Abstract
Adenoid cystic carcinoma is among the most frequent malignancies in the salivary and lacrimal glands and has a grave prognosis characterized by frequent local recurrences, distant metastases, and tumor-related mortality. Conversely, adenoid cystic carcinoma of the breast is a rare type of triple-negative (estrogen and progesterone receptor, HER2) and basal-like carcinoma, which in contrast to other triple-negative and basal-like breast carcinomas has a very favorable prognosis. Irrespective of site, adenoid cystic carcinoma is characterized by gene fusions involving MYB, MYBL1, and NFIB, and the reason for the different clinical outcomes is unknown. In order to identify the molecular mechanisms underlying the discrepancy in clinical outcome, we characterized the phenotypic profiles, pattern of gene rearrangements, and global microRNA expression profiles of 64 salivary gland, 9 lacrimal gland, and 11 breast adenoid cystic carcinomas. All breast and lacrimal gland adenoid cystic carcinomas had triple-negative and basal-like phenotypes, while salivary gland tumors were indeterminate in 13% of cases. Aberrations in MYB and/or NFIB were found in the majority of cases in all three locations, whereas MYBL1 involvement was restricted to tumors in the salivary gland. Global microRNA expression profiling separated salivary and lacrimal gland adenoid cystic carcinoma from their respective normal glands but could not distinguish normal breast adenoid cystic carcinoma from normal breast tissue. Hierarchical clustering separated adenoid cystic carcinomas of salivary gland origin from those of the breast and placed lacrimal gland carcinomas in between these. Functional annotation of the microRNAs differentially expressed between salivary gland and breast adenoid cystic carcinoma showed these as regulating genes involved in metabolism, signal transduction, and genes involved in other cancers. In conclusion, microRNA dysregulation is the first class of molecules separating adenoid cystic carcinoma according to the site of origin. This highlights a novel venue for exploring the biology of adenoid cystic carcinoma.
Similar content being viewed by others
Introduction
Adenoid cystic carcinoma is among the most frequent malignancies in the salivary and lacrimal gland, and is clinically characterized by frequent local invasion, distant metastatic spread, and a dismal prognosis, with 10-year disease-specific survival rates of 75% and 52%, respectively [1, 2]. In the breast, adenoid cystic carcinoma is rare and belongs to the basal-like (estrogen receptor and human epidermal growth factor receptor 2 (HER2) negative, CK5/6 and/or epidermal growth factor receptor (EGFR) positive) and triple-negative (negative for estrogen receptor, progesterone receptor, and HER2) subtypes [3]. The phenotypic classification of breast carcinoma (i.e., luminal, basal-like, and HER2 phenotypes) carries significant prognostic and therapeutic implications in breast cancer and has been shown to apply to at least a subset of salivary and lacrimal gland carcinomas [3,4,5,6]. The phenotypic profile of adenoid cystic carcinoma has not been characterized, which constitutes a significant gap in the context of phenotypic correlations between tumors of the salivary gland, lacrimal gland, and breast. Triple-negative breast carcinoma is generally regarded as a clinically aggressive group of malignancies without available targeted therapies, but in contrast to this and the behavior of adenoid cystic carcinoma in salivary and lacrimal gland, adenoid cystic carcinoma in the breast is a notable exception since recurrences and tumor-related deaths are exceptionally rare for these tumors [7, 8]. In addition to having morphology in common, the genetic landscape is highly similar in adenoid cystic carcinoma regardless of site, including pattern of copy-number alterations, low mutational burden, and the MYB-NFIB gene fusion [9,10,11,12,13,14,15,16]. This phenomenon of similarities in morphology and genetics but discrepancy in clinical behavior has earned adenoid cystic carcinoma the title as the Dr. Jekyll and Mr. Hyde of exocrine gland carcinomas [3].
The occurrence of salivary-like carcinomas in the lacrimal gland and breast is well known and includes several entities besides adenoid cystic carcinoma, namely, acinic cell carcinoma, mucoepidermoid carcinoma, secretory carcinoma, ductal carcinoma, and other more rare types, which collectively is highly suggestive of a common pathogenesis irrespective of site [5, 17,18,19]. Indeed, it is firmly established that histologically identical tumors are genetically similar in all three sites and that, despite the rarity of some type of salivary-like breast carcinomas, the aggressiveness is largely similar, with the notable exception of adenoid cystic carcinoma [5, 9,10,11, 15, 19,20,21].
Despite the inadequacy of genetic studies to explain the differences in clinical course of adenoid cystic carcinoma according to site, intrinsic differences in tumor biology remain the most likely explanations. Epigenetic mechanisms have been shown to be potent regulators of gene expression and involved in numerous malignancies of different organs [22, 23]. Among these, microRNAs (miRNAs) are short non-coding RNAs ∼22 nucleotides in length, functioning as potent post-transcriptional regulators of gene expression by causing either degradation or translational blockage of target mRNAs [24, 25]. MicroRNAs have been demonstrated as prognostic markers in salivary gland adenoid cystic carcinoma, whereas the involvement in lacrimal gland and breast adenoid cystic carcinoma is unknown [26, 27].
Searching for molecular mechanisms to explain the differences in clinical behavior between adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast, we characterize their phenotypic profile, distribution of genetic rearrangements, and global miRNA expression profile. For the first time, these findings shed light on the molecular background for this discordance in clinical behavior.
