Abstract
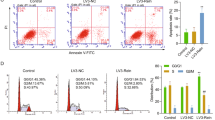
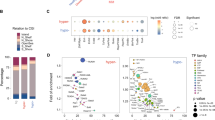
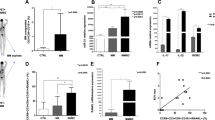
Myeloma bone disease is a devastating complication of multiple myeloma (MM) and is caused by dysregulation of bone remodeling processes in the bone marrow microenvironment. Previous studies showed that microRNA-138 (miR-138) is a negative regulator of osteogenic differentiation of mesenchymal stromal cells (MSCs) and that inhibiting its function enhances bone formation in vitro. In this study, we explored the role of miR-138 in myeloma bone disease and evaluated the potential of systemically delivered locked nucleic acid (LNA)-modified anti-miR-138 oligonucleotides in suppressing myeloma bone disease. We showed that expression of miR-138 was significantly increased in MSCs from MM patients (MM-MSCs) and myeloma cells compared to those from healthy subjects. Furthermore, inhibition of miR-138 resulted in enhanced osteogenic differentiation of MM-MSCs in vitro and increased the number of endosteal osteoblastic lineage cells (OBCs) and bone formation rate in mouse models of myeloma bone disease. RNA sequencing of the OBCs identified TRPS1 and SULF2 as potential miR-138 targets that were de-repressed in anti-miR-138-treated mice. In summary, these data indicate that inhibition of miR-138 enhances bone formation in MM and that pharmacological inhibition of miR-138 could represent a new therapeutic strategy for treatment of myeloma bone disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Manier S, Salem KZ, Park J, Landau DA, Getz G, Ghobrial IM. Genomic complexity of multiple myeloma and its clinical implications. Nat Rev Clin Oncol. 2017;14:100–13.
Arnulf B, Lecourt S, Soulier J, Ternaux B, Lacassagne MN, Crinquette A, et al. Phenotypic and functional characterization of bone marrow mesenchymal stem cells derived from patients with multiple myeloma. Leukemia. 2007;21:158–63.
Corre J, Mahtouk K, Attal M, Gadelorge M, Huynh A, Fleury-Cappellesso S, et al. Bone marrow mesenchymal stem cells are abnormal in multiple myeloma. Leukemia. 2007;21:1079–88.
Garderet L, Mazurier C, Chapel A, Ernou I, Boutin L, Holy X, et al. Mesenchymal stem cell abnormalities in patients with multiple myeloma. Leuk Lymphoma. 2007;48:2032–41.
Garayoa M, Garcia JL, Santamaria C, Garcia-Gomez A, Blanco JF, Pandiella A, et al. Mesenchymal stem cells from multiple myeloma patients display distinct genomic profile as compared with those from normal donors. Leukemia. 2009;23:1515–27.
Xu S, Evans H, Buckle C, De Veirman K, Hu J, Xu D, et al. Impaired osteogenic differentiation of mesenchymal stem cells derived from multiple myeloma patients is associated with a blockade in the deactivation of the Notch signaling pathway. Leukemia. 2012;26:2546–9.
Ambros V, Lee RC, Lavanway A, Williams PT, Jewell D. MicroRNAs and other tiny endogenous RNAs in C. elegans. Curr Biol. 2003;13:807–18.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Lawrie CH. MicroRNAs in hematological malignancies. Blood Rev. 2013;27:143–54.
Pichiorri F, Suh SS, Ladetto M, Kuehl M, Palumbo T, Drandi D, et al. MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci USA. 2008;105:12885–90.
Gutierrez NC, Sarasquete ME, Misiewicz-Krzeminska I, Delgado M, De Las Rivas J, Ticona FV, et al. Deregulation of microRNA expression in the different genetic subtypes of multiple myeloma and correlation with gene expression profiling. Leukemia. 2010;24:629–37.
Bi C, Chng WJ. MicroRNA: important player in the pathobiology of multiple myeloma. BioMed Res Int. 2014;2014:521586.
Guo L, Zhao RC, Wu Y. The role of microRNAs in self-renewal and differentiation of mesenchymal stem cells. Exp Hematol. 2011;39:608–16.
