Abstract
In more than 30% of B-cell precursor acute lymphoblastic leukaemia (B-ALL), chromosome 21 sequence is overrepresented through aneuploidy or structural rearrangements, exemplified by intrachromosomal amplification of chromosome 21 (iAMP21). Although frequent, the mechanisms by which these abnormalities promote B-ALL remain obscure. Intriguingly, we found copy number neutral loss of heterozygosity (CN-LOH) of 12q was recurrent in iAMP21-ALL, but never observed in B-ALL without some form of chromosome 21 gain. As a consequence of CN-LOH 12q, mutations or deletions of the adaptor protein, SH2B3, were converted to homozygosity. In patients without CN-LOH 12q, bi-allelic abnormalities of SH2B3 occurred, but only in iAMP21-ALL, giving an overall incidence of 18% in this sub-type. Review of published data confirmed a tight association between overrepresentation of chromosome 21 and both CN-LOH 12q and SH2B3 abnormalities in B-ALL. Despite relatively small patient numbers, preliminary analysis linked 12q abnormalities to poor outcome in iAMP21-ALL (p = 0.03). Homology modelling of a leukaemia-associated SH2 domain mutation and in vitro analysis of patient-derived xenograft cells implicated the JAK/STAT pathway as one likely target for SH2B3 tumour suppressor activity in iAMP21-ALL.
Similar content being viewed by others
Introduction
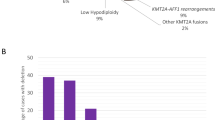
Intrachromosomal amplification of chromosome 21 (iAMP21-ALL) defines a distinct, high-risk sub-group comprising 2% of precursor B-lineage acute lymphoblastic leukaemia (B-ALL) [1, 2]. Originating through breakage-fusion-bridge cycles and chromothripsis, rearrangements of chromosome 21, giving rise to iAMP21, although chaotic, result in common regions of genomic amplification and increased gene expression [3, 4]. The precise mechanism by which iAMP21 promotes leukaemia is currently unknown, but is assumed to involve increased expression of genes or functional non-coding sequences within the chromosome 21 amplified regions. Other structural amplifications of chromosome 21 (OSA21), such as isochromosome 21, i(21q), are rare but gains of one or more copies of whole chromosome 21 (wc21) are extremely common. Notably chromosome 21 is the only chromosome always overrepresented in high hyperdiploid, hypodiploid and near-haploid ALL [5, 6], which together account for about 30% of B-ALL. Somatically acquired trisomy or tetrasomy 21 may also be present as sole aneuploidies [7] and, individuals with constitutional trisomy 21 (Down syndrome, DS) are at an ~20-fold increased risk of developing B-ALL [8, 9].
Deletions of RB1 and EBF1, mutations affecting the RAS pathway, CRLF2 activating rearrangements, gain of the X chromosome and partial deletion of chromosome 7 are all enriched in iAMP21-ALL [4, 10,11,12]. Suggesting a mechanistic link between iAMP21 and overrepresentation of whole chromosome 21, some of these secondary abnormalities also occur at high frequency in hyperdiploid or DS-ALL [13, 14].
Duplication of part of the maternal or paternal genome with concurrent loss of equivalent sequence from the other parental chromosome, known as copy number neutral loss of heterozygosity (CN-LOH), can have pathological consequences, through imbalance of expression of imprinted loci or conversion to homozygosity of inherited or acquired mutations and copy number abnormalities (CNA). In B-ALL, only deletions of the CDKN2A/B genes have been clearly linked to CN-LOH 9p [15,16,17,18,19,20], but other regions were recurrent, suggesting that they harbour imprinted or mutated genes or CNA that contribute to ALL. Here, we investigated CN-LOH in B-ALL with whole or partial gain of chromosome 21. We uncovered a highly specific association between CN-LOH 12q and the presence of iAMP21/OSA21, or less frequently gain of wc21. We further identified SH2B3 abnormalities as a target of CN-LOH 12q and found that, with rare exceptions, previously published cases of B-ALL with SH2B3 abnormalities also harboured whole or partial amplification of chromosome 21. Preliminary analysis of outcome linked these 12q abnormalities to an increased risk of relapse in iAMP21-ALL.
Materials and methods
Patient and PDX material
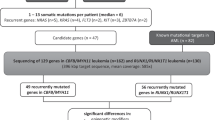
Bone marrow or peripheral blood diagnostic, matched remission and/or relapse samples were obtained from patients in the United Kingdom, Germany or the USA. Ethical approval and consent was obtained for all patients in accordance with the declaration of Helsinki. Cytogenetic results were reviewed by the Leukaemia Research Cytogenetics Group [21]. PDX were generated from stored viable bone marrow cells from patients 78, 72, 62 and 69 as previously described [22]. Patient samples were obtained as DNA or viable cells from the reference laboratories or the Bloodwise Childhood Leukaemia Cell Bank, UK. DNA was extracted from viable patient cells or from blasts isolated from PDX spleens and purified over FICOL, using the DNeasy Extraction kit (Qiagen, Manchester, UK). Study specific IDs and references, for patients with whole or partial overrepresentation of chromosome 21, that have been previously reported [4, 10, 11, 22, 23] are indicated in supplementary table S1.
Definition of chromosome 21 CN status
The majority of cases of iAMP21-ALL had been diagnosed on the basis of cytogenetic identification of an abnormal chromosome 21 and ≥3 additional RUNX1 FISH signals, with probes hybridising to the RUNX1 genomic region [24]. iAMP21 was confirmed in these cases from the characteristic chromosome 21 SNP 6.0 array profiles, as previously described [3]. In a minority of cases, where suitable material for cytogenetic or FISH analysis was unavailable or <3 extra copies of RUNX1 were present but flanking regions were more highly amplified, iAMP21 was identified by SNP 6.0 array CN profile alone (supplementary figure 4). Constitutional or somatically acquired aneuploidy 21 was identified by cytogenetic analysis and confirmed by SNP array as increased whole chromosome 21 CN. A minority of aneuploidies were defined on the basis chromosome 21 CN profile alone. Other cases, collectively referred to as other structural amplifications of 21 (OSA21), included those identified by cytogenetics as isochromosome 21q, i(21q), or isodicentric 21, idic(21), and less clearly defined abnormalities, which failed to meet our definitions of iAMP21 or aneuploidy. The techniques used to define chromosome 21 CN abnormalities are shown in Supplementary Table 1b for each case.