Materials and methods
Patient material
The Biospecimen Reporting for Improved Study Quality (BRISQ) was used as a guideline for this study [28]. Patients diagnosed with adenoid cystic carcinoma of the salivary gland, lacrimal gland, or breast were identified by searching the Danish Pathology Registry and the Eye Pathology Institute files [29]. Material was fixed in 10% neutral-buffered formalin immediately following surgery and subsequently embedded in paraffin. Material from 64 salivary gland adenoid cystic carcinomas (13 from parotid, 28 from submandibular, 3 from sublingual, nine from sinonasal tract, and 11 from oral, pharyngeal, and laryngeal minor salivary/ glands), nine lacrimal gland adenoid cystic carcinomas, and 11 breast adenoid cystic carcinomas stored at room temperature were sectioned and reviewed for diagnostic accuracy [30]. Sinonasal carcinomas were tested for presence of human papillomavirus (HPV) to exclude HPV-related multiphenotypic carcinoma and palatal carcinomas and tested for PRKD1 c.2130 mutation to exclude polymorphous adenocarcinoma, respectively [31, 32]. Adjacent normal gland from 10, seven, and six salivary gland, breast, and lacrimal gland adenoid cystic carcinomas, respectively, were included for miRNA expression profiling. Clinicopathologic details and outcomes were collected from patient files and from the Danish Patient Registry, the Danish Cancer Registry, and the Danish Pathology Registry, as previously described [1]. Staging was performed according to the American Joint Committee on Cancer (AJCC), 8th edition [33]. The Regional Ethics Committee (H-6-2014-086) and the Danish Data Protection Agency (Journal no. REG-94-2014) approved this protocol.
Histology, immunohistochemistry, phenotypic profiling, and survival statistics
From all cases, 4-μm-thick formalin-fixed and paraffin-embedded sections were cut and stained with hematoxylin and eosin (H&E) according to standard protocols, or subjected to immunohistochemical staining on the Ventana Benchmark Ultra platform (Ventana Medical Systems, Tucson, AZ) as previously described [34]. Primary antibodies included androgen receptor (clone AR441, 1:400, Dako (Glostrup, Denmark)), CK5/6 (clone D5/16, 1:20, Dako), EGFR (clone E30, 1:200, Dako), estrogen receptor (clone SP1, ready-to-use (RTU), Roche (Basel, Switzerland)), HER2 (clone 4B5, RTU, Roche), ki-67 (clone MIB1, 1:100, Dako), MYB (clone ER769Y, 1:150, AbCam (Cambridge, UK)), and progesterone receptor (clone 1E2, RTU, Roche). Positive controls as suggested on datasheets were used on each slide, and the expected reaction was confirmed. Negative controls were included on all slides. HER2 and EGFR was evaluated according to ASCO guidelines [35]. MYB positivity was defined as nuclear reaction in at least 5% of tumor cells [36]. CK 5/6 was regarded as positive when >10% of cells were positive while androgen receptor, estrogen receptor, and progesterone receptor were regarded as positive when Allred score was >2 as previously described [37, 38]. The HER2 phenotype was defined as membranous HER2 positivity of 3+ or 2+ with concurrent HER2 gene amplification regardless of other markers. Luminal phenotype was defined as HER2-negativity but with expression of estrogen receptor and/or progesterone receptor. Luminal androgen receptor-positive was defined as negativity for HER2, estrogen receptor, and progesterone receptor and positivity for androgen receptor. Basal-like phenotype was defined as negativity for HER2, estrogen receptor, progesterone receptor, and androgen receptor along with either 1) EGFR at 3+ or 2+ with concurrent EGFR gene amplification and/or 2) positivity for CK5/6. Cases negative for all markers were categorized as indeterminate [6, 39]. Nottingham grade was determined for breast carcinomas and architectural grade as described by Ro et al. on H&E-stained sections [40, 41]. Comparison of overall survival between basal-like and indeterminate phenotypes was performed using the Kaplan–Meier method and compared by the log-rank test in SPSS v.22.0 (SPSS, Chicago, IL) with a p value of ≤0.05 considered significant.
Fluorescence in situ hybridization
The 4-μm-thick formalin-fixed and paraffin-embedded sections from each case were placed onto positively charged slides. Hematoxylin and eosin stained slides were examined for determination of areas for counting. Fluorescence in situ hybridization (FISH) on all salivary gland cases and lacrimal gland and breast cases was performed using break-apart probes for MYB (ZytoVision GmbH, Bremerhaven, Germany), MYBL1 (custom probe oligonucleotides chr8:67076230-67474559 and chr8:67526335-68426199 (Agilent Technologies, Santa Clara, CA)/[Empire Genomics, Buffalo, NY]), and NFIB (custom probe oligonucleotides chr9:13740671-14140560 and chr9:14340306-14740560 (Agilent Technologies)/(Empire Genomics)) and an MYB-NFIB fusion probe (custom probe oligonucleotides chr6:135271234-135771043 and chr9:13990266-14490285 (Agilent Technologies)/(Cytotest, Rockville, MD), according to the manufacturers’ protocol as previously described [20, 31]. Nuclei were counterstained with 4′,6-diamidino-2-phenylindole (DAPI) II (ZytoVision). Break-apart signal in ≥10% of cells was considered to represent rearrangement and fused signals in ≥20% of cells was considered to represent fusion [42]. Amplification was defined as ≥3 signals/cell in ≥10% of cells. In case of equivocal (2+) EGFR immunohistochemistry, the EGFR (ZytoVision) numeric probe was employed according to ASCO guidelines as previously described [5, 35]. One hundred nuclei were counted, and only nuclei where the entire nuclear membrane could be visualized were scored.
