Abstract
Although the over-expression of angiogenic factors is reported in diffuse large B-cell lymphoma (DLBCL), the poor response to anti-VEGF drugs observed in clinical trials suggests that angiogenesis in these tumours might be driven by VEGF-independent pathways. We show that sphingosine kinase-1 (SPHK1), which generates the potent bioactive sphingolipid sphingosine-1-phosphate (S1P), is over-expressed in DLBCL. A meta-analysis of over 2000 cases revealed that genes correlated with SPHK1 mRNA expression in DLBCL were significantly enriched for tumour angiogenesis meta-signature genes; an effect evident in both major cell of origin (COO) and stromal subtypes. Moreover, we found that S1P induces angiogenic signalling and a gene expression programme that is present within the tumour vasculature of SPHK1-expressing DLBCL. Importantly, S1PR1 functional antagonists, including Siponimod, and the S1P neutralising antibody, Sphingomab, inhibited S1P signalling in DLBCL cells in vitro. Furthermore, Siponimod, also reduced angiogenesis and tumour growth in an S1P-producing mouse model of angiogenic DLBCL. Our data define a potential role for S1P signalling in driving an angiogenic gene expression programme in the tumour vasculature of DLBCL and suggest novel opportunities to target S1P-mediated angiogenesis in patients with DLBCL.
Similar content being viewed by others
Introduction
Despite the use of rituximab and chemotherapy (R-CHOP), the survival of patients with diffuse large B-cell lymphoma (DLBCL) remains poor with a significant proportion of patients being refractory or relapsing after therapy [1, 2]. Gene expression profiling has identified two clinically distinct DLBCL subgroups, the activated B cell (ABC) and germinal centre B cell (GCB) forms, characterised by different, therapeutically tractable, molecular abnormalities. Gene expression profiling has also identified tumour microenvironment-derived molecular signatures, including the Stromal-2 signature that is reported to be enriched for angiogenesis-associated genes [3].
Although there may be therapeutic benefits in targeting angiogenesis in DLBCL, the results of clinical trials of bevacizumab, a monoclonal antibody against VEGFA, have been disappointing. Although this drug was well tolerated when delivered as a single agent in relapsed DLBCL [4] or in combination with R-CHOP in first-line treatment [5], a phase III study of RA-CHOP (MAIN trial) was stopped due to increased cardiotoxicity in the absence of any significant impact on progression-free survival [6]. One explanation for the limited clinical efficacy of bevacizumab in DLBCL is that angiogenesis in DLBCL does not depend on VEGF; a contention supported by studies showing that VEGF expression does not correlate with micro-vessel density in DLBCL [7, 8].
S1P is a bioactive sphingolipid metabolite which promotes cell growth and survival and has been shown to be a potent inducer of angiogenesis [9]. SPHK1, one of the two isoenzymes responsible for the production of S1P, is over-expressed in different cancers [10, 11]. Secreted S1P is a ligand for a family of five G-protein-coupled S1P receptors (S1PR1-5). Signalling through S1PR1 and/or S1PR3 has been shown to stimulate DNA synthesis, chemotactic motility and tube formation in endothelial cells and to induce angiogenesis in vivo [12,13,14,15]. As with other potent bioactive mediators, S1P levels are tightly regulated and controlled by the balance between its generation and its degradation by S1P lyase and S1P phosphatases [16]. Although it has been suggested that SPHK1 might play a role in haematological malignancies [17], it’s role in DLBCL remains to be established [18]. Furthermore, the effects of S1P signalling on the DLBCL microenvironment, including its influence on the tumour vasculature, have not been explored.
In the present study, we have shown that the over-expression of SPHK1 correlates with an angiogenic transcriptional programme in DLBCL. We defined an endothelial cell transcriptional signature of S1P signalling and used this to show that the expression of S1P target genes in these cells was correlated with that of SPHK1 in primary DLBCL. Moreover, Siponimod, a small-molecule functional antagonist of S1PR1 [19], reversed S1P signalling and reduced angiogenesis and tumour growth in an S1P-producing mouse model of DLBCL. Our data suggest novel opportunities to target S1P-mediated angiogenesis in patients with DLBCL.
Materials and methods
Cells and tissues
Tonsils and DLBCL samples were obtained with informed consent and ethical approval (REC_RG_HBRC_12-071). DLBCL cases were reviewed by haematopathologists (ZR, YLH, UZ). Isolation of tonsillar germinal centre (GC) and blood-derived B cells was described before [20,21,22]. Endothelial cells (EC) were isolated from umbilical cords (HUVEC) under informed consent (REC_RG_HBRC_14-180) using collagenase treatment [23] and cultured in M199 media supplemented with 5% fetal bovine serum (FBS), 1% glutamine, 1% penicillin/streptomycin (ThermoFisher Scientific, Waltham, MA, USA) and 1% EC-growth supplement (Caltag Medsystems, Buckingham, UK) at 37 °C/5% CO2. HT, Karpas-442, OCI-LY1, OCI-LY7, SUDHL4, SUDHL5, SUDHL6 are EBV-negative GC-DLBCL lines, Farage is an EBV-positive GC-DLBCL line. OCI-LY3 and U2932 are EBV-negative ABC-DLBCL lines. Lines were from DSMZ (Braunschweig, Germany), OCI (Ontario, Canada) or ATCC (Manassas, VA, USA) and were cultured in RPMI1640 or IMDM (OCI-LY1, OCI-LY7) media (ThermoFisher Scientific) supplemented with 10% FBS and 1% penicillin/streptomycin.
Mouse xenografts and flow cytometry
3 × 106 SUDHL6 cells were injected subcutaneously into NSG mice (Charles River Laboratories, Wilmington, MA, USA). After 17 days (when tumour volume averaged 63 mm3) mice were randomised into two groups (each n = 4) and treated orally with either vehicle (0.1% DMSO in 10% 2-hydroxypropyl-β-cyclodextrin; Cayman Chemical, MI, USA) or 6 mg/kg Siponimod (Selleckchem.com, Munich, Germany) every 48 h. Mice were culled when average tumour volumes in control mice reached 400 mm3 (28 days). Organs were weighed, minced and incubated with Liberase DL/Liberase TL and DNASEI (Roche, Basel, Switzerland) [24]. Cell suspensions were labelled with mouse CD31 and CountBright absolute counting beads (Thermofisher Scientific) and analysed by flow cytometry on LSRII and FACS diva 8 (BD, Franklin Lakes, NJ, USA). Details of the other mouse models tested are in Supplementary Materials and Methods. All mouse experiments were done according to UK Home Office guidelines.