Techniques described in the supplementary methods
Detailed methods for; SNP6.0 and methylation array analysis, Sequencing of SH2B3, homology modelling, western blots, STAT and ERK activation assays and statistical analysis are provided in the supplementary methods.
Results
The landscape of CN-LOH in B-ALL patients with whole or partial gain of chromosome 21
From analysis of SNP6.0 arrays, we first determined the frequency and genomic location of CN-LOH in B-ALL patients with iAMP21-ALL (n = 49), OSA21-ALL (n = 11) and two (n = 5) or one (n = 34) extra entire copy of chromosome 21, either as a constitutional or acquired single aneuploidy or within a hyperdiploid or hypodiploid karyotype. Excluding two patients with multiple regions, that we attributed to consanguinity [25], five cases showed whole chromosome CN-LOH (wcCN-LOH), 27 involved interstitial regions (iCN-LOH) and 19 were terminal (tCN-LOH) (Fig. 1 and Supplementary Table 1a/b). In agreement with previous reports [16, 26], tCN-LOH were of somatic and iCN-LOH of germline origin as deduced from those cases with available matched, non-leukaemic samples. Notably, eight tCN-LOH involved the long arm of chromosome 12 (12q), six occurring in iAMP21-ALL and two in OSA21-ALL (Fig. 2a). Other recurrent regions involved the short arms of chromosomes 6 and 9 (6p, 9p) or whole chromosome 9 (wc9).
The landscape of CN-LOH in B-ALL with whole or partial overrepresentation of chromosome 21. Identification (ID) numbers and chromosome 21 CNA are shown in the left hand columns for patients carrying regions of CN-LOH identified by analysis of SNP 6.0 arrays. Chromosome numbers are indicated in the top panel and CN-LOH status for each patient by box colour as shown in the key. For patients with an available remission sample, the origin was defined as somatic (not present in remission) or germline (present in remission) with iCN-LOH always identified as germline and all but one tCN-LOH defined as somatic. Chromosomal position of CN-LOH is indicated by; p (segment/s of the short arm), q (segment/s of the long arm) pq (segments of both short and long arms but not the whole chromosome), wc (whole chromosome). Multiple regions of CN-LOH that were predominantly interstitial, seen in patients 41 and 93, are indicative of consanguinity. tCN-LOH 6p (n = 2), 9p (n = 4) and 12q (n = 8) and wcCN-LOH 9 (n = 2) were recurrent
Co-occurrence of 12q CN-LOH, amplification of chromosome 21 and deletion of SH2B3. a On the left SNP 6.0 genotype profiles show the relative signal intensity of the B allele probes for chromosome 12. Normal genomic regions are characterized by three levels of intensity corresponding to the AA, AB and BB genotypes and regions of CN-LOH by loss of the AB genotype. Arrows mark the proximal boundary of CN-LOH which were terminal in each case. To the right are shown patient IDs and chromosome 21 log 2 CN profiles, black lines mark the mean signal intensity for a CN of 2. All patients showed evidence for structural amplification of chromosome 21, as indicated, six abnormalities were defined as iAMP21. b Heat maps of CN for a region of 12q showing clonal deletion in two patients with 12q CN-LOH. Both deletions were bi-allelic with the smaller of the two confined to exons of SH2B3 and ATXN2. c Suggesting they target a common underlying genomic abnormality, allelotype and CN heat maps for a presentation sample from patient 78 (P 78) were consistent with the presence of competing sub-clones carrying 12q CN-LOH and a mono-allelic deletion that included the SH2B3/ATXN2 region. In a PDX derived from the same sample (PDX 78), CN-LOH was lost while the deletion became clonal. To the right a detailed view of dot-plots of SNP 6.0 CN within the deleted region, suggests the presence of a somatically acquired bi-allelic micro-deletion centred on exon 2 of SH2B3 in both patient and PDX samples but not the remission sample (R 78). Black lines indicate a copy number of 2. Arrows indicate the position of SH2B3 exon 2. d Sequence amplified from patient 78 using SH2B3 intron 1 and 2 primers that defines breakpoints for a 17.4 Kb deletion. e SNP6.0 CN heat maps of deletions that included the SH2B3 region found in three iAMP21 patients without CN-LOH 12q. In patient 61 the deletion is mono-allelic while in patients 3 and 72 nested deletions resulted in bi-allelic loss of SH2B3
tCN-LOH 12q is unique to a background of whole or partial amplification of chromosome 21
We then screened 203 B-ALL patients with normal chromosome 21 copy number (CN) and found tCN-LOH 9p in 10 cases but none involving 12q or 6p (Supplementary Table 2). To validate findings within our own institution, we investigated CN-LOH 12q and 6p among 974 patients from a broad range of B-ALL subtypes from six published studies (Table 1) [6, 15,16,17, 19, 20]. In patients with disclosed chromosome 21 CN status, 6p CN-LOH was associated with both partially amplified (n = 3) and apparently normal chromosomes 21 (n = 3). Among cases with CN-LOH 12q, one was described as iAMP21-ALL [17] and two co-occurred with amplifications of 21q to CN > 5 [15], thus were likely iAMP21-ALL. The remaining case harboured tetrasomy 21 [16]. So, taken together with our data, CN-LOH 12q was exclusive to a background of whole or partial chromosome 21 CN gain. The incidence, while only 0.9% among B-ALL patients overall, was 13% among our cohort of iAMP21/OSA21-ALL. The overall incidence of CN-LOH 6p was 1.1% and, not exclusive to, but possibly enriched in association with whole or partial gain of chromosome 21, as it occurred at a rate of 3% among our cases with and never without these abnormalities. A 70.3 Mb common region of 12q tCN-LOH was defined from our cases, with breakpoints clustered between the centromere and chromosome band 12q14 (Fig. 2a and Supplementary Table 1b).
