Abstract
Cutaneous T-cell lymphomas (CTCLs) represent different subtypes of lymphoproliferative disorders with no curative therapies for the advanced forms of the disease (namely mycosis fungoides and the leukemic variant, Sézary syndrome). Molecular events leading to CTCL progression are heterogeneous, however recent DNA and RNA sequencing studies highlighted the importance of NF-κB and β-catenin pathways. We here show that the kinase TAK1, known as essential in B-cell lymphoma, is constitutively activated in CTCL cells, but tempered by the MYPT1/PP1 phosphatase complex. Blocking PP1 activity, both pharmacologically and genetically, resulted in TAK1 hyperphosphorylation at residues T344, S389, T444, and T511, which have functional impact on canonical NF-κB signaling. Inhibition of TAK1 precluded NF-κB and β-catenin signaling and induced apoptosis of CTCL cell lines and primary Sézary syndrome cells both in vitro and in vivo. Detection of phosphorylated TAK1 at T444 and T344 is associated with the presence of lymphoma in a set of 60 primary human samples correlating with NF-κB and β-catenin activation. These results identified TAK1 as a potential biomarker and therapeutic target for CTCL therapy.
Similar content being viewed by others
Introduction
Cutaneous T-cell lymphomas (CTCL) are lymphoid malignant neoplasms included as peripheral T-cell non-Hodgkin’s lymphomas that primarily manifest in the skin. The most frequent CTCL, mycosis fungoides (MF) and the leukemic variant Sézary syndrome (SS), are characterized by proliferation of T-helper cells with mature phenotype (CD3+, CD4+, and CD45RO+). MF is characterized by a clinical multistage development starting with erythematous scaly patches that are followed by infiltrated plaques and final transformation into the tumor stage. In SS, the disease is clinically characterized by erythroderma associated with peripheral blood involvement manifested by circulating malignant lymphoid cells with cerebriform nuclei (Sézary cells). Tumor-stage MF and SS are considered aggressive forms of the disease and usually have unfavorable prognosis. Till date, there are no targeted therapies that provide curative alternative for advanced CTCL patients. Interferon, oral retinoids (bexarotene), and non-specific histone deacetylase inhibitors are currently prescribed as therapeutic options, but most cases achieve response rates of about 30% (reviewed in [1]).
Although the pathogenic mechanisms implicated in CTCL progression are fairly unknown, several reports have suggested a relevant role for STAT3, Notch and β-catenin pathways in this group of disorders [2,3,4,5,6,7]. Recently, whole-genome/exome DNA and RNA sequencing of tumor-stage MF and SS has clearly identified alterations in elements upstream of TAK1 and IKK such as CARD11 and TNFR2, which suggest a pivotal role for NF-κB signaling in CTCL [8,9,10,11]. Although this pathway has been mainly associated to B-cell lymphoma [12,13,14,15,16,17,18], there are several reports indicating that particular NF-κB elements can also contribute to T-cell lymphoma [19, 20]. In fact, NF-κB is an essential regulator of normal T-cell homeostasis and function [21,22,23], whereas inactivation of the pathway leads to a blockage in the differentiation and survival of mature T cell compartment [24,25,26] and precludes tumor progression in a mouse model of Notch-induced Acute T-cell Leukemia [27].
Phosphorylation of IκBα by IKKβ is the critical step on NF-κB activation, which is initiated, in a stimulus-dependent manner, by the TAK1 kinase downstream of the ubiquitin-ligase elements TRAF6 and Ubc13. Treatment of primary and transformed T cells with PP2A or PP1 inhibitors has been found to increase the amount of phosphorylated IκBα leading to NF-κB activation [28, 29], thus indicating the existence of constitutive IKK activity that is counteracted by phosphatases in this particular cell lineage. Various phosphatases have been identified that negatively regulate IKK, thus guaranteeing precise and transient cellular responses to extracellular stimuli in particular cell types. That is the case of CUEDC2/PP1 [30] and PP4R1 [31] phosphatase complexes. One step upstream in the pathway, PP1 through GADD34 repressed TAK1 kinase in macrophages [32] by dephosphorylation of its regulatory S412 residue [33], thus preventing excessive activation of TLR pathway during inflammatory immune responses. Whether TAK1 and NF-κB play a critical role in human T-cell lymphoma has not convincingly been addressed.
Here, we study the contribution and potential therapeutic relevance of TAK1 and NF-κB signaling in CTCL. Our results indicate that TAK1 is constitutively activated in human CTCL cells although attenuated by PP1-mediated dephosphorylation of specific residues. However, the remaining TAK1 activity is sufficient and required to maintain NF-κB and β-catenin activation and its inhibition has potent anti-tumor effects leading to reduced proliferation and increased apoptosis of lymphoma cells.
Materials and Methods
Cell cultures and cell lines
Cutaneous T-cell Lymphoma cells included 2 MF (HH and MYLA) and 2 SS (HUT78 and SeAx) cell lines that were tested as mycoplasma free. Patient-derived SS samples (identified as SZ #1–4) were obtained from fresh peripheral blood mononuclear cells of selected SZ patients with high tumor burden (representing 90–95% of CD4+ SZ cell population according to current morphologic and/or phenotypic diagnostic criteria). Mononuclear cells were recovered by Ficoll (GE Healthcare, Princeton, NJ) gradient separation. CD4+ T cells were purified by positive selection using immune-magnetic beads (Miltenyi Biotec, Bergisch Gladbach, Germany), cultured at a density of 1 × 106 cells/ml and then activated and expanded in vitro using anti-CD3/CD28 beads (Life Technologies, Norway).