S1P measurements
For intracellular S1P measurements, cell pellets were snap-frozen in liquid nitrogen. For secreted S1P measurements, SUDHL4 cells were cultured in serum-free RPMI (without phenol-red) supplemented with 1% tissue-culture grade fatty acid-free BSA (Sigma-Aldrich., St Louis, MO, USA) for indicated times. Supernatants were harvested into pre-chilled HPLC grade methanol (Sigma-Aldrich) supplemented with Halt™ Protease and Phosphatase Inhibitor Cocktail (ThermoFisher Scientific). S1P levels were quantified by liquid chromatography-electrospray ionization-tandem mass spectrometry (LC-ESI-MS/MS; 4000 QTRAP, AB Sciex, Framingham, MA, USA) as previously described [25].
Treatment of cells
S1P (Sigma-Aldrich) was prepared as in Supplementary Materials and Methods and as before [26]. Prior to treatments, HUVEC were cultured in full media depleted of EC-growth supplement for 16 h and stimulated with 0.5 µM S1P (or control/vehicle) for 5 min to detect ERK1/2 activation or for 4 h to detect S1P transcriptional targets. For S1P inhibition, Sphingomab or isotype/control antibodies (LPath Inc., San Diego, CA, USA) were incubated with S1P (at 150 μg/ml per μM of S1P) for 1 h prior to S1P stimulation. For S1PR1 inhibition, HUVEC were treated with 100 nM S1PR1 functional antagonists Siponimod/BAF312, Ozanimod/RPC1063 or Ponesimod/ACT-128800 (Selleckchem.com) for 1 h prior to S1P stimulation.
Quantitative RT-PCR (Q-PCR)
RNA was extracted with RNeasy Mini kit including genomic DNA removal with RNase-Free DNase Set (Qiagen, Manchester, UK). cDNA was generated with qScript™ cDNA SuperMix (QuantaBio, Beverly, MA, USA). Gene transcripts were quantified with commercial gene expression assays (ThermoFisher Scientific; Supplementary Table 1a) [21]. The 2−ΔΔCT method was used to quantify target expression relative to housekeeping control. The normalized values are shown relative to the reference sample that was set to a relative quantity value of 1.
Protein analysis
Immunoblotting was by standard methodology [20]. Detection was with enhanced chemiluminescence (ECL; GE Healthcare, Chalfont St Giles, UK) on a ChemiDoc MP (Bio-Rad, Watford, UK). Immunohistochemistry (IHC) was on 4 µm FFPE sections and HUVEC grown on microscope slides, using either citrate or EDTA buffer antigen retrieval and 0.3% H2O2 and 5× casein blocking [26,27,28]. Slides were incubated either overnight at 4 °C or for 1 h at room temperature in primary antibodies in 0.05% PBS/Tween and visualised with species-specific ImmPRESS kits, followed by diaminobenzidine or ImmPACT NovaRED (Vector Laboratories, Peterborough, UK) with haematoxylin counterstain. DLBCL were stained for BCL6, CD10 and IRF4, and defined as either GCB or non-GCB type by Hans algorithm [29]. A subset of DLBCL was also stained for SPHK1, S1PR1, CXCL12, SELE, COL1A1 and MAP1B. Mouse tumours were stained for Ki67, cPARP and CXCL12. For CXCL12, a Mouse-on-Mouse block (Vector Laboratories) was performed prior to incubation in casein. cPARP- and Ki67- positive cells were enumerated in at least 10 high power fields per tumour. Photomicrographs were acquired on a Nikon Eclipse E400 microscope with ×20 or ×40 objectives (NA 0.40 and 0.65) at room temperature and a Nikon DS-Fi-1 camera. Supplementary Table 1b lists antibodies used for immunoblotting and IHC.
RNAseq analysis
RNAseq was performed on 4 biological replicates of HUVEC treated with S1P or vehicle/control for 4 h. Quality control (RNA integrity number >7), library construction and sequencing was performed by BGI Tech Solutions (Hong Kong). RNA was hybridised to Illumina HiSeq 2000 platform and data obtained using 10 M clean reads.
Analysis of gene expression
RNAseq data were aligned to the hg19 human genome using Rsubread aligner [30] and assigned to genes with featureCounts function. Read counts normalised between samples were converted to counts-per-million (CPM) reads using the edgeR package in R [31]. RNAseq data for 32 ABC and 54 GCB DLBCL were from the controlled access area of NIH database of genotypes and phenotypes (dbGaP; accession code phs000532.v5.p2) [32]. The RNAseq data for 4 GC B cell samples were from GEO (GSE45982) [33]. Differentially expressed genes were identified using edgeR with p < 0.05 and read CPM > 1 in at least half of the samples. Microarray data for 11 DLBCL and 10 GC B cell samples were from GEO (GSE12453) [34] and analysed with MAS5 algorithm of the Affymetrix Expression Console to generate expression levels for each probe set (GCOS Signal). The MAS5 TGT was set to 100. The series matrix expression and clinical data reported in Lenz et al. [3] were downloaded from GEO (GSE10846). Meta-analysis of 11 different DLBCL datasets was performed as before [35]. Spearman test was used to correlate gene expression between samples.
Statistics
Statistical tests are indicated in relevant sections. All experiments were performed at least in triplicate, and for B cells on at least three separate donors. Tests were considered statistically significant if p < 0.05.
Results
SPHK1 is over-expressed in the tumour cells of DLBCL
To explore the contribution of S1P signalling to angiogenesis in DLBCL, we examined the expression of SPHK1, an enzyme which generates S1P from sphingosine. We measured SPHK1 mRNA in published datasets, comparing global gene expression in DLBCL with that in normal GC B cells [32, 34]. SPHK1 mRNA levels were significantly higher in DLBCL compared to GC B cells (Fig. 1a) and in both ABC- and GCB- DLBCL subtypes when analysed separately (Fig. 1b).
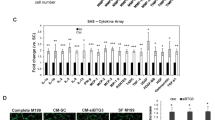
SPHK1 is over-expressed in primary DLBCL. a SPHK1 mRNA expression in the re-analysis of a published microarray dataset [34] and b SPHK1 mRNA expression in the re-analysis of published RNAseq data [32]. c IHC for SPHK1 protein expression in normal GC B cells and also in the tumour cells of representative cases of DLBCL. Arrows indicate tumour-associated EC which did not express SPHK1 (lower middle and lower right panels). GC germinal centre, MZ mantle zone. Original magnifications ×200 and ×600. d Upper panel: qPCR analysis for the expression of SPHK1 mRNA in DLBCL cell lines. Data are in triplicate and each is representative of three separate biological replicates. Lower panel: Immunoblotting of DLBCL cell lines for phosphorylated and total SPHK1 (pSPHK1, tSPHK1). β-tubulin is a loading control. e Intracellular S1P levels in primary B cells and DLBCL cell lines. Data show means (± SEM) of four biological replicates. Statistics were based on a comparison of individual cell lines with B cells#2. *Denotes p < 0.05, **p < 0.01, ***p < 0.001 (Student’s t-test). f S1P levels secreted by SUDHL4 cells incubated in serum-free media for indicated times measured by mass spectrometry. Data show means (± SEM) of three biological replicates. *Denotes p < 0.05
We performed IHC using an antibody we previously showed is specific for SPHK1 [26]. We found that SPHK1 was present in normal GC B cells, and in the tumour cells of all 32 cases of DLBCL examined; including cases of both GCB and non-GCB type defined by the Hans algorithm [29] (Fig. 1c; Supplementary Table 2). The intensity of SPHK1 expression in tumour cells was variable but was generally higher than in background of non-malignant cells, including macrophages and small reactive lymphocytes (not shown). Furthermore, in contrast to the strong staining observed in tumour cells, SPHK1 was either only weakly detectable or undetectable in EC of most (27/31) evaluable cases (Supplementary Table 2; Fig. 1c).
