Abstract
The aim of this study was to define the miRNA profile of small intestinal neuroendocrine tumors and to search for novel molecular subgroups and prognostic biomarkers. miRNA profiling was conducted on 42 tumors from 37 patients who underwent surgery for small intestinal neuroendocrine tumors. Unsupervised hierarchical clustering analysis of miRNA profiles identified two groups of tumor metastases, denoted cluster M1 and M2. The smaller cluster M1 was associated with shorter overall survival and contained tumors with higher grade (WHO grade G2/3) and multiple chromosomal gains including gain of chromosome 14. Tumors of cluster M1 had elevated expression of miR-1246 and miR-663a, and reduced levels of miR-488-3p. Pathway analysis predicted Wnt signaling to be the most significantly altered signaling pathway between clusters M1 and M2. Analysis of miRNA expression in relation to tumor proliferation rate showed significant alterations including downregulation of miR-137 and miR-204-5p in tumors with Ki67 index above 3%. Similarly, tumor progression was associated with significant alterations in miRNA expression, e.g. higher expression of miR-95 and miR-210, and lower expression of miR-378a-3p in metastases. Pathway analysis predicted Wnt signaling to be altered during tumor progression, which was supported by decreased nuclear translocation of β-catenin in metastases. Survival analysis revealed that downregulation of miR-375 was associated with shorter overall survival. We performed in situ hybridization on biopsies from an independent cohort of small intestinal neuroendocrine tumors using tissue microarrays. Expression of miR-375 was found in 578/635 (91%) biopsies and survival analysis confirmed that there was a correlation between downregulation of miR-375 in tumor metastases and shorter patient survival. We conclude that miRNA profiling defines novel molecular subgroups of metastatic small intestinal neuroendocrine tumors and identifies miRNAs associated with tumor proliferation rate and progression. miR-375 is highly expressed in small intestinal neuroendocrine tumors and may be used as a prognostic biomarker.
Similar content being viewed by others
Introduction
Neuroendocrine tumors are the most frequent neoplasms of the small intestine. They are derived from endocrine cells of the intestinal mucosa and usually secrete large amounts of serotonin. Although the tumors are of low grade and proliferate slowly, the majority of patients have metastatic disease at the time of diagnosis. For these patients, curative surgery cannot be performed. Adjuvant therapy is recommended and includes long-acting somatostatin analogs, hepatic artery embolization, and somatostatin receptor-targeted radiotherapy [1, 2]. Despite adjuvant therapy, the 5-year survival is < 60% in patients with liver metastases [3]. However, the prognosis is highly variable and difficult to predict. Novel biomarkers are needed to identify molecular subtypes of tumors, to monitor disease progression, and to predict response to therapy [4].
Efforts have been made to characterize the genetic and epigenetic alterations responsible for the development and progression of small intestinal neuroendocrine tumors [5,6,7,8,9]. Profiling of somatic copy number alterations has shown that loss of chromosome 18 is the most frequent genomic event in small intestinal neuroendocrine tumors. Multiple gains of whole chromosomes, notably gain of chromosomes 4, 5, 7, 14, and 20, have been observed in a small group of tumors. These tumors were associated with shorter disease-free survival and shorter overall survival [6, 10]. Sequencing of whole exome, however, failed to identify driver mutations and showed that tumors were mutationally silent [7, 8]. Recurrent mutations were observed in CDKN1B, but they were only present in 8.5% of tumors without any correlation to clinical characteristics or patient survival [8, 11]. Profiling of DNA methylation, on the other hand, showed small intestinal neuroendocrine tumors to be highly epigenetically dysregulated and there was a correlation between methylation patterns and progression-free survival [9]. Epigenetic dysregulation frequently occurs in cancers, causing alterations in gene expression without modifying the underlying DNA sequence. MicroRNAs (miRNAs) are short non-coding RNA molecules that function as epigenetic regulators by suppressing mRNA translation and reducing mRNA stability. miRNAs control almost one-third of the human transcriptome, and as perfect base pairing is not required each miRNA is able to regulate hundreds of mRNAs [12, 13]. Altered expression of miRNAs has been demonstrated in virtually all tumors, contributing to carcinogenesis and tumor progression [14,15,16,17]. In tumors, aberrantly expressed miRNAs can either act as oncomiRs or tumor suppressors by targeting genes involved in proliferation, cell cycle control, apoptosis, invasion, and drug resistance [18].
In this study, we characterized the epigenome of small intestinal neuroendocrine tumors by determining their miRNA profiles. Tumors were collected from patients treated at a single center with long-term follow-up. These tumors had previously been subjected to profiling of copy number alterations and gene expression patterns. To date, this is the largest study of miRNA profiles in patients with long-term follow-up, which has allowed us to identify subgroups of tumors with clinical relevance as well as biomarkers of patient outcome.
Material and methods
Tumor material used for miRNA profiling
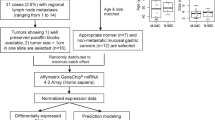
Tumor biopsies from 37 patients who underwent surgery for small intestinal neuroendocrine tumors (ileal carcinoids) at Sahlgrenska University Hospital, Gothenburg, Sweden, between 1991 and 2009, were included. Fresh tumor tissue was obtained at surgery from a total of 16 primary tumors, 24 liver metastases, 2 lymph node metastases, and 6 normal small intestinal mucosa. In 5 cases, paired samples of primary tumor and liver metastasis were analyzed. The diagnostic criteria used for small intestinal neuroendocrine tumors were according to the World Health Organization guidelines (WHO, 2010) [19]. The histopathological evaluation was performed on hematoxylin and eosin-stained sections. Immunohistochemical staining was also performed for chromogranin A, synaptophysin, Ki67, serotonin, CDX2, and cytokeratin 8/18. All patients were diagnosed with well-differentiated small intestinal neuroendocrine tumors and had metastatic disease at the time of diagnosis (TNM stage IIIB or IV) [20]. Mean follow-up time after operation was 70.4 months (median 60.5, range 5‒201 months). The clinical characteristics of patients are given in Table 1. Copy number alterations and expression profiles of the cohort have been published previously [5, 6]. We obtained consent from the patients and also approval from the Regional Ethical Review Board in Gothenburg for the use of clinical materials for research purposes.