RNA extraction
To maximize tumor content, a one 1 mm core was obtained from each tumor after identification of a representative tumor area from an H&E slide. The QiaCube (Qiagen, Valencia, CA) was used for automated isolation of miRNA with the miRNeasy FFPE Kit (Qiagen) according to the manufacturer’s instructions [43]. Total RNA concentration was measured using the NanoDrop ND-1000 (Thermo Scientific, Wilmington, DE), and ranged from 294 to 1648 ng/μl with A260/A280 ratio ranging from 1.7 to 2.7, indicating high nucleic acid purity. RNA quality, as estimated by RNA integrity number (RIN), was measured on the 10 oldest samples using the Agilent 2100 Bioanalyzer platform (Agilent Technologies, Santa Clara, CA) using the Agilent RNA 6000 Nano Kit (Agilent) with RIN ranging from 0.9 to 1.6 (median 1.1).
MicroRNA array
The Affymetrix miRNA 4.1 array platform (Affymetrix, Santa Clara, CA) was used, covering all entries in Sanger miRbase database v.20, including 2,578 mature and 2,025 immature human miRNAs with a dynamic range of 4 logs. For miRNA analysis, 300 μg of total RNA was labeled with the FlashTag Biotin HSR RNA Labeling kit (Affymetrix) according to the manufacturer’s instructions. To eliminate batch variation, all cases were run in a single batch. Array plates were washed, stained, and scanned on the GeneTitan Instrument (Affymetrix).
MicroRNA data analysis
Raw global miRNA expression data were normalized by using the quantile normalization method implemented in the free R package preprocessCore [44]. The normalized data were then log-transformed with base 2 to ensure normal or approximately normal distribution before statistical analysis. For each miRNA, linear regression models were fitted, regressing the log-transformed miRNA expression level on the tumor indicator variable (tumor = 1, normal = 0) for each of the three tissue types, or on tumor type variable (salivary, lacrimal, breast, with breast as reference). Correction for multiple testing was done by estimating the Benjamini–Hochberg adjusted p values (false discovery rate, FDR). MiRNAs with FDR ≤ 0.05 were considered to be statistically significant. All statistical analysis were performed using the free R software (https://cran.r-project.org/).
Pathway analysis and functional annotation
Pathway analysis of miRNAs was performed with DIANA-miRPath v3.0 software available from http://snf-515788.vm.okeanos.grnet.gr/. [45] The DIANA-TarBase v7.0 was selected as miRNA target prediction algorithm [46]. For functional annotation, the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were used and enrichment score for KEGG pathways presented by –ln (p value) [47].
Results
Patient demographics and clinicopathologic characteristics
Stage at diagnosis in salivary gland adenoid cystic carcinoma ranged from I to IV, and all lacrimal gland cases presented with N0M0 disease with T-stage ranging from 2a to 4c (Table 1). In the breast, all cases were stage I or II, except in one patient with stage III (case 7) due to multiple ipsilateral axillary lymph node metastases (pN2a). Radical surgery was performed in the majority of patients with breast adenoid cystic carcinoma, while this was the case in 23% and 0% of salivary gland and lacrimal gland adenoid cystic carcinoma, respectively. Median follow-up was 81, 92, and 123 months for breast, salivary gland, and lacrimal gland cases, respectively (Table 1). Perineural invasion (PNI) was identified in 30%, 85%, and 100% of breast, salivary gland, and lacrimal gland carcinomas, respectively. In the salivary gland, 9/64 (14%) experienced local recurrence, 16/64 (25%) distant metastases, and 27/64 (42%) died of disease. In patients with lacrimal gland adenoid cystic carcinoma, 5/9 (56%) experienced each of these events. None of the breast carcinomas recurred.
Histology and phenotypic profiling
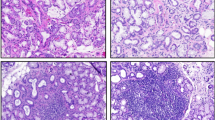
Irrespective of site, all tumors displayed the histological hallmark features of adenoid cystic carcinoma with combinations of tubular, cribriform, and solid areas (Fig. 1a–c). Architectural grade ranged from I to III at all three sites, with grade III (i.e., “solid histology”) in 1/11 (9%), 7/62 (11%), and 2/9 (22%) of breast, salivary gland, and lacrimal gland tumors, respectively (Table 1). Immunohistochemistry identified dual populations of luminal and abluminal cells in varying proportions in tumors at all three sites (Fig. 1d–f). The results of phenotypic profiling are summarized in Table 1 and further specified in Table S1. The majority of cases at all sites were CK5/6 positive, most intensely in the luminal cells (Fig. 1d, e). Androgen receptor and progesterone receptor were consistently negative, while 1/64 (2%) carcinomas from the salivary gland expressed estrogen receptor (Fig. 2a–c). EGFR was positive (3+) in 6/64 (9%) of salivary gland tumors and in none of the lacrimal gland or breast tumors (Fig. 2d). All EGFR 2+ cases (five in the salivary gland) had intact EGFR copy number by FISH (Fig. 2d, insert). HER2 was consistently negative (Fig. 2e, f). This categorized 55/64 (86%) of salivary gland adenoid cystic carcinomas as basal-like, 8/64 (13%) as indeterminate, and 1/64 (2%) as luminal type due to estrogen receptor expression (Case 32, Fig. 2b insert). There was no difference in overall survival between basal-like and indeterminate phenotypes (p = 0.59) (Figure S2). All lacrimal gland and breast adenoid cystic carcinomas were basal-like.