SPHK1 mRNA and protein were also expressed in DLBCL lines (Fig. 1d). Because the catalytic activity of SPHK1 is enhanced by its phosphorylation at Ser225 we used an antibody specific for this phosphorylation site [36, 26]. Immunoblotting with this antibody revealed that SPHK1 was phosphorylated in all cell lines examined (Fig. 1d; lower panel and Supplementary Fig. 1A). S1P levels were significantly higher in all six DLBCL cell lines examined compared to normal B cells (Fig. 1e). Because S1P is a ligand for S1P receptors present on the cell surface, it was of interest to determine whether DLBCL cells such as SUDHL4, which express SPHK1 and produce S1P intracellularly, can also secrete it. We found that DLBCL cells readily export S1P outside the cells reaching a maximum level of around 1 nM within 2 min (Fig. 1f).
Finally, we stained the DLBCL cases for S1PR1. We found that S1PR1 was expressed on endothelial cells in all 32 cases and in tumour cells in 20/32 cases (Supplementary Table 2, Supplementary Fig. 1B upper panels). In keeping with this, Q-PCR revealed that some, but not all, DLBCL cell lines expressed S1PR1 (Supplementary Fig. 1B, lower panel). In primary tumours, there was no relationship between tumour cell expression of S1PR1 and disease subtype.
SPHK1 expression is associated with an angiogenic transcriptome in DLBCL
We next performed a meta-analysis of 11 primary DLBCL gene expression datasets comprising over 2000 cases of DLBCL [35]. For each dataset, genes were ordered by their variance across the patient samples and the top 80% were used to calculate the Spearman’s rank correlations for their expression against that of SPHK1. The resultant p-values and correlation matrices were merged across the 11 datasets by taking the median values. A SPHK1-correlated gene set was created by taking all genes which were present in six or more datasets with a median p < 0.05. This identified 2236 genes positively correlated and 1658 genes negatively correlated with SPHK1 expression in DLBCL (Supplementary Table 3). We noted that genes positively correlated with SPHK1 included well known angiogenesis-associated genes, such as VEGF and VEGFR (r = 0.28, p = 0.0001; and r = 0.19, p = 0.0042, respectively). A gene ontology (GO) analysis [37] also showed a significant enrichment of angiogenesis and vasculature functions among genes positively correlated with SPHK1 (Fig. 2a). To provide further confirmation of a relationship between SPHK1 expression and angiogenesis, we utilised a published ‘tumour vascular gene signature’ generated from a meta-analysis of more than 1000 primary human cancers [38]. We found that genes positively correlated with SPHK1 expression in DLBCL were significantly enriched for tumour vascular signature genes (odds ratio (OR) = 7.92, p < 0.0001), whereas genes negatively correlated with SPHK1 were significantly depleted for tumour vascular signature genes (OR = 0.37, p < 0.0001; Fig. 2b).
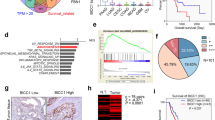
SPHK1 expression is associated with an angiogenic transcriptome in primary DLBCL. a Enrichment of angiogenesis-related terms (red arrows) in a GO analysis of genes positively correlated with SPHK1 expression in primary DLBCL. The top 10 significant GO categories are shown ordered by p-value. b Tumour vascular signature genes [38] were significantly enriched among genes positively correlated with SPHK1 expression in primary DLBCL (left panel), but significantly depleted among those negatively correlated with SPHK1 expression in primary DLBCL (right panel) (Fisher’s exact test for both comparisons). c Re-analysis of gene expression from primary DLBCL [32] reveals a statistically significant positive correlation between SPHK1 expression and the expression of EC marker genes CD34, CDH5, PECAM1 and VWF (r = Spearman rank correlation coefficient)
Repeating the meta-analysis, we found that tumour vascular signature genes were significantly enriched among genes positively correlated with SPHK1 expression in GCB and ABC subtypes when analysed separately (both p < 0.0001; Supplementary Fig. 2). As a further check on the association between SPHK1 and angiogenesis, we used two further gene signatures of angiogenesis (the “Hallmark” angiogenic signature, M5944, and the signature for the GO term ‘angiogenesis’). For both alternative angiogenic gene sets, we again found a significant overlap with genes positively correlated with SPHK1 (both p < 0.0001, not shown). Finally, we directly compared the mRNA expression of SPHK1 with that of classical EC markers in a further published RNAseq dataset reported by Morin et al. [32], not included in our original meta-analysis. This revealed a significant positive correlation between the expression of SPHK1 and that of CD34, CDH5, PECAM1/CD31 and VWF (Fig. 2c; Supplementary Fig. 3).
We also used the same 11 primary DLBCL datasets described above to perform a meta-analysis of genes correlated with S1PR1 in DLBCL. In keeping with the observed strong expression of S1PR1 in the tumour vasculature of DLBCL, we found that genes positively correlated with S1PR1 in primary tumours were also significantly enriched for tumour vascular signature genes (OR = 7.92, p < 0.0001; not shown). Furthermore, we also found that S1PR1 was positively correlated with CD34, (Spearman test, r = 0.48; p < 0.0001), CDH5 (r = 0.42; p < 0.0001), PECAM1 (r = 0.46; p < 0.0001), and VWF (r = 0.44; p < 0.0001) in the dataset reported by Morin et al. [32].
We conclude that the expression of both SPHK1 and S1PR1 is associated with an angiogenic transcriptome in DLBCL.