Clustering analysis of small intestinal neuroendocrine tumors from 37 patients. Unsupervised hierarchical clustering analysis was performed on miRNA profiles from tumors and normal small intestinal mucosa. Tumors were separated into 3 distinct groups, denoted cluster P, cluster M1, and cluster M2. The majority of the primary tumors clustered together in cluster P, whereas metastases were separated in 2 different clusters, a smaller cluster M1 and a larger cluster M2. Types of tissue analyzed are indicated: M, metastasis; N, normal small intestinal mucosa; P, primary tumor. Paired tumor samples (primary and metastases) were available from 5 patients, indicated by a, b, c, d, and e (for details, see Table 1)
Tissue microarray used for immunohistochemistry and in situ hybridization
All patients who underwent surgery for small intestinal neuroendocrine tumors at Sahlgrenska University Hospital from 1986 to 2013 were included in a tissue microarray. Formalin-fixed and paraffin-embedded tumor tissue from this cohort was retrieved from the archives of the Department of Clinical Pathology and Genetics, Sahlgrenska University Hospital, Gothenburg. The diagnosis was confirmed by reviewing hematoxylin and eosin-stained sections and immunohistochemical stainings. Sufficient tumor material for construction of tissue microarray was available from 412 patients. From each tumor, 1.0 mm core biopsies were obtained. Each recipient block contained a total of 121 core biopsies including tumor tissue and normal tissues from stomach, small intestine, and colon. When available, core biopsies were taken from primary tumor, lymph node metastases, liver metastases, and other distant metastases. A total of 8 recipient blocks were constructed, derived from 846 tumors. For comparative purposes, we also analyzed a different set of tissue microarrays containing adenocarcinomas of the stomach, colon, and pancreas, as well as gastrointestinal stromal tumors. The quality of the constructed tissue microarray was evaluated on hematoxylin and eosin-stained sections and on immunohistochemical stainings for chromogranin A, synaptophysin, serotonin, and Ki67. We obtained approval from the Regional Ethical Review Board in Gothenburg, Sweden, for the use of clinical materials for research purpose.
Immunohistochemistry and scoring
Full sections of normal small intestine and tumor tissue as well as sections of tissue microarray were placed on positively charged glass slides. Sections were subjected to antigen retrieval using EnVision FLEX Target Retrieval Solution (high pH) in a Dako PT-Link. Immunohistochemical staining was performed in a Dako Autostainer Link using EnVision FLEX according to the manufacturer’s instructions (DakoCytomation). The following primary antibodies were used: anti-chromogranin A (MAB319; Chemicon), anti-synaptophysin (SY38, M0776; Dako), anti-Ki67 (MIB1; Dako), anti-serotonin (H209; Dako), anti-cytokeratin 8 and 18 (NCL5D3; Leica), anti-CDX2 (EPR2764Y, Abcam), anti-β-catenin (14/Beta-catenin, BD Transduction Laboratories), and anti-FOXM1 (sc-271746, Santa Cruz Biotechnology). Positive and negative controls were included in each run. Stained sections were scanned using Leica SCN 4000 at × 40 magnification. The fraction of Ki67- and forkhead box M1 (FOXM1)-positive cells was estimated by counting 500–2,000 tumor cells per tumor using printouts and manual counting [21]. The scoring of β-catenin staining was evaluated independently by two board certified surgical pathologists (ON and AB) using an Olympus BX51 light microscope. The total staining intensity of tumor cells was scored as weak or strong and the staining of tumor cell nuclei was scored as negative or positive for β-catenin. The intensity of tumor cell staining was scored as strong when it had the same intensity as crypt cells of the normal small intestinal mucosa.
miRNA profiling
We used tumor biopsies from 43 previously reported patients [5, 6]. Of these patients, 6 were excluded due to lack of high-quality miRNA or poor quality of the array hybridization experiments. The remaining tumor biopsies from 37 patients (16 primary tumors, 2 lymph node metastases, and 24 liver metastases) were included in the study. The purity of tumor biopsies was assessed by light microscopy using hematoxylin and eosin-stained sections, and was found to contain at least 70% tumor cells, except for 2 primary tumor samples (50‒60% tumor cells). Biopsies of normal small intestinal mucosa from 6 patients undergoing surgery for adenocarcinoma of the right colon were used as controls. RNA was isolated from fresh-frozen biopsies using the miRNeasy Mini Kit (Qiagen) according to the manufacturer’s protocol. cDNA synthesis, labeling and hybridization were performed according to the One-Color Microarray-Based miRNA Complete labeling and hybridization protocol (v2.1) (Agilent Technologies). Labeled samples were hybridized to 8 × 15 K human miRNA microarrays (V3) (G4470C; Agilent Technologies) containing 866 human and 89 human viral miRNAs. Arrays were scanned using an Agilent G2565BA Microarray Scanner (Agilent Technologies). Images were read and processed using Feature Extraction software v10.7.1.1 (Agilent Technologies).
In situ hybridization of miR-375
Tissue microarray and tissue blocks from normal small intestine were cut in 6 μm-thick sections and placed on positively charged glass slides. Slides were deparaffinized in xylene and hydrated in alcohol. In situ hybridization was performed according to the manufacturer’s instructions (Exiqon). Briefly, the sections were permeabilized with proteinase K (15 μg/ml; Sigma-Aldrich), then washed with phosphate-buffered saline (PBS), dehydrated in alcohol, and air-dried. miRCURY LNA detection probes double-labeled with digoxigenin (Exiqon) targeting miR-375 and scramble-miRNA (negative control) were denatured for 4 min and diluted to 40 nM in hybridization buffer (Exiqon). The sections were hybridized to the denatured probes in a Dako Hybridizer for 1 h at 54 °C. Nonspecifically bound probe was removed by washing with sodium sulphate citrate (Sigma-Aldrich). To visualize bound probe, the slides were blocked with sheep serum (Sigma-Aldrich) then incubated with sheep anti-digoxigenin-AP antibody (Roche) for 60 min, washed with PBS-Tween, and incubated with AP substrate BCIP/NBT (Dako). Endogenous AP activity was blocked with Levamisol (Fluka). Sections were counterstained with nuclear fast red (Sigma). The specificity of the miR-375 probe was evaluated on small intestinal mucosa and normal pancreas (positive control). The endocrine cells in the mucosa and the islets of Langerhans were specifically and strongly labeled by the probe (Supplementary figure S1). The intensity of the in situ hybridization signal was evaluated independently by two board certified surgical pathologists (ON and AB) using an Olympus BX51 light microscope. The staining intensity of tumor cells was scored as follows: 0 = no staining; 1 = weak staining; 2 = intermediate staining; and 3 = strong staining, where score 3 was the same level of staining intensity as in neuroendocrine cells of the small intestinal mucosa. Core biopsies containing less than one-third tumor cells were excluded. Sections adjacent to in situ hybridization sections were stained with hematoxylin and eosin, to verify the presence of tumor cells in each core. Only tumor biopsies that were positive for synaptophysin, chromogranin A, and serotonin were included.
Quantitative real time PCR analysis of miR-375
RNA was isolated from 20 biopsies of small intestinal neuroendocrine tumors (also present on tissue microarray), 11 biopsies of normal small intestinal mucosa, and 5 biopsies of colorectal adenocarcinoma. The purity of the biopsies was assessed by light microscopy using hematoxylin and eosin-stained sections and biopsies that contained 50–90% tumor cells were included. Targeted reversed transcription and amplification of miR-375 and miR-16 (endogenous control) was performed using TaqMan MicroRNA Reverse Transcription Kit and a sequence specific TaqMan MicroRNA Assays (Life Technologies) for human miR-375 (Assay ID: 000564) and miR-16 (Assay ID: 000391) according to manufacturer’s instructions. The PCR reactions were performed using a 7500 Fast Real-Time PCR System (Applied Biosystems). The samples were analyzed in triplicate. The miR-375 expression was calculated by using the comparative cycle threshold method (2-ΔΔCT) using miR-16 as an endogenous control and relative to normal small intestinal mucosa [22, 23].
