Abstract
Objective
To test associations between grades 3 or 4 (severe) intraventricular hemorrhage (IVH) and single nucleotide polymorphisms (SNPs) associated with coagulation, inflammation, angiogenesis, and organ development in an exploratory study.
Study design
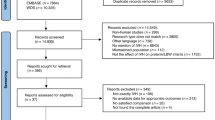
Extremely low-birthweight (ELBW) infants enrolled in the Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network’s (NRN) Cytokines Study were included if they had cranial ultrasound (CUS) and genotyping data available in the NRN Anonymized DNA Repository and Database. Associations between SNPs and IVH severity were tested with multivariable logistic regression analysis.
Result
One hundred thirty-nine infants with severe IVH and 687 infants with grade 1 or 0 IVH were included. One thousand two hundred seventy-nine SNPs were genotyped. Thirteen were preliminarily associated with severe IVH including five related to central nervous system (CNS) neuronal and neurovascular development.
Conclusion
Genetic variants for CNS neuronal and neurovascular development may be associated with severe IVH in premature infants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Change history
16 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41372-020-00859-w
References
Sherlock RL, Anderson PJ, Doyle LW, Victorian Infant Collaborative Study G. Neurodevelopmental sequelae of intraventricular haemorrhage at 8 years of age in a regional cohort of ELBW/very preterm infants. Early Hum Dev. 2005;81:909–16.
Stoll BJ, Hansen NI, Bell EF, Walsh MC, Carlo WA, Shankaran S, et al. Trends in care practices, morbidity, and mortality of extremely preterm neonates, 1993-2012. JAMA. 2015;314:1039–51.
Bhandari V, Bizzarro MJ, Shetty A, Zhong X, Page GP, Zhang H, et al. Familial and genetic susceptibility to major neonatal morbidities in preterm twins. Pediatrics. 2006;117:1901–6.
Petaja J, Hiltunen L, Fellman V. Increased risk of intraventricular hemorrhage in preterm infants with thrombophilia. Pediatr Res. 2001;49:643–6.
Ramenghi LA, Fumagalli M, Groppo M, Consonni D, Gatti L, Bertazzi PA, et al. Germinal matrix hemorrhage: intraventricular hemorrhage in very-low-birth-weight infants: the independent role of inherited thrombophilia. Stroke. 2011;42:1889–93.
Gopel W, Gortner L, Kohlmann T, Schultz C, Moller J. Low prevalence of large intraventricular haemorrhage in very low birthweight infants carrying the factor V Leiden or prothrombin G20210A mutation. Acta Paediatr. 2001;90:1021–4.
Ment LR, Aden U, Bauer CR, Bada HS, Carlo WA, Kaiser JR, et al. Genes and environment in neonatal intraventricular hemorrhage. Semin Perinatol. 2015;39:592–603.
Kenet G, Kuperman AA, Strauss T, Brenner B. Neonatal IVH-mechanisms and management. Thromb Res. 2011;127 Suppl 3:S120–2.
Piotrowski A, Dabrowska-Wojciak I, Mikinka M, Fendler W, Walas W, Sobala W, et al. Coagulation abnormalities and severe intraventricular hemorrhage in extremely low birth weight infants. J Matern Fetal Neonatal Med. 2010;23:601–6.
McDonald MM, Johnson ML, Rumack CM, Koops BL, Guggenheim MA, Babb C, et al. Role of coagulopathy in newborn intracranial hemorrhage. Pediatrics. 1984;74:26–31.
Setzer ES, Webb IB, Wassenaar JW, Reeder JD, Mehta PS, Eitzman DV. Platelet dysfunction and coagulopathy in intraventricular hemorrhage in the premature infant. J Pediatr. 1982;100:599–605.
Fulia F, Cordaro S, Meo P, Gitto P, Gitto E, Trimarchi G, et al. Can the administration of antithrombin III decrease the risk of cerebral hemorrhage in premature infants? Biol Neonate. 2003;83:1–5.
Dani C, Poggi C, Ceciarini F, Bertini G, Pratesi S, Rubaltelli FF. Coagulopathy screening and early plasma treatment for the prevention of intraventricular hemorrhage in preterm infants. Transfusion. 2009;49:2637–44.
Tran TT, Veldman A, Malhotra A. Does risk-based coagulation screening predict intraventricular haemorrhage in extreme premature infants? Blood Coagul Fibrinolysis. 2012;23:532–6.
Beverley DW, Pitts-Tucker TJ, Congdon PJ, Arthur RJ, Tate G. Prevention of intraventricular haemorrhage by fresh frozen plasma. Arch Dis Child. 1985;60:710–3.
Poralla C, Hertfelder HJ, Oldenburg J, Muller A, Bartmann P, Heep A. Elevated interleukin-6 concentration and alterations of the coagulation system are associated with the development of intraventricular hemorrhage in extremely preterm infants. Neonatology. 2012;102:270–5.