Schoolmeesters A, Eklund T, Leake D, Vermeulen A, Smith Q, Force Aldred S, et al. Functional profiling reveals critical role for miRNA in differentiation of human mesenchymal stem cells. PLoS ONE. 2009;4:e5605.
Zeng Y, Qu X, Li H, Huang S, Wang S, Xu Q, et al. MicroRNA-100 regulates osteogenic differentiation of human adipose-derived mesenchymal stem cells by targeting BMPR2. FEBS Lett. 2012;586:2375–81.
Wu T, Xie M, Wang X, Jiang X, Li J, Huang H. miR-155 modulates TNF-alpha-inhibited osteogenic differentiation by targeting SOCS1 expression. Bone. 2012;51:498–505.
Bae Y, Yang T, Zeng HC, Campeau PM, Chen Y, Bertin T, et al. miRNA-34c regulates Notch signaling during bone development. Hum Mol Genet. 2012;21:2991–3000.
Schaap-Oziemlak AM, Raymakers RA, Bergevoet SM, Gilissen C, Jansen BJ, Adema GJ, et al. MicroRNA hsa-miR-135b regulates mineralization in osteogenic differentiation of human unrestricted somatic stem cells. Stem Cells Dev. 2010;19:877–85.
Huang S, Wang S, Bian C, Yang Z, Zhou H, Zeng Y, et al. Upregulation of miR-22 promotes osteogenic differentiation and inhibits adipogenic differentiation of human adipose tissue-derived mesenchymal stem cells by repressing HDAC6 protein expression. Stem Cells Dev. 2012;21:2531–40.
Tang X, Lin J, Wang G, Lu J. MicroRNA-433-3p promotes osteoblast differentiation through targeting DKK1 expression. PLoS ONE. 2017;12:e0179860.
Miyaki S, Nakasa T, Otsuki S, Grogan SP, Higashiyama R, Inoue A, et al. MicroRNA-140 is expressed in differentiated human articular chondrocytes and modulates interleukin-1 responses. Arthritis Rheum. 2009;60:2723–30.
Tuddenham L, Wheeler G, Ntounia-Fousara S, Waters J, Hajihosseini MK, Clark I, et al. The cartilage specific microRNA-140 targets histone deacetylase 4 in mouse cells. FEBS Lett. 2006;580:4214–7.
Lin EA, Kong L, Bai XH, Luan Y, Liu CJ. miR-199a, a bone morphogenic protein 2-responsive MicroRNA, regulates chondrogenesis via direct targeting to Smad1. J Biol Chem. 2009;284:11326–35.
Eskildsen T, Taipaleenmaki H, Stenvang J, Abdallah BM, Ditzel N, Nossent AY, et al. MicroRNA-138 regulates osteogenic differentiation of human stromal (mesenchymal) stem cells in vivo. Proc Natl Acad Sci USA. 2011;108:6139–44.
Qu B, Xia X, Wu HH, Tu CQ, Pan XM. PDGF-regulated miRNA-138 inhibits the osteogenic differentiation of mesenchymal stem cells. Biochem Biophys Res Commun. 2014;448:241–7.
Yan J, Zhang C, Zhao Y, Cao C, Wu K, Zhao L, et al. Non-viral oligonucleotide antimiR-138 delivery to mesenchymal stem cell sheets and the effect on osteogenesis. Biomaterials. 2014;35:7734–49.
D’Souza S, del Prete D, Jin S, Sun Q, Huston AJ, Kostov FE, et al. Gfi1 expressed in bone marrow stromal cells is a novel osteoblast suppressor in patients with multiple myeloma bone disease. Blood. 2011;118:6871–80.
Reagan MR, Mishima Y, Glavey SV, Zhang Y, Manier S, Lu ZN, et al. Investigating osteogenic differentiation in multiple myeloma using a novel 3D bone marrow niche model. Blood. 2014;124:3250–9.
Zhou Y, Chen L, Barlogie B, Stephens O, Wu X, Williams DR, et al. High-risk myeloma is associated with global elevation of miRNAs and overexpression of EIF2C2/AGO2. Proc Natl Acad Sci USA. 2010;107:7904–9.