Statistical analysis
The miRNA microarrays were analyzed using R software v3.1.3 (www.r-project.org) and the LIMMA package [24]. After quality assessment and removal of control probes, each array was background-corrected using the normexp method [25]. Normalization over all microarrays was done using the quantile–quantile method [26]. Replicates of identical probes (4 to 8 spots/probe) in each microarray were merged by taking the average of the normalized values. Microarrays from the same biopsy were also averaged. Differentially expressed miRNAs were identified and ranked using the moderated t-statistics [24]. In the analysis of all tumors against normal samples, the 5 paired samples (having both a primary tumor and a liver metastasis in the study) contributed only with primary tumors. miRNAs associated with good or poor survival were identified using the Cox proportional hazards model, adjusting for age and gender, where the survival times were calculated from the date of surgery. miRNA target scan was performed using TargetScan Human version 6.2 followed by biological pathway enrichment analysis using Fisher’s exact test in Partek Flow software, version 5.0 (Partek Inc., St. Louis, MO, USA). All P-values were adjusted for multiple testing using Benjamini–Hochberg false discovery rate.
The miRNA expression profiles were clustered using unsupervised hierarchical clustering with complete linkage and the Euclidean distance as metric. Associations between clusters and clinical characteristics were assessed using Fisher’s exact test and the ordinary t-test. For the metastases, the clustering of miRNA expression profiles were compared with the clustering of the mRNA profiles published previously using a permutation test [5]. Only cases represented by samples from the same tumor site in both studies were included in the test. First, within each study all possible pairs of such samples were annotated according to whether or not the two samples were situated in the same cluster. A test statistic, defined as the proportion of sample pairs having the same annotation in both studies was then calculated. A null distribution was calculated based on 100,000 permutations of clusters from the miRNA study. A corresponding P-value was derived by comparing the observed value of the test statistic to the null distribution. Expression data are available at the Gene Express Omnibus database (http://www.ncbi.nlm.nih.gov/geo/) with accession number GSE103317.
Survival analysis, testing the difference in survival time between groups of samples (formed either by clustering or by levels of miR-375 expression as measured on the tissue microarray), was performed using both Cox proportional hazards model (taking age and gender into account) and Kaplan‒Meier estimates together with a log-rank test. The miR-375 expression levels from the tissue microarray were stratified based on their scores, whereby samples with score 3 were classified as ‘high’ and samples with a score of 0‒2 were classified as ‘low’.
Student’s t-test was used to assess significance of differences in miR-375 expression measured by quantitative real-time PCR (qRT-PCR) (validation experiments). Fisher’s exact test was used to assess associations with β-catenin staining, including one tumor (chosen at random) per patient from the tissue microarray. A permutations test was used to evaluate association between nuclear expression of FOXM1 and copy number alteration.
Results
Cluster analysis of miRNA profiles differentiated primary small intestinal neuroendocrine tumors from their metastases and identified two groups of metastatic tumors
Small intestinal neuroendocrine tumors can be subdivided according to tumor grade, alterations in copy number, and gene expression pattern. In order to determine whether miRNA profiles can contribute to the molecular subtyping of tumors, we performed unsupervised hierarchical clustering analysis. miRNA profiles from all tumor biopsies and biopsies from normal intestinal mucosa were included in the analysis. Three tumor clusters were identified (Fig. 1). The largest cluster, designated M2, comprised 17 of the 42 tumors and consisted almost exclusively of metastases. The two smaller clusters, designated M1 and P, contained metastases only (10/42 tumors) (M1) or primary tumors only (13/42 tumors) (P). Two primary tumors clustered together with the normal small intestinal mucosa. Paired tumor samples (primary and metastases) from five patients segregated into different clusters. All paired primaries were found in cluster P, whereas all their paired metastases were found either in cluster M1 or cluster M2.
Histopathological and molecular characteristics of small intestinal neuroendocrine tumor clusters defined by miRNA profiles. Data on somatic copy number alterations, gene expression profiles, tumor grade, and tumor site are given for each patient. Tumors with higher grades (G2/G3) were more frequent in cluster M1, whereas lower-grade tumors (G1) predominated in cluster M2 and cluster P. Cluster M1 also had a higher proportion of tumors with multiple gains, including gain of chromosome 14, whereas clusters M2 and P had a higher proportion of tumors with loss of chromosome 18. Loss of CDKN1B, which is frequently mutated in small intestinal neuroendocrine tumors, was observed in one tumor from each cluster. Cluster M1 was enriched with tumors belonging to mRNA expression groups B and C, whereas cluster M2 was enriched with tumors belonging to mRNA expression group A. Details of somatic copy number alterations and RNA expression profiles of tumors have been published previously [5, 6]
Metastatic tumors in clusters M1 and M2 differ with respect to tumor grade, copy number alterations, and patient survival
In order to evaluate the clinical relevance of clusters M1 and M2 for metastatic tumors, we repeated the clustering analysis on metastases only, and obtained identical subgrouping of tumors. We then compared the clinicopathological characteristics of patients in clusters M1 and M2, and found that they were similar with respect to age, gender, and tumor stage (Table 2). However, Kaplan–Meier analysis showed significantly shorter overall survival of patients in cluster M1 than of patients in cluster M2 (P-value = 0.029) (Fig. 2). Using a Cox model, adjusting for gender and age, confirmed significantly shorter survival for patients in cluster M1 (hazard ratio (HR) = 3.4, P-value = 0.018). Cluster M1 had a significant overrepresentation of tumors with increased proliferation rate compared with cluster M2 measured as a higher proportion of tumors with higher grade (G2/G3) (P-value = 0.0011) and Ki67 index > 3% (P-value = 0.00023). Cluster M1 also had significant overrepresentation of tumors with gain of chromosome 14 (P-value = 0.0013). To determine whether clusters M1 and M2 were correlated to subgroups identified by mRNA expression profiles in our previous study [5], we performed a permutation test on tumors with both miRNA and mRNA profiles available. The test showed a significant association between tumor clusters obtained from miRNA profiles and from mRNA profiles (P-value = 0.0005). Tumors in cluster M1 were enriched in mRNA clusters B and C, while tumors in cluster M2 were enriched in mRNA cluster A (Table 2 and Fig. 3).