SeAx tumor model in NSG mouse
We injected 4 × 105 SeAx cells into the dermal space under the central dorsal surface of the ears above the cartilage plane subcutaneous in the ear of 2 months old NSG mice. These cells generated tumors that infiltrate the adjacent lymph nodes and were detectable by palpation at 7–10 days after the injection. At the time of tumor detection, we randomly defined 2 experimental groups that were treated by intra-peritoneal injection either with DMSO (control; n = 6 animals) or 15 mg/kg (5Z)-7-oxozeaenol (TAK1 inhibitor; n = 6 animals) every other day. After 5 rounds of treatment, the animals were killed and the tumors were collected, measured, and processed for subsequent analysis. All experiments involving animals were performed according to the regulations of the Catalan Government and the Ethical Committee at Institut Municipal d’Investigacions Mèdiques (IMIM).
Antibodies against active TAK1
Antibodies targeting phosphorylated TAK1 protein (residues T344, S389, T444, and T511 corresponding to human isoform A) were generated by Abyntek (Bizkaia, Spain) by immunizing rabbits with the corresponding phosphor-peptides and depleting serum from antibodies recognizing non-phosphorylated peptides. Commercial antibodies used in this work are specified in supplemental material.
Generation of TAK1 point mutants and cell transfection
TAK1 point mutants were generated using sequence overlap extension with specific mutagenic primers. Star-Prime polymerase (Takara) was used following manufacturer’s instructions, and TAK1 inserts were sequenced to confirm the mutations of interest and the absence of additional undesired changes. The cells were transfected using polyethylenimine at a DNA/reagent ratio of 1:5.
Gel filtration assay on Superdex200 columns
CTCL cells were lysed in 100 μl PBS containing 0.5% Triton X-100, 1 mM EDTA, 100 mM Na-orthovanadate, 0.25 mM phenylmethanesulfonylfluoride (PMSF), and complete protease inhibitor cocktail (Roche, Basel, Switzerland), centrifuged at 13.000 r.p.m. Supernatants were loaded on Superdex200 gel filtration column (GE Healthcare). One drop (40 μl) per fraction was collected and analyzed by western blot.
Mass spectrometry analysis
Immunoprecipitated proteins were reduced (10 mM DTT, 1 h at 37 °C), alkylated (20 mM IAA, 30 min, RT), and digested with trypsin (o/n, 37 °C) prior to being analyzed in a LTQ-Orbitrap Velos Pro mass spectrometer (see Supplementary materials for details). Raw data were analyzed using Proteome Discoverer software suite (v1.4) and Mascot search engine (v2.5). The data were searched against the human protein database derived from SwissProt with a precursor ion mass tolerance of 7 ppm, and up to three missed cleavages. The fragment ion mass tolerance was set to 0.5 Da. Oxidation (M), Phosphorylation (STY), Acetylation (Protein N-term), and Methylation (K), were defined as variable modifications, whereas carbamidomethylation (C) was set as fixed modification. Identified peptides were filtered by FDR <5%. Differentially phosphorylated TAK1 peptides were selected based on its presence in calyculin A1-treated samples and absence in control samples in at list 2 out of 3 independent experiments performed. TAK1 interactors were defined as those detected in 2 out of 3 IP experiments in control conditions.
Tissue samples and sample preparation
All human tissues used in this study were obtained from Parc de Salut MAR Biobank (MARBiobanc, Barcelona) with the consent of its ethical Committee, and following Spanish Ethical regulations and guidelines of the Declaration of Helsinki. Patient identity remained anonymous. Formalin-fixed, paraffin-embedded tissue blocks of human CTCL were from MARBiobanc archives. Areas of invasive lymphoma lesions were identified on the corresponding hematoxylin-eosin-stained slides.
Statistical methods for biomarker correlations
Data are expressed as frequencies and percentages for categorical variables and ordinal variables or are specified otherwise. The associated risk (odds ratios (OR) and 95%CI) for the grade of biomarkers intensity and for grade >0 with lymphoma was assessed by means of ordinal and binary logistic regression models, respectively. To study correlations, we used Spearman coefficients. All analyses were performed using SAS 9.2 software (SAS Institute, Cary, NC, USA) and the level of significance was established at the two-sided 5% level.
Image analysis
Immunohistochemistry of cutaneous lymphoma sections was evaluated by two investigators in an Olympus BX61 microscope. Immunofluorescence images were taken in a confocal Leica SP5 TCS upright microscope and the Leica Application Suite Advanced Fluorescence software.
Results and discussion
PP1 activity counteracts the TAK1-dependent activation of NF-κB signaling in CTCL cells
Recent sequencing projects in human CTCL identified mutations in CARD11 [8, 10] and TNFR2 [9] that might support constitutive NF-κB activity. In a project focused in the identification of NF-κB regulators, we precipitated the TAK1 kinase from MYLA cells and performed mass spectrometry analysis. We identified the canonical TAK1-associated proteins TAB1, TAB2, nd TAB3, as expected, but also peptides corresponding to the phosphatase PP1 and its regulatory subunit PPP1E12A/MYPT1 (Table 1A). To test whether PP1 was affecting NF-κB activity in CTCL, we incubated HH, MYLA, and SeAx cells with the specific PP1 inhibitor calyculin A1. Treating cells for 40 min led to accumulation of phosphorylated IKK and IκBα, and a shift in the electrophoretic mobility of TAK1, as determined by western blot (WB) (Fig. 1a), associated to the accumulation of nuclear p65-NF-κB (Figure S1A). As a specificity control, PP1 knockdown using 3 different shRNAs also increased IKK and IκBα phosphorylation in HH, MYLA, and SeAx cells (Figure S1B). However, the effect of PP1 knockdown was less than the observed with calyculin A1 treatment, suggesting that TAK1 hyperphosphorylation after PP1 inhibition is temporary and then attenuated by negative feedback loops, or other phosphatase activities, such as PP2A also contribute to TAK1 regulation.
