Abstract
Aberrant activation of the JAK3-STAT signaling pathway is a characteristic feature of many hematological malignancies. In particular, hyperactivity of this cascade has been observed in natural killer/T-cell lymphoma (NKTL) cases. Although the first-in-class JAK3 inhibitor tofacitinib blocks JAK3 activity in NKTL both in vitro and in vivo, its clinical utilization in cancer therapy has been limited by the pan-JAK inhibition activity. To improve the therapeutic efficacy of JAK3 inhibition in NKTL, we have developed a highly selective and durable JAK3 inhibitor PRN371 that potently inhibits JAK3 activity over the other JAK family members JAK1, JAK2, and TYK2. PRN371 effectively suppresses NKTL cell proliferation and induces apoptosis through abrogation of the JAK3-STAT signaling. Moreover, the activity of PRN371 has a more durable inhibition on JAK3 compared to tofacitinib in vitro, leading to significant tumor growth inhibition in a NKTL xenograft model harboring JAK3 activating mutation. These findings provide a novel therapeutic approach for the treatment of NKTL.
Similar content being viewed by others
Introduction
Natural killer/T-cell lymphoma (NKTL) is a rare and aggressive non-Hodgkin lymphoma more prevalent in Asian and Native American populations [1, 2]. The disease affects predominantly in males and is associated with the Epstein–Barr virus infection [3, 4]. NKTL usually presents as extranodal disease in the upper aerodigestive tract and is characterized by a prominent necrosis and cytotoxic phenotype [4]. Despite the use of multi-agent chemotherapy and involved-field radiotherapy, the outcome of NKTL remains poor with 5-year overall survival of 40 and 10% for nasal and non-nasal NKTL, respectively [5,6,7]. In addition, many patients develop resistance to chemotherapy, especially those with late-stage disease [8]. To improve the clinical outcomes, the current strategy is to intensify treatment with combination chemotherapy, chemo-radiotherapy, and stem cell transplantation; however, at the expense of increased toxicity [9,10,11,12]. With no targeted therapies available, the recent research efforts has focused on understanding the molecular mechanisms underlying NKTL pathogenesis to improve patient outcome and reduce the toxicities associated with the treatments.
Janus kinase 3 (JAK3) belongs to a family of cytoplasmic non-receptor tyrosine kinases. JAK family members include JAK1, JAK2, JAK3, and TYK2, all of which play a role in different cytokine and growth factor receptor-mediated signaling pathways [13]. JAK3 intracellularly associates with the common gamma chain (also known as interleukin-2 receptor subunit gamma or (IL-2Rγ)) to initiate the signaling upon cytokine binding [14]. Although most JAK family members are ubiquitously expressed, JAK3 is mainly restricted to the hematopoietic lineage where it plays a vital role in lymphoid cell development and homeostasis [15]. Recent studies have indicated that constitutive activation of the JAK-STAT signaling is a characteristic feature of many hematological neoplasms. JAK3 activating mutations that result in persistent activation of the JAK-STAT signaling have been described in various leukemias and lymphomas, including NKTL, monomorphic epitheliotropic intestinal T-cell lymphoma, T-cell acute lymphoblastic leukemia, and hepatosplenic T-cell lymphoma [16,17,18,19,20,21,22,23]. The aberrant activation of JAK3 in hematological malignancies suggests JAK3 as a potential target for cancer therapy. However, targeting JAK3 selectively with small-molecule inhibitors has been challenging due to the highly conserved amino acid composition of the ATP binding pocket among the JAK family members [24].
The cysteine residue at position 909 of JAK3 (Cys909) is one of the three amino acids uniquely present in the ATP binding pocket of JAK3, providing an opportunity to achieve high JAK3 selectivity against JAK1, JAK2, and TYK2. By employing chemistry that allows covalent cysteine binding and utilizing a specific scaffold for stabilization [24,25,26], we produced a highly selective JAK3 inhibitor PRN371. In this study, we evaluated the therapeutic effect of PRN371 in NKTL cells in vitro and in vivo to investigate the potential of JAK3-targeted therapy in NKTL.