SPHK1 expression is associated with angiogenesis in both Stromal-1 and Stromal-2 DLBCL subtypes
Given that angiogenesis was reported to be associated with the Stromal-2 signature in DLBCL [3], we next explored the correlation of SPHK1 expression with the expression of stromal signature genes [3]. First, we studied the extent to which Stromal-2 genes overlapped with the angiogenic gene signatures. As expected, we found a significant overlap of Stromal-2 signature genes, but only with two of the three angiogenic gene sets (Fig. 3a; upper panel). Consistent with this, we found that SPHK1 expression was positively correlated with Stromal-2 gene expression (calculated as the average expression across genes in the Stromal-2 signature [3] (Spearman correlation = 0.46 and 0.47 for CHOP- and R-CHOP-treated patients, respectively; p < 0.0001, not shown). However, when we repeated this analysis, this time using Stromal-1 signature genes [3], we were surprised to find that SPHK1 expression was even more highly correlated with Stromal-1 gene expression (Spearman correlation = 0.80 and 0.77, respectively, p < 0.0001) and included a highly significant overlap with all three angiogenic gene sets (Fig. 3b; lower panel). In contrast, we observed that S1PR1 expression was positively correlated only with Stromal-2 gene expression (Spearman correlation = 0.34 and 0.45 for CHOP- and R-CHOP-treated patients, respectively; both p < 0.0001), and not with Stromal-1-gene expression (Spearman correlation r = 0.02, p = 0.807; and r = 0.09, p = 0.212, for CHOP- and R-CHOP-treated patients, respectively). We conclude that genes positively correlated with SPHK1 are enriched in both stromal signatures of DLBCL, most likely reflecting the contribution of SPHK1 to angiogenesis in both stromal subgroups.
An in vitro transcriptional signature of S1P signalling in endothelial cells
With the aim of revealing a more direct association between S1P signalling and angiogenesis in DLBCL, we next defined a transcriptional signature of S1P signalling in EC. To do this we used human umbilical vein endothelial cells (HUVEC) which we showed express the same pattern of S1P receptors as the EC of DLBCL, marked by the high expression of S1PR1 and the absence of S1PR2 and S1PR3 in most cases (Fig. 4a). We showed that S1P treatment of HUVEC induced the robust phosphorylation of ERK1/2, a well-defined downstream target of S1P signalling in EC [12] (Fig. 4b).
Identification of S1P target genes in endothelial cells. a Representative images of IHC for S1PR1, S1PR2 and S1PR3 in primary DLBCL and HUVEC. DLBCL-associated EC (left and middle panels) were positive for S1PR1 and negative for S1PR2 and S1PR3 in most cases (black arrows). Red blood cells stained strongly for S1PR2 (red arrows) and served as an internal positive control. S1PR3-positive tumour cells (bottom left) and/or squamous epithelium (not shown) were internal positive controls for S1PR3 expression. HUVEC (right panel) were positive for S1PR1 and negative for S1PR2 and S1PR3. Original magnifications ×200 (tissues) and ×600 (cells). b S1P treatment of HUVEC increased ERK1/2 phosphorylation, shown here for 4 separate donors compared with vehicle only treated cells (H1-H4). β-tubulin is a loading control. The same protein lysates were used to detect phosphorylated and total ERK1/2 (pERK1/2, tERK1/2) in separate blots. c Q-PCR for S1P transcriptional targets in HUVEC of four donors following treatment with S1P (dark grey bars) compared with vehicle (light grey bars) normalised to 1. *Denotes p < 0.05, **p < 0.01, ***p < 0.001 (Student’s t-test). d Immunoblotting for pERK1/2, tERK1/2 and β-tubulin in HUVEC representative of three donors treated with S1P in the presence of Sphingomab (SmAb) or isotype/control antibody (CmAb). The same protein lysates were used to detect pERK1/2 and tERK1/2 in separate blots. e Q-PCR for S1P targets following treatment of HUVEC with S1P in the presence of either SmAb or CmAb. Data shown are representative of three donors. *Denotes p < 0.05, **p < 0.01, ***p < 0.001 (Student’s t-test)
We next used RNAseq to describe the global transcriptional consequences of S1P signalling in HUVEC. Treatment of HUVEC with S1P was followed by the up-regulation of 115 genes and the down-regulation of 126 genes (Supplementary Table 4). We used Q-PCR to validate a subset of the genes which were both upregulated by S1P and positively correlated with SPHK1 in our meta-analysis, including IL-8, ICAM1, SELE, PDGFA, ANGPTL4 and CXCL12 (Fig. 4c).
To confirm the specificity of these effects, we repeated these experiments in the presence of Sphingomab, a monoclonal antibody which depletes extracellular S1P [39]. Sphingomab greatly suppressed S1P-induced phosphorylation of ERK1/2 (Fig. 4d) and the upregulation of its downstream transcriptional targets (Fig. 4e). In contrast, S1P induced the phosphorylation of ERK1/2 and the up-regulation of its target genes in the presence of the isotype control antibody.
Genes upregulated by S1P signalling and correlated with SPHK1 mRNA are expressed in the tumour vasculature of DLBCL
A gene ontology analysis revealed that genes upregulated by S1P in HUVEC were significantly enriched for GO terms associated with angiogenesis, and for GO terms associated with the known effects of S1P in EC, including protection from apoptosis, leukocyte adhesion and migration (Fig. 5a). Furthermore, genes up-regulated (OR = 4.30, p < 0.0001), but not those downregulated (OR = 0.78, p = 0.48), by S1P were significantly enriched among genes positively correlated with SPHK1 in DLBCL (Fig. 5b, left panel). Similarly, genes downregulated (OR = 2.58, p < 0.0001), but not those up-regulated (OR = 0.73, p = 0.48), by S1P were significantly enriched in genes negatively correlated with SPHK1 (Fig. 5b, right panel). The enrichment of upregulated S1P targets among genes positively correlated with SPHK1 was also evident when the GCB and ABC subtypes were analysed separately (Supplementary Fig. 4).
Genes upregulated by S1P and correlated with SPHK1 mRNA are expressed in DLBCL tumour vasculature. a GO analysis of genes upregulated by S1P in HUVEC revealed a significant enrichment of GO terms associated with angiogenesis. b Genes upregulated (left panel), but not those downregulated (not shown) in S1P-treated HUVEC were enriched among genes positively correlated with SPHK1 expression in primary DLBCL. Genes downregulated (right panel), but not those upregulated (not shown) in S1P-treated HUVEC were enriched among genes negatively correlated with SPHK1 expression in primary DLBCL. c Representative examples of the expression of the S1P target genes, CXCL12, SELE, COL1A1 and MAP1B, in the tumour vasculature of DLBCL. Original magnifications ×200
To provide direct evidence of the expression of S1P target genes in the tumour vasculature of DLBCL we performed IHC for CXCL12 and SELE, as well as COL1A1 and MAP1B, two further genes up-regulated in S1P-treated HUVEC, positively correlated with SPHK1 expression in DLBCL, and that had available antibodies that work robustly in FFPE tissues. We found that all four markers were expressed in the EC of both subtypes of DLBCL. In contrast, tumour cells were either negative (CXCL12, COL1A1, SELE) or showed only weak expression (MAP1B) (Supplementary Table 5). In summary, these data provide evidence of an S1P-regulated transcriptional programme in the tumour vasculature of DLBCL.