CN-LOH 12q does not unmask an imprinted gene
To explore the possibility that CN-LOH caused unbalanced expression of an imprinted locus on 12q, we compared methylation array data from seven of the cases with CN-LOH and an equal number of iAMP21-ALL patients without evidence of a 12q abnormality. Specific genes; DCN, WIF1, FBRSL1 and E2F2, had been predicted to be imprinted and fell within the 12q common region of CN-LOH (www.geneimprint.com/Catalogue _of Parent of Origen Effects). However no evidence was found for differential methylation, between the two groups, of probes associated with these genes (Supplementary Figure 1). Furthermore analysis of the entire 12q common region of CN-LOH revealed no significant blocks of differential methylation between cases with and without CN-LOH. Therefore we concluded that CN-LOH 12q was not driven by loss of imprinting in these cases.
Deletions of the SH2B3 gene are coincident with CN-LOH 12q
We next investigated regions of CN-LOH 12q for coincident CNA and identified two with clonal deletions that focused attention on a region containing SH2B3 and ATXN2 (Fig. 2b and Supplementary table 3). In a third patient, and the patient-derived xenografts (PDX), sub-clones with 12q deletion or CN-LOH fluctuated in level, suggesting that competing sub-clones targeted a lesion on the second allele through different mechanisms. Close examination of presentation and PDX array data revealed evidence for an intragenic deletion of SH2B3 centred on exon 2, not present in a remission sample (Fig. 2c). Loss of exon 2 was confirmed by PCR amplification of sequence from this sample, but not a normal control, using SH2B3 intron 1 and 2 primers, which defined the breakpoints of a 17.4 Kb deletion (Fig. 2d). Albeit at low frequencies, mutations and CNAs of SH2B3 have previously been described in B-ALL [27,28,29,30], so we further investigated this gene in iAMP21 and other sub-types of ALL.
Deletions of SH2B3 in the absence of CN-LOH are recurrent in iAMP21-ALL
We then analysed SNP6.0 arrays for 12q CN status in patients from our cohort with or without whole or partial chromosome 21 gain, without 12q CN-LOH. We identified a clonal mono-allelic and two bi-allelic deletions affecting SH2B3 in three cases of iAMP21-ALL (Fig. 2e and Table S3). Deletion of SH2B3 was never seen outside iAMP21-ALL. The combined incidence of 12q CN-LOH and/or deletion of the region including SH2B3 was 9/49 (18%) in iAMP21-ALL, in marked contrast to non-iAMP21-ALL, where among 253 patients only two were seen (0.8%) and both cases carried OSA21. The association between amplification of chromosome 21 and CN-LOH/deletion 12q was highly significant in our cohort (p ≤ 0.01). To investigate a larger cohort with less bias towards chromosome 21 abnormalities, we screened publicly available SNP array data from 648 presentation B-ALL [28], identifying just two with SH2B3 deletions, one with tetrasomy 21 and the other with no recorded evidence of a chromosome 21 CN abnormality (Supplementary Figure 2 and Supplementary table 4). These data confirmed that, in the absence of iAMP21/OSA21, deletion of the SH2B3 region is extremely rare in B-ALL.
Mutations of SH2B3 are associated with CN-LOH and heterozygous deletion of 12q in iAMP21-ALL
Although rare, mutations and deletions of SH2B3 have previously been described in B-ALL [27,28,29,30,31,32,33], T-ALL [27, 34] and a range of myeloid neoplasms (reviewed in [35]). In addition, polymorphisms of SH2B3 have been associated with skewed haematological parameters and predisposition to autoimmune disease or myeloid malignancy, the most common of these is rs3184504, which results in a R262W substitution that affects PH domain function [36, 37]. To investigate involvement of polymorphisms or acquired mutations, we carried out Illumina sequencing of coding exons of SH2B3 in patients with CN-LOH and/or deletion of 12q (n = 9), as well as iAMP21-ALL patients without identified 12q abnormalities (n = 6) (Table 2). In four cases with bi-allelic deletion, as expected, no sequence abnormalities were found, but frameshift mutations accompanied 12q CN-LOH without deletion (n = 2) or mono-allelic deletion without 12q CN-LOH (n = 1). Of two remaining patients with 12q CN-LOH, one carried an R392W polymorphism (rs770836648). This variant, which has a reported frequency of 0.000016 in normal populations [38], was also acquired in two patients with myeloproliferative disorders [39]. In our case, R392W was evidently of somatic origin because, although no remission sample was available, it was identified in 49% of reads, while allelic bias due to CN-LOH was 74%, as indicated by the read count for the rs3184504 C allele. Based on the crystal structure of the mouse SH2B1 SH2 domain [40], we constructed a model of the equivalent human SH2B3 sequence. This model strongly predicted that substitution of the aromatic ring of tryptophan for the amino group of the conserved 392 arginine residue would disrupt normal interaction with the autophosphorylation site of JAK2 (Fig. 3a–d) and likely equivalent sites on other kinases.