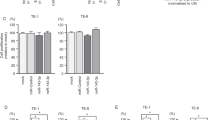
Differential expression of miRNAs in small intestinal neuroendocrine tumor clusters M1 and M2. (a) Average expression levels of miRNAs in clusters M1 and M2. (b) The top 10 most significant differentially expressed miRNAs in cluster M1 compared with cluster M2. (c) Pathway analysis of differentially expressed miRNAs between clusters M1 and M2. The top 10 most significantly altered biological pathways are given. Wnt signaling was predicted to be the most differentially regulated pathway between clusters
Pathway analysis of miRNA targets predicted significant alterations in Wnt signaling between metastatic tumors in clusters M1 and M2
In order to identify miRNAs that were differentially expressed in the two metastatic subgroups, we compared the expression of miRNAs in cluster M1 with that of miRNAs in cluster M2. We found that 76 miRNAs were significantly upregulated and 65 were significantly downregulated in cluster M1 compared with cluster M2 (adjusted P-value < 0.05) (Fig. 4 and Supplementary Table S1). We then selected significantly regulated miRNA with a log2 fold change of < − 1 or > 1, and searched for them in the miRCancer database (http://mircancer.ecu.edu). We found that 20/32 (62.5%) of the differentially expressed miRNAs were reported to regulate important properties of malignant tumors. The two most upregulated miRNAs in cluster M1, miR-1246, and miR-663a, which are highly expressed in several types of cancers, have been implicated as oncomiRs regulating tumor cell proliferation, invasiveness, and formation of metastases [27,28,29,30,31]. The most significantly downregulated miRNA in cluster M1, miR-488-3p, has been shown to act as a tumor suppressor in gastric cancer [32]. To predict the molecular pathways affected by significantly regulated miRNAs, we performed a miRNA target scan followed by biological pathway enrichment analysis. Wnt signaling was predicted to be the most significantly affected signaling pathway (adjusted P-value = 3.2 × 10−9) followed by the axon guidance (adjusted P-value = 2.1 × 10−6) and phosphatidylinositol signaling pathways (adjusted P-value = 5.8 × 10−6) (Fig. 4c). Four of the significantly regulated miRNAs (miR-1246, miR-663a, miR-483-5p, and miR-1290) have been shown to control Wnt signaling by targeting different components of the signaling cascade [33,34,35,36].
Differential expression of miRNAs in metastases compared to primary tumors of small intestinal neuroendocrine tumors. We analyzed miRNA profiles from paired samples from 5 patients. (a) Comparison of average expression levels of miRNA in liver metastases and in primary tumors. (b) The top 10 most significant differentially expressed miRNA in metastases compared with primary tumors. *miRNA previously shown to be downregulated in metastases compared with primary tumors [52,53,54]. (c) Pathway analysis of differentially expressed miRNAs between metastases and primary tumors. The top 10 most significant differentially regulated biological pathways are shown
miRNA profiles associated with proliferation rate in small intestinal neuroendocrine tumors
Small intestinal neuroendocrine tumors are graded into three categories (G1–G3) based on their proliferation rate measured by mitotic counts and Ki67 index (WHO 2010). Tumor grade significantly influences prognosis and may predict response to therapy [3, 37]. In order to identify miRNAs that may control tumor cell proliferation, we compared tumor metastases with Ki67 index > 3% with metastases with Ki67 index < 3% and found 48 differentially expressed miRNAs (adjusted P-value < 0.05; Supplementary Table S2). Among these we found downregulation of miRNAs, which function as tumor suppressors, e.g., miR-137, miR-204-5p, miR-486-5p, and miR-30c, and upregulation of miRNAs that function as oncogenes, e.g., miR-21 and miR1290. A majority of the differentially expressed miRNAs has been shown to regulate tumor proliferation, but a substantial proportion of identified miRNAs have also been implicated in invasive behavior and drug resistance [38,39,40,41,42]. To search for molecular pathways affected by differentially expressed miRNAs, we performed a miRNA target scan followed by biological pathway enrichment analysis. Wnt signaling was predicted to be the most significantly affected pathway (adjusted P-value = 2.9 × 10−6) followed by the phosphatidylinositol signaling system (adjusted P-value = 1.1 × 10−5) and axon guidance (adjusted P-value = 1.1 × 10−5) (Supplementary Table S3). We also compared tumor metastases of lower grade (G1) with those of higher grade (G2/3) and found differential expression of the same miRNA that were identified when tumors were compared with respect to Ki67 index. However, differential expression of miRNA between tumor grade did not reach statistical significance. In order to evaluate whether tumor differentiation or secretory function influenced the expression of miRNAs we compared tumors with high expression of CDX2 or serotonin with those with low expression. This comparison did not show any significantly regulated miRNAs (Supplementary Table S8).
miRNA profiles associated with copy number alterations in small intestinal neuroendocrine tumors
We have previously shown that small intestinal neuroendocrine tumors carrying multiple chromosomal gains, notably gain of chromosome 14, have worse prognosis and are characterized by activation of the oncogenic transcription factor FOXM1 [5, 6]. To extend the molecular characterization of this subgroup of tumors, we compared the miRNA profiles of metastases with gain of chromosome 14 with those with no gain of chromosome 14. Altogether, 44 miRNAs were differentially regulated between the two tumor groups (adjusted P-value < 0.05) (Supplementary Table S4). Regulated miRNAs were distributed over all chromosomes, with only two significantly upregulated miRNAs located on chromosome 14 (miR-1260a, log2 fold change = 1.14; and miR-345-5p, log2 fold change = 0.72). Five of the regulated miRNAs (miR-320a, miR-320b, miR-320c, miR-320d, and miR-21) directly regulate the expression of FOXM1 [43,44,45]. Alterations in the expression levels of these 5 miRNAs were consistent with activation of FOXM1 in cluster M1. Staining for FOXM1 protein in the metastases of the microarray cohort confirmed higher nuclear expression of FOXM1 in biopsies with gain of chromosome 14 (P-value = 3.0 × 10−5) (Supplementary Table S8). We performed an miRNA target scan on all significantly regulated miRNAs, followed by biological pathway enrichment analysis and found that Wnt signaling was the most significantly regulated pathway (adjusted P-value = 2.5 × 10−06) (Supplementary Table S5). Three of the most significantly regulated miRNAs (miR-1290, miR-501-5p, and let-7e-5p) have been shown to directly regulate Wnt signaling [36, 46, 47].
miRNA profiles associated with disease progression in small intestinal neuroendocrine tumors
In order to identify miRNAs associated with disease progression, we compared the miRNA profiles of paired tumor biopsies (primaries and metastases) from 5 patients. Unsupervised clustering analysis grouped all paired primary tumors in cluster P and all paired metastases in cluster M1 or cluster M2 (Fig. 1), indicating that the miRNA profiles of primary tumors and their metastases differ substantially. We then performed a pairwise analysis of metastases and primary tumors, and searched for differentially expressed miRNAs. A total of 25 miRNAs were found to be differentially expressed, 14 of which were upregulated and 11 downregulated in the metastases (adjusted P-value < 0.05) (Supplementary Table S6). The 10 most significantly regulated miRNAs included 2 upregulated miRNAs, miR-95 and miR-210, which are known to be oncomiRs regulating radio-resistance and renewal of tumor-initiating cells [27, 48]. miRNAs found to be downregulated in metastases included mir-378-3p, miR-1-3p, and miR-133b, which are known to act as tumor suppressors (Fig. 5) [49,50,51]. Comparison with previous studies indicated good agreement in differentially expressed miRNAs during tumor progression [52,53,54]. miRNA target scan on all significantly regulated miRNAs followed by biological pathway enrichment analysis predicted that endocytosis (adjusted P-value = 8.7 × 10−7), Wnt signaling (adjusted P-value = 8.7 × 10−7), and axon guidance (adjusted P-value = 9.5 × 10−7) were the three most differentially regulated pathways (Fig. 5c).