PP1 inhibition promotes TAK1- and IKKβ-dependent activation of NF-κB signaling in CTCL. a Mass spectrometry analysis of anti-P-TAK1 precipitates from MYLA cell lysates to identify elements associated with TAK1 complex. b Western blot (WB) analysis of indicated CTCL cells treated with calyculin A1 (CALY) for 40 min. c–d Western blot analysis of the indicated CTCL cells treated with 2 different TAK1 (S, from Sigma and M, from Millipore) c or IKKβ inhibitors d for 16 h. e WB analysis of the indicated cell lines to evaluate the effect of different treatments on IκBα stabilization and apoptosis as determined by cleaved-caspase 3 detection. f Quantification of the number of cells obtained after culturing them in the indicated conditions. g WB of nuclear extracts from different CTCL cells treated for 36 h as indicated, and quantification by band densitometry of the differences detected. h Quantification of the number of cells obtained after culturing cells with the indicated inhibitors. In 1 h, & indicated synergy in the combination treatment (p < 0.01) as determined by PKF115*BAY65 interaction term, which was confirmed in a mixed model analysis that included the terms cell line and treatment, and taking into account the replica count factor from 4 independent experiments performed (p = 0.0145). In all the experiments n.s. non-significant; * p < 0.05; ** p < 0.01; *** p < 0.001, and **** p < 0.0001 obtained by two-sided t-test analysis. Graphs represent the mean and s.d
Accumulation of phosphorylated IKK and IκBα following calyculin A1 treatment was abrogated by pre-incubation with the TAK1 inhibitor (5Z)-7-oxozeaenol (Fig. 1b) and the IKKβ inhibitor BAY65-8072 [34] (Fig. 1c), supporting the concept that NF-κB was constitutively activated in CTCL cells although buffered by PP1-mediated dephosphorylation of specific elements, most likely TAK1.
Next, we tested the consequences of inhibiting TAK1, IKKβ, and ROCK, which regulates the MYPT1/PP1 complex, on CTCL survival. We observed a distinctive pro-apoptotic effect of TAK1 inhibition in HH and SeAx cells, as determined by cleaved-caspase 3 detection (Fig. 1d), which was associated with its higher impact on CTCL cell growth (70–80% growth reduction after 36 h) (Fig. 1e). Because TAK1, IKKβ, and ROCK inhibitors induced similar IκBα stabilization (Fig. 1d), we speculated that NF-κB was not the only pathway contributing to CTCL cell survival downstream of TAK1. Further analysis demonstrated that 36 h of treatment with TAK1 inhibitor imposed a variable, but significant reduction in the nuclear levels of p65-NF-κB and β-catenin in all tested CTCL cell lines (Fig. 1f) that was associated to transcriptional repression of canonical NF-κB and β-catenin target genes c-Myc, Axin2, A20, and Nfkbia (Figure S1C). In addition, knocking-down TAK1 with 2 different shRNAs promoted accumulation of IκBα, β-catenin destabilization, and abrogation of P-IκBα when compared with the scramble shRNA controls (Figure S1D).
To study the possibility that TAK1 controls CTCL cell survival through simultaneous activation of NF-κB and β-catenin signaling, we treated CTCL cells with the NF-κB inhibitor, the β-catenin inhibitor PKF115-584, or both. Inhibition of β-catenin and IKKβ reduced growth of CTCL cells lines (Fig. 1g) and primary SS cells (Figure S1E), but the effect was less than TAK1 inhibition. Inhibition of NF-κB and β-catenin pathways together showed a synergistic effect compared to either treatment alone (Fig. 1g). Cell growth arrest imposed by the combination treatment was associated with increased apoptosis as determined by cleaved-caspase 3 detection (Figure S1F). As control, PKF115-584 and NF-κB inhibitors reduced transcription of Axin2 and Nfkbia, respectively, both in single treatments and in combination (Figure S1G).
Of note, β-catenin was previously identified as pro-tumorigenic factor in T cell neoplasms [7, 35,36,37,38], a target of TAK1 in colorectal cancer cells [39], and a required co-factor for both Notch-induced and Notch-independent acute T cell leukemia [35,36,37]. Together these results point-out β-catenin as an essential element for oncogenic transformation all along the T cell differentiation pathway. Moreover, the identification of β-catenin as an enhancer of NF-κB pro-survival function downstream of TAK1 could be clinically exploitable.
Regulation of canonical NF-κB by PP1 in CTCL lays downstream of the ROCK/MYPT1 pathway
It was previously shown that TNFα induced MYPT1 phosphorylation at T696 leading to PP1 inactivation and cytoskeleton reorganization in muscle cells [40,41,42]. Since both MYPT1 and PP1 were identified in TAK1 precipitates from MYLA cells, we investigated whether TNFα modified MYPT1 and PP1 activity and this affected NF-κB signaling in CTCL. In MYLA cells, we found that TNFα treatment induced MYPT1 phosphorylation at T696 starting at 5 min with a maximum at 20 min, linked to increased phosphorylation of IκBα and IKK and IκBα degradation (Fig. 2a). In Superdex S200 columns, TNFα treatment led to a significant reduction in the amount of PP1 and MYPT1 that coeluted in the TAK1- and IKK-containing fractions, and we did not detect phosphorylated MYPT1 coeluting with TAK1 in any experimental condition (Fig. 2b). These results further support the concept that MYPT1 phosphorylation precluded PP1 activity on TAK1 and NF-κB.
