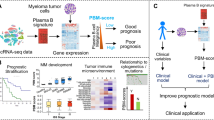
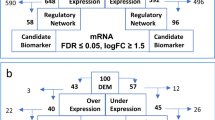
Abstract
Sequencing studies have shed some light on the pathogenesis of progression from smouldering multiple myeloma (SMM) and symptomatic multiple myeloma (MM). Given the scarcity of smouldering samples, little data are available to determine which translational programmes are dysregulated and whether the mechanisms of progression are uniform across the main molecular subgroups. In this work, we investigated 223 SMM and 1348 MM samples from the University of Arkansas for Medical Sciences (UAMS) for which we had gene expression profiling (GEP). Patients were analysed by TC-7 subgroup for gene expression changes between SMM and MM. Among the commonly dysregulated genes in each subgroup, PHF19 and EZH2 highlight the importance of the PRC2.1 complex. We show that subgroup specific differences exist even at the SMM stage of disease with different biological features driving progression within each TC molecular subgroup. These data suggest that MMSET SMM has already transformed, but that the other precursor diseases are distinct clinical entities from their symptomatic counterpart.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Misund K, Keane N, Stein CK, Asmann YW, Day G, Welsh S, et al. MYC dysregulation in the progression of multiple myeloma. Leukemia. 2020;34:322–6.
Bustoros M, Sklavenitis-Pistofidis R, Park J, Redd R, Zhitomirsky B, Dunford AJ, et al. Genomic profiling of smoldering multiple myeloma identifies patients at a high risk of disease progression. JCO. 2020;38:2380–9.
Boyle EM, Deshpande S, Tytarenko R, Ashby C, Wang Y, Bauer MA, et al. The molecular make up of smoldering myeloma highlights the evolutionary pathways leading to multiple myeloma. Nat Commun. 2021;12:293.
Weinhold N, Heuck C, Rosenthal A, Thanendrarajan S, Stein C, Van Rhee F, et al. The clinical value of molecular subtyping multiple myeloma using gene expression profiling. Leukemia. 2016;30:423–30.
Khan R, Dhodapkar M, Rosenthal A, Heuck C, Papanikolaou X, Qu P, et al. Four genes predict high risk of progression from smoldering to symptomatic multiple myeloma (SWOG S0120). Haematologica. 2015;100:1214–21.
Dhodapkar MV, Sexton R, Waheed S, Usmani S, Papanikolaou X, Nair B, et al. Clinical, genomic, and imaging predictors of myeloma progression from asymptomatic monoclonal gammopathies (SWOG S0120). Blood. 2014;123:78–85.
Stein CK, Pawlyn C, Chavan S, Rasche L, Weinhold N, Corken A, et al. The varied distribution and impact of RAS codon and other key DNA alterations across the translocation cyclin D subgroups in multiple myeloma. Oncotarget. 2017;8:27854–67.
Blighe K, Rana S, Lewis M. Enhanced Volcano: Publication-ready volcano plots with enhanced colouring and labeling. R package version; 1.8.0. https://github.com/kevinblighe/EnhancedVolcano.
Mason MJ, Schinke C, Eng CLP, Towfic F, Gruber F, Dervan A, et al. Multiple Myeloma DREAM challenge reveals epigenetic regulator PHF19 as marker of aggressive disease. Leukemia. 2020;34:1866–74.
Alaterre E, Raimbault S, Goldschmidt H, Bouhya S, Requirand G, Robert N, et al. CD24, CD27, CD36 and CD302 gene expression for outcome prediction in patients with multiple myeloma. Oncotarget. 2017;8:98931–44.
Franqui-Machin R, Hao M, Bai H, Gu Z, Zhan X, Habelhah H, et al. Destabilizing NEK2 overcomes resistance to proteasome inhibition in multiple myeloma. J Clin Invest. 2018;128:2877–93.
Mikulasova A, Ashby C, Tytarenko RG, Qu P, Rosenthal A, Dent JA, et al. Microhomology-mediated end joining drives complex rearrangements and overexpression of MYC and PVT1 in multiple myeloma. Haematologica. 2020;105:1055–66. https://doi.org/10.3324/haematol.2019.217927.
Wang Y, Ren F, Chen P, Liu S, Song Z, Ma X. Identification of a six-gene signature with prognostic value for patients with endometrial carcinoma. Cancer Med. 2018;7:5632–42.
Zhang Y, Manjunath M, Yan J, Baur BA, Zhang S, Roy S, et al. The cancer-associated genetic variant Rs3903072 modulates immune cells in the tumor microenvironment. Front Genet. 2019;10:754.
Cho SH, Goenka S, Henttinen T, Gudapati P, Reinikainen A, Eischen CM, et al. PARP-14, a member of the B aggressive lymphoma family, transduces survival signals in primary B cells. Blood. 2009;113:2416–25.
Acknowledgements
We thank all the patients and their families for their contributions to this study. BAW and GJM received grant support through a Translational Research Programme award from the Leukemia & Lymphoma Society (6602-20 and 6600-20).
Author information
Authors and Affiliations
Contributions
Designed the study: EMB, BAW, GJM, FVR. Analysed the data: EMB, AR, AH. Interpreted the data: EMB, BAW, FED, HG, YuW, GJM. Acquired the data: PF, MR, CA, MB, SKJ, CPW, YanW, CDS, ST, MZ, BB, MVD, FVR, GLM, FED. Wrote the manuscript: EMB, FED, BW. Reviewed the manuscript: all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Boyle, E.M., Rosenthal, A., Ghamlouch, H. et al. Plasma cells expression from smouldering myeloma to myeloma reveals the importance of the PRC2 complex, cell cycle progression, and the divergent evolutionary pathways within the different molecular subgroups. Leukemia 36, 591–595 (2022). https://doi.org/10.1038/s41375-021-01379-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-021-01379-y
This article is cited by
-
WDR26 and MTF2 are therapeutic targets in multiple myeloma
Journal of Hematology & Oncology (2021)
-
Insights into high-risk multiple myeloma from an analysis of the role of PHF19 in cancer
Journal of Experimental & Clinical Cancer Research (2021)