Results
PRN371 is a highly selective JAK3 inhibitor
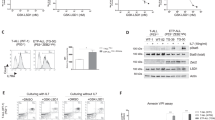
PRN371 is an acrylamide-based inhibitor that targets the JAK3 non-catalytic Cys909 residue (Fig. 1a). To assess its overall kinase selectivity, we evaluated PRN371 against a panel of 251 kinases at a concentration of 0.1 μM. Only four kinases, including JAK3, ARK5, CLK4, and JAK2, were inhibited by greater than 90% (Fig. 1b and Supplementary Table 1), and importantly the most potent results were observed for JAK3 (>98% inhibition). To further evaluate the selectivity of PRN371 specifically against the JAK kinases, we carried out cell-free enzymatic assays with each family member. PRN371 demonstrated a clear JAK3 selectivity over other JAK kinases (Fig. 1c). The compound was 282-fold, 378-fold, and 1194-fold more potent in repressing JAK3 activity compared to TYK2, JAK2, and JAK1, respectively. In contrast, tofacitinib, a pan-JAK inhibitor, blocked the activities of JAK1-3 at similar concentrations. Furthermore, using cell-based assays, we demonstrated that PRN371 inhibited IL-2-stimulated JAK1/3-STAT5 phosphorylation in human peripheral blood mononuclear cells (PBMCs) (IC50 = 99 nM) and IL-4-stimulated JAK1/3-STAT6 phosphorylation in Ramos B-cells (IC50 = 26 nM; Fig. 1d). However, IL-6-stimulated JAK2-STAT3 activity in PBMCs was not inhibited up to 3 μM concentration, indicating the specificity of PRN371 in inhibiting JAK3 signaling. Similarly to the in vitro kinase assay, tofacitinib suppressed JAK-STAT phosphorylation equally in all conditions tested. These findings indicate that PRN371 has a strong cellular selectivity against JAK3 over other members of the JAK family and suggest that this drug may be used in JAK3-targeted therapy.
In vitro kinase spectrum of PRN371. a Chemical structure of PRN371. b Kinase selectivity profile of PRN371. PRN371 was profiled at a concentration of 0.1 μM against a panel of 251 kinases. The kinases inhibited by greater than 90 and 98% are indicated by small and large red dots, respectively. The yellow dots represent the kinases that contain a conserved Cys residue with JAK3. The figure is reproduced by the courtesy of Cell Signaling Technology, Inc. TK: c In vitro kinase assays carried out with JAK family members and PRN371 or tofacitinib. PRN371 selectivity is expressed in reference to JAK3. d The efficacy and selectivity of PRN371 and tofacitinib in cell-based assays. Cells were stimulated with IL-2, IL-4, or IL-6 and treated with the drugs to evaluate the IC50 value of induced phosphorylation. ND not determined, PBMC peripheral blood mononuclear cells
PRN371 effectively inhibits the growth of JAK3-driven cancer cells
To investigate whether PRN371’s in vitro selectivity translates into differential efficacy in a cellular context, we evaluated its ability to inhibit cell growth in two cell lines that harbor JAK3-activating mutations, NK-S1 and CMK [16, 17, 27]. PRN371 dose dependently suppressed the growth of both cell lines, whereas no effect was detected in K562 and HEL92, which are driven by BCR-ABL gene fusion and JAK2-activating mutation, respectively (Fig. 2a, b). Similar effects were observed with tofacitinib (Fig. S1A, 1B). Importantly, blocking of constitutively activated JAK3 by PRN371 in NK-S1 and CMK cells led to the downregulation of its downstream effector proteins, phosphorylated STAT3 and STAT5 (Fig. 2c). This effect was not observed in K562 and HEL92 cells, further supporting their resistance to PRN371 treatment (Fig. 2d). To investigate whether the growth inhibition by PRN371 is specifically through STAT3/5 activity, we ectopically expressed STAT3 wild-type or mutant (D661Y) in NK-S1 and treated these cells with PRN371. Compared to cells with empty vector or wild-type STAT3, constitutive activation of mutant STAT3 rescued the inhibition of STAT3/5 phosphorylation, leading to resistance of NK-S1 to PRN371 treatment (Fig. S1C, 1D). Taken together, these results demonstrate that PRN371 effectively suppresses the viability of JAK3-driven cancer cells, mainly by targeting the JAK3-STAT3/5 signaling pathway.
PRN371 effectively blocks JAK3-mediated signaling in JAK3-driven cancer cells. a IC50 of PRN371 and tofacitinib in JAK3-dependent and JAK3-independent cells was determined by cell viability assays using 2-fold dilution series ranging from 10 nM to 10 μM concentration. b Dose-dependent response of JAK3-driven cells to PRN371 treatment. Cells were treated with the indicated concentration for 96 h. Cell viability is normalized to the control (DMSO) and presented as mean ± s.d. The effect of PRN371 on the downstream effector proteins of the JAK3-STAT pathway in c JAK3-dependent and d JAK3-independent cells. Cells were treated with the indicated concentration for 2 h
PRN371 effectively inhibits the growth of NKTL cells via suppressing the JAK3-STAT pathway
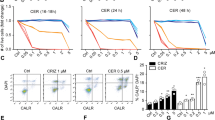
Frequent somatic mutations and/or hyperactivity of JAK3 have been demonstrated in NKTL and the suppression of JAK3 by pan-JAK inhibitor tofacitinib was shown to reduce cell viability by inducing apoptosis [17, 18]. To establish the potential of PRN371 use in NKTL, we first profiled the endogenous activity of JAK3-STAT pathway in a panel of nine NKTL cell lines. The majority of cell lines displayed hyper-activation of the signaling cascade as demonstrated by high expression of phospho-JAK3 and its downstream effector proteins phospho-STAT3/5 (Fig. 3a). Importantly, no hyper-activation was observed in normal NK cells, presenting an opportunity to specifically target the malignant cells. To determine the efficacy of PRN371 in NKTL, we treated all nine cell lines with various concentrations to establish the IC50 (Fig. 3b). The potency of PRN371 was in the nanomolar range (IC50 values) in all NKTL cells. The observed efficacies were similar to those of tofacitinib-treated cells, indicating that both drugs induced a strong growth inhibition on the NKTL cells (Fig. 3b). Interestingly, NKTL cell lines harboring STAT3-activating mutations (Supplementary Table 2) such as NYKS, SNK6, SNT8, and HANK1 (labeled with asterisk) [28, 29] are generally less sensitive to PRN371 treatment, suggesting a potential resistant mechanism that is consistent with the rescued experiments with STAT3-activating mutations (Fig. S1C, 1D). In addition, PRN371, similarly to tofacitinib, also dose-dependently suppressed the downstream signaling in the JAK3 wild-type cells, KAI-3 and NK92, which are dependent on IL-2 signaling (Fig. 3c, d). Surprisingly, PRN371 treatment resulted in upregulation of JAK3 phosphorylation in both NKTL cells, suggesting PRN371 can lead to an increase in JAK3 activation loop phosphorylation, despite inhibition of JAK3 downstream effectors. Taken together, these results suggest that PRN371 inhibits cell proliferation and suppresses JAK3-STAT3/5 signaling in NKTL cells with hyperactivated JAK3-STAT-mediated signaling cascade.