Regulation of canonical NF-κB by PP1 in CTCL cells lays downstream of the ROCK/MYPT1 pathway. a WB analysis of total cell lysates from TNFα-treated MYLA cells at the indicated time points. b WB of untreated or TNFα-treated MYLA cell lysates separated in Superdex S200 columns. c WB analysis of MYLA cells treated with TNFα at different time points in the absence or presence of ROCK inhibitor. d Same experiment as in C from cells pretreated for 16 h with the proteasome inhibitor MG132. e Quantification of the relative levels obtained by densitometry analysis of the images of IκBα and ubiquitinated P-IκBα from the experiments shown in C (upper panel) and D (lower panel). The average and standard deviation values from three independent biological replicates performed are shown. f Quantification of the number of cells obtained after 96 h of culture in the indicated conditions. In all experiments n.s. non-significant; * p < 0.05; ** p < 0.01, and *** p < 0.001 obtained by two-sided t-test. In (f), & indicates synergy in the combination treatment (p < 0.05). Graphs represent the mean and s.d
Inactivating phosphorylation of MYPT1 is primarily dependent on the activity of ROCK kinases [43,44,45]. We tested the possibility that ROCK inhibition prevented MYPT1 phosphorylation and NF-κB signaling after TNFα treatment in CTCL cells. Incubation of MYLA cells with the ROCK inhibitor Y-27632 suppressed TNFα-induced MYPT1 phosphorylation at T696 and also reduced IKK phosphorylation and IκBα degradation following TNFα treatment (Fig. 2c, d). Comparable results were obtained following ROCK1 knockdown with specific shRNA (Figure S2). In addition, Y-27632 treatment resulted in decreased P-IκBα polyubiquitination induced by TNFα after pre-incubation with the proteasome inhibitor MG132 (Fig. 2e). In all tested CTCL lines, ROCK inhibition reduced cell growth and showed additive or synergistic effects with the β-catenin inhibitor PKF115-584 (Fig. 2f), suggesting that ROCK/MYPT1 primarily regulate NF-κB-linked TAK1 activity in these cells.
Our results indicate that the MYPT1/PP1 complex, either downstream of ROCK1 or in response to TNFα signaling regulates canonical NF-κB in CTCL cells and contributes to promote tumor cell survival. Thus, we speculate that in vivo, the presence of a tumor-associated inflammatory environment, or processes that involve ROCK activation such as cellular migration, cell adhesion, or cytoskeleton reorganization might impose MYPT/PP1 inactivation, leading to TAK1 phosphorylation and activation of the pro-tumorigenic pathways NF-κB and β-catenin. Interestingly, activation of the IKK kinase complex and NF-κB in response to cellular attachment and cytoskeleton reorganization was previously observed in prostate cancer cells [46], although the underlying mechanisms have never been addressed.
PP1 regulates TAK1 phosphorylation at non-canonical regulatory sites
To study the possibility that PP1 was a direct regulator of the TAK1 and IKK, we separated freshly isolated CTCL extracts on Superdex S200 gel filtration columns, which are used to identify multi-protein complexes. We detected canonical IKKα, IKKβ, and IKKγ/NEMO subunits eluting at fractions 71–81, showing partial overlap with their upstream kinase TAK1 (eluting at fractions 77–91). Interestingly, PP1 that was largely eluting at fractions 91–105 (corresponding to low molecular weight complexes) was also detected at small amounts in fractions 73–83, co-distributing with IKK and TAK1 (Fig. 3a, left panels). Moreover, PP1 protein from MYLA cells coprecipitated with P-TAK1 (S412) and IKK (Fig. 3a, right panels), indicative of physical association. To confirm that electrophoretic mobility shift of TAK1 imposed by calyculin A1 treatment (see Fig. 1b) was due to increased phosphorylation of the kinase, we treated these lysates with alkaline phosphatase (AP). Incubation of calyculin A1-treated CTCL cell lysates with AP partially, but not totally, reversed the mobility shift of TAK1 (Fig. 3b), indicating that PP1 inhibition promoted the accumulation of both AP-sensitive and AP-resistant TAK1 phosphor-residues, although we cannot formally discard the existence of other post-translation modifications in TAK1. To identify phosphor-residues that were substrates of PP1 activity in CTCL, we precipitated P-TAK1 (S412) from MYLA cells untreated or treated with calyculin A1 for 40 min (Figure S3) and performed mass spectrometry analysis of the precipitates. Analysis of the data indicated the presence of differentially phosphorylated peptides corresponding to TAK1 at positions T344, S389, T444, and T511 (Table 1B), whereas we failed to detect any increase in the phosphorylated peptides containing T184 and T187 (at the catalytic domain of TAK1) or S412, in three independent experiments.
PP1 binds to and regulates TAK1 phosphorylation at non-canonical sites. a Western blot (WB) of CTCL (MYLA) cell lysates separated in a Superdex S200 column (left panels) and analysis of MYLA cell lysates immunoprecipitated (IP) with antibodies against active TAK1 (P-S412) or non-related control IgG. 1/10 of the total lysate is shown as input control (right panels). b Lysates from calyculin A1-treated MYLA cells where left untreated or treated with calf intestinal phosphatase (CIP) for 30 min at 37° C and analyzed by WB. c Mass spectrometry analysis of anti-P-TAK1 precipitates to identify TAK1 peptides differentially phosphorylated following calyculin A1 treatment,
Functional impact of phosphorylation at T344, S389, T444, and T511 on TAK1 activity
To study the functional contribution of phosphorylated TAK1 residues T344, S389, T444, and T511, we generated specific point mutants of the protein. Replacement of these particular S or T into A reduced the capacity of TAK1 to induce IκBα phosphorylation and degradation in HEK-293 cells, similar to the inactive T184/187A mutant (Fig. 4a).