Expression of β-catenin protein in small intestinal neuroendocrine tumors. The staining intensity and nuclear translocation of β-catenin were analyzed by immunohistochemistry on tissue microarray. Representative images of different patterns of β-catenin staining. (a) A primary tumor with strong membranous and cytoplasmic staining as well as nuclear translocation of β-catenin. (b) A liver metastasis with strong membranous and cytoplasmic staining of β-catenin but no nuclear translocation. (c) A primary tumor with weak membranous staining of β-catenin and nuclear translocation. (d) A primary tumor with weak membranous staining of β-catenin and no nuclear translocation. (e) Nuclear localization of β-catenin occurs when the Wnt canonical pathway is activated. β-Catenin was significantly more frequent translocated to the nucleus in primary tumors compared to metastasis (P-value < 1 × 10−15)
β-Catenin protein is downregulated in small intestinal neuroendocrine tumors during tumor progression
Nuclear translocation of β-catenin is a hallmark of activated canonical Wnt signaling pathway. To validate altered Wnt/β-catenin signaling in small intestinal neuroendocrine tumors we analyzed a separate cohort of small intestinal neuroendocrine tumors by immunohistochemical staining of the β-catenin protein on a tissue microarray. We analyzed the intensity and cellular localization of β-catenin and found positive but variable staining intensity in all samples (Fig. 6a, b, c, d). β-Catenin staining of tumor metastases was lower than that of primary tumors (P-value = 0.0005) (Fig. 6e). Nuclear localization of β-catenin was found in 31% of the primary tumors but was lost in tumor metastases (P-value < 1 × 10−15) (Fig. 6e).
Differential expression of miRNAs in small intestinal neuroendocrine tumors compared with normal small intestinal mucosa. (a) Average expression levels of miRNAs in tumors compared with normal small intestinal mucosa. (b) The top 10 most significant differentially expressed miRNAs in tumors compared with small intestinal mucosa. miR-7-5p and miR-375 were the two most significantly upregulated miRNA in tumors. *Indicates miRNA previously shown to be deregulated in small intestinal neuroendocrine tumors compared with normal small intestine [52,53,54]. (c) Pathway analysis of differentially expressed miRNAs in tumors. The top 10 most significantly affected biological pathways are shown
miR-7-5p and miR-375 are highly expressed in small intestinal neuroendocrine tumors
To search for potential biomarkers of small intestinal neuroendocrine tumors, we compared the miRNA expression profiles of small intestinal neuroendocrine tumors with that of normal small intestinal mucosa. A total of 155 miRNAs were found to be differentially expressed, with 75 miRNAs being upregulated and 80 downregulated in tumor biopsies (adjusted P-value < 0.05 and log2 fold change < − 1 or > 1) (Supplementary Table S7). miR-7-5p and miR-375 were the most significantly upregulated miRNAs (adjusted P-values = 1.0 × 10−19 and 4.2 × 10−19). The relative expression of miR-7-5p and miR-375 in tumor biopsies compared with normal small intestinal mucosa was 5.7 and 4.6 log2 fold change, respectively. The average expression values of miR-7-5p and miR-375 in tumor biopsies were high (Fig. 7a). To validate the expression of miR-375 in small intestinal neuroendocrine tumors we performed qRT-PCR analysis of 20 cases from an independent cohort of tumors. qRT-PCR measurements showed on average 4.1 log2 fold (17 times) higher expression in small intestinal neuroendocrine tumors compared with normal small intestinal mucosa, which was in good agreement with array data (Supplementary Figure S2). We also validated differentially expressed miRNAs, by comparing our data with those of a recent profiling study on small intestinal neuroendocrine tumors [53]. We could confirm upregulation of 27 out of the 29 miRNAs previously reported to be upregulated in tumor biopsies. To predict the signaling pathways that are deregulated in tumors, we performed a miRNA target scan on all significantly regulated miRNAs, followed by biological pathway enrichment analysis, and found Wnt signaling to be the most significantly affected pathway (adjusted P-value = 7.0 × 10−9) followed by Hippo signaling (adjusted P-value = 9.1 × 10−6) (Fig. 7c). Three of the most significantly regulated miRNAs (miR-375, miR-153, and miR-29c) have been shown to regulate Wnt signaling, whereas miR-375 has been implicated in the regulation of Hippo signaling [55,56,57].
In situ expression of miR-375. (a) In situ hybridization of normal small intestinal mucosa showed high expression of miR-375 in enteroendocrine cells located both at the base of the crypt and on the villus. In addition, miR-375 was also expressed in Paneth cells at a lower level. (b, c, d) In situ hybridization of small intestinal neuroendocrine tumors showed high expression of miR-375 in tumor cells. MiR-375 was localized to the cytoplasm of tumor cells, with no expression in surrounding stromal cells. Representative images from a primary tumor (b), a lymph node metastasis (c), and a liver metastasis (d)
miR-375 is highly expressed in small intestinal neuroendocrine tumors and its downregulation in tumor metastases is associated with shorter patient survival
The epigenetic alterations that contribute to the aggressive behavior of small intestinal neuroendocrine tumors are unknown. To address this question, we analyzed the relation between individual miRNA expression values and patient survival using a Cox proportional hazards model. Searching miRNA expression in all tumor metastases (n = 26), we found that downregulation of miR-375 showed the most significant correlation with patient survival (HR = 0.036, adjusted P-value = 0.093) (Table 3). In order to characterize the in situ expression of miR-375 and its correlation with patient survival, we performed in situ hybridization in normal small intestinal mucosa and in small intestinal neuroendocrine tumors using a miR-375-specific probe (Fig. 8). In normal intestinal mucosa, miR-375 was highly expressed in enteroendocrine cells located both in the crypts and on villi. In addition, miR-375 was also present in the Paneth cells, but with a lower level of expression. In tumor tissue, the miR-375 expression was localized to the cytoplasm of the tumor cells with no expression in the stroma cells (Fig. 8b, c, d). Using a tissue microarray from an independent cohort of small intestinal neuroendocrine tumors, we found expression of miR-375 in 578/635 biopsies (91.0%) by in situ hybridization (Fig. 9 and Table 4). Kaplan–Meier analysis showed that patients with high expression of miR-375 in liver metastases (score 3, n = 15) had significantly longer survival (P-value = 0.016) than patients with low expression (scores 0, 1, or 2; n = 45) (Fig. 10). Using a Cox model, adjusting for gender and age, we confirmed longer survival for patients with high expression of miR-375 (HR = 0.32, P-value = 0.026). There was also a trend (not significant) that patients with high expression of miR-375 in lymph node metastases had better survival than patients with low expression.