Histological features of fusion-positive adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast. a Adenoid cystic carcinoma of the submandibular gland with tubular and cribriform areas with accumulation of basal membrane material in pseudoluminae. Normal ducts and acinar cells are seen in the upper right corner (asterisk). b Adenoid cystic carcinoma of the lacrimal gland with normal ducts and acinar cells in the lower left corner (asterisk) and c adenoid cystic carcinoma of the breast surrounding a normal duct (asterisk). d Cytokeratin 5/6 in salivary gland and e breast adenoid cystic carcinoma with intense cytoplasmic reaction in luminal cells, and f nuclear p63 reaction in abluminal cells lining pseudoluminae
Phenotypic profiling of adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast. a Androgen receptor was uniformly negative in adenoid cystic carcinoma irrespective of site (a salivary gland specimen is shown). b Estrogen receptor was negative in all cases irrespective of site (a breast specimen is shown), with the exception of one salivary gland case (inset) with intense expression in 60% of tumor cells. c Progesterone receptor was uniformly negative in tumors of all three sites (a salivary gland specimen is shown). d Membranous EGFR expression (3+) was identified exclusively in a subset of salivary gland carcinomas, and (inset) all cases with 2+ scores showed intact EGFR copy number status with fluorescent in situ hybridization. e HER2 was consistently negative in adenoid cystic carcinoma at all three sites, with a salivary gland (f) and breast specimen shown
MYB, MYBL1, and NFIB gene status in adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast
Concurrent rearrangement of MYB and NFIB was the most frequent finding in tumors at all sites, with MYB–NFIB fusion found in all of these (Table 2, Fig. 3a–c). The second most frequent finding was concurrent rearrangement of MYBL1 and NFIB, which was seen exclusively in salivary gland adenoid cystic carcinoma (Table 2, Fig. 3d). More rare events included MYB amplification and isolated rearrangements of MYB (MYB-X), NFIB (NFIB-X), and MYBL1 (MYBL1-X) (Table 2, Fig. 3e, g). There was no difference in overall survival between patients with MYB and MYBL1 involvement (p = 0.92) (Figure S3). Among non-salivary sites, one lacrimal gland case had MYB amplification and two breast cases had isolated NFIB rearrangement (NFIB-X). All cases without detectable genetic abnormalities were intensely positive for MYB protein (Fig. 3f, Table 2).
Genetic characterization of adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast. a Fluorescent in situ hybridization using break-apart probes for MYB and b NFIB showing separate red (arrows) and green (arrowheads) signals, indicating rearrangement in a breast adenoid cystic carcinoma. c The fusion probe demonstrates the presence of the MYB–NFIB fusion in the same specimen. d Break-apart probe demonstrating MYBL1 rearrangement that was identified exclusively in a subset of salivary gland carcinomas. e In a subset of non-rearranged cases, numerous green signals were found which is consistent with amplification of the 5′ part of the MYB gene. f In the non-rearranged carcinomas, all cases overexpressed MYB protein. g Overview of the genetic spectrum in salivary gland, lacrimal gland, and breast adenoid cystic carcinoma. MYB and NFIB involvement was seen in adenoid cystic carcinoma of all three organs, while MYBL1 was exclusively found in the salivary gland
MicroRNA expression profiling of adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast
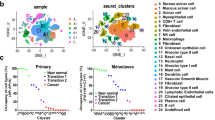
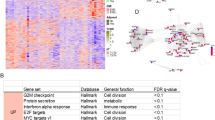
After establishing morphological and genetic similarities between adenoid cystic carcinoma irrespective of site, the miRNA expression profiles of the tumor and normal tissue were compared. This identified 22 upregulated and 25 downregulated miRNAs in salivary gland adenoid cystic carcinoma (Table S2), and eight upregulated miRNAs in lacrimal gland adenoid cystic carcinoma. Among these differentially expressed miRNAs, hsa-miR-455-5p and hsa-miR-181d-5p were upregulated in carcinomas of both salivary and lacrimal gland (Fig. 4, Table S2, and Table S3). In contrast, adenoid cystic carcinoma of the breast did not separate from normal breast until increasing FDR to <0.27 (Fig. 4, Table S4). Comparing adenoid cystic carcinoma from the salivary gland and breast identified 31 upregulated and 62 downregulated miRNAs (Table 3). Hierarchical clustering identified two main clusters, with the largest cluster being composed exclusively of salivary gland carcinomas (51/64, 80%) (Fig. 5). The other main cluster divided into two subclusters, with one being composed exclusively of 8 breast adenoid cystic carcinomas (8/11, 73%) and the other harboring all 9 (9/9, 100%) lacrimal gland carcinomas, plus the remaining 13 (13/64, 20%) salivary gland carcinomas and 3 (3/11, 27%) breast carcinomas (Fig. 5). Among the miRNAs identified by hierarchical clustering as being differentially expressed between salivary gland and breast adenoid cystic carcinoma, none were differentially expressed between the corresponding normal glands, excluding tissue-specific differences as the cause for the differences found. Functional annotation of miRNAs differentially expressed between salivary gland and breast adenoid cystic carcinoma identified their involvement in several different biological processes, including the Kyoto Encyclopedia of Genes and Genomes (KEGG) categories: metabolism, signal transduction, and cancers (Table S5).