Phosphorylation at T344, S389, T444 and T511 residues of TAK1 functionally impacts on its activity on NF-κB. a WB analysis of HEK-293T cells transiently expressing the indicated TAK1 kinase mutants. Inactive T184-187A is shown as negative control. b WB analysis of lysates from untreated or calyculin A1-treated HH cells immunoprecipitated (IP) with antibody against IKKγ/NEMO or control IgG. 1/10 of the total lysate is shown as input control. c Western blot (WB) with different anti-P-TAK1 antibodies of HH cell lysates separated in a Superdex S200 column
We then produced polyclonal antibodies against individual TAK1 phosphor-residues and determined their specificity on lysates from HA-TAK1-transfected HEK-293T cells. Antibodies against phosphorylated T344 and S389 completely failed to detect TAK1 T344A and S389A mutants, respectively, whereas they recognized all other constructs in WB assay. Antibodies to phosphorylated T444 and T511 showed reduced affinity for the corresponding mutants. Interestingly, antibody to T444 failed to detect the inactive T184/187A TAK1 mutant, in contrast to anti-P-TAK1 (T412) that recognized all TAK1 constructs independent on their functional capacity (Figure S4A). As expected, antibodies against phosphorylated TAK1 (T344, S389, T444, T511, and S412) better detect endogenous TAK1 from calyculin A1-treated precipitates (Figure S4B), which was reversed by phosphatase treatment of the lysates (Figure S4C). Interestingly, all P-TAK1 antibodies recognized TAK1 that associates with the regulatory NF-κB element IKKγ/NEMO from unstimulated and calyculin A1-treated cells (Fig. 4b). As shown in Fig. 4b, c and S4B, antibodies against phosphorylated TAK1 including the commercial anti-T184-187, all recognized additional bands other than the expected 75 kDa band suggesting that several P-TAK1 species carrying variable combinations of phosphor-residues, or other post-translational modifications, can coexist in CTCL cells, or different TAK1 isoforms are differentially phosphorylated at specific S/T residues. Interestingly, although all phosphorylated TAK1 species were associated with the IKK subunit NEMO in the IP assay (Fig. 4b), TAK1 phosphorylated at S412 was eluting at separate fractions compared with TAK1 phosphorylated at T344, S389, T444, and T511 residues in the Superdex S200 columns (Fig. 4c). Together these results indicated that CTCL cells contain different TAK1 phosphorylated species all associated to the IKK complex, but suggested that they are functionally different.
Inhibition of TAK1 reduces CTCL tumors in vivo
Then, we investigated the therapeutic efficacy of TAK1 inhibition using a previously described in vivo model for CTCL [47] (Fig. 5a). SeAx cells were injected in the ear of NSG mice, which were treated 7 days after the injection either with DMSO (control; n = 6 animals) or 15 mg/kg of (5Z)-7-oxozeaenol (TAK1 inhibitor; n = 6 animals) every other day for 10 days. The animals were randomly included in either group of treatment. After killing the animals, the tumors obtained from TAK1 inhibitor-treated animals showed extensive areas of apoptosis and necrosis when compared with control-treated animals, as determined by evaluation of hematoxylin/eosin staining of tissue sections (Fig. 5b) and cleaved-caspase 3 staining (Figure S5A). Necrotic areas were characterized by the presence of cells with eosinophilic cytoplasm and pyknotic nucleus, cells in cariorexe with irregular chromatin distribution that accumulated in the nuclear membrane, and cells with fragmented nuclei or dissolution of the chromatin with disappearance of the nuclear structure. In contrast, growing areas of both groups of tumors were composed of large anaplastic cells, with irregular nucleus, hyperchromatism, and presence of abundant mitotic figures. Tumor cells in the growing areas were positive for the human pan-hematopoietic marker CD45 (Figure S5B) and displayed high levels of P-TAK1 (T444), which were reduced in the treatment group (Figure S5C). In the growing areas, TAK1 inhibition led to a general decrease in tumor cellularity that was accompanied by a slight, but significant reduction in cell proliferation as determined by ki67 staining (Fig. 5c). Comparable effects on tumor proliferation and apoptosis were obtained in three independent experiments performed and using an alternative treatment regime (Figure S5D).
Effects of ROCK, β-catenin, and TAK1 inhibition on CTCL cells in vitro. a Scheme of the procedure used to generate the CTCL in vivo model in NSG mice. b Representative hematoxylin & eosin (H&E) staining of tumor sections from control or TAK1-inhibited tumors (left panels). Necrotic (N) areas are enclosed using dashed lines to distinguish from growing (G) areas. Quantification of the longest cross-section of the necrotic and growing areas in control and TAK1-inhibited tumors (thickness), and the ratio between the necrotic and growing area in each tumor is shown (right panels). Asterisks denote the significance of the differences with the corresponding controls. c Representative ki67 staining in the growing area of control and treated tumors and quantification of the total and ki67 positive cells counted in 15 independent 20 × fields. Graphs represent the mean and s.d. Statistical significance was * p < 0.05, ** p < 0.01, and *** p < 0.001 obtained by two-sided t-test
These results indicate that inhibition of TAK1 in vivo reduced growth of CTCL-derived tumors likely associated to increased apoptosis and reduced proliferation. However, we consistently detected the presence of growing areas in both experimental groups (control and treatment) that show detectable levels of P-TAK1, suggest that the capacity of the inhibitor to eradicate lymphoma cells was related with the accessibility of the drug to particular tumor areas. In agreement with this hypothesis, we detected low vascularization in the growing areas of TAK1 inhibitor-treated tumors as determined by CD31 staining (Figure S5E).
