB2 (Calcium Voltage-Gated Channel Auxiliary Subunit Beta 2), HOXA9 (Homeobox A9), HOXD1 (Homeobox D1), HOXD8 (Homeobox D8) and RNLS (Renalase, FAD dependent amine oxidase; also refer to Supplementary Figure S5). KRT7 (Cytokeratin 7) was also represented among the 100 most relevant CpG sites.
Discussion
The main focus of this study was to find further evidence of the pulmonary origin of pulmonary enteric adenocarcinoma, to molecularly characterize this non-small cell lung cancer subtype and to generate a methylation-based procedure to reliably distinguish pulmonary enteric adenocarcinomas from pulmonary metastases of colorectal cancer as well as esophageal and gastric adenocarcinomas.
Our histological and immunohistochemical evaluation nicely illustrated the known overlap of pulmonary enteric adenocarcinomas and metastatic colorectal cancers and demonstrated the diagnostic problems in clinical workup of these cases. Indeed, not a single of our pulmonary enteric adenocarcinomas was considered as such by all five investigators. All five investigators are from the same institution, so likely the interpretation may vary even considerably more between different institutions. Interestingly, we also observed that pulmonary metastases of colorectal cancers can potentially mimic primary lung tumors, as one metastatic colorectal cancer sample was classified as a primary pulmonary adenocarcinoma by all pathologists due to heterogenous growth patterns and CK7 expression.
There is little data regarding the mutational profile of pulmonary enteric adenocarcinomas. The few existing case series mainly focused on KRAS, EGFR, and ALK alterations. To get a more comprehensive view on the molecular changes that could characterize pulmonary enteric adenocarcinomas, we used a targeted next-generation sequencing panel covering 22 genes associated with lung and colorectal cancer. In accordance with previous studies, we observed a slightly higher frequency of KRAS mutations compared to common non-small cell lung cancers [10].
We also investigated predictive markers for possible treatment strategies. However, we observed no ALK, MET, EGFR or ROS1 alterations. According to previous studies, EGFR mutations are a rare event in pulmonary enteric adenocarcinomas and there is only one study reporting a single case with ALK rearrangement [3, 14, 16, 44]. Although there is only limited data, alterations eligible for tyrosine kinase inhibitor therapy seem to be less common than in other non-small cell lung cancer subtypes.
Regarding copy number alterations, pulmonary enteric adenocarcinomas showed chromosomal aberrations that are more common in pulmonary than in colorectal adenocarcinomas. Interestingly, the frequency of gains of chromosome 20 in pulmonary enteric adenocarcinoma was more in accordance with typical colorectal cancer alterations. Furthermore, loss of chromosome 3p might be a characteristic event in pulmonary enteric adenocarcinomas.
On the basis of publicly available methylation data from a reference cohort of 600 samples derived from The Cancer Genome Atlas dataset, we were able to train a robust machine learning-based algorithm that successfully classified a validation cohort consisting of 680 pulmonary, colorectal and upper gastrointestinal adenocarcinomas samples without any misclassification. Additionally, all 15 pulmonary enteric adenocarcinoma specimens were classified as primary lung tumors. In conclusion, this study makes headway in proving that pulmonary enteric adenocarcinoma is a rare but actually existing non-small cell lung cancer subtype rather than a metastatic lesion. The fact that the methylation signature is in accordance with common lung adenocarcinomas rules out other explanations for the unusual morphological and immunohistochemical features of this tumor entity, such as a missed or a regressed colorectal cancer.
In theory, the analysis of the methylation profile of metastases might be complicated by several potential confounding factors. On the one hand, samples may be contaminated with adjacent benign tissue of the affected organ; on the other hand, the microenvironment at the metastatic site may induce changes in the methylation profile of the tumor cells that differ from the primary tumor. To address this concern, we also analyzed a set of four pulmonary metastases from colorectal carcinomas. All samples were classified as colorectal tumors by the random forest algorithm, providing evidence that the algorithm is not biased by these potential confounding factors.
In t-distributed stochastic neighbor embedding analysis, there was no evidence for a pulmonary enteric adenocarcinoma or metastatic colorectal cancer subgroup as all samples mixed with other primary pulmonary or colorectal cancers, respectively. However, interestingly, mucinous pulmonary adenocarcinomas accumulated in the upper gastrointestinal cancer cluster, suggesting that these tumors are epigenetically more related to esophageal or gastric adenocarcinomas than other primary lung cancers. Still, these tumors were consistently classified as pulmonary adenocarcinomas by the random forest classifier.