Scoring of miR-375 expression on tissue microarray of small intestinal neuroendocrine tumors. The tissue microarray was subjected to in situ hybridization using a miR-375-specific probe. The intensity of the miR-375 hybridization signal was scored as follows: 0 = no staining; 1 = weak staining; 2 = intermediate staining; and 3 = strong staining. Each biopsy on the tissue microarray was confirmed to contain tumor tissues by evaluating consecutive sections stained with hematoxylin and eosin, and for chromogranin A and synaptophysin
Downregulation of miR-375 in liver metastases correlates to shorter overall survival. Kaplan‒Meier analysis was performed on patients with liver metastases grouped according to their in situ expression of miR-375: high expression (score 3, n = 15) and low expression (score 0‒2, n = 45). The survival analysis was performed on a cohort of patients that was independent from that analyzed with miRNA profiling
Discussion
Large efforts have been made during the past years, to improve our knowledge of the molecular background of small intestinal neuroendocrine tumors. Although the mutational landscape of these tumors has been described, the molecular mechanisms that drive the tumorigenesis and disease progression are still poorly understood. Sequencing of the whole genome and exome have shown that small intestinal neuroendocrine tumors are genetically relatively stable [7]; however, epigenetic deregulation has emerged as important mechanism of tumor initiation and progression in small intestinal neuroendocrine tumors. Major epigenetic alterations in small intestinal neuroendocrine tumors include promoter methylation and miRNA deregulation [53, 58, 59]. To further investigate the role of miRNAs in small intestinal neuroendocrine tumors we have performed a miRNA profiling study on a large cohort of patients treated with surgery at a single center. We analyzed a total of 42 tumors with known somatic copy number alterations, transcriptional profiles, and long-term follow-up. Data from this study allowed us to identify miRNA alterations not previously reported, e.g., alterations associated with novel groups of metastatic tumors, specific miRNAs related to tumor proliferation rate, and miRNA associated with patient survival.
First, we performed an unsupervised clustering analysis to search for tumor subgroups defined by their miRNA expression. This analysis separated primary tumors from metastases and identified two groups of metastases. Comparing the clinicopathological characteristics of the two metastatic clusters we found that patients with metastases in cluster M1 had the shortest postoperative survival and this cluster was enriched with higher-grade tumors. Analysing the miRNA profiles of the two clusters, we were able to identify a number of differentially expressed miRNAs previously reported to be associated with malignant behavior of tumors, including well-known oncomiRs such as miR-1246 and miR-663a, and tumor suppressor miRNA miR-488-3p [27,28,29,30,31,32]. To search for miRNAs that specifically control tumor proliferation rate, we compared miRNA expression in metastases with high Ki67 index versus low Ki67 index (cutoff 3%). This comparison generated a number of significantly regulated miRNAs, some of which have previously been implied to control tumor proliferation as well as invasive growth and metastasis, e.g., miR-137 and miR-204-5p [38,39,40,41,42, 53]. The molecular pathways that are controled by these miRNAs are not known and need to be further elucidated. Also, the potential usefulness of proliferation-associated miRNAs as prognostic and predictive biomarkers for small intestinal neuroendocrine tumors should be explored.
Deregulation of miRNAs is an important event in the progression of tumors. To study deregulation of miRNAs in small intestinal neuroendocrine tumors during tumor progression, we compared paired samples of primary and metastatic tumors. The results of this analysis were in good agreement with those from previous studies showing high expression of miR-122-5p in metastases and high expression of miR-133b, miR-145-5p, and miR-1-3p in primary tumors [52,53,54]. In order to better understand the cellular effects of deregulated miRNAs during progression of small intestinal neuroendocrine tumors, we performed a combined target prediction and pathway analysis. Using this approach, we identified the Wnt signaling pathway to be one of the most significantly affected pathway when primary tumors were compared with metastatic tumors. Wnt, Hippo, Notch, Hedgehog, and transforming growth factor-β/bone morphogenetic protein signaling pathways are highly conserved regulatory networks that control the proliferation and differentiation of epithelial cells in the gastrointestinal mucosa [60]. In order to assess Wnt signaling in small intestinal neuroendocrine tumors, we measured the activity of the canonical Wnt signaling pathway by analyzing nuclear translocation of β-catenin. We found that β-catenin protein was preferentially translocated to the nuclei in primary tumors with virtually no translocation of β-catenin in tumor metastases. The difference in Wnt signaling activity may be due to deregulation of miRNA expression. Alternatively, the difference in Wnt signaling may be due to tumor microenvironment, providing a stronger signal for Wnt activation in primary tumors compared to their metastases. It has also been shown that the β-catenin promoter is hypermethylated in metastases of small intestinal neuroendocrine tumors, providing yet another explanation for alterations of β-catenin activity during tumor progression [58, 61].
In order to identify candidate biomarkers for small intestinal neuroendocrine tumors, we compared tumors to normal intestinal mucosa and found that miR-375 and miR-7-5p were highly expressed in tumors. This finding was in agreement with a recent miRNA profiling study [53]. To further validate this observation, we performed in situ hybridization and confirmed that miR-375 was highly expressed in small intestinal neuroendocrine tumor cells with no expression in stromal cells. We also analyzed non-endocrine tumors of the gastrointestinal tract and pancreas, and found them to be negative for miR-375 (Supplementary Table S9). Analysis of normal small intestinal mucosa showed that miR-375 was specifically expressed in the enteroendocrine cells located in the crypt and on the villus. These findings are in agreement with a recent report identifying miR-375 as a regulator of the enteroendocrine cell lineage [62]. Furthermore, endocrine organs, e.g. pituitary and adrenal gland as well as the endocrine pancreas have been shown to express miR-375 and miR-7-5p at high levels [63,64,65]. Collectively, these observations suggest that expression of miR-375 in small intestinal neuroendocrine tumors represents a specific marker of neuroendocrine differentiation and suggest that small intestinal neuroendocrine tumors originate from the enteroendocrine cell linage. Interestingly, survival analysis of the miRNA profiles also identified miR-375 as the strongest predictor of overall patient survival. We therefore analyzed the in situ expression of miR-375 in an independent cohort of tumors and confirmed that downregulation of miR-375 in liver metastases was associated with shorter survival. These data are consistent with the idea of miR-375 acting as a tumor suppressor in small intestinal neuroendocrine tumors, and is in agreement with observations from other tumors identifying miR-375 as a tumor suppressor and a biomarker of prognosis [66]. However, the functional role of miR-375 in small intestinal neuroendocrine tumors need to be established since the effect of miRNAs depend on the cellular context [18]. Although miR-375 acts as a tumor suppressor in the majority of tumors, oncogenic effects have been described in breast and prostate cancers [66]. Of great interest is the observation that miR-375 is specifically expressed in neuroendocrine cells of the pituitary, pancreas, and intestine. The expression of miR-375 is regulated by the transcription factors NeuroD1 and Pdx1, and also by methylation of the promoter region of the miR-375 gene [66,67,68]. NeuroD1 has been shown to be an important regulator of the enteroendocrine cell linage, and is expressed at the late stage of differentiation when cells have become restricted to the enteroendocrine cell fate. Recently, miR-375 was identified as a regulator of the enteroendocrine lineage during renewal and differentiation of small intestinal epithelium [62]. This observation raises the possibility that miR-375 might regulate the neuroendocrine differentiation of gastrointestinal neuroendocrine tumors, including small intestinal neuroendocrine tumors. Further investigations are warranted to define the role of miR-375 in determining differentiation and cell linage in neuroendocrine tumors. Moreover, miR-375 should be evaluated as a diagnostic marker for small intestinal neuroendocrine tumors as well as a biomarker for prognostication and stratification of patients in clinical trials.