PRN371 effectively reduces the viability of NKTL cells by suppressing JAK3-mediated signaling. a Immunoblots of total and phosphorylated JAK3, STAT5, and STAT3 demonstrating constitutive activation of the JAK3-STAT signaling in NKTL. b IC50 of PRN371 and tofacitinib was determined by cell viability assays using 2-fold dilution series ranging from 10 nM to 10 μM concentration. The asterisk indicates cell lines that harbor STAT3-activating mutations. Western blot analysis of total and phosphorylated JAK3, STAT5, and STAT3 levels in NK92 and KAI-3 cells treated with the indicated concentrations of c PRN371 or d tofacitinib. Cells were cultured overnight without IL-2 and stimulated with IL-2 for 2 h with or without the indicated drug
PRN371 inhibits NKTL cell growth by inducing apoptosis and blocking colony formation ability
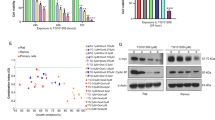
We next performed various cellular assays to understand the mechanism of PRN371-induced growth inhibition in NKTL cells. Treatment of JAK3-driven NK-S1 cell line with PRN371 resulted in potent G1 cell cycle arrest and cell death (Fig. 4a, upper panel). Annexin V staining revealed that, similarly to tofacitinib, PRN371-induced cell death was due to increased apoptosis (Fig. 4a, lower panel). In addition, both PRN371 and tofacitinib also induced cell death in JAK3 wild-type NKTL cells but not in JAK3-independent non-NKTL cells (Fig. 4b), further support that both drugs act via activated JAK3-STAT signaling. Moreover, both PRN371 and tofacitinib also reduced the colony formation ability of both JAK3 mutant and JAK3 wild-type NKTL cells (Fig. 4c). Although PRN371 is less effective in NKTL cells harboring STAT3-activating mutations (Fig. 3b), our findings demonstrated that it might still have partial effect, most probably via downstream STAT5 (Fig. S1C).
In vitro characterization of PRN371-induced growth inhibition in NKTL cells. a Cell cycle analysis of NK-S1 cells treated with 1.0 μM of PRN371 or tofacitinib for 72 h. Staining was performed using propidium-iodide and counted using flow cytometry (upper panel). The apoptotic cells were detected using Annexin V staining (lower panel). b Sub-G1 population analysis in NKTL and non-NKTL cells treated with 1.0 μM PRN371 or tofacitinib for 72 h. c Colony formation assay in NKTL cells treated with 1.0 μM PRN371 or tofacitinib for 14 days. DMSO-treated cells served as controls in all experiments
PRN371 exerts durable inhibition of JAK3 activity
We next proceeded to determine the durability of JAK3 inhibition by PRN371 in comparison to tofacitinib using a wash-out experiment. Pre-treatment of JAK3-driven NK-S1 cells with 1.0 μM PRN371 for 2 h was sufficient to completely inhibit STAT3/5 phosphorylation up to 4 h after washout of the drug (Fig. 5a). Although PRN371 and tofacitinib demonstrated comparable efficacy, NK-S1 cells treated similarly with tofacitinib showed recovery of phosphorylated STAT3/5 activity within half an hour post inhibitor washout, an effect expected for a non-covalent inhibitor. Similar results were also observed in JAK3 wild-type NKTL cells, indicating that the effect is independent of JAK3 mutation status (Fig. 5b). Together, these findings demonstrate that despite similar anti-tumorigenic activity in NKTL cells, PRN371 has a longer-lasting effect on JAK3 inhibition due to the covalent nature of the target engagement.