Sarma K, Levasseur P, Aristarkhov A, Lee JT. Locked nucleic acids (LNAs) reveal sequence requirements and kinetics of Xist RNA localization to the X chromosome. Proc Natl Acad Sci USA. 2010;107:22196–201.
Kumar R, Singh SK, Koshkin AA, Rajwanshi VK, Meldgaard M, Wengel J. The first analogues of LNA (locked nucleic acids): phosphorothioate-LNA and 2’-thio-LNA. Bioorg Med Chem Lett. 1998;8:2219–22.
Campbell MA, Wengel J. Locked vs. unlocked nucleic acids (LNA vs. UNA): contrasting structures work towards common therapeutic goals. Chem Soc Rev. 2011;40:5680–9.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20.
Jiang L, Liu X, Kolokythas A, Yu J, Wang A, Heidbreder CE, et al. Downregulation of the Rho GTPase signaling pathway is involved in the microRNA-138-mediated inhibition of cell migration and invasion in tongue squamous cell carcinoma. Int J Cancer. 2010;127:505–12.
Li J, Chen Y, Qin X, Wen J, Ding H, Xia W, et al. MiR-138 downregulates miRNA processing in HeLa cells by targeting RMND5A and decreasing Exportin-5 stability. Nucleic Acids Res. 2014;42:458–74.
Zhang Y, Roccaro AM, Rombaoa C, Flores L, Obad S, Fernandes SM, et al. LNA-mediated anti-miR-155 silencing in low-grade B-cell lymphomas. Blood. 2012;120:1678–86.
Oyajobi BO, Franchin G, Williams PJ, Pulkrabek D, Gupta A, Munoz S, et al. Dual effects of macrophage inflammatory protein-1alpha on osteolysis and tumor burden in the murine 5TGM1 model of myeloma bone disease. Blood. 2003;102:311–9.
Gai Z, Gui T, Muragaki Y. The function of TRPS1 in the development and differentiation of bone, kidney, and hair follicles. Histol Histopathol. 2011;26:915–21.
Piscopo DM, Johansen EB, Derynck R. Identification of the GATA factor TRPS1 as a repressor of the osteocalcin promoter. J Biol Chem. 2009;284:31690–703.
Zaman G, Staines KA, Farquharson C, Newton PT, Dudhia J, Chenu C, et al. Expression of Sulf1 and Sulf2 in cartilage, bone and endochondral fracture healing. Histochem Cell Biol. 2016;145:67–79.
Zhang Y, Xie RL, Gordon J, LeBlanc K, Stein JL, Lian JB, et al. Control of mesenchymal lineage progression by microRNAs targeting skeletal gene regulators Trps1 and Runx2. J Biol Chem. 2012;287:21926–35.
Roodman GD. Pathogenesis of myeloma bone disease. Leukemia. 2009;23:435–41.
Saad F, Lipton A, Cook R, Chen YM, Smith M, Coleman R. Pathologic fractures correlate with reduced survival in patients with malignant bone disease. Cancer. 2007;110:1860–7.
Sha HH, Wang DD, Chen D, Liu SW, Wang Z, Yan DL, et al. MiR-138: A promising therapeutic target for cancer. Tumour Biol. 2017;39:1010428317697575.
Jiang B, Mu W, Wang J, Lu J, Jiang S, Li L, et al. MicroRNA-138 functions as a tumor suppressor in osteosarcoma by targeting differentiated embryonic chondrocyte gene 2. J Exp Clin Cancer Res. 2016;35:69.
Xiao L, Zhou H, Li XP, Chen J, Fang C, Mao CX, et al. MicroRNA-138 acts as a tumor suppressor in non small cell lung cancer via targeting YAP1. Oncotarget. 2016;7:40038–46.
Ma F, Zhang M, Gong W, Weng M, Quan Z. MiR-138 suppresses cell proliferation by targeting Bag-1 in gallbladder carcinoma. PLoS ONE. 2015;10:e0126499.