Impact of the R392W substitution and other SH2B3 abnormalities. a 104 amino acid sequence comprising 92% of the SH2 domain of human SH2B3 aligned to the mouse SH2B1 sequence. Black arrows indicate three arginine residues required to coordinate pTyr813 of JAK2 within the mouse SH2B1 SH2 binding pocket. Red arrow shows the position of the R392W arginine to tryptophan substitution identified in patient 25. b Ribbon diagram based on the published co-crystalized structure of the mouse SH2B1 SH2 domain and an eleven residue phosphopeptide surrounding tyrosine pTyr813 within the activation loop of JAK2 [40]. The structure has a canonical SH2 domain architecture with a central β-sheet (magenta) flanked by two α-helices (cyan) that form two binding pockets for the JAK2 phosphopeptide (shown as a stick representation with carbon atoms in green, oxygen atoms in red, nitrogen atoms in dark blue and phosphate atoms in orange). c A similar canonical SH2 domain structure is seen in a homology model of 104 residues of the SH2 domain of human SH2B3 based on the mouse SH2B1 crystal structure. Overall sequence identity between the two SH2 domains was 71%, with a structural homology of 1 Å RMSD when superposed. d Detailed view of superposed models of the phosphotyrosine binding pocket of SH2 domains of mouse SH2B1 (grey ribbons) and human SH2B3 (magenta ribbons) demonstrating close structural homology for this region. The JAK2 phosphopeptide and three arginine residues (R534, R555 and R560) that coordinate pTyr813 within the binding pocket through salt bridges (broken black lines) are indicated by stick representations, with carbon atoms of arginine in grey and other atoms coloured as in 3B. The R392W substitution in human SH2B3 is also shown as a stick representation with carbon atoms coloured in magenta. Tryptophan in this position is unable to form salt bridges with pTyr813, the R392W substitution therefore results in loss of two of four critical interactions with phosphotyrosine and is predicted to destabilise binding of SH2B3 to activated JAK. e Summary of abnormalities of SH2B3 identified in this study. At the top the SH2B3 linear protein structure is represented with major functional domains marked; DIME (dimerization), PH (Pleckstrin homology), SH2 (Src homology 2). For each patient, genomic abnormalities of the SH2B3 coding region, as indicated in the key, are aligned with the domain representation. Numbers on the left are patient ids, genomic position of CN-LOH and deletion are indicated on the right. The codon position and consequence of somatically acquired mutations are as labelled, with arrows marking their positions on the SH2B3 protein. Through combinations of mutation, deletion and CN-LOH, SH2B3 abnormalities were all homozygous and predicted to result in absence of a coding transcript (patients 3, 12, 28, 72 and 78), production of truncated and likely unstable transcripts / proteins (patients 44, 61 and 88) or production of a protein with functionally impaired SH2 domain (patient 25)
Lastly, we postulated a predisposing role for R262W, as re3184504 was homozygous for the T allele in patient 45, the only case where 12q CN-LOH but no SH2B3 mutation or deletion was identified. We also noted that rs3184504 was never heterozygous in the six patients without 12q abnormalities. However, homozygosity of R262W in patient 45 was unrelated to 12q CN-LOH, as further sequencing demonstrated it was also homozygous in germline tissue and sequencing of 16 additional iAMP21-ALL cases identified the T/T, T/C and C/C genotypes in two, nine and five patients, respectively. Thus it is it unlikely that rs3184504 contributes to iAMP21-ALL and we saw no evidence for selection of other inherited SH2B3 variants.
Collectively, CN and sequencing data linked 12q CN-LOH to bi-allelic loss of SH2B3 function, in all but one case (Fig. 3e) and, although rare in B-ALL overall, we reasoned that the incidence of SH2B3 abnormalities was highly elevated in iAMP21-ALL. This finding prompted us to examine SH2B3 abnormalities previously reported in B-ALL in the context of information relating to chromosome 21 CN status (Table 3). Remarkably, we discovered that in the two largest series described to date [28, 31], among 16 cases with SH2B3 abnormalities, 12 carried previously unreported iAMP21/OSA21 and four had gain of WC21 (personal communications from M. Devidas on behalf of the Children’s Oncology Group, USA). In other studies [27, 29, 30, 32, 41], we found only a single example where chromosome 21 CN status was defined and an SH2B3 abnormality was not associated with iAMP21/OSA21/wc21 gain. This case was also exceptional because an SH2B3 mutation was inherited and unmasked by germline iCN-LOH [27].
STAT5 is activated by IL7 stimulation in iAMP21 cells with SH2B3 deletion
In two studies, SH2B3 abnormalities co-occurred recurrently with potentially synergistic IL7R and/or FLT3 activating mutations [28, 31], prompting us to investigate IL7R/JAK/STAT, FLT3/STAT and FLT3/RAS/RAF/ERK signalling pathways in iAMP21-ALL. PDX cells were used for this purpose as diagnostic patient cells were not available. We first demonstrated that SH2B3 protein was expressed in cells derived from two patients without a 12q abnormality, but not from cells derived from a patient carrying a bi-allelic deletion of SH2B3. On stimulation with IL7, we observed a clear dose dependent activation of STAT5, but not STAT1, 3, 4 or 6, in cells with SH2B3 inactivation. In contrast, IL7 stimulation of cells without evidence of SH2B3 inactivation showed no significant response to any STAT, while FLT3 ligand failed to activate either STATs or ERK1/2 in any of the cells tested (Fig. 4 and Supplementary Figure 3).
IL7 stimulates activation of STAT5 in iAMP21-ALL cells with SH2B3 deletion. a Immuno-blot with anti-SH2B3 confirmed loss of expression in PDX of patient 72 that carry a bi-allelic deletion of SH2B3. The full length protein was detected in patients 62 and 69 PDX that showed no evidence for SH2B3 CN or coding sequence abnormalities. MWM (molecular weight marker) with 70 Kd band shown. b Treatment of PDX cells carrying SH2B3 bi-allelic deletion with IL7 resulted in a dose related increase in STAT5 phosphorylation (four flow cytometric immunophenotyping experiments with cells from patient 72) *(P < 0.01) **(P < 0.05). c Analysis of PDX cells expressing wild-type SH2B3 failed to demonstrate any significant increase in STAT5 phosphorylation with IL7 treatment (four flow cytometric immunophenotyping experiments with cells from patient 62 [n = 2] and 69 [n = 2]). P values are for paired t-tests, error bars are standard error of the mean.
12q abnormalities may predict poor outcome in iAMP21-ALL
We were able to investigate the outcome of 26 iAMP21-ALL patients from this study who had been treated on UKALL97/99 (n = 5) or UKALL2003 (n = 21) clinical trials [1, 42]. After a median follow-up time of >5 years, there was evidence of an association with outcome: 5/7 (71%) patients with CN-LOH or deletion of 12q had relapsed and/or died compared with 4/19 (21%) patients without 12q abnormality (p = 0.03). Importantly, iAMP21 patients treated on ALL97/99 had a significantly higher rate of relapse compared with those treated on ALL2003 (70% vs. 16% at 5 years) due to the intensification of treatment received by these patients on UKALL2003 [1]. While prognosis overall of iAMP21 patients treated on ALL97 was worse than in UKALL 2003, there remained a strong association with 12q abnormality across both trials. This is illustrated by the fact that none of the three ALL 97 treated patients without 12q abnormality relapsed while 3/5 ALL 2003 patients with the abnormality relapsed/died.