The durability of JAK3 inhibition in vitro and in vivo, and the efficacy of PRN371 in vivo. Wash-out experiment of NK-S1 a and KAI-3 b cells treated with 1.0 µM of PRN371 or tofacitinib for 2 h and harvested at different time points as indicated. DMSO treatment served as control. c Immunoblots of JAK3-STAT signaling activity in Nod/Scid mice bearing NK-S1 tumors treated with vehicle or 25 or 50 mg/kg of PRN371. The tumors were harvested at indicated time points after the treatment. d Mice bearing NK-S1 tumors were treated with the indicated concentrations of PRN371 for 14 days. Tumor growth is presented as mean ± s.d.
PRN371 is effective in an in vivo NKTL model
Due to the short half-life of tofacitinib, the reported in vivo effects have been derived from continuous administration of the drug via a subcutaneous mini-osmotic pump [18, 30]. To assess the clinical potential of PRN371, we determined its exposure following oral administration and intraperitoneal (i.p.) injection (50 mg/kg). PRN371 was quickly absorbed, reaching peak plasma concentration of 1000 ng/ml and 1 × 104 ng/ml for oral and i.p. route, respectively, at 30 min (Fig. S2A). The circulating levels of PRN371 was 10-fold higher when administered intraperitoneally compared to oral route, and was maintained at higher exposure levels for the 24 h. Thus, the i.p. route is preferred for administration of the drug. To access the potential toxicity with PRN371, the inhibitor was administered intraperitoneally into mice for 14 consecutive days. The liver function tests were performed to evaluate the liver enzymes, such as alanine aminotransferase and aspartate aminotransferase, in both mice groups treated with vehicle and PRN371. PRN371 did not induce hepatic toxicity after a prolong exposure (Fig. S2B). In addition, full blood count test indicated that there were no significant changes in white blood cell and hemoglobin between mice treated with vehicle and PRN371 (Fig. S2C).
To determine the ability of PRN371 to penetrate the tumor and exert its anti-JAK3 inhibition in vivo, we assessed the pharmacodynamics of PRN371 in NK-S1 subcutaneous xenograft model. Mice bearing an average tumor size of 300 mm3 were injected intraperitoneally with 50 mg/kg PRN371 and tumor biopsies were collected 2, 8, and 24 h post treatment. Consistent with the in vitro results, PRN371-treated tumors showed suppression of phosphor-STAT5 occurred within 2 h after treatment and persisted up to 24 h, while downregulation of phospho-STAT3 became evident 8 h post treatment (Fig. 5c), confirming the ability of PRN371 to block the JAK3-STAT3/5 signaling in vivo and the effect persisted more than 24 h.
Lastly, we evaluated the anti-tumorigenic potential of PRN371 in NK-S1 subcutaneous xenograft model. The inhibitor was administered intraperitoneally into tumor-bearing mice (average tumor volume 200–300 mm3) for 14 consecutive days. Treatment with both 25 and 50 mg/kg PRN371 decreased the tumor volume as early as on the 6th day and significant difference in tumor size was observed after 8 days of treatment (p < 0.05, Student’s t-test, Fig. 5d). Importantly, there was no significant reduction in body weight, change in liver and kidney functions, and total blood counts, indicating tolerable side effects (Fig. S2B–D). Collectively, these data suggest that inhibition of JAK3 by PRN371 is an effective strategy for the treatment of NKTL.
Discussion
NKTL is an aggressive T-cell malignancy with an unmet medical need. The discovery of frequent somatic mutations in genes associated with the JAK-STAT pathway has raised the possibility for a targeted approach to improve clinical outcome and engender hopes for a targeted therapy that will emulate the success of radiation in early, localized NKTL treatment. Our sequencing effort in this lymphoma has led to the discovery of frequent JAK3 activating mutation, thus the development of JAK3-specific inhibitor.
In this study, we demonstrated for the first time the preclinical evaluation of a novel chemical entity, PRN371, a small molecule designed as a potent and selective JAK3 inhibitor, which specifically binds to cysteine 909 pocket binding site of JAK3 kinase. Both in vitro and in vivo data showed that PRN371 is highly selectivity for JAK3 over other members of the JAK family and has high efficacy for anti-tumor activity. Compared with tofacitinib, a pan-JAK inhibitor, PRN371 showed high selectivity for JAK3 inhibition, suggesting that PRN371 may have a more favorable anti-tumor activity and fewer side effects in targeted therapy than those experienced clinically with tofacitinib. Furthermore, its covalent and irreversible engagement of target cysteine enabled a more durable drug response than non-covalent inhibitors such as tofacitinib. We noted that PRN371 induces the activation of an auto-feedback loop and increases the phosphorylation of JAK3. Similar findings are observed in most of type I JAK inhibitors, suggesting that PRN371 may function as a type I JAK inhibitor. This type I binding mode can result in an increase in JAK activation loop phosphorylation [31,32,33]. Importantly, the auto-feedback loop effect did not affect the efficacy of PRN371 as demonstrated by its ability to downregulate phosphorylation of STAT3 and STAT5 protein.