S1PR1 inhibition reverses S1P signalling in vitro and reduces angiogenesis and tumour growth in a mouse model of angiogenic DLBCL
Finally, we explored the effects of inhibiting S1P signalling in a mouse model of DLBCL. We initially studied ERK1/2 phosphorylation in vitro following the stimulation of HUVEC with S1P, in the presence or absence of Siponimod, Ozanimod and Ponesimod, three functional antagonists of S1PR1 [19, 40, 41]. We found that all three drugs decreased ERK1/2 phosphorylation in HUVEC, confirming their ability to reduce the effects of S1P on downstream signalling (Fig. 6a). We next studied the impact of inhibiting S1P signalling in a mouse model of angiogenic DLBCL, focussing on Siponimod, a potent functional antagonist of S1PR1 and S1PR5 (EC50:0.39 and 0.98 nM, respectively) with > 1000-fold selectivity for S1PR1 versus S1PR2, S1PR3 and S1PR4 [42]. For these experiments we used SUDHL6 cells, not only because they produce high levels of S1P (Fig. 1e), but also because as xenografts they induce levels of angiogenesis that are almost 8-fold-higher than those found in xenografts of OCI-LY1 cells, and nearly 60-fold higher than those observed in the A20 syngeneic mouse model of DLBCL (Supplementary Fig. 5A). We treated established xenografts of SUDHL6 cells with Siponimod for up to two weeks before culling the animals and measuring mouse EC numbers in tumours by flow cytometry. We found that compared to controls, treatment of Siponimod significantly reduced the numbers of CD31-positive mouse EC in tumour tissues (Fig. 6b) as well as reducing their expression of the S1P target gene, CXCL12 (Supplementary Fig. 5B). Furthermore, Siponimod led to a significant reduction in tumour volumes in treated animals at the later time points (Fig. 6c) that was associated with increased numbers of apoptotic cells as measured by immunohistochemistry for cleaved PARP (Supplementary Fig. 5C). In contrast, we did not observe significantly reduced tumour volumes following Siponimod treatment of xenografts of the poorly angiogenic OCI-LY1 and A20 lines (Supplementary Fig. 5D, E). We conclude that the inhibition of S1P signalling can reduce angiogenesis and tumour growth in a mouse model of angiogenic DLBCL.
S1PR1 inhibition decreases S1P-induced signalling and reduces angiogenesis in a mouse model of DLBCL. a Immunoblot of HUVEC pre-treated for 1 h with 100 nM of S1PR1 functional antagonists, Siponimod, Ozanimod and Ponesimod and stimulated with S1P or control (S1P vehicle) for 5 min. S1P or control only treated cells received the equivalent dose of DMSO (drug solvent) to S1PR1 inhibitor-treated cells (0.01 µl DMSO per 1 mL of media). β-tubulin was used as a loading control. The same protein lysates were used to detect pERK1/2 and tERK1/2 in separate blots. Data are representative of three separate donors. b Flow cytometry analysis of numbers of CD31-positive mouse cells/per mg tumour of SUDHL6 xenografts in NSG mice treated with Siponimod compared with drug vehicle (4 mice for each group). c Tumour volumes in days after injection with SUDHL6 cells in Siponimod-treated versus vehicle-treated animals. Arrows indicate treatment days. *Denotes p < 0.05 in Student’s t-test
Discussion
Although previous studies have shown that targeting VEGF or its receptors can decrease tumour vascularization and reduce the growth of DLBCL-derived cell lines in vivo, a phase III trial of patients with aggressive NHL receiving R-CHOP plus bevacizumab showed cardiotoxicity without improvement in progression-free survival [6]. We speculated that DLBCL-associated angiogenesis could be driven by VEGF-independent signalling; a contention supported by other studies which reveal no correlation between lymphoma cell VEGF expression and micro-vessel density in primary DLBCL [7, 8]. Here, we have focused on the potential contribution of the potent bioactive sphingolipid metabolite, S1P. We have shown that SPHK1, the major enzyme responsible for the production of S1P, which can be secreted, is over-expressed in DLBCL. SPHK1 is strongly implicated in tumour angiogenesis in other tumours [43], and is important for the generation of S1P that drives tumour-induced hemangiogenesis and lymphangiogenesis in murine models of breast cancer [44]. The underlying mechanisms responsible for the over-expression of SPHK1 in DLBCL are not known but do not usually involve amplification and/or mutation; our review of five separate cohorts of DLBCL has revealed that the frequency of amplification and/or mutation of the SPHK1 gene is very low (<1%; not shown; TCGA Research Network: http://cancergenome.nih.gov/; [32, 45,46,47]).
SPHK1 can be activated by growth factors and cytokines [48,49,50,51,52], and its catalytic activity is enhanced by ERK1/2-mediated phosphorylation at Ser225 [36]. The pSer225 antibody used in our study showed that SPHK1 was constitutively phosphorylated at Ser225 in DLBCL cell lines. In keeping with this, we showed that S1P production was increased in DLBCL cell lines compared to normal B cells, and furthermore that this S1P was secreted into the extracellular environment.
Our meta-analysis of DLBCL revealed a strong correlation between the expression of SPHK1 and that of angiogenesis meta-signature and EC-classifier genes. One explanation for this finding could be expression of SPHK1 in tumour-associated EC. However, we observed SPHK1 expression predominantly in cancer cells, consistent with the notion that SPHK1 expression is upregulated primarily in the tumour cell population [18].
Although the meta-analysis and our in situ expression data suggested that angiogenesis in DLBCL is driven by tumour-derived SPHK1, these approaches do not directly implicate S1P. Interactions between tumour and EC mediated by S1P have been shown to be important for angiogenesis in many solid cancers, including those of the breast, prostate, liver and kidney [53,54,55]. To investigate the involvement of S1P in DLBCL-associated angiogenesis, we defined a transcriptional signature of S1P signalling in HUVEC [23]. Gene ontology analysis revealed that genes up-regulated in this transcriptional signature were enriched for angiogenesis-associated GO terms, as well as for GO terms reflecting known functions of S1P [9, 12, 13, 56, 57]. We used this transcriptional signature to show that genes upregulated by S1P were enriched among genes correlated with SPHK1, an association that was also evident separately for both COO subtypes. IHC revealed that S1P target genes were expressed in the tumour vasculature of SPHK1-expressing DLBCL, with little or no tumour cell expression, suggesting that the enrichment of S1P target genes observed in SPHK1-expressing DLBCL was primarily a consequence of their expression in tumour-associated EC.