References
Niederle B, Pape UF, Costa F, et al. ENETS Consensus Guidelines update for neuroendocrine neoplasms of the jejunum and ileum. Neuroendocrinology. 2016;103:125–38.
Pavel M, O’Toole D, Costa F, et al. ENETS Consensus Guidelines update for the management of distant metastatic disease of intestinal, pancreatic, bronchial neuroendocrine neoplasms (NEN) and NEN of unknown primary site. Neuroendocrinology. 2016;103:172–85.
Ahmed A, Turner G, King B, et al. Midgut neuroendocrine tumours with liver metastases: results of the UKINETS study. Endocr Relat Cancer. 2009;16:885–94.
Modlin IM, Moss SF, Chung DC, Jensen RT, Snyderwine E. Priorities for improving the management of gastroenteropancreatic neuroendocrine tumors. J Natl Cancer Inst. 2008;100:1282–9.
Andersson E, Arvidsson Y, Swärd C, et al. Expression profiling of small intestinal neuroendocrine tumors identifies subgroups with clinical relevance, prognostic markers and therapeutic targets. Mod Pathol. 2016;29:616–29.
Andersson E, Swärd C, Stenman G, Ahlman H, Nilsson O. High-resolution genomic profiling reveals gain of chromosome 14 as a predictor of poor outcome in ileal carcinoids. Endocr Relat Cancer. 2009;16:953–66.
Banck MS, Kanwar R, Kulkarni AA, et al. The genomic landscape of small intestine neuroendocrine tumors. J Clin Invest. 2013;123:2502–8.
Francis JM, Kiezun A, Ramos AH, et al. Somatic mutation of CDKN1B in small intestine neuroendocrine tumors. Nat Genet. 2013;45:1483–6.
Karpathakis A, Dibra H, Pipinikas C, et al. Prognostic impact of novel molecular subtypes of small intestinal neuroendocrine tumor. Clin Cancer Res. 2016;22:250–8.
Hashemi J, Fotouhi O, Sulaiman L, et al. Copy number alterations in small intestinal neuroendocrine tumors determined by array comparative genomic hybridization. BMC Cancer. 2013;13:505.
Crona J, Gustavsson T, Norlén O, et al. Somatic mutations and genetic heterogeneity at the CDKN1B locus in small intestinal neuroendocrine tumors. Ann Surg Oncol. 2015;22:1428–35.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Mendell JT, Olson EN. MicroRNAs in stress signaling and human disease. Cell. 2012;148:1172–87.
Calin GA, Ferracin M, Cimmino A, et al. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med. 2005;353:1793–801.
He H, Jazdzewski K, Li W, et al. The role of microRNA genes in papillary thyroid carcinoma. Proc Natl Acad Sci USA. 2005;102:19075–80.
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Volinia S, Calin GA, Liu CG, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA. 2006;103:2257–61.
Svoronos AA, Engelman DM, Slack FJ. OncomiR or tumor suppressor? The Duplicity of MicroRNAs in Cancer. Cancer Res. 2016;76:3666–70.
Bosman TF CF, Rhuban RH (eds). WHO Classification of Tumors of the Digestive Systems Vol 4th edn, pp 102–107 (IARC: Lyon, France, 2010).
Sobin L, Gospodarowicz M, Wittekind C (eds). TNM Classification of Malignant Tumors. Vol 7th edn, pp 310 (Wiley-Blackwell: Oxford, UK, 2010)
Tang LH, Gonen M, Hedvat C, Modlin IM, Klimstra DS. Objective quantification of the Ki67 proliferative index in neuroendocrine tumors of the gastroenteropancreatic system: a comparison of digital image analysis with manual methods. Am J Surg Pathol. 2012;36:1761–70.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–8.
Li SC, Khan M, Caplin M, et al. Somatostatin analogs treated small intestinal neuroendocrine tumor patients circulating microRNAs. PLoS ONE. 2015;10:e0125553.
Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article3.
Ritchie ME, Silver J, Oshlack A, et al. A comparison of background correction methods for two-colour microarrays. Bioinformatics. 2007;23:2700–7.
Bolstad BM, Irizarry RA, Astrand M, Speed TP. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19:185–93.
Huang W, Li H, Luo R. The microRNA-1246 promotes metastasis in non-small cell lung cancer by targeting cytoplasmic polyadenylation element-binding protein 4. Diagn Pathol. 2015;10:127.
Kim G, An HJ, Lee MJ, et al. Hsa-miR-1246 and hsa-miR-1290 are associated with stemness and invasiveness of non-small cell lung cancer. Lung Cancer. 2016;91:15–22.
Liu ZY, Zhang GL, Wang MM, Xiong YN, Cui HQ. MicroRNA-663 targets TGFB1 and regulates lung cancer proliferation. Asian Pac J Cancer Prev. 2011;12:2819–23.
Neerincx M, Sie DL, van de Wiel MA, et al. MiR expression profiles of paired primary colorectal cancer and metastases by next-generation sequencing. Oncogenesis. 2015;4:e170.
Yi C, Wang Q, Wang L, et al. MiR-663, a microRNA targetingp21(WAF1/CIP1), promotes the proliferation and tumorigenesis of nasopharyngeal carcinoma. Oncogene. 2012;31:4421–33.
Zhao Y, Lu G, Ke X, et al. miR-488 acts as a tumor suppressor gene in gastric cancer. Tumour Biol. 2016;37:8691–8.
Chai S, Ng KY, Tong M, et al. Octamer 4/microRNA-1246 signaling axis drives Wnt/beta-catenin activation in liver cancer stem cells. Hepatology. 2016;64:2062–76.
Miao CG, Shi WJ, Xiong YY, et al. MicroRNA-663 activates the canonical Wnt signaling through the adenomatous polyposis coli suppression. Immunol Lett. 2015;166:45–54.
Song Q, Xu Y, Yang C, et al. miR-483-5p promotes invasion and metastasis of lung adenocarcinoma by targeting RhoGDI1 and ALCAM. Cancer Res. 2014;74:3031–42.
Wu J, Ji X, Zhu L, et al. Up-regulation of microRNA-1290 impairs cytokinesis and affects the reprogramming of colon cancer cells. Cancer Lett. 2013;329:155–63.
Sorby H, Welin S, Langer SW, et al. Predictive and prognostic factors for treatment and survival in 305 patients with advanced gastrointestinal neuroendocrine carcinoma (WHO G3): the NORDIC NEC study. Ann Oncol. 2013;24:152–60.
Hu Y, Xie H, Liu Y, et al. miR-484 suppresses proliferation and epithelial-mesenchymal transition by targeting ZEB1 and SMAD2 in cervical cancer cells. Cancer Cell Int. 2017;17:36.
Lin M, Shi C, Lin X, et al. sMicroRNA-1290 inhibits cells proliferation and migration by targeting FOXA1 in gastric cancer cells. Gene. 2016;582:137–42.
Shen H, Wang L, Ge X, et al. MicroRNA-137 inhibits tumor growth and sensitizes chemosensitivity to paclitaxel and cisplatin in lung cancer. Oncotarget. 2016;7:20728–42.