Wang Q, Tang H, Yin S, Dong C. Downregulation of microRNA-138 enhances the proliferation, migration and invasion of cholangiocarcinoma cells through the upregulation of RhoC/p-ERK/MMP-2/MMP-9. Oncol Rep. 2013;29:2046–52.
Yuan Y, Zhang GQ, Chai W, Ni M, Xu C, Chen JY. Silencing of microRNA-138-5p promotes IL-1beta-induced cartilage degradation in human chondrocytes by targeting FOXC1: miR-138 promotes cartilage degradation. Bone Jt Res. 2016;5:523–30.
Cozzolino F, Torcia M, Aldinucci D, Rubartelli A, Miliani A, Shaw AR, et al. Production of interleukin-1 by bone marrow myeloma cells. Blood. 1989;74:380–7.
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9:654–9.
Carroll SH, Ravid K. Differentiation of mesenchymal stem cells to osteoblasts and chondrocytes: a focus on adenosine receptors. Expert Rev Mol Med. 2013;15:e1.
Komori T. Regulation of bone development and extracellular matrix protein genes by RUNX2. Cell Tissue Res. 2010;339:189–95.
DeLise AM, Fischer L, Tuan RS. Cellular interactions and signaling in cartilage development. Osteoarthr Cartil. 2000;8:309–34.
McBeath R, Pirone DM, Nelson CM, Bhadriraju K, Chen CS. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev Cell. 2004;6:483–95.
Strzelecka-Kiliszek A, Mebarek S, Roszkowska M, Buchet R, Magne D, Pikula S. Functions of Rho family of small GTPases and Rho-associated coiled-coil kinases in bone cells during differentiation and mineralization. Biochim Biophys Acta. 2017;1861:1009–23.
Dai Y, Yang Y, MacLeod V, Yue X, Rapraeger AC, Shriver Z, et al. HSulf-1 and HSulf-2 are potent inhibitors of myeloma tumor growth in vivo. J Biol Chem. 2005;280:40066–73.
Acknowledgements
This work was supported in part by NCI R01 CA181683-01A1 and R01 CA205954-01, Leukemia and Lymphoma Society and Multiple Myeloma Research Foundation. This work was also supported by the Danish Council for Independent Research, the Danish Cancer Society, Direktoer Ib Henriksens Foundation, the Lundbeck Foundation, the Niels Bohr Foundation, the Novo Nordisk Foundation, and the Dagmar Marshalls Foundation. ST was supported by a postdoctoral fellowship from the Uehara Memorial Foundation, Japan. We thank Ms. Catherine A. Sypher for technical assistance with animal experiments and Ms. Suzan Lazo for technical assistance with flow cytometric analysis.
Author contributions
ST, MBL, KZS, MRR, SK, and IMG conceived and designed the experiments. ST, MBL, and KZS performed in vitro studies. MR and DH collected clinical samples. ST, OZ, and YK performed in vivo studies. MBL, JP, and AP performed bioinformatics analysis. ST, MBL, SM, ST, KK, CAT, and MC analyzed the data. KN performed bone histomorphometric analysis. ST and MBL wrote the manuscript with input from AMR, RB, SK, and IMG.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Tsukamoto, S., Løvendorf, M.B., Park, J. et al. Inhibition of microRNA-138 enhances bone formation in multiple myeloma bone marrow niche. Leukemia 32, 1739–1750 (2018). https://doi.org/10.1038/s41375-018-0161-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0161-6
This article is cited by
-
Knockdown of lncRNA BDNF-AS inhibited the progression of multiple myeloma by targeting the miR-125a/b-5p-BCL2 axis
Immunity & Ageing (2022)
-
Roles of miRNA dysregulation in the pathogenesis of multiple myeloma
Cancer Gene Therapy (2021)
-
Chitosan-miRNA functionalized microporous titanium oxide surfaces via a layer-by-layer approach with a sustained release profile for enhanced osteogenic activity
Journal of Nanobiotechnology (2020)
-
Inhibition of lung cancer growth and metastasis by DHA and its metabolite, RvD1, through miR-138-5p/FOXC1 pathway
Journal of Experimental & Clinical Cancer Research (2019)
-
Epigenetics of Multiple Myeloma Bone Disease
Current Molecular Biology Reports (2019)