Villamor-Martinez E, Fumagalli M, Mohammed Rahim O, Passera S, Cavallaro G, Degraeuwe P, et al. Chorioamnionitis is a risk factor for intraventricular hemorrhage in preterm infants: a systematic review and meta-analysis. Front Physiol. 2018;9:1253.
Ryckman KK, Dagle JM, Kelsey K, Momany AM, Murray JC. Replication of genetic associations in the inflammation, complement, and coagulation pathways with intraventricular hemorrhage in LBW preterm neonates. Pediatr Res. 2011;70:90–5.
Szpecht D, Szymankiewicz M, Seremak-Mrozikiewicz A, Gadzinowski J. The role of genetic factors in the pathogenesis of neonatal intraventricular hemorrhage. Folia Neuropathol. 2015;53:1–7.
Carlo WA, McDonald SA, Tyson JE, Stoll BJ, Ehrenkranz RA, Shankaran S, et al. Cytokines and neurodevelopmental outcomes in extremely low birth weight infants. J Pediatr. 2011;159:919–25 e913.
Papile LA, Burstein J, Burstein R, Koffler H. Incidence and evolution of subependymal and intraventricular hemorrhage: a study of infants with birth weights less than 1,500 gm. J Pediatr. 1978;92:529–34.
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet. 2006;38:904–9.
Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP, et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics. 2010;26:2336–7.
Hartel C, Konig I, Koster S, Kattner E, Kuhls E, Kuster H, et al. Genetic polymorphisms of hemostasis genes and primary outcome of very low birth weight infants. Pediatrics. 2006;118:683–9.
Aden U, Lin A, Carlo W, Leviton A, Murray JC, Hallman M, et al. Candidate gene analysis: severe intraventricular hemorrhage in inborn preterm neonates. J Pediatr. 2013;163:1503–6.
Guerrero JA, Rivera J, Quiroga T, Martinez-Perez A, Anton AI, Martinez C, et al. Novel loci involved in platelet function and platelet count identified by a genome-wide study performed in children. Haematologica. 2011;96:1335–43.
Ken-Dror G, Drenos F, Humphries SE, Talmud PJ, Hingorani AD, Kivimaki M, et al. Haplotype and genotype effects of the F7 gene on circulating factor VII, coagulation activation markers and incident coronary heart disease in UK men. JTH. 2010;8:2394–403.
Bilguvar K, DiLuna ML, Bizzarro MJ, Bayri Y, Schneider KC, Lifton RP, et al. COL4A1 mutation in preterm intraventricular hemorrhage. J Pediatr. 2009;155:743–5.
Volpe JJ, Inder T, Darras B, de Vries LS, du Plessis A, Neil J, et al. Volpe’s neurology of the newborn. In: Volpe J, editor. 6th ed. Philadelphia, PA: Elsevier; 2017.
Baier RJ. Genetics of perinatal brain injury in the preterm infant. Front Biosci. 2006;11:1371–87.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs4240872#frequency_tab.
Velez DR, Fortunato SJ, Williams SM, Menon R. Interleukin-6 (IL-6) and receptor (IL6-R) gene haplotypes associate with amniotic fluid protein concentrations in preterm birth. Hum Mol Genet. 2008;17:1619–30.
Clark EAS, Weiner SJ, Rouse DJ, Mercer BM, Reddy UM, Iams JD, et al. Genetic variation, magnesium sulfate exposure, and adverse neurodevelopmental outcomes following preterm birth. Am J Perinatol. 2018;35:1012–22.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs10847980#publications.
Haataja R, Karjalainen MK, Luukkonen A, Teramo K, Puttonen H, Ojaniemi M, et al. Mapping a new spontaneous preterm birth susceptibility gene, IGF1R, using linkage, haplotype sharing, and association analysis. PLoS Genet. 2011;7:e1001293.
Parodi A, Morana G, Severino MS, Malova M, Natalizia AR, Sannia A, et al. Low-grade intraventricular hemorrhage: is ultrasound good enough? J Matern Fetal Neonatal Med. 2015;28(Suppl 1):2261–4.
Electronic Databased online: U.S. National Library of Medicine, National Institutes of Health. NCBI Insights [database]. National Center for Biotechnology Information: Bethesda, MD. https://ncbiinsights.ncbi.nlm.nih.gov/2017/05/08/dbsnps-human-build-150-has-doubled-the-amount-of-refsnp-records/.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs367024#clinical_significance.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs747004#clinical_significance.
Electronic Database online: Crown Human Genome Center. GeneCards [database]. Weizmann Institute of Science: Israel. https://www.genecards.org/cgi-bin/carddisp.pl?gene=IGF1R.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs12442623#clinical_significance.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs1513643#clinical_significance.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs1810225#clinical_significance.