Wang Z, Yang H, Ren L. MiR-21 promoted proliferation and migration in hepatocellular carcinoma through negative regulation of Navigator-3. Biochem Biophys Res Commun. 2015;464:1228–34.
Yin Y, Zhang B, Wang W, et al. miR-204-5p inhibits proliferation and invasion and enhances chemotherapeutic sensitivity of colorectal cancer cells by downregulating RAB22A. Clin Cancer Res. 2014;20:6187–99.
Song G, Sharma AD, Roll GR, et al. MicroRNAs control hepatocyte proliferation during liver regeneration. Hepatology. 2010;51:1735–43.
Wan LY, Deng J, Xiang XJ, et al. miR-320 enhances the sensitivity of human colon cancer cells to chemoradiotherapy in vitro by targeting FOXM1. Biochem Biophys Res Commun. 2015;457:125–32.
Vishnubalaji R, Hamam R, Yue S, et al. MicroRNA-320 suppresses colorectal cancer by targeting SOX4, FOXM1, and FOXQ1. Oncotarget. 2016;7:35789–802.
Fan D, Ren B, Yang X, Liu J, Zhang Z. Upregulation of miR-501-5p activates the wnt/beta-catenin signaling pathway and enhances stem cell-like phenotype in gastric cancer. J Exp Clin Cancer Res. 2016;35:177.
Vinas JL, Ventayol M, Brune B, et al. miRNA let-7e modulates the Wnt pathway and early nephrogenic markers in mouse embryonic stem cell differentiation. PLoS ONE. 2013;8:e60937.
Ullmann P, Qureshi-Baig K, Rodriguez F, et al. Hypoxia-responsive miR-210 promotes self-renewal capacity of colon tumor-initiating cells by repressing ISCU and by inducing lactate production. Oncotarget. 2016;7:65454–70.
Anton R, Chatterjee SS, Simundza J, Cowin P, Dasgupta R. A systematic screen for micro-RNAs regulating the canonical Wnt pathway. PLoS ONE. 2011;6:e26257.
Dai J, Wu H, Zhang Y, et al. Negative feedback between TAp63 and Mir-133b mediates colorectal cancer suppression. Oncotarget. 2016;7:87147–60.
Zhang GJ, Zhou H, Xiao HX, Li Y, Zhou T. MiR-378 is an independent prognostic factor and inhibits cell growth and invasion in colorectal cancer. BMC Cancer. 2014;14:109.
Li SC, Essaghir A, Martijn C, et al. Global microRNA profiling of well-differentiated small intestinal neuroendocrine tumors. Mod Pathol. 2013;26:685–96.
Miller HC, Frampton AE, Malczewska A, et al. MicroRNAs associated with small bowel neuroendocrine tumours and their metastases. Endocr Relat Cancer. 2016;23:711–26.
Ruebel K, Leontovich AA, Stilling GA, et al. MicroRNA expression in ileal carcinoid tumors: downregulation of microRNA-133a with tumor progression. Mod Pathol. 2010;23:367–75.
Hua HW, Jiang F, Huang Q, Liao Z, Ding G. MicroRNA-153 promotes Wnt/beta-catenin activation in hepatocellular carcinoma through suppression of WWOX. Oncotarget. 2015;6:3840–7.
Jiang J, Yu C, Chen M, et al. Reduction of miR-29c enhances pancreatic cancer cell migration and stem cell-like phenotype. Oncotarget. 2015;6:2767–78.
Xu L, Wen T, Liu Z, et al. MicroRNA-375 suppresses human colorectal cancer metastasis by targeting Frizzled 8. Oncotarget. 2016;7:40644–56.
Fotouhi O, Adel Fahmideh M, Kjellman M, et al. Global hypomethylation and promoter methylation in small intestinal neuroendocrine tumors: an in vivo and in vitro study. Epigenetics. 2014;9:987–97.
Karpathakis A, Dibra H, Pipinikas C, et al. Progressive epigenetic dysregulation in neuroendocrine tumour liver metastases. Endocr Relat Cancer. 2017;24:L21–L5.
Jeon MK, Klaus C, Kaemmerer E, Gassler N. Intestinal barrier: molecular pathways and modifiers. World J Gastrointest Pathophysiol. 2013;4:94–9.
Kim JT, Li J, Jang ER, et al. Deregulation of Wnt/beta-catenin signaling through genetic or epigenetic alterations in human neuroendocrine tumors. Carcinogenesis. 2013;34:953–61.
Knudsen LA, Petersen N, Schwartz TW, Egerod KL. The microRNA repertoire in enteroendocrine cells: identification of miR-375 as a potential regulator of the enteroendocrine lineage. Endocrinology. 2015;156:3971–83.
Landgraf P, Rusu M, Sheridan R, et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell. 2007;129:1401–14.
Ludwig N, Leidinger P, Becker K, et al. Distribution of miRNA expression across human tissues. Nucleic Acids Res. 2016;44:3865–77.
He J, Cao Y, Su T, et al. Downregulation of miR-375 in aldosterone-producing adenomas promotes tumour cell growth via MTDH. Clin Endocrinol (Oxf). 2015;83:581–9.
Yan JW, Lin JS, He XX. The emerging role of miR-375 in cancer. Int J Cancer. 2014;135:1011–8.
Avnit-Sagi T, Kantorovich L, Kredo-Russo S, Hornstein E, Walker MD. The promoter of the pri-miR-375 gene directs expression selectively to the endocrine pancreas. PLoS ONE. 2009;4:e5033.
Keller DM, McWeeney S, Arsenlis A, et al. Characterization of pancreatic transcription factor Pdx-1 binding sites using promoter microarray and serial analysis of chromatin occupancy. J Biol Chem. 2007;282:32084–92.
Acknowledgements
The expert technical assistance of Pauline Brattberg and Linda Inge is greatly appreciated. This study was supported by the Swedish Cancer Society, Sahlgrenska Academy (the government ALF agreement), BioCARE- a National Strategic Research program at University of Gothenburg, Chalmers University of Technology, The Sahlgrenska University Funds, and the Assar Gabrielsson Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest:
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Arvidsson, Y., Rehammar, A., Bergström, A. et al. miRNA profiling of small intestinal neuroendocrine tumors defines novel molecular subtypes and identifies miR-375 as a biomarker of patient survival. Mod Pathol 31, 1302–1317 (2018). https://doi.org/10.1038/s41379-018-0010-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-018-0010-1
This article is cited by
-
Exposure to nonanoic acid alters small intestinal neuroendocrine tumor phenotype
BMC Cancer (2023)
-
The Quest for Circulating Biomarkers in Neuroendocrine Neoplasms: a Clinical Perspective
Current Treatment Options in Oncology (2023)
-
SMAD4 haploinsufficiency in small intestinal neuroendocrine tumors
BMC Cancer (2021)
-
Profiling of Serum Exosome MiRNA Reveals the Potential of a MiRNA Panel as Diagnostic Biomarker for Alzheimer’s Disease
Molecular Neurobiology (2021)
-
Loss of copy of MIR1-2 increases CDK4 expression in ileal neuroendocrine tumors
Oncogenesis (2020)