Discussion
Supporting some of our key findings, two previous reports noted an association between gross abnormalities of 12q affecting the SH2B3 region and iAMP21-ALL [41, 43]. However, these studies were inconclusive as sequencing was not performed to confirm the involvement of SH2B3. Also the numbers of iAMP21-ALL cases analysed were insufficient to accurately assess the overall incidence, proportions affected by different types of lesions or influence on outcome. We studied a larger cohort of patients, in which we defined underlying SH2B3 sequence abnormalities and performed preliminary analysis of outcome and also extended our investigation to include published data. We first established that in B-ALL, tCN-LOH 12q was exclusive to B-ALL with iAMP21/OSA21 or, at low incidence, with additional copies of wc21. Although existence of a second target for CN-LOH 12q cannot be formally ruled out, sequencing and copy number analysis identified bi-allelic inactivation of SH2B3 as one consequence of these abnormalities. As further support for SH2B3 as a relevant target, we showed the same pattern of high incidence in association with iAMP21-ALL and low incidence with aneuploidy 21 when SH2B3 was inactivated in the absence of CN-LOH 12q. Abnormality affecting SH2B3 without any identifiable whole/partial chromosome 21 gain was extremely rare, with only a single case reported within the literature and a second identified through analysis of archival array data.
In our patients, all SH2B3 lesions were bi-allelic, with CN-LOH being the dominant mechanism and deletion less frequently involved in conversion to homozygosity. We concluded that high copy number gain of specific sequences on chromosome 21, as seen in iAMP21-ALL, confers strong pressure for complete loss of SH2B3 function, while gain of one or two copies is associated with a similar though weaker selective environment. The absence of reported tCN-LOH 12q, in B-ALL without chromosome 21 gain, may be explained by the extreme rarity of SH2B3 lesions among these cases. In haematological malignancies other than B-ALL, abnormalities of SH2B3 have not been associated with 12q CN-LOH, suggesting less selective pressure for homozygous inactivation. Consistent with lineage specific tumour suppressor thresholds, Tp53-/- SH2B3± mice developed B-ALL in a manner that was strictly dependent on loss of the wild type SH2B3 allele [44], while SH2B3± mice developed a milder form of the myeloproliferative disease seen in SH2B3 -/- mice [45]. We also note that in B-ALL, SH2B3 abnormalities were mostly whole gene deletions or truncating mutations, while in myeloid malignancies, substitutions affecting the PH domain predominated (reviewed in [35]).
SH2B3 is one of a three member family of SH2B adaptor proteins that share common motifs, including N-terminal dimerization, pleckstrin homology (PH) and Src homology 2 (SH2) domains [46]. Of the three, SH2B3 has the most prominent role in haematopoietic development, where it negatively regulates various receptor and non-receptor kinases, through direct interaction of the SH2 domain with phosphorylated tyrosine (reviewed in [47]). Critically, it functions in negative feedback loops controlling cell growth, development and survival signals because activated target kinases also induce SH2B3 expression and activation through phosphorylation [44, 48, 49]. Our data clearly pose the question of why SH2B3 loss of function should confer a strong selective advantage to B-ALL, specifically when certain chromosome 21 sequences are highly amplified through complex rearrangements. The mechanism by which iAMP21 promotes B-ALL is currently unknown and, in the absence of representative cell lines or genetically engineered animal models, challenging to investigate. Although usually defined at the cytogenetic level by ≥3 additional copies of RUNX1, there is little evidence that RUNX1 overexpression drives iAMP21-ALL. Despite higher RUNX1 copy number in iAMP21-ALL, it was not shown to be over-expressed relative to high hyperdiploid-ALL [23]. Furthermore B-ALL chromosome 21 copy number profiles are occasionally iAMP21-like, but not highly amplified for RUNX1 compared with flanking sequences (Supplementary Figure 4). A second gene in the region, DYRK1A [3], is more consistently amplified and has been implicated in multiple signalling pathways contributing to cell growth and survival and cancer (reviewed in [50]). In common with B-ALL, the incidence of acute megakaryoblastic leukaemia (AMKL) is highly elevated in individuals with DS and Dyrk1A dosage increase was shown to co-operate with Gata1 and Mpl mutations to promote AMKL [51]. Interestingly, although we found no examples of SH2B3 abnormalities in DS-ALL, they were reported in DS-AMKL but not non-DS-AMKL [52, 53]. Therefore also in this highly specific haematological malignancy, SH2B3 abnormalities may be preferentially selected on a background of chromosome 21 gain, in this instance with DYRK1A amplification demonstrated to be relevant. Although it has not been shown that DYRK1A amplification contributes to iAMP21-ALL, it does play a critical role in the large to small pre-B cell transition stage, where it induces quiescence through destabilisation of cyclin D3. However, while pre-B cells in Dyrk1a deficient mice had elevated levels of cyclin D3, they also displayed impaired proliferation, suggesting involvement in other, cyclin D3 independent, aspects of B-cell differentiation [54]. In iAMP21-ALL, co-operating abnormalities, such as loss of SH2B3 function, might therefore promote leukaemia by fine-tuning a mixture of oncogenic and tumour suppressor signals orchestrated by amplified DYRK1A. Like DYRK1A, GSK-3β is a multifunctional serine/threonine kinase implicated in diverse signalling pathways and both tumour promoting and tumour suppressor roles (reviewed in [55]). Interactions between GSK-3β and DYRK1A signalling almost certainly occur in in B cells; for instance GSK-3β also targets cyclin D for degradation [56] and is essential for the phosphorylation and nuclear exclusion of NFAT transcription factors, an event primed by DYRK1A and implicated as an important consequence of increased wc21 CN in DS-AMKL [51]. GSK-3β activity is regulated by posttranslational phosphorylation and in endothelial cells phosphorylation mediated by integrin was shown to depend on SH2B3 [57]. Loss of SH2B3 function could therefore modulate signals orchestrated by increased dose of DYRK1A through altered GSK-3β activity, although it remains to be determined whether this contributes to B-ALL.