MicroRNA expression profiling separates most breast from salivary gland adenoid cystic carcinoma irrespective of phenotypic profile and genetic status. Unsupervised hierarchical clustering of adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast clustered the majority of salivary gland carcinomas (51/64) and breast carcinomas (8/11) at each end of the spectrum. All lacrimal gland cases (9/9) clustered between the pure salivary gland and breast clusters. The lacrimal gland cluster included the remaining subset of salivary gland (11/64) and breast carcinomas (3/11). Clustering of cases occurred irrespective of architectural grade (solid), phenotypic profile, and gene status
Discussion
While being by far most common in the salivary gland, adenoid cystic carcinoma is found at other sites including the lacrimal gland, lung, breast, and skin [9, 10, 48]. Adenoid cystic carcinoma of the salivary gland, lacrimal gland, and lung has a protracted but relentless clinical course, contrasting with cutaneous and breast adenoid cystic carcinoma which are indolent neoplasms [1, 3, 9, 48]. The contrast in clinical outcome between adenoid cystic carcinoma of the salivary and lacrimal gland on one side and breast on the other is confirmed in the present series (Table 1). While it could be speculated that this paradox is simply due to a morphological overlap of different entities, combined immunohistochemical phenotyping and pattern of genetic rearrangements uniformly rejects this theory. Hence, the genetic cause of the majority of adenoid cystic carcinoma irrespective of site is unquestioned, but does not explain the site-dependent difference in clinical behavior. Through recent years, several studies on the genetics of especially salivary gland adenoid cystic carcinoma have demonstrated a low mutational burden as compared to other cancers [12, 14, 49,50,51,52]. However, none of these have been shown to be independent prognostic factors, nor do they apply to the majority of patients with this malignancy.
The indolent nature of breast adenoid cystic carcinoma not only stand out in this type of malignancy across different anatomical sites but also among triple-negative and basal-like breast carcinomas, which are otherwise known to be associated with an aggressive clinical course [4, 7]. In the breast, phenotypic classification places adenoid cystic carcinoma in the basal-like and triple-negative categories, and gene expression profiling is in accordance with adenoid cystic carcinoma being similar to other basal-like carcinomas [15, 53, 54]. Our results are in agreement with the exclusively basal-like and triple-negative phenotype of breast adenoid cystic carcinoma, and this finding also applied to lacrimal gland tumors. However, salivary gland tumors were triple negative in all cases, but despite being basal-like in the majority of these, a substantial proportion was negative for all markers and thereby of indeterminate phenotype (Fig. 5, Table S4). In the breast, indeterminate phenotypes are more aggressive than basal-like phenotypes, but this was not the case between these two groups of salivary gland adenoid cystic carcinomas (Figure S1) [4]. Hence, the significance of phenotypic profiling in salivary gland tumors, if any, is not straightforward and deserves further investigation.
While gene fusions in salivary gland adenoid cystic carcinoma have been studied in great detail, the fusion status of much fewer cases from the lacrimal gland and breast have been reported [9, 10, 15, 55, 56]. In agreement with previous studies, we identified a spectrum of genetic abnormalities in salivary gland carcinomas, including two cases with isolated MYB rearrangement of which one was estrogen receptor-positive [57]. To the best of our knowledge, an estrogen receptor-positive salivary gland adenoid cystic carcinoma has not previously been described. In contrast, a much narrower selection of genetic rearrangements was found in lacrimal gland and breast adenoid cystic carcinomas. Lacrimal gland abnormalities were restricted to MYB–NFIB fusions and MYB amplification, while a somewhat larger proportion of breast adenoid cystic carcinomas had normal MYB and NFIB genes or had isolated NFIB rearrangement (NFIB-X). To our knowledge, isolated NFIB rearrangement have not previously been described in breast adenoid cystic carcinoma [10, 15, 56]. Also, MYBL1 rearrangements seem to be unique to salivary gland adenoid cystic carcinoma, and, in agreement with the literature, we did not find a difference in overall survival between cases with MYB and MYBL1 involvement (Figure S2) [58]. An intriguing observation is that, among the different genetic aberrations identified, NFIB was the most consistently involved (Figs. 3 and 5). The function of NFIB is poorly characterized but is crucial for salivary gland embryonic development and paradoxically serves as a metastatic driver in some cancers while having a tumor suppressor function others [59,60,61,62,63]. It is interesting that the salivary gland and breast are the two tissues in which NFIB is expressed in the highest level among a large selection of human tissues in the Genotype-Tissue Expression project (GTEx) (Figure S3) [64]. Although shown to be involved in other types of salivary gland malignancies, further clarification of NFIB function in these organs is warranted [65].
Post-transcriptional regulation of gene expression by miRNAs is known to be involved in a number of human malignancies, but the two previous studies comparing miRNA expressions in salivary gland adenoid cystic carcinoma and normal salivary gland have identified widely different patterns of miRNA dysregulation [26, 66, 67]. Also, only little agreement is found between these studies and our findings, and the general discrepancy in miRNA expression between studies is well known [68]. To improve transparency, we conducted this study according to BRISQ guidelines and sought to minimize the influence of external factors by sampling pure tumor tissue, assuring the quality of extracted miRNA, use of a comprehensive and validated array, and inclusion of a large number of samples. The fact that miRNA expression could not distinguish between breast adenoid cystic carcinoma and normal breast tissue, while pronounced dysregulation was found between tumor tissue and normal tissue from both salivary and lacrimal gland, goes well in hand with the differences in malignant potential (Fig. 4). Indeed, despite characterizing only nine cases from the lacrimal gland, an overlap in tumor-specific miRNA expression was found between lacrimal and salivary gland adenoid cystic carcinoma. These two miRNAs (hsa-miR-455-5p, hsa-miR-181d-5p) have been repeatedly found to be overexpressed in other malignancies [69,70,71]. In unsupervised hierarchical clustering, global miRNA expression profiling separated salivary gland from breast adenoid cystic carcinoma with high accuracy and placed lacrimal gland tumors in between these two (Fig. 5). Importantly, this separation was not caused by tissue-specific expression but by differences between the tumors, and the plausibility of these differentially expressed miRNAs as being responsible for the difference in clinical behavior is strengthened by their involvement in the regulation of genes in several KEGG orthology categories frequently involved in cancer (Table S5). In addition to being the first class of molecules able to separate adenoid cystic carcinoma according to site of origin, this lends support to miRNA as having a role in the clinical behavior of these tumors. The mechanism for this miRNA dysregulation is currently unknown, as none of these are located to chromosomal regions involved in recurrent copy number alterations in adenoid cystic carcinoma, and needs to be further studied (Table 3) [72].