Despite controversy in JAK3 mutation frequencies being reported among different NKTL cohorts [17, 18, 28, 29], the signaling cascade as a whole is still hyper activated. There is considerable cross-talk between various signaling pathways in this disease and further complicated by somatic mutations in other genes associated with the JAK-STAT pathway, such as STAT3 and JAK1 [28, 29]. We demonstrated that PRN371 is effective in the majority of NKTL cells including most STAT3 mutants suggesting that PRN371 might not be just acting on the JAK3-STAT3/5 pathway, but other pathways downstream of JAK3 such as the EZH2 pathway, which recently been demonstrated in NKTL [34]. SNK6 showed resistance to PRN371 treatment most probably because it harbors D661Y activating mutation in the STAT3 gene, suggesting that patient selection is necessary in future clinical trial. This may be an important area for future investigation to shed light on the mechanisms of SH2 binding sites involved with STAT3 activation as conformational structure and plasticity of STAT3 have been known to play important roles in downstream events [35, 36]. Complete inhibition of tumor growth was not achieved in this study, which may be attributed to PRN371 under-dosing which occurred from day 10 onward due to volume loss from repeated i.p. administrations in the study animals. Since several signaling pathways are known to be altered in this cancer, we suspect that single agent therapy might not be able to achieve complete remission and a combination therapy approach targeting several pathways or in combination with radiotherapy might be more appropriate for treating advance malignancy.
Taken together, PRN371 is a potent inhibitor of JAK3 that exhibits anti-cancer properties against NKTL both in vitro and in vivo. The identification of NKTL cell lines harboring STAT3-activating mutations correlating to PRN371 resistance highlighted the importance of genetic screening, suggesting a selected subset of patients with appropriate molecular stratification may use PRN371 as an alternative treatment to current chemotherapy. Translation of these findings into early clinical trials should shed light on the potential clinical impact and validity of such a strategy.
Materials and methods
Cell lines and reagents
NK-S1 was generated in-house [27]. KAI-3, KHYG1, and CMK cell lines were purchased from JCRB Cell Bank, and NK92, K562, HEL92, and Ramos from ATCC. SNK1, SNK6, SNT8, and NKYS were generously provided by Dr. Norio Shimizu and HANK1 by Dr. Yoshitoyo Kagami. NK-S1 and K562 were cultured in DMEM medium (Life Technologies) supplemented with 10% FBS (Hyclone) and 10% equine serum (Life Technologies). HANK1 and SNT8 were maintained in Artemis medium-2 (ScyMed Inc.) supplemented with 2% human plasma serum (Clontech) and 100 IU/ml or 700 IU/ml IL-2 (Miltenyi Biotec), respectively. All remaining cell lines were cultured in RPMI1640 medium (Life Technologies) supplemented with 10% FBS. 100 IU/ml IL-2 was included into KAI-3, KHYG1, and NKSY growth media, and 10% equine serum and 100 IU/ml IL-2 into NK92, SNK1, and SNK6 growth media. All cultures were routinely checked for mycoplasma contamination. NK-cells from health donor are separated with NK cell isolation kit (Miltenyi Biotec). PBMCs are isolated by the Ficoll-Paque method from human whole blood. Tofacitinib (CP-690550) was purchased from Selleck Chemicals.
In vitro kinase assay
LabChip 3000 Drug Discovery System (Caliper Life Sciences) was utilized for the kinase assay to separate the phosphorylated peptides from the unphosphorylated ones. The reaction mixture for each kinase assay consisted of enzyme, substrate, and cofactors, specifically: 100 mM Hepes pH 7.5, 0.1% BSA, 0.01% Triton X-100, 1 mM DTT, 10 mM MgCl2, 10 μM sodium orthovanadate, 10 μM beta-glycerophosphate, a specific concentration of ATP optimized for each kinase, 1% DMSO (from test compound), 1 μM peptide substrate, the enzyme, and either 1 μM or 0.1 μM of test compound. The reaction was allowed to proceed for several hours at 25 °C and was terminated by addition of 20 mM EDTA. The reaction mix was run through capillary electrophoresis, excited by the blue laser (480 nM) and fluorescence intensity was measured both for the substrate and product peaks in each well of the microtiter plate. The kinase activity was evaluated as the product to sum ratio (PSR) given by P/(S + P), where P is the product height and S is the substrate height. Negative control (0% inhibition; no inhibitor) and positive control (100% inhibition; activity in 20 mM EDTA) were determined in replicates of four. The percent inhibition for each compound concentration was calculated using the following equation: Pinh = (PSR0%−PSRinh)/(PSR0%−PSR100%), where PSRinh is the PSR with inhibitor, PSR0% is the PSR with no inhibitor, and PSR100% is the PSR for fully inhibited samples. Experiments were conducted in duplicate at each compound concentration.