Another likely arena for cooperation between abnormalities of chromosomes 21 and 12 involves the JAK/STAT pathway, because DYRK1A has been reported to phosphorylate STAT3 [58] and SH2B3 has been shown to negatively regulate JAK signalling [27, 44]. The only non-frameshift mutation identified among our cases was demonstrated, by homology modelling, to result in loss of interactions that repress activated JAK. Moreover, we found that iAMP21-ALL PDX cells with SH2B3 bi-allelic deletions were more sensitive to IL7 induced STAT5 activation than those with wild type SH2B3. Evidence is also accumulating that in B-ALL, loss of SH2B3 function cooperates with abnormalities in addition to those of chromosome 21. Further implicating the JAK/STAT pathway, IL7R activating mutations were report in 8 of 16 cases with SH2B3 abnormalities [28, 31]. However the emerging picture is complex because other JAK-STAT pathway activating abnormalities appear to correlate negatively with SH2B3 inactivation. For example JAK mutations were absent and only a single CRLF2 activating rearrangement co-occurred with an SH2B3 abnormality [28]. Among our iAMP21 patients tested for PAR1 deletions, that result in the P2YR8-CRLF2 and consequent CRLF2 over-expression, none were apparent in cases with (n = 7), but they were seen in 5 out of 26 cases (19%) without 12q abnormalities (data not shown). It may be that a background chromosome 21 gain promotes selection for activation of JAK/STAT, but not beyond a certain threshold. Cross talk between other pathways influenced by the various JAK/STAT activating rearrangements, may also favour selection of certain combinations but not others. SH2B3 is known to interact with other receptor and non-receptor kinases, such as FLT3 and LCK, that play a role in early B-cell development [49, 59]. We were unable to demonstrate ERK or STAT activation in SH2B3 deleted PDX cells stimulated with FLT3 ligand nevertheless abnormalities activating FLT3 appear to be co-selected with those inactivating SH2B3 [28, 31] and a role in dysregulation of the RAS/RAF/MEK/ERK pathway should not be discounted. Although JAK-STAT and RAS pathway activation clearly contributes to iAMP21-ALL [11], these pathways are also commonly activated in patients without chromosome 21 abnormalities. The near-exclusive relationship we observed may therefore be driven by other SH2B3 interactions involving pathways dysregulated by high expression of DYRK1A and/or other genes amplified through iAMP21-ALL rearrangements. Notwithstanding these considerations, CN-LOH/deletion 12q may function as useful markers for cases that will potentially respond to JAK/STAT pathway inhibitors, indeed blast counts were reduced in PDX from one patient with IL7R activating mutation and SH2B3 deletion treated with ruxolitinib [33].
Importantly, we showed 12q abnormalities to be associated with an increased rate of treatment failure in iAMP21-ALL. Given the number of patients involved and level of significance, this finding is preliminary, but if confirmed in larger studies, may have important implications as iAMP21-ALL patients are classified as high risk [1] and currently at risk of the adverse consequences of intensive chemotherapy. Lastly further analysis of the tumour suppressor role of SH2B3 in B-ALL may help to define critical signalling events elicited by increased doses of chromosome 21 gene expression in iAMP21-ALL, as well as other important sub-types with gain of wc21.
References
Moorman AV, Robinson H, Schwab C, Richards SM, Hancock J, Mitchell CD, et al. Risk-directed treatment intensification significantly reduces the risk of relapse among children and adolescents with acute lymphoblastic leukemia and intrachromosomal amplification of chromosome 21: a comparison of the MRC ALL97/99 and UKALL2003 trials. J Clin Oncol. 2013;31:3389–96.
Heerema NA, Carroll AJ, Devidas M, Loh ML, Borowitz MJ, Gastier-Foster JM, et al. Intrachromosomal amplification of chromosome 21 is associated with inferior outcomes in children with acute lymphoblastic leukemia treated in contemporary standard-risk children’s oncology group studies: a report from the children’s oncology group. J Clin Oncol. 2013;31:3397–402.
Li Y, Schwab C, Ryan SL, Papaemmanuil E, Robinson HM, Jacobs P, et al. Constitutional and somatic rearrangement of chromosome 21 in acute lymphoblastic leukaemia. Nature. 2014;508:98–102.
Rand V, Parker H, Russell LJ, Schwab C, Ensor H, Irving J, et al. Genomic characterization implicates iAMP21 as a likely primary genetic event in childhood B-cell precursor acute lymphoblastic leukemia. Blood. 2011;117:6848–55.
Safavi S, Paulsson K. Near-haploid and low-hypodiploid acute lymphoblastic leukemia: two distinct subtypes with consistently poor prognosis. Blood. 2017;129:420–3.
Paulsson K, Forestier E, Lilljebjorn H, Heldrup J, Behrendtz M, Young BD, et al. Genetic landscape of high hyperdiploid childhood acute lymphoblastic leukemia. Proc Natl Acad Sci USA. 2010;107:21719–24.
Hertzberg L, Betts DR, Raimondi SC, Schafer BW, Notterman DA, Domany E, et al. Prediction of chromosomal aneuploidy from gene expression data. Genes Chromosomes Cancer. 2007;46:75–86.
Hasle H, Clemmensen IH, Mikkelsen M. Risks of leukaemia and solid tumours in individuals with Down’s syndrome. Lancet. 2000;355:165–9.
Maloney KW. Acute lymphoblastic leukaemia in children with Down syndrome: an updated review. Br J Haematol. 2011;155:420–5.
Harrison CJ, Moorman AV, Schwab C, Carroll AJ, Raetz EA, Devidas M, et al. An international study of intrachromosomal amplification of chromosome 21 (iAMP21): cytogenetic characterization and outcome. Leukemia. 2014;28:1015–21.
Ryan SL, Matheson E, Grossmann V, Sinclair P, Bashton M, Schwab C, et al. The role of the RAS pathway in iAMP21-ALL. Leukemia. 2016;30:1824–31.
Schwab CJ, Chilton L, Morrison H, Jones L, Al-Shehhi H, Erhorn A, et al. Genes commonly deleted in childhood B-cell precursor acute lymphoblastic leukemia: association with cytogenetics and clinical features. Haematologica. 2013;98:1081–8.
Buitenkamp TD, Izraeli S, Zimmermann M, Forestier E, Heerema NA, van den Heuvel-Eibrink MM, et al. Acute lymphoblastic leukemia in children with Down syndrome: a retrospective analysis from the Ponte di Legno study group. Blood. 2014;123:70–77.
Paulsson K, Lilljebjorn H, Biloglav A, Olsson L, Rissler M, Castor A, et al. The genomic landscape of high hyperdiploid childhood acute lymphoblastic leukemia. Nat Genet. 2015;47:672–6.