The MYPT1–TAK1 pathway is activated in human CTCL
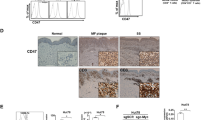
Finally, we studied the relative contribution of activated TAK1 to human CTCL. By IHC of MF biopsies, we detected P-TAK1 (T444) in both the lymphocytic component and adjacent keratinocytes (Fig. 6a, left panels) that was associated with the presence of phosphorylated (inactivated) MYPT1 (T696) (Fig. 6a, right panels), P-IκBα, and active β-catenin (Figures S6A and S6B). Cells carrying P-MYPT1 mainly corresponded to CD3-positive transformed T-cells (Figure S6C). We next determined P-TAK1 (T444) (Fig. 6b), P-IκBα, and β-catenin levels in a cohort of 60 human samples including CTCL lesions (MF and SS) (n = 27), 2 indolent CD30 + lymphomas, and inflammatory diseases such as psoriasis and lupus (n = 31) (Supplementary Table 1). We detected variable levels of β-catenin (cytoplasmic and nuclear) and/or P-IκBα in 55 of the analyzed samples, and 36 were annotated as double positive (Supplementary Table 1). Of note that cytoplasmic accumulation of β-catenin had been previously reported in human lymphoma samples where nuclear β-catenin was not detected [38]. Detection of P-TAK1 (T444) associated with higher risk of lymphoma (OR [95%CI]: 2.16 [1.05–4.43], p = 0.035; 2.99 [1.01–8.84], p = 0.048; for the ordinal and binary analyses, respectively). We did not observe significant associations for the rest of studied biomarkers and presence of lymphoma in the biopsies (Supplementary Table 2). Nuclear β-catenin correlated with total β-catenin and P-TAK1 (ρ = 0.620; p < 0.001 and ρ = 0.317; p = 0.017, respectively) in all samples, and with total β-catenin in control samples (ρ = 0.610 p < 0.001). Most importantly, P-TAK1 levels in the lymphoma samples significantly correlated with β-catenin, nuclear β-catenin, and P-IκBα (ρ = 0.435 p = 0.026, ρ = 0.406 p = 0.044, and ρ = 0.553 p = 0.003, respectively), and nuclear β-catenin with total β-catenin (ρ = 0.643 p < 0.001) (Supplementary Table 3). The prevalence of activated TAK1 in the lymphoma lesions was confirmed on available samples using a second anti-P-TAK1 (T344) antibody (Supplementary Table 1). Moreover, the robust association between P-TAK1 detection in the tumor component and in the adjacent epithelial (EPI) cells of the skin (see Fig. 6a, b) suggested that CTCL cells might also impose an additional immunosuppressive effect by the paracrine activation of TGFβ (downstream of TAK1) in the tumor stroma, as previously proposed [20]. In this scenario, lymphoma cells would escape from the inhibitory effect of TGFβ signaling by means of TGFβ receptor inactivation [48] or upregulation of the inhibitory subunit SMAD7 [49].
The MYPT1-TAK1 pathway is activated in human cutaneous T-cell lymphomas. a Immunofluorescence analysis of P-TAK1 (left panels) and P-MYPT1 (right panels) in human primary CTCL frozen samples. Representative images from eight different samples analyzed. b Immunohistochemistry analysis of P-TAK1 (T444) in two representative (1 MF and 1 SS) paraffin-embedded human primary CTCL samples. Dashed lines in A and B delineate the boundary between epidermal cells (EPI) and the dermis where lymphoma cells are located. c Model for TAK1 inhibition as therapy for CTCL. Based on our data, NF-κB is activated in CTCL, but partially counteracted by the PP1/MYPT1 phosphatase activity. Cytokines (TNFα) or other factors promote a ROCK1-dependent MYPT1 phosphorylation leading to TAK1 activation. Active TAK1 regulates NF-κB and β-catenin signaling, leading to increased cell survival. Thus, inactivation of TAK1 kinase activity promotes apoptosis in CTCL cells
Collectively, our results support the concept that CTCL cells display basal TAK1 activation, which is attenuated by the action of the MYPT1/PP1 phosphatase complex. Remaining active TAK1 is sufficient to sustain NF-κB and β-catenin signaling in CTCL, thus promoting cell survival and proliferation. We have here identified various residues where phosphorylation is required for TAK1 activation in lymphoma cells, and we generated specific antibodies against them to detect activated TAK1 in human samples. Our data indicate that patient stratification based on P-TAK1 status and, alternatively, on the simultaneous activation of β-catenin and NF-κB in the tumor component might provide a solid principle for therapeutic prescription. Thus, patients carrying lymphoma cells with specific phosphorylation of TAK1 associated with active NF-κB and β-catenin would benefit from treatments targeting the TAK1 kinase or, alternatively, the combined treatment with ROCK/NF-κB and β-catenin inhibitors (Fig. 6c), which would also allow modulating therapy toxicity by adjusting the administration protocols. Several inhibitors related to these pathways are currently being investigated as anti-cancer agents for other types of tumors. The actual possibility of stratifying patients based on P-TAK1 patterns might facilitate their future application. Further investigation will involve the implementation of SS patient-derived in vivo models to validate our results and directly test therapeutic strategies based on P-TAK1 status. We are also currently searching for other targetable TAK1 substrates, or upstream regulators, that might be relevant for preventing tumor initiation of progression.
References
Zinzani PL, Bonthapally V, Huebner D, Lutes R, Chi A, Pileri S. Panoptic clinical review of the current and future treatment of relapsed/refractory T-cell lymphomas: cutaneous T-cell lymphomas. Crit Rev Oncol/Hematol. 2016;99:228–240.
Vieyra-Garcia PA, Wei T, Naym DG, Fredholm S, Fink-Puches R, Cerroni L, et al. STAT3/5-Dependent IL9 Overexpression contributes to neoplastic cell survival in mycosis fungoides. Clin Cancer Res 2016;22(13):3328–39.
van der Fits L, Qin Y, Out-Luiting JJ, Vermeer KG, Whittaker S, van Es JH, et al. NOTCH1 signaling as a therapeutic target in Sezary syndrome. J Invest Dermatol. 2012;132(12):2810–7.
McKenzie RC, Jones CL, Tosi I, Caesar JA, Whittaker SJ, Mitchell TJ. Constitutive activation of STAT3 in Sezary syndrome is independent of SHP-1. Leukemia. 2012;26(2):323–31.
Kamstrup MR, Gjerdrum LM, Biskup E, Lauenborg BT, Ralfkiaer E, Woetmann A, et al. Notch1 as a potential therapeutic target in cutaneous T-cell lymphoma. Blood. 2010;116(14):2504–12.
Gallardo F, Sandoval J, Diaz-Lagares A, Garcia R, D’Altri T, Gonzalez J, et al. Notch1 pathway activation results from the epigenetic abrogation of notch-related MicroRNAs in Mycosis Fungoides. J Invest Dermatol. 2015;135(12):3144–52.
Bellei B, Cota C, Amantea A, Muscardin L, Picardo M. Association of p53 Arg72Pro polymorphism and beta-catenin accumulation in mycosis fungoides. Br J Dermatol. 2006;155(6):1223–9.
da Silva Almeida AC, Abate F, Khiabanian H, Martinez-Escala E, Guitart J, Tensen CP, et al. The mutational landscape of cutaneous T cell lymphoma and Sezary syndrome. Nat Genet. 2015;47(12):1465–70.