Cell viability assays
For each assay, 2000 cells were seeded on a 96-well plate and treated with indicated concentrations (10 nM to 10 μM) of PRN371 or tofacitinib for 96 h. Cell viability was measured using CellTiter-Glo Luminescent Cell Viability Assay (Promega) following manufacturer’s instructions. All experiments were performed in triplicate. IC50 values were calculated.
Cell cycle and apoptosis assays
For both assays, 2 × 105 cells were seeded on a 6-well plate and treated with 1.0 μM PRN371 or tofacitinib for 72 h. For the cell cycle analysis, the cells were fixed with 70% ethanol and stained with 50 μg/ml propidium iodide (Sigma-Aldrich). For the apoptosis assay, the cells were washed with 1× PBS and stained with Annexin V-FITC (BD Bioscience). The stained cells were analyzed by FACScalibur (BD Bioscience) and quantified using CellQuest software (BD Bioscience).
Colony formation assay
A total of 1 × 104 cells were suspended in DMEM containing 0.2% methylcellulose (Sigma-Aldrich) and 10% FBS, and layered on top of DMEM containing 0.6% agar, 10% FBS, and 1.0 μM PRN371 or tofacitinib on a 6-well plate. After 4 weeks, the colonies were stained with iodonitrotetrazolium chloride (Sigma-Aldrich) overnight. The experiment was performed in triplicates and images were acquired from randomly selected areas using Nikon Eclipse microscope image system.
Western blot analysis
A total of 2 × 105 cells were seeded on a 6-well plate, treated with 1.0 μM PRN371 or tofacitinib for 2 h and harvested for protein extraction. Cell lysis, protein separation, transfer, and visualization were performed as previously described [21]. Protein concentration was measured by Quick StartTM Bradford Protein Assay (Bio-Rad) and 15 μg were loaded on each polyacrylamide gel. The used antibodies are listed in Supplementary Table 3.
Wash-out assay
2.5 × 106 NK-S1 and KAI-3 cells were seeded on a 6-well plate and treated with 1.0 μM PRN371 or tofacitinib for 2 h. Cells were washed twice with 1× PBS and resuspended in complete growth media without the inhibitor. The cells were harvested at the time of washout or 0.5, 1, 2, or 4 h after the washout.
In vivo studies
For pharmacokinetic analysis, female NOD/SCID mice were administered 40 mg/kg PRN371 formulated in 6% Capmul/14% Cremophore EL using three mice per time point. Blood plasma exposure from each animal was analyzed by LC/MS/MS analysis methods using a Shimadzu LC20AD HLPC system connected to a Sciex API4000 QTrap mass spectrometer. Sciex Analyst software (version 1.6) was used for LC/MS/MS instrument control and acquisition. For efficacy studies, 5–7-week-old female NOD/SCID mice (InVivos) were kept under standard laboratory conditions according to the National Advisory Committee for Laboratory Animal Research guidelines. All experiments were approved by the SingHealth Institutional Animal Care and Use Committee. 5 × 106 NK-S1 cells were suspended in 0.1 ml of 1× PBS and injected subcutaneously into the left flank of each animal. For the pharmacodynamic experiment, 24 tumor-bearing mice (average tumor volume 300 mm3) were divided into four groups. One group was treated with 1× PBS and the remaining three groups with 50 mg/kg of PRN371. Mice were sacrificed and tumor biopsies collected 2, 8, and 24 h post treatment. For the anti-tumorigenicity experiment, tumor-bearing mice (average tumor volume 200 mm3) were used. PRN371 (25 or 50 mg/kg) was administered by intraperitoneal injection for 14 consecutive days. 1× PBS was used as the vehicle control. Tumor volume was monitored 2–3 times per week and body weight was measured daily. Tumor volume (V) was calculated as V = width2 × length × 0.537. Randomization was performed by equally dividing tumor-bearing mice of similar tumor burden into different groups for drug treatment. No statistical method was used to predetermine sample size and no experimental samples were excluded in this study.
Statistical analysis
All statistical analyses were performed with GraphPad Prism version 6.0 and statistical difference for tumor weight was calculated by two-sided Student’s t-test. Statistical significance was considered as p < 0.05. For animal experiments, no blinding was done for treatment and analyses.
References
Kwong YL, Anderson BO, Advani R, Kim WS, Levine AM, Lim ST. Management of T-cell and natural-killer-cell neoplasms in Asia: consensus statement from the Asian Oncology Summit 2009. Lancet Oncol. 2009;10:1093–101.
Niu SQ, Yang Y, Li YY, Wen G, Wang L, Li ZM, et al. Primary site and regional lymph node involvement are independent prognostic factors for early-stage extranodal nasal-type natural killer/T cell lymphoma. Chin J Cancer. 2016;35:34.
Li L, Zhang Y, Guo BB, Chan FK, Tao Q. Oncogenic induction of cellular high CpG methylation by Epstein-Barr virus in malignant epithelial cells. Chin J Cancer. 2014;33:604–8.