A major limitation of our study is the relatively low number of pulmonary enteric adenocarcinoma samples that were available for analysis. However, this could not be overcome due to the rare incidence of this subtype. When methylation profiling is considered in a diagnostic setting, the relatively high costs of consumables and technical equipment as well as currently not developed technical expertise have to be considered. This currently limits the use of this technology outside of specialized institutes. However, a broader establishment of DNA methylation analysis in the following years seems likely, considering the huge success and dynamics of this method in the classification of brain tumors [20]. Another limitation of this technique is the fact that it is crucial to ensure a high tumor content, as benign tissue (e.g. immune cells, fibroblasts, non-cancerous epithelium) may interfere with the methylation signature. Whereas non-tumorous areas can simply be excluded when evaluating immunohistochemical stainings, some samples might not be suitable for methylation analysis if the amount of tumor cells compared to normal cells is too low. For brain tumors, the dropout because of low tumor cell content was 4% of cases in a prospective experimental diagnostic implementation [20].
However, the methodology used in this study also has some major and revolutionary advances. Array-based and genome-wide methylation analyses deliver reproducible and detailed epigenetic tumor profiles that are relatively resistant to batch effects. Furthermore, the analysis can be performed on formalin-fixed paraffin embedded tissue. In contrast to immunohistochemistry, these tests investigate thousands of loci and genes. In combination with machine learning algorithms, these large datasets can be used to derive highly valuable information which can be used to solve specific diagnostic problems, as shown in this study, or to even classify whole tumor entities [20].
An R script with an example input file that uses the classifier generated in this study to predict the molecular subtype of inputted 450 K or EPIC methylation array data is publicly available at https://github.com/aennecken/PEAD. Although this pulmonary enteric adenocarcinoma classifier is a research tool and proof of principle and should be used with caution, in the future, this algorithm could enable pathologists to diagnose pulmonary enteric adenocarcinoma solely based on tissue analysis with numerous potential benefits for patients and physicians. In theory, the described method could even prevent unnecessary and potentially hazardous examinations and minimize radiation exposure. It could potentially shorten the time that is needed to make a definitive diagnosis, which is crucial to decide on the optimal surgical strategy (wedge or segmental resection vs. lobectomy) or the need of additional systemic therapy. Furthermore, the algorithm could be essential to distinguish metachronous metastasis from a secondary tumor in patients with a history of colorectal cancer. Another advantage of this methodology is the fact that it can also be performed on biopsy samples. The conventional interpretation of biopsy samples might be especially hard because only a small proportion of the tumor can be examined and the tumor cells may be altered due to sampling artifacts.
In summary, this study describes a robust classifier based on DNA methylation data that could in future potentially enable pathologist to make the diagnosis of pulmonary enteric adenocarcinoma on the basis of tissue analysis alone, independently from complex and time-consuming clinical examinations.
References
Tot T. Cytokeratins 20 and 7 as biomarkers: usefulness in discriminating primary from metastatic adenocarcinoma. Eur J Cancer. 2002;38:758–63.
Tsao M, Fraser R. Primary pulmonary adenocarcinoma with enteric differentiation. Cancer. 1991;68:1754–7.
Wang CX, Liu B, Wang YF, Zhang RS, Yu B, Lu ZF, et al. Pulmonary enteric adenocarcinoma: a study of the clinicopathologic and molecular status of nine cases. Int J Clin Exp Pathol. 2014;7:1266–74.
Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JHM, Beasley MB, et al. The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol. 2015;10:1243–60.
Travis WD, Brambilla E, Noguchi M, Nicholson AG, Geisinger KR, Yatabe Y, et al. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol. 2011;6:244–85.
Guo M, Tomoshige K, Meister M, Muley T, Fukazawa T, Tsuchiya T, et al. Gene signature driving invasive mucinous adenocarcinoma of the lung. EMBO Mol Med. 2017;9:462–81.
Rossi G, Murer B, Cavazza A, Losi L, Natali P, Marchioni A, et al. Primary mucinous (so-called colloid) carcinomas of the lung: a clinicopathologic and immunohistochemical study with special reference to CDX-2 homeobox gene and MUC2 expression. Am J Surg Pathol. 2004;28:442–52.
Bayrak R, Yenidünya S, Haltas H. Cytokeratin 7 and cytokeratin 20 expression in colorectal adenocarcinomas. Pathol Res Pract. 2011;207:156–60.
Lin L, Zhuang W, Wang W, Xu C, Chen R, Guan Y, et al. Genetic mutations in lung enteric adenocarcinoma identified using next-generation sequencing. Int J Clin Exp Pathol. 2017;10:9583–90.