Recent studies exploring the transcriptional landscape of DLBCL have defined tumour microenvironment-derived gene signatures predicting clinical outcome [2, 3]. The ‘Stromal-1’ signature, associated with a favourable outcome, is characterised by expression of extracellular matrix- and macrophage-associated genes. In contrast, the ‘Stromal-2’ signature has an unfavourable outcome, and is reportedly enriched for angiogenesis-associated genes [3]. We formally tested the relationship between angiogenesis and these stromal signatures [3]. As expected, we found a significant enrichment of angiogenesis-associated genes within the Stromal-2 signature. However, we were surprised to find an even stronger enrichment of angiogenesis genes within the Stromal-1 signature. Both stromal signatures were enriched for genes positively correlated with SPHK1 expression, suggesting that SPHK1 contributes to angiogenesis in both stromal subtypes.
Our data suggest novel therapeutic opportunities for patients with SPHK1-expressing DLBCL insofar as the S1PR1 functional antagonists, Siponimod, Ozanimod and Ponesimod, were all able to block S1P signalling in EC. Although we observed the same effect with the S1P-specific monoclonal antibody, Sphingomab, we suggest that S1PR1 is a preferred target; S1P-blocking agents will affect signalling through all S1P receptors. Potentially unwanted side-effects of blocking S1P could include the inhibition of S1PR2 signalling which we and others have reported to be tumour suppressive in DLBCL [58, 59, 60]. Moreover, we showed that Siponimod, a highly potent functional antagonist of S1PR1 that is effective and safe in patients with multiple sclerosis [61], can reduce angiogenesis and tumour growth of SUDHL6 xenografts which produce S1P and which we used as a mouse model of angiogenic DLBCL. Targeting tumour cell over-expression of S1PR1, which is observed in a subset of poor prognosis DLBCL, is also potentially advantageous, since S1PR1 has been shown to be important for the proliferation and survival of DLBCL [62, 63]. S1P/S1PR1 signalling also promotes the accumulation of Treg while inhibiting CD8+T cell recruitment and activation in xenograft models of breast cancer and melanoma [64]. Therefore, blocking S1PR1 might also promote anti-tumour immunity. Taken together our observations suggest that specific S1PR1 antagonists should be investigated for their therapeutic potential in DLBCL patients [40, 65, 66].
References
Coiffier B, Thieblemont C, Van Den Neste E, Lepeu G, Plantier I, Castaigne S, et al. Long-term outcome of patients in the LNH-98.5 trial, the first randomized study comparing rituximab-CHOP to standard CHOP chemotherapy in DLBCL patients: a study by the Groupe d’Etudes des Lymphomes de l’Adulte. Blood. 2010;116:2040–5.
Sehn LH, Gascoyne RD. Diffuse large B-cell lymphoma: optimizing outcome in the context of clinical and biologic heterogeneity. Blood. 2015;125:22–32.
Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H, et al. Stromal gene signatures in large-B-cell lymphomas. New Engl J Med. 2008;359:2313–23.
Stopeck AT, Unger JM, Rimsza LM, Bellamy WT, Iannone M, Persky DO, et al. A phase II trial of single agent bevacizumab in patients with relapsed, aggressive non-Hodgkin lymphoma: Southwest oncology group study S0108. Leuk Lymphoma. 2009;50:728–35.
Ganjoo KN, An CS, Robertson MJ, Gordon LI, Sen JA, Weisenbach J, et al. Rituximab, bevacizumab and CHOP (RA-CHOP) in untreated diffuse large B-cell lymphoma: safety, biomarker and pharmacokinetic analysis. Leuk Lymphoma. 2006;47:998–1005.
Seymour JF, Pfreundschuh M, Trnĕný M, Sehn LH, Catalano J, Csinady E, et al. R-CHOP with or without bevacizumab in patients with previously untreated diffuse large B-cell lymphoma: final MAIN study outcomes. Haematologica. 2014;99:1343–9.
Gratzinger D, Zhao S, Tibshirani RJ, Hsi ED, Hans CP, Pohlman B, et al. Prognostic significance of VEGF, VEGF receptors, and microvessel density in diffuse large B cell lymphoma treated with anthracycline-based chemotherapy. Lab Invest. 2008;88:38–47.
Jorgensen JM, Sorensen FB, Bendix K, Nielsen JL, Olsen ML, Funder AM, et al. Angiogenesis in non-Hodgkin’s lymphoma: clinico-pathological correlations and prognostic significance in specific subtypes. Leuk Lymphoma. 2007;48:584–95.
Takuwa Y, Du W, Qi X, Okamoto Y, Takuwa N, Yoshioka K. Roles of sphingosine-1-phosphate signaling in angiogenesis. World J Biol Chem. 2010;1:298–306.
Pyne NJ, Pyne S. Sphingosine 1-phosphate and cancer. Nat Rev Cancer. 2010;10:489–503.
Maceyka M, Harikumar KB, Milstien S, Spiegel S. Sphingosine-1-phosphate signaling and its role in disease. Trends Cell Biol. 2012;22:50–60.
Lee OH, Kim YM, Lee YM, Moon EJ, Lee DJ, Kim JH, et al. Sphingosine 1-phosphate induces angiogenesis: its angiogenic action and signaling mechanism in human umbilical vein endothelial cells. Biochem Biophys Res Commun. 1999;264:743–50.
Wang F, Van Brocklyn JR, Hobson JP, Movafagh S, Zukowska-Grojec Z, Milstien S, et al. Sphingosine 1-phosphate stimulates cell migration through a G(i)-coupled cell surface receptor. Potential Involv angiogenesis J Biol Chem. 1999;274:35343–50.
English D, Welch Z, Kovala AT, Harvey K, Volpert OV, Brindley DN, et al. Sphingosine 1-phosphate released from platelets during clotting accounts for the potent endothelial cell chemotactic activity of blood serum and provides a novel link between hemostasis and angiogenesis. FASEB J. 2000;14:2255–65.
Liu F, Verin AD, Wang P, Day R, Wersto RP, Chrest FJ, et al. Differential regulation of sphingosine-1-phosphate- and VEGF-induced endothelial cell chemotaxis. Involvement of G(ialpha2)-linked Rho kinase activity. Am J Respir Cell Mol Biol. 2001;24:711–9.
Spiegel S, Milstien S. Sphingosine-1-phosphate: an enigmatic signalling lipid. Nat Rev Mol Cell Biol. 2003;4:397–407.
Wallington-Beddoe CT, Bradstock KF, Bendall LJ. Oncogenic properties of sphingosine kinases in haematological malignancies. Br J Haematol. 2013;161:623–38.
Bayerl MG, Bruggeman RD, Conroy EJ, Hengst JA, King TS, Jimenez M, et al. Sphingosine kinase 1 protein and mRNA are overexpressed in non-Hodgkin lymphomas and are attractive targets for novel pharmacological interventions. Leuk Lymphoma. 2008;49:948–54.
Pan S, Gray NS, Gao W, Mi Y, Fan Y, Wang X, et al. Discovery of BAF312 (Siponimod), a potent and selective S1P receptor modulator. ACS Med Chem Lett. 2013;4:333–7.