Electronic Database online: e!Ensembl West [database]. European Molecular Biology Laboratory’s European Bioinformatics Institute: Hinxton, UK. http://useast.ensembl.org/Homo_sapiens/Variation/Citations?db=core;r=17:83257441-83257441;v=rs8072199;vdb=variation;vf=362660739.
Salam MT, Bastain TM, Rappaport EB, Islam T, Berhane K, Gauderman WJ, et al. Genetic variations in nitric oxide synthase and arginase influence exhaled nitric oxide levels in children. Allergy. 2011;66:412–9.
Folsom TD, Fatemi SH. The involvement of Reelin in neurodevelopmental disorders. Neuropharmacology. 2013;68:122–35.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. Gene [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/gene/?term=5649.
Chen N, Bao Y, Xue Y, Sun Y, Hu D, Meng S, et al. Meta-analyses of RELN variants in neuropsychiatric disorders. Behav Brain Res. 2017;332:110–9.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variations [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs1886233#publications.
Electronic Database online: Crown Human Genome Center. GeneCards [database]. Weizmann Institute of Science: Israel. https://www.genecards.org/cgi-bin/carddisp.pl?gene=GRIN3A.
Electronic Database online: U.S. National Library of Medicine, National Institutes of Health. dbSNP, Short genetic variataions [database]. National Center for Biotechnology Information: Bethesda, MD. https://www.ncbi.nlm.nih.gov/snp/rs13298667#publications.
Electronic Database online: Crown Human Genome Center. GeneCards [database]. Weizmann Institute of Science: Israel. https://www.genecards.org/cgi-bin/carddisp.pl?gene=TFAP2B.
Waleh N, Hodnick R, Jhaveri N, McConaghy S, Dagle J, Seidner S, et al. Patterns of gene expression in the ductus arteriosus are related to environmental and genetic risk factors for persistent ductus patency. Pediatr Res. 2010;68:292–7.
Dagle JM, Ryckman KK, Spracklen CN, Momany AM, Cotten CM, Levy J, et al. Genetic variants associated with patent ductus arteriosus in extremely preterm infants. J Perinatol. 2019;39:401–8.
Acknowledgements
We are indebted to our medical and nursing colleagues and the infants and their parents who agreed to take part in this study. Data collected at participating NRN sites were transmitted to RTI International, the data coordinating center (DCC) for the NRN, which stored, managed, and analyzed the data for this study. On behalf of the network, AD (DCC PI) and GPP (DCC Statistician) had full access to all the data in the study and take responsibility for the integrity of the data and accuracy of the data analysis.
Funding
The study was supported by the Children’s Miracle Network (CT). The National Institutes of Health (General Clinical Research Center grants M01 RR30, M01 RR32, M01 RR39, M01 RR70, M01 RR80, M01 RR633, M01 RR750, M01 RR997, M01 RR6022, M01 RR7122, M01 RR8084, M01 RR16587, UL1 RR24979) and the Eunice Kennedy Shriver National Institute of Child Health and Human Development (grants U01 HD36790, U10 HD21364, U10 HD21373, U10 HD21385, U10 HD21397, U10 HD21415, U10 HD27851, U10 HD27853, U10 HD27856, U10 HD27871, U10 HD27880, U10 HD27881, U10 HD27904, U10 HD34216, U10 HD40461, U10 HD40492, U10 HD40498, U10 HD40689, U10 HD53109) provided grant support for the Neonatal Research Network’s Glutamine trial and Genomics secondary study. JCM received assistance for the GENEVA study from the National Human Genome Research Institute (U01 HG4423). MEH received funding from the National Eye Institute (2R01EY015130, 5R01EY017011) and a Department Grant from Research to Prevent Blindness to the Department of Ophthalmology and Visual Science. The funding agencies provided overall oversight for study conduct, but all data analyses and interpretation were independent of the funding agencies.
Author information
Authors and Affiliations
Consortia
Contributions
CDT and CMC conceptualized and designed the analysis and drafted, reviewed, and revised the manuscript. SWE and GPP conducted the data analysis and reviewed and revised the manuscript. EASC, MMD, MEH, RFG, JMD, JCM, BBP, and AD collected data and critically reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to report. The contents of this report represent the views of the authors and do not represent the views of the Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network or the National Institutes of Health.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Members of the Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network and their affiliations are presented in Supplementary information.
Rights and permissions
About this article
Cite this article
Thornburg, C.D., Erickson, S.W., Page, G.P. et al. Genetic predictors of severe intraventricular hemorrhage in extremely low-birthweight infants. J Perinatol 41, 286–294 (2021). https://doi.org/10.1038/s41372-020-00821-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41372-020-00821-w