Zhao L, Huang S, Liu J, Zhao J, Li Q, Wang HQ. Clinicopathological, radiographic, and oncogenic features of primary pulmonary enteric adenocarcinoma in comparison with invasive adenocarcinoma in resection specimens. Medicine. 2017;96:e8153.
Hatanaka K, Tsuta K, Watanabe K, Sugino K, Uekusa T. Primary pulmonary adenocarcinoma with enteric differentiation resembling metastatic colorectal carcinoma: a report of the second case negative for cytokeratin 7. Pathol Res Pract. 2011;207:188–91.
Chen M, Liu P, Yan F, Xu S, Jiang Q, Pan J, et al. Distinctive features of immunostaining and mutational load in primary pulmonary enteric adenocarcinoma: implications for differential diagnosis and immunotherapy. J Transl Med. 2018;16:81.
Inamura K, Satoh Y, Okumura S, Nakagawa K, Tsuchiya E, Fukayama M, et al. Pulmonary adenocarcinomas with enteric differentiation: histologic and immunohistochemical characteristics compared with metastatic colorectal cancers and usual pulmonary adenocarcinomas. Am J Surg Pathol. 2005;29:660–5.
Matsushima J, Yazawa T, Suzuki M, Takahashi Y, Ota S, Nakajima T, et al. Clinicopathological, immunohistochemical, and mutational analyses of pulmonary enteric adenocarcinoma: usefulness of SATB2 and β-catenin immunostaining for differentiation from metastatic colorectal carcinoma. Hum Pathol. 2017;64:179–85.
Sun WW, Xu ZH, Wang CF, Wu F, Cao JM, Cui PJ, et al. Pulmonary enteric adenocarcinoma with pancreatic metastasis: A case report. Oncol Lett. 2017;13:4651–6.
Bonanno L, Attili I, Nannini N, Del Bianco P, Frega S, Pasello G, et al. P3.01-017 Primary lung adenocarcinomas with enteric differentiation: a retrospective analysis. J Thorac Oncol. 2017;12:S1128–9.
Nottegar A, Tabbò F, Luchini C, Brunelli M, Bria E, Veronese N, et al. Pulmonary adenocarcinoma with enteric differentiation: immunohistochemistry and molecular morphology. Appl Immunhisto Mol Morpho. 2018;26.
Lokk K, Modhukur V, Rajashekar B, Märtens K, Mägi R, Kolde R, et al. DNA methylome profiling of human tissues identifies global and tissue-specific methylation patterns. Genome Biol. 2014;15:r54.
Chen Y1, Breeze CE, Zhen S, Beck S, Teschendorff AE. Tissue-independent and tissue-specific patterns of DNA methylation alteration in cancer. Epigenetics Chromatin. 2016;9:10.
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D, et al. DNA methylation-based classification of central nervous system tumours. Nature. 2018;555:469–74.
Moran S, Martínez-Cardús A, Sayols S, Musulén E, Balañá C, Estival-Gonzalez A, et al. Epigenetic profiling to classify cancer of unknown primary: a multicentre, retrospective analysis. Lancet Oncol. 2016;17:1386–95.
Grossman RL, Heath AP, Ferretti V, Varmus HE, Lowy DR, Kibbe WA, et al. Toward a shared vision for cancer genomic data. N Eng J Med. 2016;375:1109–12.
Qu X, Sandmann T, Frierson H Jr, Fu L, Fuentes E, Walter K, et al. Integrated genomic analysis of colorectal cancer progression reveals activation of EGFR through demethylation of the EREG promoter. Oncogene. 2016;35:6403–15.
Blueprint consortium. Quantitative comparison of DNA methylation assays for biomarker development and clinical applications. Nat Biotechnol. 2016;34:726–37.
Zhou W, Laird PW, Shen H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 2017;45:e22.
Kettunen E, Hernandez-Vargas H, Cros MP, Durand G, Le Calvez-Kelm F, Stuopelyte K, et al. Asbestos-associated genome-wide DNA methylation changes in lung cancer. Int J Cancer. 2017;141:2014–29.
Kajiura K, Masuda K, Naruto T, Kohmoto T, Watabnabe M, Tsuboi M, et al. Frequent silencing of the candidate tumor suppressor TRIM58 by promoter methylation in early-stage lung adenocarcinoma. Oncotarget. 2017;8:2890–905.
Kerr KM. Pulmonary adenocarcinomas: classification and reporting. Histopathology. 2009;54:12–27.