Kawamata N, Ogawa S, Zimmermann M, Kato M, Sanada M, Hemminki K, et al. Molecular allelokaryotyping of pediatric acute lymphoblastic leukemias by high-resolution single nucleotide polymorphism oligonucleotide genomic microarray. Blood. 2008;111:776–84.
Lundin KB, Olsson L, Safavi S, Biloglav A, Paulsson K, Johansson B. Patterns and frequencies of acquired and constitutional uniparental isodisomies in pediatric and adult B-cell precursor acute lymphoblastic leukemia. Genes Chromosomes Cancer. 2016;55:472–9.
Baughn LB, Biegel JA, South ST, Smolarek TA, Volkert S, Carroll AJ, et al. Integration of cytogenomic data for furthering the characterization of pediatric B-cell acute lymphoblastic leukemia: a multi-institution, multi-platform microarray study. Cancer Genet. 2015;208:1–18.
Kuiper RP, Schoenmakers EF, van Reijmersdal SV, Hehir-Kwa JY, van Kessel AG, van Leeuwen FN, et al. High-resolution genomic profiling of childhood ALL reveals novel recurrent genetic lesions affecting pathways involved in lymphocyte differentiation and cell cycle progression. Leukemia. 2007;21:1258–66.
Ninomiya S, Tyybakinoja A, Borze I, Raty R, Saarinen-Pihkala UM, Usvasalo A, et al. Integrated analysis of gene copy number, copy neutral LOH, and microRNA profiles in adult acute lymphoblastic leukemia. Cytogenet Genome Res. 2012;136:246–55.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD, et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature. 2007;446:758–64.
Moorman AV, Ensor HM, Richards SM, Chilton L, Schwab C, Kinsey SE, et al. Prognostic effect of chromosomal abnormalities in childhood B-cell precursor acute lymphoblastic leukaemia: results from the UK Medical Research Council ALL97/99 randomised trial. Lancet Oncol. 2010;11:429–38.
Sinclair PB, Blair HH, Ryan SL, Buechler L, Cheng J, Clayton J, et al. Dynamic clonal progression in xenografts of acute lymphoblastic leukemia with intrachromosomal amplification of chromosome 21. Haematologica. 2018;103:634–44.
Strefford JC, van Delft FW, Robinson HM, Worley H, Yiannikouris O, Selzer R, et al. Complex genomic alterations and gene expression in acute lymphoblastic leukemia with intrachromosomal amplification of chromosome 21. Proc Natl Acad Sci USA. 2006;103:8167–72.
Harrison CJ, Haas O, Harbott J, Biondi A, Stanulla M, Trka J, et al. Detection of prognostically relevant genetic abnormalities in childhood B-cell precursor acute lymphoblastic leukaemia: recommendations from the Biology and Diagnosis Committee of the International Berlin-Frankfurt-Munster study group. Br J Haematol. 2010;151:132–42.
Kearney HM, Kearney JB, Conlin LK. Diagnostic implications of excessive homozygosity detected by SNP-based microarrays: consanguinity, uniparental disomy, and recessive single-gene mutations. Clin Lab Med. 2011;31:595–613. ix
O’Keefe C, McDevitt MA, Maciejewski JP. Copy neutral loss of heterozygosity: a novel chromosomal lesion in myeloid malignancies. Blood. 2010;115:2731–9.
Perez-Garcia A, Ambesi-Impiombato A, Hadler M, Rigo I, LeDuc CA, Kelly K, et al. Genetic loss of SH2B3 in acute lymphoblastic leukemia. Blood. 2013;122:2425–32.
Roberts KG, Li Y, Payne-Turner D, Harvey RC, Yang YL, Pei D, et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N Engl J Med. 2014;371:1005–15.
Olsson L, Albitar F, Castor A, Behrendtz M, Biloglav A, Paulsson K, et al. Cooperative genetic changes in pediatric B-cell precursor acute lymphoblastic leukemia with deletions or mutations of IKZF1. Genes Chromosomes Cancer. 2015;54:315–25.
Lindqvist CM, Lundmark A, Nordlund J, Freyhult E, Ekman D, Almlof JC, et al. Deep targeted sequencing in pediatric acute lymphoblastic leukemia unveils distinct mutational patterns between genetic subtypes and novel relapse-associated genes. Oncotarget. 2016;7:64071–88.
Reshmi SC, Harvey RC, Roberts KG, Stonerock E, Smith A, Jenkins H, et al. Targetable kinase gene fusions in high-risk B-ALL: a study from the Children’s Oncology Group. Blood. 2017;129:3352–61.
Russell LJ, Jones L, Enshaei A, Tonin S, Ryan SL, Eswaran J, et al. Characterisation of the genomic landscape of CRLF2-rearranged acute lymphoblastic leukemia. Genes Chromosomes Cancer. 2017;56:363–72.
Roberts KG, Morin RD, Zhang J, Hirst M, Zhao Y, Su X, et al. Genetic alterations activating kinase and cytokine receptor signaling in high-risk acute lymphoblastic leukemia. Cancer Cell. 2012;22:153–66.
Zhang J, Ding L, Holmfeldt L, Wu G, Heatley SL, Payne-Turner D, et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature. 2012;481:157–63.
Maslah N, Cassinat B, Verger E, Kiladjian JJ, Velazquez L. The role of LNK/SH2B3 genetic alterations in myeloproliferative neoplasms and other hematological disorders. Leukemia. 2017;31:1661–70.
Soranzo N, Spector TD, Mangino M, Kuhnel B, Rendon A, Teumer A, et al. A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nat Genet. 2009;41:1182–90.
Lesteven E, Picque M, Conejero Tonetti C, Giraudier S, Varin-Blank N, Velazquez L, et al. Association of a single-nucleotide polymorphism in the SH2B3 gene with JAK2V617F-positive myeloproliferative neoplasms. Blood. 2014;123:794–6.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P, et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood. 2013;122:3616–27.
Hu J, Hubbard SR. Structural basis for phosphotyrosine recognition by the Src homology-2 domains of the adapter proteins SH2-B and APS. J Mol Biol. 2006;361:69–79.