In conclusion, we present a comparative study of adenoid cystic carcinoma of the salivary gland, lacrimal gland, and breast and corroborate the histological and genetic similarity between these lesions despite the large variations in clinical behavior. We show that global miRNA expression profiling separates these tumors according to site of origin, and that differentially expressed miRNAs are involved in regulation of cellular processes that could be responsible for a more aggressive clinical course in salivary gland adenoid cystic carcinoma as compared to those in the breast. These findings are helpful in directing studies for prognostication, intensity of treatment and monitoring of patients with salivary gland adenoid cystic carcinoma.
References
Bjørndal K, Krogdahl A, Therkildsen MH, et al. Salivary adenoid cystic carcinoma in Denmark 1990–2005: outcome and independent prognostic factors including the benefit of radiotherapy. Results of the Danish Head and Neck Cancer Group (DAHANCA). Oral Oncol. 2015;51:1138–42.
Andreoli MT, Aakalu V, Setabutr P. Epidemiological trends in malignant lacrimal gland tumors. Otolaryngol Head Neck Surg. 2015;152:279–83.
Marchiò C, Weigelt B, Reis-Filho JS. Adenoid cystic carcinomas of the breast and salivary glands (or “The strange case of Dr Jekyll and Mr Hyde” of exocrine gland carcinomas). J Clin Pathol. 2010;63:220–28.
Voduc KD, Cheang MC, Tyldesley S, et al. Breast cancer subtypes and the risk of local and regional relapse. J Clin Oncol. 2010;28:1684–91.
Andreasen S, Grauslund M, Heegaard S. Lacrimal gland ductal carcinomas: clinical, morphological and genetic characterization and implications for targeted treatment. Acta Ophthalmol. 2017;95:299–6.
Di Palma S, Simpson RH, Marchió C, et al. Salivary duct carcinomas can be classified into luminal androgen receptor-positive, HER2 and basal-like phenotypes. Histopathology. 2012;61:629–43.
Badve S, Dabbs DJ, Schnitt SJ, et al. Basal-like and triple-negative breast cancers: a critical review with an emphasis on the implications for pathologists and oncologists. Mod Pathol. 2011;24:157–67.
Miyai K, Schwartz MR, Divatia MK, et al. Adenoid cystic carcinoma of breast: recent advances. World J Clin Cases. 2014;2:732–41.
von Holstein SL, Fehr A, Persson M, et al. Adenoid cystic carcinoma of the lacrimal gland: MYB gene activation, genomic imbalances, and clinical characteristics. Ophthalmology. 2013;120:2130–38.
Martelotto LG, De Filippo MR, Ng CK, et al. Genomic landscape of adenoid cystic carcinoma of the breast. J Pathol. 2015;237:179–89.
Persson M, Andrén Y, Mark J, et al. Recurrent fusion of MYB and NFIB transcription factor genes in carcinomas of the breast and head and neck. Proc Natl Acad Sci USA. 2009;106:18740–44.
Stephens PJ, Davies HR, Mitani Y, et al. Whole exome sequencing of adenoid cystic carcinoma. J Clin Invest. 2013;123:2965–68.
Persson M, Andrén Y, Moskaluk CA, et al. Clinically significant copy number alterations and complex rearrangements of MYB and NFIB in head and neck adenoid cystic carcinoma. Genes Chromosomes Cancer. 2012;51:805–17.
Ho AS, Kannan K, Roy DM, et al. The mutational landscape of adenoid cystic carcinoma. Nat Genet. 2013;45:791–98.
Wetterskog D, Lopez-Garcia MA, Lambros MB, et al. Adenoid cystic carcinomas constitute a genomically distinct subgroup of triple-negative and basal-like breast cancers. J Pathol. 2012;226:84–96.
Horlings HM, Weigelt B, Anderson EM, et al. Genomic profiling of histological special types of breast cancer. Breast Cancer Res Treat. 2013;142:257–69.
Andreasen S, Esmaeli B, von Holstein SL, et al. An update on tumors of the lacrimal gland. Asia-Pac J Ophthalmol. 2017;6:159–72.
Pia-Foschini M, Reis-Filho JS, Eusebi V, et al. Salivary gland-like tumours of the breast: surgical and molecular pathology. J Clin Pathol. 2003;56:497–6.
Dalin MG, Desrichard A, Katabi N, et al. Comprehensive molecular characterization of salivary duct carcinoma reveals actionable targets and similarity to apocrine breast cancer. Clin Cancer Res. 2016;22:4623–33.
Skálová A, Vanecek T, Sima R, et al. Mammary analogue secretory carcinoma of salivary glands, containing the ETV6-NTRK3 fusion gene: a hitherto undescribed salivary gland tumor entity. Am J Surg Pathol. 2010;34:599–8.