Jurmeister P, Lenze D, Berg E, Mende S, Schäper F, Kellner U et al. Parallel screening for ALK, MET and ROS1 alterations in non-small cell lung cancer with implications for daily routine testing. Lung Cancer. 2015;87:122–9.
von Laffert M, Stenzinger A, Hummel M, Weichert W, Lenze D, Warth A, et al. ALK-FISH borderline cases in non-small cell lung cancer: Implications for diagnostics and clinical decision making. Lung Cancer. 2015;90:465–71.
Cappuzzo F, Marchetti A, Skokan M, Rossi E, Gajapathy S, Felicioni L, et al. Increased MET gene copy number negatively affects survival of surgically resected non-small-cell lung cancer patients. J Clin Oncol. 2009;27:1667–74.
Tanaka A, Sueoka-Aragane N, Nakamura T, Takeda Y, Mitsuoka M, Yamasaki F, et al. Co-existence of positive MET FISH status with EGFR mutations signifies poor prognosis in lung adenocarcinoma patients. Lung Cancer. 2012;75:89–94.
Vollbrecht C, Lehmann A, Lenze D, Hummel M. Validation and comparison of two NGS assays for the detection of EGFR T790M resistance mutation in liquid biopsies of NSCLC patients. Oncotarget. 2018;9:18529–39.
Hovestadt V, Zapatka M. Conumee: enhanced copy-number variation analysis using Illumina DNA methylation arrays. 2017. R package version 1.9.0. http://bioconductor.org/packages/conumee/. Accessed on: 27 Aug 2018.
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. 2017. https://www.R-project.org/.
RStudio Team. RStudio: Integrated Development for R. RStudio, Inc., Boston. 2016. http://www.rstudio.com/.
Maksimovic J, Phipson B, Oshlack A. A cross-package bioconductor workflow for analysing methylation array data. F1000. 2016;5:1281.
Aryee MJ, Jaffe AE, Corrada-Bravo H, Ladd-Acosta C, Feinberg AP, Hansen KD, et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics. 2014;30:1363–9.
Fortin JP, Labbe A, Lemire M, Zanke BW, Hudson TJ, Fertig EJ, et al. Functional normalization of 450k methylation array data improves replication in large cancer studies. Genome Biol. 2014;15:503.
Chen YA, Lemire M, Choufani S, Butcher DT, Grafodatskaya D, Zanke BW, et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics. 2013;8:203–9.
Oytam Y, Sobhanmanesh F, Duesing K, Bowden JC, Osmond-McLeod M, Ross J. Risk-conscious correction of batch effects: maximising information extraction from high-throughput genomic datasets. BMC Bioinforma. 2016;17:332.
Krijthe JH. Rtsne: T-distributed stochastic neighbor embedding using a Barnes-Hut implementation. 2015. https://github.com/jkrijthe/Rtsne. Accessed: 27 Aug 2018.
Kuhn M. Caret: Classification and regression training. 2016. R package version 6.0-71. https://CRAN.R-project.org/package=caret. Accessed: 27 Aug 2018.
Nottegar A, Tabbò F, Luchini C, Guerrera F, Gaudiano M, Bria E, et al. Pulmonary adenocarcinoma with enteric differentiation: dissecting oncogenic genes alterations with DNA sequencing and FISH analysis. Exp Mol Pathol. 2017;102:276–9.
Acknowledgements
We gratefully acknowledge the excellent technical assistance of Daniel Teichmann, Alexandra Förster, Vera Arnemann and Didier Nana-Kouegoua.
Funding
Maximilian von Laffert and Leonille Schweizer are participants in the Berlin Institute of Health (BIH) Charité Clinician Scientist Program funded by the Charité–Universitätsmedizin Berlin and the Berlin Institute of Health. The work of Philipp Jurmeister is supported by the European Fund for Regional Development (EFRE) and the Federal State of Berlin. The funding number is 1303/2013.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jurmeister, P., Schöler, A., Arnold, A. et al. DNA methylation profiling reliably distinguishes pulmonary enteric adenocarcinoma from metastatic colorectal cancer. Mod Pathol 32, 855–865 (2019). https://doi.org/10.1038/s41379-019-0207-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-019-0207-y
This article is cited by
-
EpiDiP/NanoDiP: a versatile unsupervised machine learning edge computing platform for epigenomic tumour diagnostics
Acta Neuropathologica Communications (2024)
-
Frequent nuclear β-catenin expression in pulmonary enteric-type adenocarcinoma according to the current World Health Organization criteria
Virchows Archiv (2023)
-
Lungenmetastasen – Onkologische Bedeutung und Therapie
Zeitschrift für Pneumologie (2023)
-
Lungenmetastasen – Onkologische Bedeutung und Therapie
best practice onkologie (2023)
-
Lungenmetastasen – Onkologische Bedeutung und Therapie
Die Onkologie (2023)