Ivanov Ofverholm I, Tran AN, Olsson L, Zachariadis V, Heyman M, Rudd E, et al. Detailed gene dose analysis reveals recurrent focal gene deletions in pediatric B-cell precursor acute lymphoblastic leukemia. Leuk Lymphoma. 2016;57:2161–70.
Moorman AV, Richards SM, Robinson HM, Strefford JC, Gibson BE, Kinsey SE, et al. Prognosis of children with acute lymphoblastic leukemia (ALL) and intrachromosomal amplification of chromosome 21 (iAMP21). Blood. 2007;109:2327–30.
Baughn LB, Meredith MM, Oseth L, Smolarek TA, Hirsch B. SH2B3 aberrations enriched in iAMP21 B lymphoblastic leukemia. Cancer Genet. 2018;226–7:30–35.
Cheng Y, Chikwava K, Wu C, Zhang H, Bhagat A, Pei D, et al. LNK/SH2B3 regulates IL-7 receptor signaling in normal and malignant B-progenitors. J Clin Invest. 2016;126:1267–81.
Velazquez L, Cheng AM, Fleming HE, Furlonger C, Vesely S, Bernstein A, et al. Cytokine signaling and hematopoietic homeostasis are disrupted in Lnk-deficient mice. J Exp Med. 2002;195:1599–611.
Rudd CE. Lnk adaptor: novel negative regulator of B cell lymphopoiesis. Sci STKE. 2001;2001:pe1.
Gery S, Koeffler HP. Role of the adaptor protein LNK in normal and malignant hematopoiesis. Oncogene. 2013;32:3111–8.
Baran-Marszak F, Magdoud H, Desterke C, Alvarado A, Roger C, Harel S, et al. Expression level and differential JAK2-V617F-binding of the adaptor protein Lnk regulates JAK2-mediated signals in myeloproliferative neoplasms. Blood. 2010;116:5961–71.
Lin DC, Yin T, Koren-Michowitz M, Ding LW, Gueller S, Gery S, et al. Adaptor protein Lnk binds to and inhibits normal and leukemic FLT3. Blood. 2012;120:3310–7.
Fernandez-Martinez P, Zahonero C, Sanchez-Gomez P. DYRK1A: the double-edged kinase as a protagonist in cell growth and tumorigenesis. Mol Cell Oncol. 2015;2:e970048.
Malinge S, Bliss-Moreau M, Kirsammer G, Diebold L, Chlon T, Gurbuxani S, et al. Increased dosage of the chromosome 21 ortholog Dyrk1a promotes megakaryoblastic leukemia in a murine model of Down syndrome. J Clin Invest. 2012;122:948–62.
Yoshida K, Toki T, Okuno Y, Kanezaki R, Shiraishi Y, Sato-Otsubo A, et al. The landscape of somatic mutations in Down syndrome-related myeloid disorders. Nat Genet. 2013;45:1293–9.
de Rooij JD, Branstetter C, Ma J, Li Y, Walsh MP, Cheng J, et al. Pediatric non-Down syndrome acute megakaryoblastic leukemia is characterized by distinct genomic subsets with varying outcomes. Nat Genet. 2017;49:451–6.
Thompson BJ, Bhansali R, Diebold L, Cook DE, Stolzenburg L, Casagrande AS, et al. DYRK1A controls the transition from proliferation to quiescence during lymphoid development by destabilizing Cyclin D3. J Exp Med. 2015;212:953–70.
Nagini S, Sophia J, Mishra R. Glycogen synthase kinases: moonlighting proteins with theranostic potential in cancer. Semin Cancer Biol 2018. https://doi.org/10.1016/j.semcancer.2017.12.010
Naderi S, Gutzkow KB, Lahne HU, Lefdal S, Ryves WJ, Harwood AJ, et al. cAMP-induced degradation of cyclin D3 through association with GSK-3beta. J Cell Sci. 2004;117(Pt 17):3769–83.
Devalliere J, Chatelais M, Fitau J, Gerard N, Hulin P, Velazquez L, et al. LNK (SH2B3) is a key regulator of integrin signaling in endothelial cells and targets alpha-parvin to control cell adhesion and migration. FASEB J. 2012;26:2592–606.
Matsuo R, Ochiai W, Nakashima K, Taga T. A new expression cloning strategy for isolation of substrate-specific kinases by using phosphorylation site-specific antibody. J Immunol Methods. 2001;247:141–51.
Li Y, He X, Schembri-King J, Jakes S, Hayashi J. Cloning and characterization of human Lnk, an adaptor protein with pleckstrin homology and Src homology 2 domains that can inhibit T cell activation. J Immunol. 2000;164:5199–206.
Acknowledgements
The Authors would like to acknowledge funding by the European Research Council (grant # 249891), Bloodwise (grant #s 11004 and 15036), the Kay Kendal Leukemia Fund and Newcastle University for funding a visiting professorship to BYD. Core infrastructure support was provided by the North of England Children’s Cancer Research Fund (NECCR). The authors would like to thank; all UK contributing laboratories for providing DNA, cytogenetic data and demographic information for leukaemia patients included in the study, the Bloodwise Childhood Leukaemia Cell Bank for providing viable leukaemia patient samples and the Childhood Oncology Group for providing additional cytogenetic information for published cases of B-ALL.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sinclair, P.B., Ryan, S., Bashton, M. et al. SH2B3 inactivation through CN-LOH 12q is uniquely associated with B-cell precursor ALL with iAMP21 or other chromosome 21 gain. Leukemia 33, 1881–1894 (2019). https://doi.org/10.1038/s41375-019-0412-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0412-1
This article is cited by
-
Whole genome sequencing provides comprehensive genetic testing in childhood B-cell acute lymphoblastic leukaemia
Leukemia (2023)
-
Disruption to the FOXO-PRDM1 axis resulting from deletions of chromosome 6 in acute lymphoblastic leukaemia
Leukemia (2023)
-
Adaptor protein LNK promotes anaplastic thyroid carcinoma cell growth via 14-3-3 ε/γ binding
Cancer Cell International (2020)
-
Gain of chromosome 21 in hematological malignancies: lessons from studying leukemia in children with Down syndrome
Leukemia (2020)
-
Parental origin of monosomic chromosomes in near-haploid acute lymphoblastic leukemia
Blood Cancer Journal (2020)