Krings G, Joseph NM, Bean GR, et al. Genomic profiling of breast secretory secretory carcinomas. Mod Pathol. 2017;30:1086–99.
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–38.
Iorio MV, Croce CM. microRNA involvement in human cancer. Carcinogenesis. 2012;33:1126–33.
Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet. 2008;9:102–14.
Ul Hussain M. Micro-RNAs (miRNAs): genomic organisation, biogenesis and mode of action. Cell Tissue Res. 2012;349:405–13.
Mitani Y, Roberts DB, Fatani H, et al. MicroRNA profiling of salivary adenoid cystic carcinoma: association of miR-17-92 upregulation with poor outcome. PLoS One. 2013;8:e66778.
Kiss O, Tőkés AM, Spisák S, et al. Breast- and salivary gland-derived adenoid cystic carcinomas: potential post-transcriptional divergencies. A pilot study based on miRNA expression profiling of four cases and review of the potential relevance of the findings. Pathol Oncol Res. 2015;21:29–4.
Moore HM, Kelly AB, Jewell SD, et al. Biospecimen reporting for improved study quality (BRISQ). J Proteome Res. 2011;10:3429–38.
Erichsen R, Lash TL, Hamilton-Dutoit SJ, et al. Existing data sources for clinical epidemiology: The Danish National Pathology Registry and Data Bank. Clin Epidemiol. 2010;2:51–56.
Andreasen S, Bjørndal K, Agander TK, et al. Tumors of the sublingual gland: a national clinicopathologic study of 29 cases. Eur Arch Otorhinolaryngol. 2016;273:3847–56.
Andreasen S, Bishop JA, Hansen TV, et al. Human papillomavirus-related carcinoma with adenoid cystic-like features of the sinonasal tract: clinical and morphological characterization of 6 new cases. Histopathology. 2017;70:880–88.
Andreasen S, Melchior LC, Kiss K, et al. The PRKD1 E710D hotspot mutation is highly specific in separating polymorphous adenocarcinoma of the palate from adenoid cystic carcinoma and pleomorphic adenoma on FNA. Cancer. 2017 [Epub ahead of print] doi:10.1002/cncy.21959
Amin MB, Edge SB, Greene FL et al. AJCC cancer staging manual. 8th ed. New York: Springer-Verlag; 2017.
Andreasen S, Therkildsen MH, Grauslund M, et al. Activation of the interleukin-6/Janus kinase/STAT3 pathway in pleomorphic adenoma of the parotid gland. APMIS. 2015;123:706–15.
Wolff AC, Hammond ME, Hicks DG, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol. 2013;31:3997–13.
Brill LB 2nd, Kanner WA, Fehr A, et al. Analysis of MYB expression and MYB-NFIB gene fusions in adenoid cystic carcinoma and other salivary neoplasms. Mod Pathol. 2011;24:1169–76.
Harvey JM, Clark GM, Osborne CK, et al. Estrogen receptor status by immunohistochemistry is superior to the ligand-binding assay for predicting response to adjuvant endocrine therapy in breast cancer. J Clin Oncol. 1999;17:1474–81.
Tan DS, Marchió C, Jones RL, et al. Triple negative breast cancer: molecular profiling and prognostic impact in adjuvant anthracycline-treated patients. Breast Cancer Res Treat. 2008;111:27–44.
Nielsen TO, Hsu FD, Jensen K, et al. Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res. 2004;10:5367–74.
Elston C, Ellis I. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer. Experience from a large study with long-term follow-up. Histopathology. 1991;19:403–10.
Ro JY, Silva EG, Gallager HS. Adenoid cystic carcinoma of the breast. Hum Pathol. 1987;18:1276–81.
Hudson JB, Collins BT. MYB gene abnormalities t(6;9) in adenoid cystic carcinoma fine-needle aspiration biopsy using fluorescence in situ hybridization. Arch Pathol Lab Med. 2014;138:403–9.
Howe K. Extraction of miRNAs from formalin-fixed paraffin-embedded (FFPE) tissues. Methods Mol Biol. 2017;1509:17–24.
Bolstad BM, Irizarry RA, Astrand M, et al. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19:185–93.
Vlachos IS, Zagganas K, Paraskevopoulou MD, et al. DIANA-miRPathv3.0: deciphering microRNA function with experimental support. Nucleic Acids Res. 2015;43:W460–66.
Vlachos IS, Paraskevopoulou MD, Karagkouni D, et al. DIANA-TarBasev7.0: Indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res. 2015;43:D153–59.
Ogata H, Goto S, Sato K, et al. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 1999;27:29–34.
Li N, Xu L, Zhao H, et al. A comparison of the demographics, clinical features, and survival of patients with adenoid cystic carcinoma of major and minor salivary glands versus less common sites within the Surveillance, Epidemiology, and End Results registry. Cancer. 2012;118:3945–53.
Liu B, Mitani Y, Rao X, et al. Spatio-temporal genomic heterogeneity, phylogeny, and metastatic evolution in salivary adenoid cystic carcinoma. J Natl Cancer Inst. 2017;109:jx033. https://doi.org/10.1093/jnci/djx033.
Rettig EM, Talbot CC Jr, Sausen M, et al. Whole-genome sequencing of salivary gland adenoid cystic carcinoma. Cancer Prev Res. 2016;9:265–74.
Kato S, Elkin SK, Schwaederle M, et al. Genomic landscape of salivary gland tumors. Oncotarget. 2015;6:25631–45.