Chan JKC, Quintanilla-Martinez L, Ferry JA, Peh S-C. Extranodal NK/T-cell lymphoma, nasal type. In: Swerdlow SH CE, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J, Vardiman JW, editors. WHO classification of tumours of haematopoietic and lymphoid tissues, 4th edn. Lyon: International Agency for Research on Cancer; 2008. pp. 285–8.
Vose J, Armitage J, Weisenburger D. International peripheral T-cell and natural killer/T-cell lymphoma study: pathology findings and clinical outcomes. J Clin Oncol. 2008;26:4124–30.
Au WY, Weisenburger DD, Intragumtornchai T, Nakamura S, Kim WS, Sng I, et al. Clinical differences between nasal and extranasal natural killer/T-cell lymphoma: a study of 136 cases from the International Peripheral T-Cell Lymphoma Project. Blood. 2009;113:3931–7.
Suzuki R, Suzumiya J, Yamaguchi M, Nakamura S, Kameoka J, Kojima H, et al. Prognostic factors for mature natural killer (NK) cell neoplasms: aggressive NK cell leukemia and extranodal NK cell lymphoma, nasal type. Ann Oncol. 2010;21:1032–40.
Tse E, Kwong YL. How I treat NK/T-cell lymphomas. Blood. 2013;121:4997–5005.
Suzuki R, Suzumiya J, Nakamura S, Kagami Y, Kameoka JI, Sakai C, et al. Hematopoietic stem cell transplantation for natural killer-cell lineage neoplasms. Bone Marrow Transplant. 2006;37:425–31.
Yamaguchi M, Tobinai K, Oguchi M, Isobe Y, Ishizawa K, Maseki N, et al. Phase I/II study of concurrent chemoradiotherapy for localized nasal NK/T-cell lymphoma: final results of JCOG0211. J Clin Oncol. 2009;27:8549.
Kim SJ, Kim K, Kim BS, Kim CY, Suh C, Huh J, et al. Phase II trial of concurrent radiation and weekly cisplatin followed by VIPD chemotherapy in newly diagnosed, stage IE to IIE, nasal, extranodal NK/T-cell lymphoma: consortium for improving survival of lymphoma study. J Clin Oncol. 2009;27:6027–32.
Yamaguchi M, Kwong YL, Kim WS, Maeda Y, Hashimoto C, Suh C, et al. Phase II study of SMILE chemotherapy for newly diagnosed stage IV, relapsed, or refractory extranodal natural killer (NK)/T-cell lymphoma, nasal type: the NK-cell tumor study group study. J Clin Oncol. 2011;29:4410–6.
Yamaoka K, Saharinen P, Pesu M, Holt VE 3rd, Silvennoinen O, O’Shea JJ. The Janus kinases (Jaks). Genome Biol. 2004;5:253.
Hofmann SR, Lam AQ, Frank S, Zhou YJ, Ramos HL, Kanno Y, et al. Jak3-independent trafficking of the common gamma chain receptor subunit: chaperone function of Jaks revisited. Mol Cell Biol. 2004;24:5039–49.
Johnston JA, Kawamura M, Kirken RA, Chen YQ, Blake TB, Shibuya K, et al. Phosphorylation and activation of the Jak-3 Janus kinase in response to interleukin-2. Nature. 1994;370:151–3.
Walters DK, Mercher T, Gu TL, O’Hare T, Tyner JW, Loriaux M, et al. Activating alleles of JAK3 in acute megakaryoblastic leukemia. Cancer Cell. 2006;10:65–75.
Koo GC, Tan SY, Tang T, Poon SL, Allen GE, Tan L, et al. Janus kinase 3-activating mutations identified in natural killer/T-cell lymphoma. Cancer Discov. 2012;2:591–7.
Bouchekioua A, Scourzic L, de Wever O, Zhang Y, Cervera P, Aline-Fardin A, et al. JAK3 deregulation by activating mutations confers invasive growth advantage in extranodal nasal-type natural killer cell lymphoma. Leukemia. 2014;28:338–48.
Cornejo MG, Kharas MG, Werneck MB, Le Bras S, Moore SA, Ball B, et al. Constitutive JAK3 activation induces lymphoproliferative syndromes in murine bone marrow transplantation models. Blood. 2009;113:2746–54.
Bains T, Heinrich MC, Loriaux MM, Beadling C, Nelson D, Warrick A, et al. Newly described activating JAK3 mutations in T-cell acute lymphoblastic leukemia. Leukemia. 2012;26:2144–6.
Nairismagi ML, Tan J, Lim JQ, Nagarajan S, Ng CC, Rajasegaran V, et al. JAK-STAT and G-protein-coupled receptor signaling pathways are frequently altered in epitheliotropic intestinal T-cell lymphoma. Leukemia. 2016;30:1311–9.
Xu N, Emelyanov AV, Fyodorov DV, Skoultchi AI. Drosophila linker histone H1 coordinates STAT-dependent organization of heterochromatin and suppresses tumorigenesis caused by hyperactive JAK-STAT signaling. Epigenetics Chromatin. 2014;7:16.