Ungewickell A, Bhaduri A, Rios E, Reuter J, Lee CS, Mah A, et al. Genomic analysis of mycosis fungoides and Sezary syndrome identifies recurrent alterations in TNFR2. Nat Genet. 2015;47(9):1056–60.
Choi J, Goh G, Walradt T, Hong BS, Bunick CG, Chen K, et al. Genomic landscape of cutaneous T cell lymphoma. Nat Genet. 2015;47(9):1011–9.
Braun FC, Grabarczyk P, Mobs M, Braun FK, Eberle J, Beyer M, et al. Tumor suppressor TNFAIP3 (A20) is frequently deleted in Sezary syndrome. Leukemia. 2011;25(9):1494–501.
Davis RE, Brown KD, Siebenlist U, Staudt LM. Constitutive nuclear factor kappaB activity is required for survival of activated B cell-like diffuse large B cell lymphoma cells. J Exp Med. 2001;194(12):1861–74.
Davis RE, Ngo VN, Lenz G, Tolar P, Young RM, Romesser PB, et al. Chronic active B-cell-receptor signalling in diffuse large B-cell lymphoma. Nature. 2010;463(7277):88–92.
Doerre S, Corley RB. Constitutive nuclear translocation of NF-kappa B in B cells in the absence of I kappa B degradation. J Immunol. 1999;163(1):269–77.
Guo X, Koff JL, Moffitt AB, Cinar M, Ramachandiran S, Chen Z, et al. Molecular impact of selective NFKB1 and NFKB2 signaling on DLBCL phenotype. Oncogene. 2017;36(29):4224–32.
Pham LV, Tamayo AT, Yoshimura LC, Lo P, Terry N, Reid PS, et al. A CD40 Signalosome anchored in lipid rafts leads to constitutive activation of NF-kappaB and autonomous cell growth in B cell lymphomas. Immunity. 2002;16(1):37–50.
Sagaert X, Van Cutsem E, De Hertogh G, Geboes K, Tousseyn T. Gastric MALT lymphoma: a model of chronic inflammation-induced tumor development. Nat Rev Gastroenterol Hepatol. 2010;7(6):336–46.
Willis TG, Jadayel DM, Du MQ, Peng H, Perry AR, Abdul-Rauf M, et al. Bcl10 is involved in t(1;14)(p22; q32) of MALT B cell lymphoma and mutated in multiple tumor types. Cell. 1999;96(1):35–45.
Chang TP, Poltoratsky V, Vancurova I. Bortezomib inhibits expression of TGF-beta1, IL-10, and CXCR4, resulting in decreased survival and migration of cutaneous T cell lymphoma cells. J Immunol. 2015;194(6):2942–53.
Chang TP, Vancurova I. NFkappaB function and regulation in cutaneous T-cell lymphoma. Am J Cancer Res. 2013;3(5):433–45.
Li L, Ruan Q, Hilliard B, Devirgiliis J, Karin M, Chen YH. Transcriptional regulation of the Th17 immune response by IKK(alpha). J Exp Med. 2011;208(4):787–96.
Chang JH, Xiao Y, Hu H, Jin J, Yu J, Zhou X, et al. Ubc13 maintains the suppressive function of regulatory T cells and prevents their conversion into effector-like T cells. Nat Immunol. 2012;13(5):481–90.
Ren H, Schmalstieg A, van Oers NS, Gaynor RB. I-kappa B kinases alpha and beta have distinct roles in regulating murine T cell function. J Immunol. 2002;168(8):3721–31.
Voll RE, Jimi E, Phillips RJ, Barber DF, Rincon M, Hayday AC, et al. NF-kappa B activation by the pre-T cell receptor serves as a selective survival signal in T lymphocyte development. Immunity. 2000;13(5):677–89.
Senftleben U, Li ZW, Baud V, Karin M. IKKbeta is essential for protecting T cells from TNFalpha-induced apoptosis. Immunity. 2001;14(3):217–30.
Schmidt-Supprian M, Courtois G, Tian J, Coyle AJ, Israel A, Rajewsky K, et al. Mature T cells depend on signaling through the IKK complex. Immunity. 2003;19(3):377–89.
Espinosa L, Cathelin S, D’Altri T, Trimarchi T, Statnikov A, Guiu J, et al. The Notch/Hes1 pathway sustains NF-kappaB activation through CYLD repression in T cell leukemia. Cancer Cell. 2010;18(3):268–81.
Sun SC, Maggirwar SB, Harhaj E. Activation of NF-kappa B by phosphatase inhibitors involves the phosphorylation of I kappa B alpha at phosphatase 2A-sensitive sites. J Biol Chem. 1995;270(31):18347–51.
Menon SD, Qin S, Guy GR, Tan YH. Differential induction of nuclear NF-kappa B by protein phosphatase inhibitors in primary and transformed human cells. Requirement for both oxidation and phosphorylation in nuclear translocation. J Biol Chem. 1993;268(35):26805–12.
Li HY, Liu H, Wang CH, Zhang JY, Man JH, Gao YF, et al. Deactivation of the kinase IKK by CUEDC2 through recruitment of the phosphatase PP1. Nat Immunol. 2008;9(5):533–41.
Brechmann M, Mock T, Nickles D, Kiessling M, Weit N, Breuer R, et al. A PP4 holoenzyme balances physiological and oncogenic nuclear factor-kappa B signaling in T lymphocytes. Immunity. 2012;37(4):697–708.
Gu M, Ouyang C, Lin W, Zhang T, Cao X, Xia Z, et al. Phosphatase holoenzyme PP1/GADD34 negatively regulates TLR response by inhibiting TAK1 serine 412 phosphorylation. J Immunol. 2014;192(6):2846–56.