Ross JS, Wang K, Rand JV, et al. Comprehensive genomic profiling of relapsed and metastatic adenoid cystic carcinomas by next-generation sequencing reveals potential new routes to targeted therapies. Am J Surg Pathol. 2014;38:235–38.
Weigelt B, Horlings HM, Kreike B, et al. Refinement of breast cancer classification bymolecular characterization of histological special types. J Pathol. 2008;216:141–50.
Shah SP, Roth A, Goya R, et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature. 2012;486:395–99.
Chen TY, Keeney MG, Chintakuntlawar AV, et al. Adenoid cystic carcinoma of the lacrimal gland is frequently characterized by MYB rearrangement. Eye. 2017;31:720–25.
Argani P, Ning Y, Cimino-Mathews A. MYB labeling by immunohistochemistry is more sensitive and specific for breast adenoid cystic carcinoma than MYB labeling by FISH. Am J Surg Pathol. 2017;41:973–79.
Fujii K, Murase T, Beppu S, et al. MYB, MYBL1, MYBL2, and NFIB gene alterations and MYC overexpression in salivary gland adenoid cystic carcinoma. Histopathology. 2017;71:823–34.
Mitani Y, Liu B, Rao PH, et al. Novel MYBL1 gene rearrangements with recurrent MYBL1-NFIB fusions in salivary adenoid cystic carcinomas lacking t(6;9) translocations. Clin Cancer Res. 2016;22:725–33.
Mellas RE, Kim H, Osinski J, et al. NFIB regulates embryonic development of submandibular glands. J Dent Res. 2015;94:312–19.
Semenova EA, Kwon MC, Monkhorst K, et al. Transcription factor NFIB is a driver of small cell lung cancer progression in mice and marks metastatic disease in patients. Cell Rep. 2016;16:631–43.
Fane ME, Chhabra Y, Hollingsworth DE, et al. NFIB mediates BRN2 driven melanoma cell migration and invasion through regulation of EZH2 and MITF. EBioMedicine. 2017;16:63–5.
Becker-Santos DD, Thu KL, English JC, et al. Developmental transcription factor NFIB is a putative target of oncofetal miRNAs and is associated with tumour aggressiveness in lung adenocarcinoma. J Pathol. 2016;240:161–72.
Stringer BW, Bunt J, Day BW, et al. Nuclear factor one B (NFIB) encodes a subtype-specific tumour suppressor in glioblastoma. Oncotarget. 2016;7:29306–20.
The GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45:580–85. https://gtexportal.org/home/gene/NFIB.
Andreasen S, Persson M, Kiss K, et al. Genomic profiling of a rare case of combined large-cell neuroendocrine carcinoma of the submandibular gland. Oncol Rep. 2016;35:2177–82.
Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10:704–14.
Gao R, Cao C, Zhang M, et al. A unifying gene signature for adenoid cystic cancer identifies parallel MYB-dependent and MYB-independent therapeutic targets. Oncotarget. 2014;5:12528–42.
Andorfer CA, Necela BM, Thompson EA, et al. MicroRNA signatures: clinical biomarkers for the diagnosis and treatment of breast cancer. Trends Mol Med. 2011;17:313–19.
Wong N, Khwaja SS, Baker CM, et al. Prognostic microRNA signatures derived from The Cancer Genome Atlas for head and neck squamous cell carcinomas. Cancer Med. 2016;5:1619–28.
Cheng CM, Shiah SG, Huang CC, et al. Up-regulation of miR-455-5p by the TGF-β–SMAD signalling axis promotes the proliferation of oral squamous cancer cells by targeting UBE2B. J Pathol. 2016;240:38–49.
Yan LX, Huang XF, Shao Q, et al. MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA. 2008;14:2348–60.
Zhang L, Mitani Y, Caulin C, et al. Detailed genome-wide SNP analysis of major salivary carcinomas localizes subtype-specific chromosome sites and oncogenes of potential clinical significance. Am J Pathol. 2013;182:2048–57.
Acknowledgements
We thank Sanni Pedersen, Heidi Ugleholdt, and Pernille Frederiksen for brilliant technical assistance. The study was funded by Einar Willumsen Foundation, Merchant Kristjan Kjær and wife Margrethe Kjær’s Foundation, The Danielsen Foundation, Resino A/S, Else and Mogens Wedell-Wedellsborg’s Foundation, Region Zealand’s Research Fund, Hans Skouby and wife Emma Skouby’s Foundation, and the A.P. Møller Foundation for the Advancement of Medical Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Andreasen, S., Tan, Q., Agander, T.K. et al. Adenoid cystic carcinomas of the salivary gland, lacrimal gland, and breast are morphologically and genetically similar but have distinct microRNA expression profiles. Mod Pathol 31, 1211–1225 (2018). https://doi.org/10.1038/s41379-018-0005-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-018-0005-y
This article is cited by
-
Dynamic Role of miRNAs in Salivary Gland Carcinomas: From Biomarkers to Therapeutic Targets
Head and Neck Pathology (2024)
-
The clinical behavior and genomic features of the so-called adenoid cystic carcinomas of the solid variant with basaloid features
Modern Pathology (2022)
-
Problematic breast tumors reassessed in light of novel molecular data
Modern Pathology (2021)
-
MLMDA: a machine learning approach to predict and validate MicroRNA–disease associations by integrating of heterogenous information sources
Journal of Translational Medicine (2019)
-
Prognostic significance of 1p36 locus deletion in adenoid cystic carcinoma of the salivary glands
Virchows Archiv (2018)