McKinney M, Moffitt AB, Gaulard P, Travert M, De Leval L, Nicolae A, et al. The genetic basis of hepatosplenic T-cell lymphoma. Cancer Discov. 2017;7:369–79.
Goedken ER, Argiriadi MA, Banach DL, Fiamengo BA, Foley SE, Frank KE, et al. Tricyclic covalent inhibitors selectively target Jak3 through an active site thiol. J Biol Chem. 2015;290:4573–89.
Serafimova IM, Pufall MA, Krishnan S, Duda K, Cohen MS, Maglathlin RL, et al. Reversible targeting of noncatalytic cysteines with chemically tuned electrophiles. Nat Chem Biol. 2012;8:471–6.
Hill RJ BA, Bradshaw JM, Brameld K, Kim EO, Li X, et al. Discovery of a highly potent, selective reversible covalent inhibitor of JAK3 kinase. Arthritis Rheum. 2012;64:2326.
Loong SL, Hwang JS, Lim ST, Yap SP, Tao M, Chong TW, et al. An Epstein-Barr virus positive natural killer lymphoma xenograft derived for drug testing. Leuk Lymphoma. 2008;49:1161–7.
Kucuk C, Jiang B, Hu X, Zhang W, Chan JK, Xiao W, et al. Activating mutations of STAT5B and STAT3 in lymphomas derived from gammadelta-T or NK cells. Nat Commun. 2015;6:6025.
Lee S, Park HY, Kang SY, Kim SJ, Hwang J, Kwak SH, et al. Genetic alterations of JAK/STAT cascade and histone modification in extranodal NK/T-cell lymphoma nasal type. Oncotarget. 2015;6:17764–76.
Ju W, Zhang M, Jiang JK, Thomas CJ, Oh U, Bryant BR, et al. CP-690,550, a therapeutic agent, inhibits cytokine-mediated Jak3 activation and proliferation of T cells from patients with ATL and HAM/TSP. Blood. 2011;117:1938–46.
Andraos R, Qian Z, Bonenfant D, Rubert J, Vangrevelinghe E, Scheufler C, et al. Modulation of activation-loop phosphorylation by JAK inhibitors is binding mode dependent. Cancer Discov. 2012;2:512–23.
Meyer SC, Keller MD, Chiu S, Koppikar P, Guryanova OA, Rapaport F, et al. CHZ868, a type II JAK2 inhibitor, reverses type I JAK inhibitor persistence and demonstrates efficacy in myeloproliferative neoplasms. Cancer Cell. 2015;28:15–28.
Wu SC, Li LS, Kopp N, Montero J, Chapuy B, Yoda A, et al. Activity of the type II JAK2 inhibitor CHZ868 in B cell acute lymphoblastic leukemia. Cancer Cell. 2015;28:29–41.
Mason CC, Khorashad JS, Tantravahi SK, Kelley TW, Zabriskie MS, Yan D, et al. Age-related mutations and chronic myelomonocytic leukemia. Leukemia. 2016;30:906–13.
Becker S, Groner B, Muller CW. Three-dimensional structure of the Stat3beta homodimer bound to DNA. Nature. 1998;394:145–51.
Qing Y, Stark GR. Alternative activation of STAT1 and STAT3 in response to interferon-gamma. J Biol Chem. 2004;279:41679–85.
Acknowledgements
This work was supported by the Singapore Ministry of Health’s National Medical Research Council (NMRC/TCR/010-NCC/2013), Guangdong Innovative and Entrepreneurial Research Team Program (2016ZT06S638, 2016ZT06S252), The Sci-Tech Project Foundation of Guangzhou City (201707020039), Tanoto Foundation as Professorship in Medical Oncology, New Century Foundation Limited, Ling Foundation, and Singapore National Cancer Centre Research Fund, ONCO ACP Cancer Collaborative Scheme. We thank Professor Balram Chowbay, Heng Kee Khiang, and Jayme Wong Sau Yeng for logistic and technical support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nairismägi, M.L., Gerritsen, M.E., Li, Z.M. et al. Oncogenic activation of JAK3-STAT signaling confers clinical sensitivity to PRN371, a novel selective and potent JAK3 inhibitor, in natural killer/T-cell lymphoma. Leukemia 32, 1147–1156 (2018). https://doi.org/10.1038/s41375-017-0004-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-017-0004-x
This article is cited by
-
Optimization of immune checkpoint inhibitor treatment planning for relapsed or refractory extranodal NK/T cell lymphoma
Annals of Hematology (2024)
-
Aberrant JAK-STAT signaling-mediated chromatin remodeling impairs the sensitivity of NK/T-cell lymphoma to chidamide
Clinical Epigenetics (2023)
-
Pathogenesis and biomarkers of natural killer T cell lymphoma (NKTL)
Journal of Hematology & Oncology (2019)
-
Identifying subpathway signatures for individualized anticancer drug response by integrating multi-omics data
Journal of Translational Medicine (2019)
-
Molecular pathogenic pathways in extranodal NK/T cell lymphoma
Journal of Hematology & Oncology (2019)