Vockerodt M, Morgan SL, Kuo M, Wei W, Chukwuma MB, Arrand JR, et al. The Epstein-Barr virus oncoprotein, latent membrane protein-1, reprograms germinal centre B cells towards a Hodgkin’s Reed-Sternberg-like phenotype. J Pathol. 2008;216:83–92.
Vrzalikova K, Vockerodt M, Leonard S, Bell A, Wei W, Schrader A, et al. Down-regulation of BLIMP1α by the EBV oncogene, LMP-1, disrupts the plasma cell differentiation program and prevents viral replication in B cells: implications for the pathogenesis of EBV-associated B-cell lymphomas. Blood. 2011;117:5907–17.
Vrzalikova K, Leonard S, Fan Y, Bell A, Vockerodt M, Flodr P, et al. Hypomethylation and over-expression of the beta isoform of BLIMP1 is induced by Epstein-Barr virus infection of B Cells; potential implications for the pathogenesis of EBV-associated lymphomas. Pathogens. 2012;1:83–101.
Jaffe EA, Nachman RL, Becker CG, Minick CR. Culture of human endothelial cells derived from umbilical veins. Identif Morphol Immunol criteria J Clin Invest. 1973;52:2745–56.
Cassetta L, Noy R, Swierczak A, Sugano G, Smith H, Wiechmann L, et al. Isolation of mouse and human tumor-associated macrophages. Adv Exp Med Biol. 2016;899:211–29.
Hait NC, Allegood J, Maceyka M, Strub GM, Harikumar KB, Singh SK, et al. Regulation of histone acetylation in the nucleus by sphingosine-1-phosphate. Science. 2009;325:1254–7.
Vrzalikova K, Ibrahim M, Vockerodt M, Perry T, Margielewska S, Lupino L, et al. S1PR1 drives a feedforward signalling loop to regulate BATF3 and the transcriptional programme of Hodgkin lymphoma cells. Leukemia. 2018;32:214–23.
Reynolds GM, Billingham LJ, Gray LJ, Flavell JR, Najafipour S, Crocker J, et al. Interleukin 6 expression by Hodgkin/Reed-Sternberg cells is associated with the presence of ‘B’ symptoms and failure to achieve complete remission in patients with advanced Hodgkin’s disease. Br J Haematol. 2002;118:195–201.
Flavell JR, Baumforth KR, Wood VH, Davies GL, Wei W, Reynolds GM, et al. Down-regulation of the TGF-beta target gene, PTPRK, by the Epstein-Barr virus encoded EBNA1 contributes to the growth and survival of Hodgkin lymphoma cells. Blood. 2008;111:292–301.
Hans CP, Weisenburger DD, Greiner TC, Gascoyne RD, Delabie J, Ott G, et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood. 2004;103:275–82.
Liao Y, Smyth GK, Shi W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 2013;41:e108.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Morin RD, Johnson NA, Severson TM, Mungall AJ, An J, Goya R, et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat Genet. 2010;42:181–5.
Beguelin W, Popovic R, Teater M, Jiang Y, Bunting KL, Rosen M, et al. EZH2 is required for germinal center formation and somatic EZH2 mutations promote lymphoid transformation. Cancer Cell. 2013;23:677–92.
Brune V, Tiacci E, Pfeil I, Döring C, Eckerle S, van Noesel CJM, et al. Origin and pathogenesis of nodular lymphocyte–predominant Hodgkin lymphoma as revealed by global gene expression analysis. J Exp Med. 2008;205:2251–68.
Care MA, Westhead DR, Tooze RM. Gene expression meta-analysis reveals immune response convergence on the IFNgamma-STAT1-IRF1 axis and adaptive immune resistance mechanisms in lymphoma. Genome Med. 2015;7:15–218.
Pitson SM, Moretti PA, Zebol JR, Lynn HE, Xia P, Vadas MA, et al. Activation of sphingosine kinase 1 by ERK1/2-mediated phosphorylation. EMBO J. 2003;22:5491–500.
Dennis G, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4:P3.
Masiero M, Simoes FC, Han HD, Snell C, Peterkin T, Bridges E, et al. A core human primary tumor angiogenesis signature identifies the endothelial orphan receptor ELTD1 as a key regulator of angiogenesis. Cancer Cell. 2013;24:229–41.
O’Brien N, Jones ST, Williams DG, Cunningham HB, Moreno K, Visentin B, et al. Production and characterization of monoclonal anti-sphingosine-1-phosphate antibodies. J Lipid Res. 2009;50:2245–57.
Olsson T, Boster A, Fernandez O, Freedman MS, Pozzilli C, Bach D, et al. Oral ponesimod in relapsing-remitting multiple sclerosis: a randomised phase II trial. J Neurol Neurosurg Psychiatry. 2014;85:1198–208.
Scott FL, Clemons B, Brooks J, Brahmachary E, Powell R, Dedman H, et al. Ozanimod (RPC1063) is a potent sphingosine-1-phosphate receptor-1 (S1P1) and receptor-5 (S1P5) agonist with autoimmune disease-modifying activity. Br J Pharm. 2016;173:1778–92.
Hale JJ, Lynch CL, Neway W, Mills SG, Hajdu R, Keohane CA, et al. A rational utilization of high-throughput screening affords selective, orally bioavailable 1-benzyl-3-carboxyazetidine sphingosine-1-phosphate-1 receptor agonists. J Med Chem. 2004;47:6662–5.
Zhang L, Wang XE, Bullock AJ, Callea M, Shah H, Song JX, et al. Anti-S1P antibody as a novel therapeutic strategy for VEGFR TKI-resistant renal cancer. Clin Cancer Res. 2015;21:1925–34.
Nagahashi M, Ramachandran S, Kim EY, Allegood JC, Rashid OM, Yamada A, et al. Sphingosine-1-phosphate produced by sphingosine kinase 1 promotes breast cancer progression by stimulating angiogenesis and lymphangiogenesis. Cancer Res. 2012;72:726–35.
Reddy A, Zhang J, Davis NS, Moffitt AB, Love CL, Waldrop A, et al. Genetic and functional drivers of diffuse large B cell lymphoma. Cell. 2017;171:481–94 e15.
Chapuy B, Stewart C, Dunford AJ, Kim J, Kamburov A, Redd RA, et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat Med. 2018;24:679–90.
Schmitz R, Wright GW, Huang DW, Johnson CA, Phelan JD, Wang JQ, et al. Genetics and pathogenesis of diffuse large B-cell lymphoma. N Engl J Med. 2018;378:1396–407.
Vann LR, Payne SG, Edsall LC, Twitty S, Spiegel S, Milstien S. Involvement of sphingosine kinase in TNF-alpha-stimulated tetrahydrobiopterin biosynthesis in C6 glioma cells. J Biol Chem. 2002;277:12649–56.