Ouyang C, Nie L, Gu M, Wu A, Han X, Wang X, et al. Transforming growth factor (TGF)-beta-activated kinase 1 (TAK1) activation requires phosphorylation of serine 412 by protein kinase A catalytic subunit alpha (PKACalpha) and X-linked protein kinase (PRKX). J Biol Chem. 2014;289(35):24226–37.
Margalef P, Colomer C, Villanueva A, Montagut C, Iglesias M, Bellosillo B, et al. BRAF-induced tumorigenesis is IKKalpha-dependent but NF-kappaB-independent. Sci Signal. 2015;8(373):ra38.
Kaveri D, Kastner P, Dembele D, Nerlov C, Chan S, Kirstetter P. beta-Catenin activation synergizes with Pten loss and Myc overexpression in Notch-independent T-ALL. Blood. 2013;122(5):694–704.
Giambra V, Jenkins CE, Lam SH, Hoofd C, Belmonte M, Wang X, et al. Leukemia stem cells in T-ALL require active Hif1alpha and Wnt signaling. Blood. 2015;125(25):3917–27.
Gekas C, D’Altri T, Aligue R, Gonzalez J, Espinosa L, Bigas A. beta-Catenin is required for T-cell leukemia initiation and MYC transcription downstream of Notch1. Leukemia. 2016;30(10):2002–10.
Bellei B, Pacchiarotti A, Perez M, Faraggiana T. Frequent beta-catenin overexpression without exon 3 mutation in cutaneous lymphomas. Mod Pathol. 2004;17(10):1275–81.
Singh A, Sweeney MF, Yu M, Burger A, Greninger P, Benes C, et al. TAK1 inhibition promotes apoptosis in KRAS-dependent colon cancers. Cell. 2012;148(4):639–50.
Birukova AA, Smurova K, Birukov KG, Usatyuk P, Liu F, Kaibuchi K, et al. Microtubule disassembly induces cytoskeletal remodeling and lung vascular barrier dysfunction: role of Rho-dependent mechanisms. J Cell Physiol. 2004;201(1):55–70.
Chen M, Ma L, Hall JE, Liu X, Ying Z. Dual regulation of tumor necrosis factor-alpha on myosin light chain phosphorylation in vascular smooth muscle. Am J Physiol Heart Circ Physiol. 2015;308(5):H398–406.
Kimura K, Ito M, Amano M, Chihara K, Fukata Y, Nakafuku M, et al. Regulation of myosin phosphatase by Rho and Rho-associated kinase (Rho-kinase). Science. 1996;273(5272):245–8.
Khasnis M, Nakatomi A, Gumpper K, Eto M. Reconstituted human myosin light chain phosphatase reveals distinct roles of two inhibitory phosphorylation sites of the regulatory subunit, MYPT1. Biochemistry. 2014;53(16):2701–9.
Aburima A, Wraith KS, Raslan Z, Law R, Magwenzi S, Naseem KM. cAMP signaling regulates platelet myosin light chain (MLC) phosphorylation and shape change through targeting the RhoA-Rho kinase-MLC phosphatase signaling pathway. Blood. 2013;122(20):3533–45.
Arthur WT, Noren NK, Burridge K. Regulation of Rho family GTPases by cell-cell and cell-matrix adhesion. Biol Res. 2002;35(2):239–46.
Hoberg JE, Yeung F, Mayo MW. SMRT derepression by the IkappaB kinase alpha: a prerequisite to NF-kappaB transcription and survival. Mol Cell. 2004;16(2):245–55.
Wu X, Sells RE, Hwang ST. Upregulation of inflammatory cytokines and oncogenic signal pathways preceding tumor formation in a murine model of T-cell lymphoma in skin. J Invest Dermatol. 2011;131(8):1727–34.
Kadin ME, Cavaille-Coll MW, Gertz R, Massague J, Cheifetz S, George D. Loss of receptors for transforming growth factor beta in human T-cell malignancies. Proc Natl Acad Sci USA. 1994;91(13):6002–6.
Nakahata S, Yamazaki S, Nakauchi H, Morishita K. Downregulation of ZEB1 and overexpression of Smad7 contribute to resistance to TGF-beta1-mediated growth suppression in adult T-cell leukemia/lymphoma. Oncogene. 2010;29(29):4157–69.
Acknowledgements
We want to thank the Bigas’ lab members for constructive discussions and suggestions. Proteomics analyses were perfomed at the CRG/UPF Proteomics Unit of the Centre de Regulació Genòmica (CRG), Universitat Pompeu Fabra (UPF), 08003 Barcelona, which is part of the Plataforma de Recursos Biomoleculares y Bioinformáticos del Instituto de Salud Carlos III (PT13/0001). This work was supported by grants from Instituto de Salud Carlos III FEDER (PT13/0002/002, PT13/0010/0005, PI13/00448 and PI16/00437), Ministerio de Economía y Competitividad (SAF2016-75613-R), Fundació la Marató de TV3 (20131210); and the “Xarxa de Bancs de tumors sponsored by Pla Director d’Oncologia de Catalunya (XBTC).
Author contributions
FG, JB, EL-A, JG, FT, MG, and SM performed experiments and analyzed data. IS, LC, MI, LFS, and RMP provided vital reagents and analyzed data. AB and LE designed the research, analyzed data, and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gallardo, F., Bertran, J., López-Arribillaga, E. et al. Novel phosphorylated TAK1 species with functional impact on NF-κB and β-catenin signaling in human Cutaneous T-cell lymphoma. Leukemia 32, 2211–2223 (2018). https://doi.org/10.1038/s41375-018-0066-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0066-4
This article is cited by
-
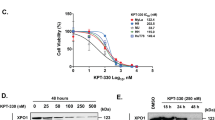
Transcriptomic and proteomic analysis of tumor suppressive effects of GZ17-6.02 against mycosis fungoides
Scientific Reports (2024)
-
Neues zu Pathogenese und molekularem Verständnis bei kutanen T-Zell-Lymphomen
Die Dermatologie (2022)
-
Protein phosphatases in TLR signaling
Cell Communication and Signaling (2021)