Osawa Y, Banno Y, Nagaki M, Brenner DA, Naiki T, Nozawa Y, et al. TNF-alpha-induced sphingosine 1-phosphate inhibits apoptosis through a phosphatidylinositol 3-kinase/Akt pathway in human hepatocytes. J Immunol. 2001;167:173–80.
Olivera A, Edsall L, Poulton S, Kazlauskas A, Spiegel S. Platelet-derived growth factor-induced activation of sphingosine kinase requires phosphorylation of the PDGF receptor tyrosine residue responsible for binding of PLCgamma. FASEB J. 1999;13:1593–600.
Young KW, Bootman MD, Channing DR, Lipp P, Maycox PR, Meakin J, et al. Lysophosphatidic acid-induced Ca2+ mobilization requires intracellular sphingosine 1-phosphate production. Potential involvement of endogenous EDG-4 receptors. J Biol Chem. 2000;275:38532–9.
Meyer zu Heringdorf D, Lass H, Alemany R, Laser KT, Neumann E, Zhang C, et al. Sphingosine kinase-mediated Ca2+ signalling by G-protein-coupled receptors. EMBO J. 1998;17:2830–7.
Ader I, Gstalder C, Bouquerel P, Golzio M, Andrieu G, Zalvidea S, et al. Neutralizing S1P inhibits intratumoral hypoxia, induces vascular remodelling and sensitizes to chemotherapy in prostate cancer. Oncotarget. 2015;6:13803–21.
Lu ZP, Zhang WY, Gao S, Jiang QL, Xiao ZL, Ye LH, et al. MiR-506 suppresses liver cancer angiogenesis through targeting sphingosine kinase 1 (SPHK1) mRNA. Biochem Bioph Res Co. 2015;468:8–13.
Mukhopadhyay P, Ramanathan R, Takabe K. S1P promotes breast cancer progression by angiogenesis and lymphangiogenesis. Breast Cancer Manag. 2015;4:241–4.
Hisano N, Yatomi Y, Satoh K, Akimoto S, Mitsumata M, Fujino MA, et al. Induction and suppression of endothelial cell apoptosis by sphingolipids: a possible in vitro model for cell-cell interactions between platelets and endothelial cells. Blood. 1999;93:4293–9.
Fernandez-Pisonero I, Duenas AI, Barreiro O, Montero O, Sanchez-Madrid F, Garcia-Rodriguez C. Lipopolysaccharide and sphingosine-1-phosphate cooperate to induce inflammatory molecules and leukocyte adhesion in endothelial cells. J Immunol. 2012;189:5402–10.
Flori M, Schmid CA, Sumrall ET, Tzankov A, Law CW, Robinson MD, et al. The hematopoietic oncoprotein FOXP1 promotes tumor cell survival in diffuse large B-cell lymphoma by repressing S1PR2 signaling. Blood. 2016;127:1438–48.
Muppidi JR, Schmitz R, Green JA, Xiao W, Larsen AB, Braun SE, et al. Loss of signalling via Galpha13 in germinal centre B-cell-derived lymphoma. Nature. 2014;516:254–8.
Vockerodt M, Vrzalikova K, Ibrahim M, Nagy E, Margielewska S, Hollows R, Lupino L, Tooze R, Care M, Simmons W, Schrader A, Perry T, Abdullah M, Foster S, Reynolds G, Dowell A, Rudzki Z, Krappmann D, Kube D, Woodman C, Wei W, Taylor G, Murray PG. Regulation of S1PR2 by the EBV oncogene LMP1 in aggressive ABC-subtype diffuse large B-cell lymphoma. J Pathol. 2019 https://doi.org/10.1002/path.5237. [Epub ahead of print].
Kappos L, Bar-Or A, Cree BAC, Fox RJ, Giovannoni G, Gold R, et al. Siponimod versus placebo in secondary progressive multiple sclerosis (EXPAND): a double-blind, randomised, phase 3 study. Lancet. 2018;391:1263–73.
Koresawa R, Yamazaki K, Oka D, Fujiwara H, Nishimura H, Akiyama T, et al. Sphingosine-1-phosphate receptor 1 as a prognostic biomarker and therapeutic target for patients with primary testicular diffuse large B-cell lymphoma. Br J Haematol. 2016;174:264–74.
Liu Y, Deng J, Wang L, Lee H, Armstrong B, Scuto A, et al. S1PR1 is an effective target to block STAT3 signaling in activated B cell-like diffuse large B-cell lymphoma. Blood. 2012;120:1458–65.
Priceman SJ, Shen S, Wang L, Deng J, Yue C, Kujawski M, et al. S1PR1 is crucial for accumulation of regulatory T cells in tumors via STAT3. Cell Rep. 2014;6:992–9.
Kappos L, Li DK, Stuve O, Hartung HP, Freedman MS, Hemmer B, et al. Safety and Efficacy of Siponimod (BAF312) in Patients With Relapsing-Remitting Multiple Sclerosis: Dose-Blinded, Randomized Extension of the Phase 2 BOLD Study. JAMA Neurol. 2016;73:1089–98.
Tran JQ, Hartung JP, Peach RJ, Boehm MF, Rosen H, Smith H, et al. Results from the first-in-human study with Ozanimod, a novel, selective sphingosine-1-phosphate receptor modulator. J Clin Pharm. 2017;57:988–96.
Acknowledgements
This work was supported by Bloodwise (LL, TP, RH, MI, CJBW, KV, PGM) and the Traynor Foundation (LL), and in part by grants RVO:61989592 and NPS I LO1304 from the Czech Ministry of Education (PGM), NIH grant R01GM043880 (SS), and by Cancer Research UK Programme Grant C7845/A17723 (MC).
Author information
Authors and Affiliations
Contributions
LL, CBJW, RS, MR, KV, RB, SS and PGM designed research; LL, KV, TP, SM, MI, GR, WS, ZR, YLH, UZ and JA performed research and analysed data; RH, MC, RT, WW and PGM contributed to statistical analysis; ZR, YLH, WS and UZ undertook pathology evaluation; LL, KV, SS and PGM wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lupino, L., Perry, T., Margielewska, S. et al. Sphingosine-1-phosphate signalling drives an angiogenic transcriptional programme in diffuse large B cell lymphoma. Leukemia 33, 2884–2897 (2019). https://doi.org/10.1038/s41375-019-0478-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0478-9
This article is cited by
-
S1PR1/S1PR3-YAP signaling and S1P-ALOX15 signaling contribute to an aggressive behavior in obesity-lymphoma
Journal of Experimental & Clinical Cancer Research (2023)
-
The targetable nanoparticle BAF312@cRGD-CaP-NP represses tumor growth and angiogenesis by downregulating the S1PR1/P-STAT3/VEGFA axis in triple-negative breast cancer
Journal of Nanobiotechnology (2021)